A Narrative Review of the Molecular Epidemiology and Laboratory Surveillance of Vaccine Preventable Bacterial Meningitis Agents: Streptococcus pneumoniae, Neisseria meningitidis, Haemophilus influenzae and Streptococcus agalactiae
Abstract
1. Introduction
2. Characteristics of S. pneumoniae, N. meningitidis, H. influenzae, and S. agalactiae Important for Vaccination and Surveillance
3. Currently Licensed Vaccines for Control of Bacterial Meningitis
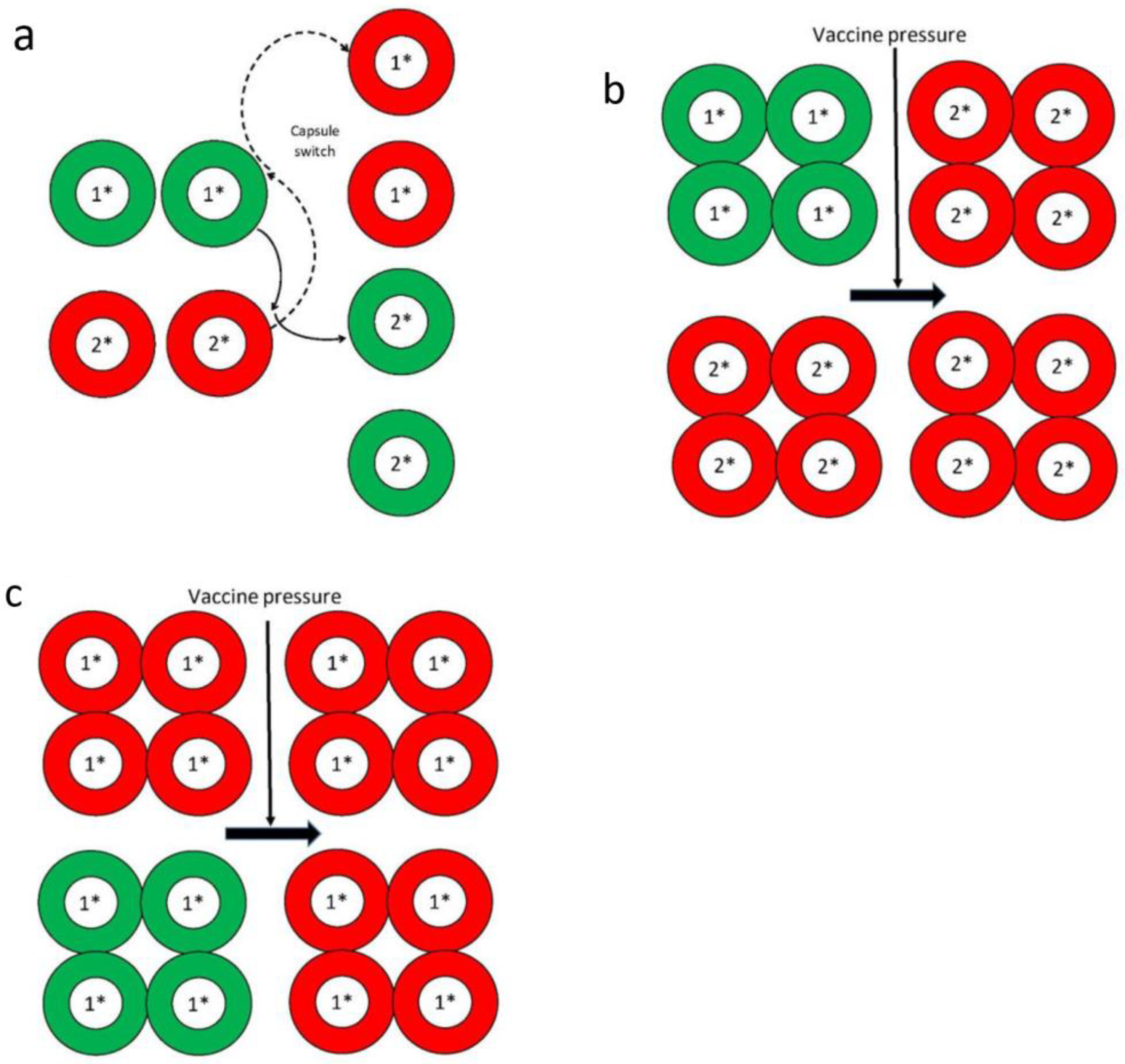
4. Effects of Vaccine Pressure, and Immune and/or Antibiotic Selection
5. Molecular Epidemiology of Invasive Pneumococcal Disease (IPD), Invasive Meningococcal Disease (IMD), and Invasive H. influenzae Disease in the Post Conjugate Vaccine Era
5.1. IPD in the Post PCV Era
5.2. IMD in the Post Conjugate Vaccine Era
5.3. Invasive H. influenzae Disease in the Post Hib Conjugate Vaccine Era
6. Laboratory Surveillance of S. pneumoniae, N. meningitidis, and H. influenzae
7. Whole Genome Sequencing (WGS) for Molecular Epidemiology and Genomic Surveillance of Vaccine-Preventable Bacterial Meningitis Agents
8. Chemoprophylaxis, Corticosteroids, and Experimental Immune Modulating Approaches for Prevention and as Adjuvant Therapeutic Agents of Bacterial Meningitis
9. Looking Ahead and What to Expect in the Post-Genomic Era of Meningitis Control
10. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Tunkel, A.R.; van der Beek, D.; Scheld, W.M. Acute meningitis. In Mandell, Douglas, and Bennett’s Principles and Practice of Infectious Diseases, 7th ed.; Mandell, G.L., Bennett, J.E., Dolin, R., Eds.; Churchill Livingstone Elsevier: Philadelphia, PA, USA, 2010. [Google Scholar]
- Kassebaum, N.J.; Zunt, J.R.; for the GBD Meningitis Collaborators. Global, regional and national burden of meningitis 1990-2016: A systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 2018, 17, 1061–1082. [Google Scholar] [CrossRef]
- Edmond, K.; Clark, A.; Korczak, V.S.; Sanderson, C.; Griffiths, U.K.; Rudan, I. Global and regional risk of disabling sequelae from bacterial meningitis: A systematic review and meta-analysis. Lancet Infect. Dis. 2010, 10, 317–328. [Google Scholar] [CrossRef]
- Wright, C.; Wordsworth, R.; Glennie, L. Counting the cost of meningococcal disease, scenarios of severe meningