Plants Specifically Modulate the Microbiome of Root-Lesion Nematodes in the Rhizosphere, Affecting Their Fitness
Abstract
1. Introduction
2. Materials and Methods
2.1. Preparation of Nematodes
2.2. Experimental Design: Microbiome Associated with RLN as Affected by Plant Species
2.2.1. Preparation of Rhizosphere and Bulk Soil Suspensions
2.2.2. Baiting of Soil Microbes Attaching to the Cuticle of P. penetrans
2.2.3. Denaturing Gradient Gelelectrophoresis (DGGE) Profiling of Nematode-Attached Microbiomes
2.2.4. Characterization of Bacterial Isolates
2.3. Biological Effects of the Nematode-Attached Microbiome
2.4. Effects of Root Exudates on Microbial Attachment to Nematodes
2.5. Data Analysis and Statistics
3. Results
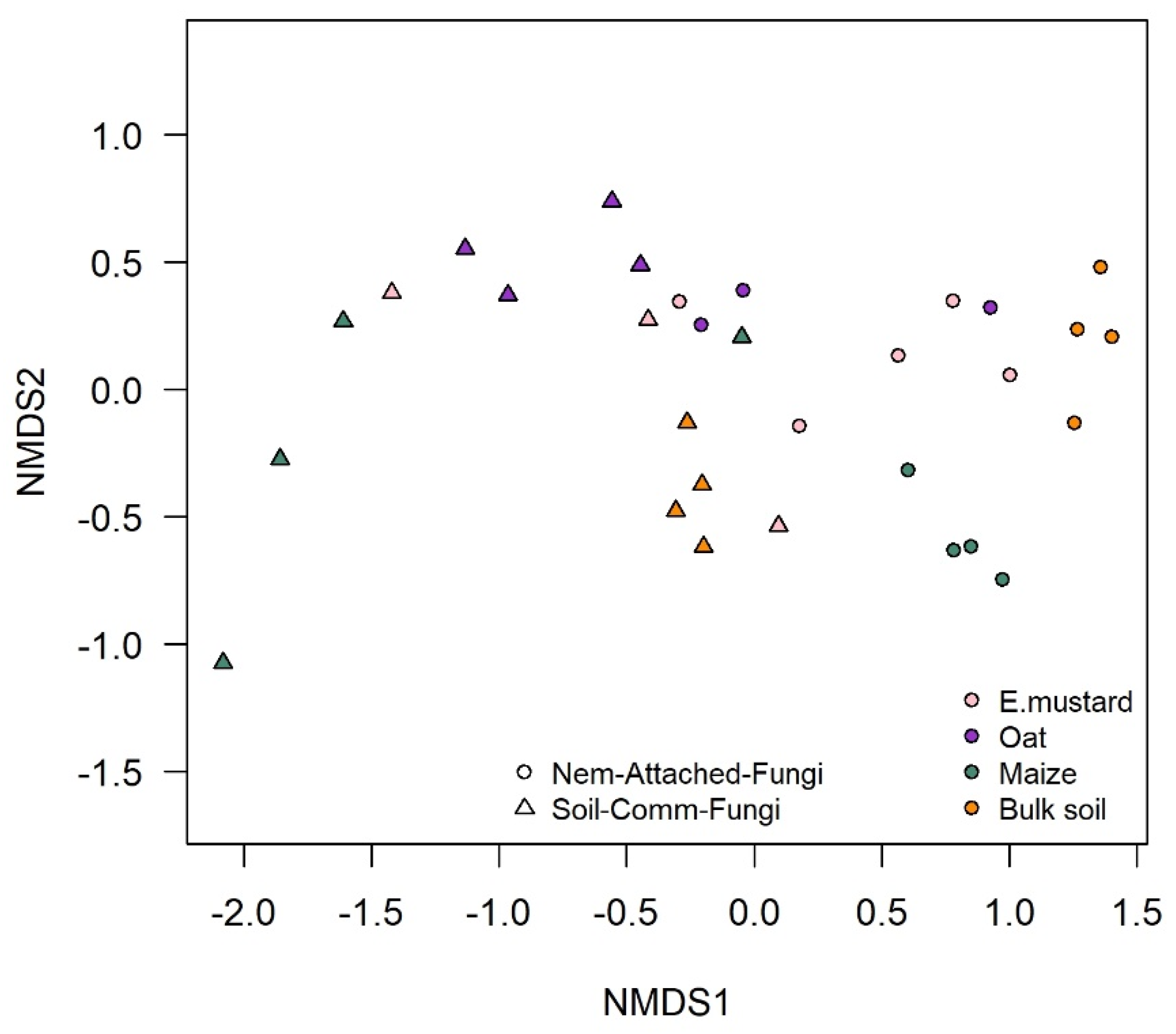
3.1. Rhizosphere Fungi Attaching to P. penetrans Depended on Plant Species
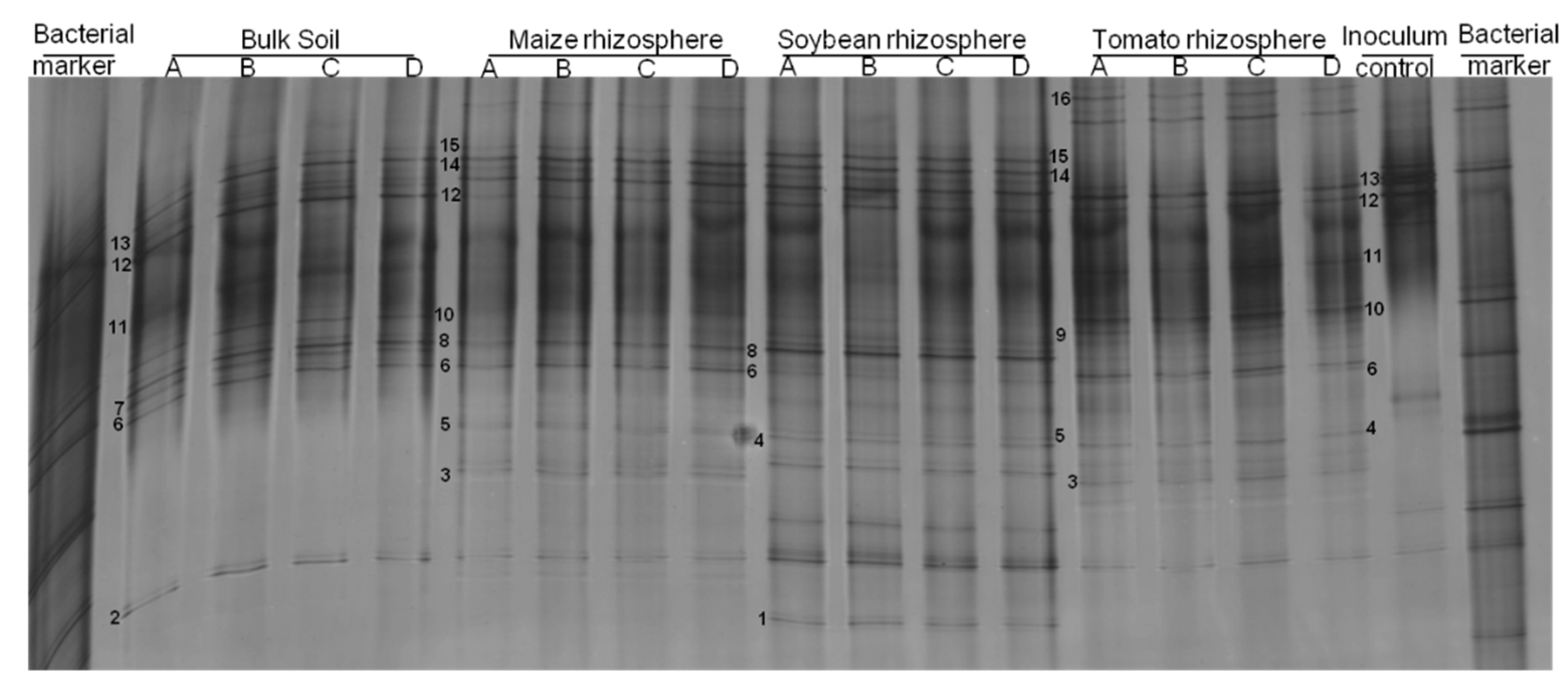
3.2. Rhizosphere Bacteria Attaching to P. penetrans Depended on Plant Species
3.3. Isolation and Characterization of Cuticle-Attached Bacteria
3.4. Effect of Cuticle-Attached Microbiomes on the Mortality of P. penetrans
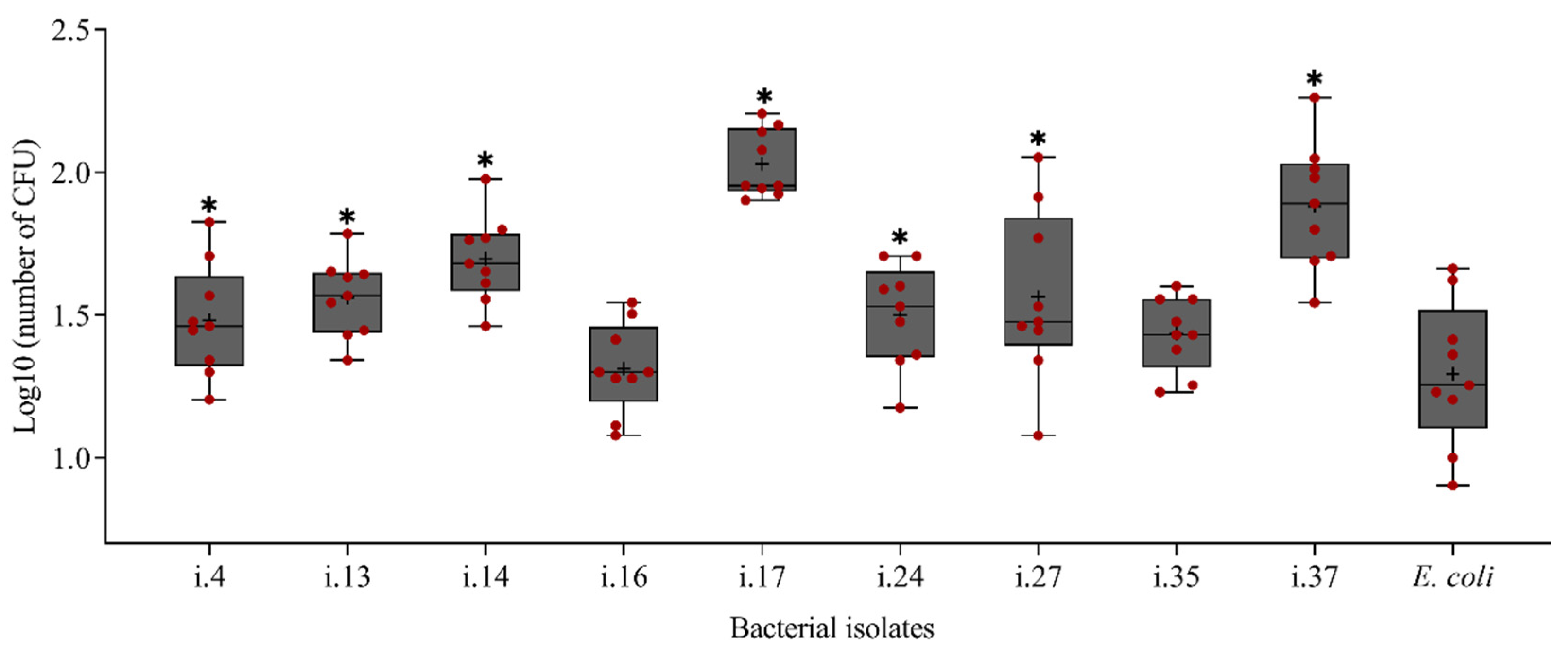
3.5. Effect of Cuticle-Attached Bacterial Isolates on the Mortality of P. penetrans
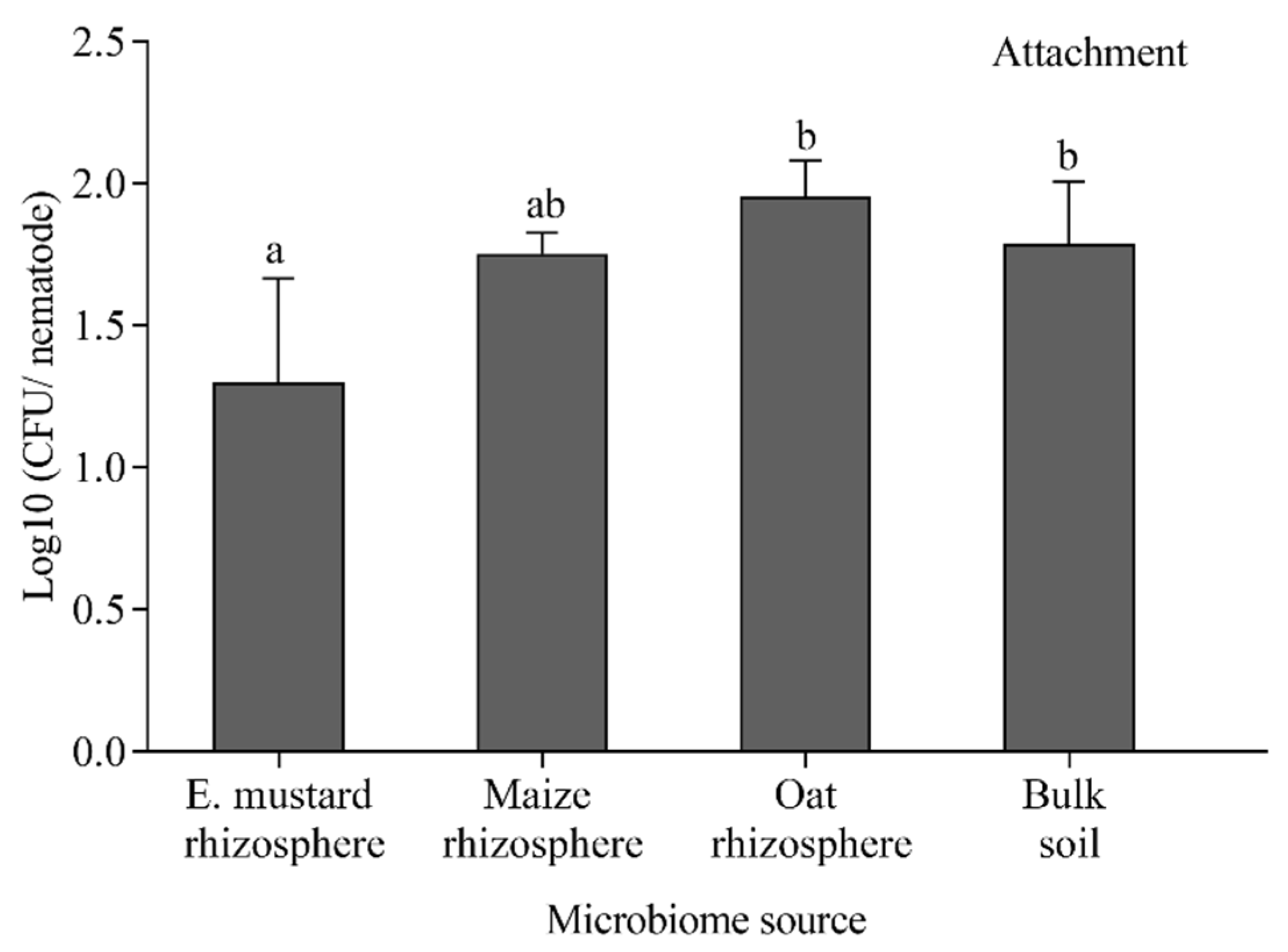
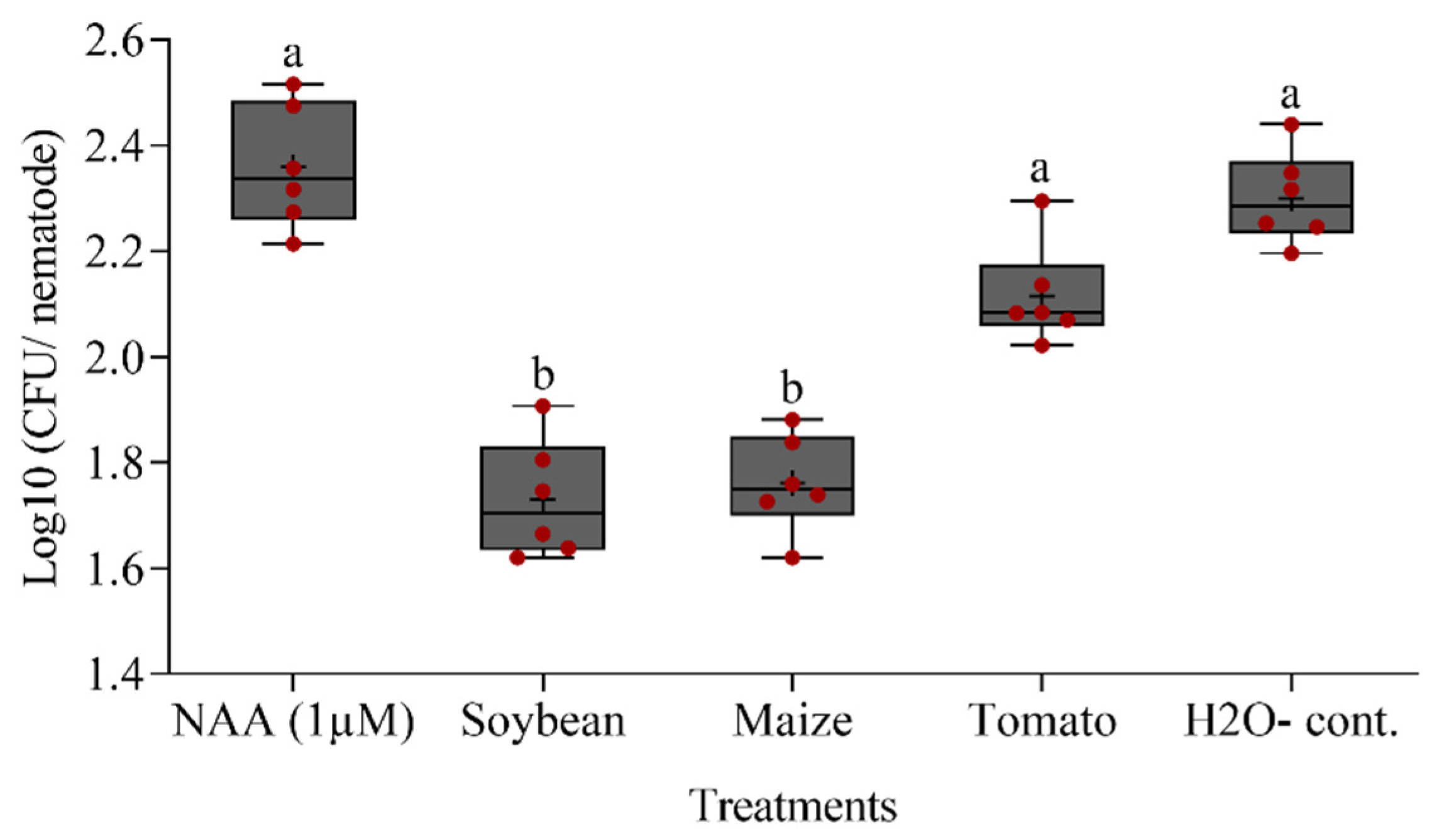
3.6. Effects of Root Exudates on the Attachment of Bacteria
4. Discussion
4.1. Plants Govern the Microbiome Associated with the Cuticle of P. penetrans in the Rhizosphere
4.2. Effects of Nematode-Attached Microbiome on Nematode Fitness
4.3. Role of Root Exudates in Microbial Attachment and Suppression of P. penetrans
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jones, J.T.; Haegeman, A.; Danchin, E.G.J.; Gaur, H.S.; Helder, J.; Jones, M.G.K.; Kikuchi, T.; Manzanilla-López, R.; Palomares-Rius, J.E.; Wesemael, W.M.L.; et al. Top 10 plant-parasitic nematodes in molecular plant pathology. Mol. Plant Pathol. 2013, 14, 946–961. [Google Scholar] [CrossRef] [PubMed]
- Fosu-Nyarko, J.; Jones, M.G. Advances in understanding the molecular mechanisms of root lesion nematode host interactions. Annu. Rev. Phytopathol. 2016, 54, 253–278. [Google Scholar] [CrossRef]
- Topalović, O.; Bredenbruch, S.; Schleker, A.S.S.; Heuer, H. Microbes attaching to endoparasitic phytonematodes in soil trigger plant defense upon root penetration by the nematode. Front. Plant Sci. 2020, 11, 138. [Google Scholar] [CrossRef] [PubMed]
- Topalović, O.; Heuer, H. Plant-nematode interactions assisted by microbes in the rhizosphere. Curr. Issues Mol. Biol. 2019, 30, 75–88. [Google Scholar] [CrossRef] [PubMed]
- Topalović, O.; Hussain, M.; Heuer, H. Plants and associated soil microbiota cooperatively suppress plant-parasitic nematodes. Front. Microbiol. 2020, 11, 313. [Google Scholar] [CrossRef] [PubMed]
- Zilber-Rosenberg, I.; Rosenberg, E. Role of microorganisms in the evolution of animals and plants: The hologenome theory of evolution. FEMS Microbiol. Rev. 2008, 32, 723–735. [Google Scholar] [CrossRef]
- Rosenberg, E.; Zilber-Rosenberg, I. The hologenome concept of evolution after 10 years. Microbiome 2018, 6, 1–14. [Google Scholar] [CrossRef]
- Sánchez-Cañizares, C.; Jorrín, B.; Poole, P.S.; Tkacz, A. Understanding the holobiont: The interdependence of plants and their microbiome. Curr. Opin. Microbiol. 2017, 38, 188–196. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Rott, M.; Schlaeppi, K.; Van Themaat, E.V.L.; Ahmadinejad, N.; Assenza, F.; Rauf, P.; Huettel, B.; Reinhardt, R.; Schmelzer, E.; et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nat. Cell Biol. 2012, 488, 91–95. [Google Scholar] [CrossRef]
- Lundberg, D.S.; Lebeis, S.L.; Paredes, S.H.; Yourstone, S.; Gehring, J.; Malfatti, S.; Tremblay, J.; Engelbrektson, A.; Kunin, V.; Del Rio, T.G.; et al. Defining the core Arabidopsis thaliana root microbiome. Nat. Cell Biol. 2012, 488, 86–90. [Google Scholar] [CrossRef] [PubMed]
- Elhady, A.; Adss, S.; Hallmann, J.; Heuer, H. Rhizosphere microbiomes modulated by pre-crops assisted plants in defense against plant-parasitic nematodes. Front. Microbiol. 2018, 9, 1133. [Google Scholar] [CrossRef] [PubMed]
- Fierer, N.; Jackson, R.B. The diversity and biogeography of soil bacterial communities. Proc. Natl. Acad. Sci. USA 2006, 103, 626–631. [Google Scholar] [CrossRef] [PubMed]
- Haichar, F.E.Z.; Santaella, C.; Heulin, T.; Achouak, W. Root exudates mediated interactions belowground. Soil Biol. Biochem. 2014, 77, 69–80. [Google Scholar] [CrossRef]
- Bertin, C.; Yang, X.; Weston, L.A. The role of root exudates and allelochemicals in the rhizosphere. Plant Soil 2003, 256, 67–83. [Google Scholar] [CrossRef]
- Bais, H.P.; Weir, T.L.; Perry, L.G.; Gilroy, S.; Vivanco, J.M. The role of root exudates in rhizosphere interactions with plants and other organisms. Annu. Rev. Plant Biol. 2006, 57, 233–266. [Google Scholar] [CrossRef]
- Adam, M.; Westphal, A.; Hallmann, J.; Heuer, H. Specific microbial attachment to root knot nematodes in suppressive soil. Appl. Environ. Microbiol. 2014, 80, 2679–2686. [Google Scholar] [CrossRef]
- Davies, K.G.; Curtis, R.H.C. Cuticle surface coat of plant-parasitic nematodes. Annu. Rev. Phytopathol. 2011, 49, 135–156. [Google Scholar] [CrossRef]
- Danks, C.; Davies, K. Carbohydrate/Protein interactions between the cuticle of infective juveniles of Meloidogyne incognita and spores of the obligate hyperparasite Pasteuria penetrans. Nematology 1993, 39, 53–64. [Google Scholar] [CrossRef]
- Grenache, D.G.; Caldicott, I.; Albert, P.S.; Riddle, D.L.; Politz, S.M. Environmental induction and genetic control of surface antigen switching in the nematode Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 1996, 93, 12388–12393. [Google Scholar] [CrossRef]
- Olsen, D.P.; Phu, D.; Libby, L.J.M.; Cormier, J.A.; Montez, K.M.; Ryder, E.F.; Politz, S.M. Chemosensory control of surface antigen switching in the nematode Caenorhabditis elegans. Genes Brain Behav. 2007, 6, 240–252. [Google Scholar] [CrossRef]
- Liu, C.; Timper, P.; Ji, P.; Mekete, T.; Joseph, S. Influence of root exudates and soil on attachment of Pasteuria penetrans to Meloidogyne arenaria. J. Nematol. 2017, 49, 304–310. [Google Scholar] [CrossRef]
- Singh, J.; Kumar, M.U.; Walia, R.K. Influence of plant root exudates on the adherence of Pasteuria penetrans endospores. Nematology 2014, 16, 121–124. [Google Scholar] [CrossRef]
- Bell, C.A.; Lilley, C.J.; McCarthy, J.; Atkinson, H.J.; Urwin, P.E. Plant-parasitic nematodes respond to root exudate signals with host-specific gene expression patterns. PLoS Pathog. 2019, 15, e1007503. [Google Scholar] [CrossRef] [PubMed]
- Duarte, A.; Maleita, C.; Abrantes, I.; Curtis, R. Tomato root exudates induce transcriptional changes of Meloidogyne hispanica genes. Phytopathol. Mediterr. 2015, 54, 104–108. [Google Scholar] [CrossRef]
- Topalović, O.; Elhady, A.; Hallmann, J.; Richert-Pöggeler, K.R.; Heuer, H. Bacteria isolated from the cuticle of plant-parasitic nematodes attached to and antagonized the root-knot nematode Meloidogyne hapla. Sci. Rep. 2019, 9, 11477. [Google Scholar] [CrossRef]
- Engelen, B.; Meinken, K.; Von Wintzingerode, F.; Heuer, H.; Malkomes, H.-P.; Backhaus, H. Monitoring impact of a pesticide treatment on bacterial soil communities by metabolic and genetic fingerprinting in addition to conventional testing procedures. Appl. Environ. Microbiol. 1998, 64, 2814–2821. [Google Scholar] [CrossRef] [PubMed]
- Weinert, N.; Meincke, R.; Gottwald, C.; Heuer, H.; Gomes, N.C.M.; Schloter, M.; Berg, G.; Smalla, K. Rhizosphere communities of genetically modified zeaxanthin-accumulating potato plants and their parent cultivar differ less than those of different potato cultivars. Appl. Environ. Microbiol. 2009, 75, 3859–3865. [Google Scholar] [CrossRef][Green Version]
- Kropf, S.; Heuer, H.; Grüning, M.; Smalla, K. Significance test for comparing complex microbial community fingerprints using pairwise similarity measures. J. Microbiol. Methods 2004, 57, 187–195. [Google Scholar] [CrossRef] [PubMed]
- Versalovic, J.; Schneider, M.; de Bruijn, F.J.; Lupski, J.R. Genomic fingerprinting of bacteria using repetitive sequence based PCR (rep-PCR). Methods Mol. Cell. Biol. 1998, 5, 25–40. [Google Scholar]
- Elhady, A.; Giné, A.; Topalovic, O.; Jacquiod, S.; Sørensen, S.J.; Sorribas, F.J.; Heuer, H. Microbiomes associated with infective stages of root-knot and lesion nematodes in soil. PLoS ONE 2017, 12, e0177145. [Google Scholar] [CrossRef]
- Quentin, M.; Eabad, P.; Efavery, B. Plant parasitic nematode effectors target host defense and nuclear functions to establish feeding cells. Front. Plant Sci. 2013, 4, 53. [Google Scholar] [CrossRef]
- Topalović, O.; Heuer, H.; Reineke, A.; Zinkernagel, J.; Hallmann, J. Antagonistic role of the microbiome from a Meloidogyne hapla-suppressive soil against species of plant-parasitic nematodes with different life strategies. Nematology 2019, 22, 75–86. [Google Scholar] [CrossRef]
- Poole, P. Shining a light on the dark world of plant root–microbe interactions. Proc. Natl. Acad. Sci. USA 2017, 114, 4281–4283. [Google Scholar] [CrossRef]
- Akhkha, A.; Kusel, J.; Kennedy, M.; Curtis, R. Effects of phytohormones on the surfaces of plant-parasitic nematodes. Parasitol. 2002, 125, 165–175. [Google Scholar] [CrossRef]
- De Mendoza, M.E.L.; Modha, J.; Roberts, M.C.; Curtis, R.; Kusel, J.R. Changes in the lipophilicity of the surfaces of Meloidogyne incognita and Haemonchus contortus during exposure to host signals. Parasitology 2000, 120, 203–209. [Google Scholar] [CrossRef]
- Ali, M.A.; Anjam, M.S.; Nawaz, M.A.; Lam, H.-M.; Chung, G. Signal transduction in plant–nematode interactions. Int. J. Mol. Sci. 2018, 19, 1648. [Google Scholar] [CrossRef] [PubMed]
- Lok, J.B. Signaling in parasitic nematodes: Physicochemical communication between host and parasite and endogenous molecular transduction pathways governing worm development and survival. Curr. Clin. Microbiol. Rep. 2016, 3, 186–197. [Google Scholar] [CrossRef] [PubMed]
- Scharf, B.E.; Hynes, M.F.; Alexandre, G.M. Chemotaxis signaling systems in model beneficial plant–bacteria associations. Plant Mol. Biol. 2016, 90, 549–559. [Google Scholar] [CrossRef] [PubMed]
- Wuichet, K.; Zhulin, I.B. Origins and diversification of a complex signal transduction system in prokaryotes. Sci. Signal. 2010, 3, ra50. [Google Scholar] [CrossRef] [PubMed]
- Burdman, S.; Dulguerova, G.; Okon, Y.; Jurkevitch, E. Purification of the major outer membrane protein of Azospirillum brasilense, its affinity to plant roots, and its involvement in cell aggregation. Mol. Plant Microbe Interact. 2001, 14, 555–561. [Google Scholar] [CrossRef]
- Maghodia, A.; Spiegel, Y.; Sela, S. Interactions between Escherichia coli and the plant-parasitic nematode Meloidogyne javanica. J. Appl. Microbiol. 2008, 105, 1810–1816. [Google Scholar] [CrossRef]
- Rosso, M.N.; Jones, J.T.; Abad, P. RNAi and functional genomics in plant parasitic nematodes. Annu. Rev. Phytopathol. 2009, 47, 207–232. [Google Scholar] [CrossRef]
- Chen, J.; Moore, W.H.; Yuen, G.Y.; Kobayashi, D.; Caswell-Chen, E.P. Influence of Lysobacter enzymogenes strain C3 on nematodes. J. Nematol. 2006, 38, 233–239. [Google Scholar] [PubMed]
- Dicklow, B.M.; Acosta, N.; Zuckerman, B.M. A novel Streptomyces species for controlling plant-parasitic nematodes. J. Chem. Ecol. 1993, 19, 159–173. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.; Qi, G.; Yin, R.; Zhang, H.; Li, C.; Zhao, X. Bacillus cereus strain S2 shows high nematicidal activity against Meloidogyne incognita by producing sphingosine. Sci. Rep. 2016, 6, 28756. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.S.; Park, Y.S.; Anees, M.; Kim, Y.C.; Kim, Y.H.; Kim, K.Y. Nematicidal activity of Lysobacter capsici YS1215 and the role of gelatinolytic proteins against root-knot nematodes. Biocontrol Sci. Technol. 2013, 23, 1427–1441. [Google Scholar] [CrossRef]
- Lee, Y.S.; Anees, M.; Hyun, H.N.; Kim, K.Y. Biocontrol potential of Lysobacter antibioticus HS124 against the root-knot nematode, Meloidogyne incognita, causing disease in tomato. Nematology 2013, 15, 545–555. [Google Scholar] [CrossRef]
- Samac, D.A.; Kinkel, L.L. Suppression of the root-lesion nematode (Pratylenchus penetrans) in alfalfa (Medicago sativa) by Streptomyces spp. Plant Soil 2001, 235, 35–44. [Google Scholar] [CrossRef]
- Shan, S.; Wang, W.; Song, C.; Wang, M.; Sun, B.; Li, Y.; Fu, Y.; Gu, X.; Ruan, W.; Rasmann, S. The symbiotic bacteria Alcaligenes faecalis of the entomopathogenic nematodes Oscheius spp. exhibit potential biocontrol of plant- and entomopathogenic fungi. Microb. Biotechnol. 2019, 12, 459–471. [Google Scholar] [CrossRef]
- Siddiqui, Z.; Mahmood, I. Role of bacteria in the management of plant parasitic nematodes: A review. Bioresour. Technol. 1999, 69, 167–179. [Google Scholar] [CrossRef]
- Wei, J.-Z.; Siehl, D.L.; Hou, Z.; Rosen, B.; Oral, J.; Taylor, C.G.; Wu, G. An Enterotoxin-Like Binary Protein from Pseudomonas protegens with Potent Nematicidal Activity. Appl. Environ. Microbiol. 2017, 83, 00942-17. [Google Scholar] [CrossRef] [PubMed]
- Ruanpanun, P.; Laatsch, H.; Tangchitsomkid, N.; Lumyong, S. Nematicidal activity of fervenulin isolated from a nematicidal actinomycete, Streptomyces sp. CMU-MH021, on Meloidogyne incognita. World J. Microbiol. Biotechnol. 2010, 27, 1373–1380. [Google Scholar] [CrossRef] [PubMed]
- Van Peer, R.; Niemann, G.J.; Schippers, B. Induced resistance and phytoalexin accumulation in biological control of Fusarium wilt of carnation by Pseudomonas sp. strain WCS417r. Phytopathology 1991, 81, 728–734. [Google Scholar] [CrossRef]
- Adam, M.; Heuer, H.; Hallmann, J. Bacterial antagonists of fungal pathogens also control root-knot nematodes by induced systemic resistance of tomato plants. PLoS ONE 2014, 9, e90402. [Google Scholar] [CrossRef]
- Reitz, M.; Oger, P.; Farrand, S.K.; Hallmann, J.; Meyer, A.; Sikora, R.A.; Niehaus, K. Importance of the O-antigen, core-region and lipid A of rhizobial lipopolysaccharides for the induction of systemic resistance in potato to Globodera pallida. Nematology 2002, 4, 73–79. [Google Scholar] [CrossRef]
- Siddiqui, I.A.; Shaukat, S.S. Systemic resistance in tomato induced by biocontrol bacteria against the root-knot nematode, Meloidogyne javanica is independent of salicylic acid production. J. Phytopathol. 2004, 152, 48–54. [Google Scholar] [CrossRef]
- Selim, M.E.; Mahdy, M.E.; Sorial, M.E.; Dababat, A.A.; Sikora, R.A. Biological and chemical dependent systemic resistance and their significance for the control of root-knot nematodes. Nematology 2014, 16, 917–927. [Google Scholar] [CrossRef]
- Martínez-Medina, A.; Fernandez, I.; Lok, G.B.; Pozo, M.J.; Pieterse, C.M.J.; Van Wees, S.C.M. Shifting from priming of salicylic acid- to jasmonic acid-regulated defences by Trichoderma protects tomato against the root knot nematode Meloidogyne incognita. New Phytol. 2017, 213, 1363–1377. [Google Scholar] [CrossRef]
- Kerry, B.R. Rhizosphere interactions and the exploitation of microbial agents for the biological control of plant-parasitic nematodes. Annu. Rev. Phytopathol. 2000, 38, 423–441. [Google Scholar] [CrossRef]
- Zhou, D.; Feng, H.; Schuelke, T.; De Santiago, A.; Zhang, Q.; Zhang, J.; Luo, C.; Wei, L. Rhizosphere microbiomes from root knot nematode non-infested plants suppress nematode infection. Microb. Ecol. 2019, 78, 470–481. [Google Scholar] [CrossRef]
- Piśkiewicz, A.M.; De Milliano, M.J.K.; Duyts, H.; Van Der Putten, W.H. Plant ectoparasitic nematodes prefer roots without their microbial enemies. Plant Soil 2008, 316, 277–284. [Google Scholar] [CrossRef]
- Yang, G.; Zhou, B.; Zhang, X.; Zhang, Z.; Wu, Y.; Zhang, Y.; Lü, S.; Zou, Q.; Gao, Y.; Teng, L. Effects of tomato root exudates on Meloidogyne incognita. PLoS ONE 2016, 11, e0154675. [Google Scholar] [CrossRef] [PubMed]





| Isolate | Source | Most Similar Sequence, Accession No. | Identity |
|---|---|---|---|
| i.1 | Bulk soil | Streptomyces violaceoruber, NR_112292.1 | 99% |
| i.3 | Bulk soil | Streptomyces atratus, NR_043490.1 | 100% |
| i.4 | Bulk soil | Bacillus marisflavi, NR_118437.1 | 99% |
| i.7 | Bulk soil | Nocardia coeliaca, NR_104776.1 | 99% |
| i.9 | Bulk soil | Mycobacterium madagascariense, NR_104690.1 | 99% |
| i.13 | Bulk soil | Microbacterium maritypicum, NR_114986.1 | 99% |
| i.14 | Bulk soil | Microbacterium mangrovi, NR_126283.1 | 99% |
| i.16 | Bulk soil | Delftia tsuruhatensis, NR_113870.1 | 99% |
| i.17 | Bulk soil | Lysobacter capsici, NR_044250.1 | 100% |
| i.23 | Maize rhizosphere | Novosphingobium aquaticum, NR_148323.1 | 99% |
| i.24 | Maize rhizosphere | Bacillus cereus, NR_074540.1 | 99% |
| i.26 | Maize rhizosphere | Pedobacter borealis, NR_044381.1 | 99% |
| i.27 | Maize rhizosphere | Pseudomonas protegens, NR_114749.1 | 99% |
| i.35 | Soybean rhizosphere | Bacillus megaterium, NR_116873.1 | 100% |
| i.37 | Soybean rhizosphere | Alcaligenes faecalis, NR_113606.1 | 99% |
| i.42 | Soybean rhizosphere | Rhizobium nepotum, NR_117203.1 | 99% |
| i.47 | Soybean rhizosphere | Bacillus megaterium, NR_116873.1 | 99% |
| i.50 | Soybean rhizosphere | Mycobacterium chubuense, NR_041902.1 | 99% |
| i.55 | Soybean rhizosphere | Bacillus aryabhattai, NR_115953.1, | 100% |
| i.63 | Tomato rhizosphere | Staphylococcus capitis, NR_113348.1 | 100% |
| Sources of Soil Suspensions for Pairwise Comparison of DGGE Fingerprints of Nematode-Attached Bacteria or Fungi | Dissimilarity (%) a | ||
|---|---|---|---|
| Bacteria | Fungi | ||
| Experiment (1) | Bulk soil vs. maize rhizosphere | 23 | 62 |
| Bulk soil vs. tomato rhizosphere | 35 | 88 | |
| Bulk soil vs. soybean rhizosphere | 40 | 65 | |
| Maize vs. tomato rhizosphere | 43 | 87 | |
| Maize vs. soybean rhizosphere | 51 | 43 | |
| Soybean vs. tomato rhizosphere | 68 | 57 | |
| Experiment (2) | Bulk soil vs. maize rhizosphere | 34 | 14 |
| Bulk soil vs. oat rhizosphere | 31 | 28 | |
| Bulk soil vs. Ethiopian mustard rhizosphere | 12 | 10 | |
| Maize vs. oat rhizosphere | 29 | 29 | |
| Maize vs. Ethiopian mustard rhizosphere | 36 | 17 | |
| Oat vs. Ethiopian mustard rhizosphere | 22 | 3 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Elhady, A.; Topalović, O.; Heuer, H. Plants Specifically Modulate the Microbiome of Root-Lesion Nematodes in the Rhizosphere, Affecting Their Fitness. Microorganisms 2021, 9, 679. https://doi.org/10.3390/microorganisms9040679
Elhady A, Topalović O, Heuer H. Plants Specifically Modulate the Microbiome of Root-Lesion Nematodes in the Rhizosphere, Affecting Their Fitness. Microorganisms. 2021; 9(4):679. https://doi.org/10.3390/microorganisms9040679
Chicago/Turabian StyleElhady, Ahmed, Olivera Topalović, and Holger Heuer. 2021. "Plants Specifically Modulate the Microbiome of Root-Lesion Nematodes in the Rhizosphere, Affecting Their Fitness" Microorganisms 9, no. 4: 679. https://doi.org/10.3390/microorganisms9040679
APA StyleElhady, A., Topalović, O., & Heuer, H. (2021). Plants Specifically Modulate the Microbiome of Root-Lesion Nematodes in the Rhizosphere, Affecting Their Fitness. Microorganisms, 9(4), 679. https://doi.org/10.3390/microorganisms9040679








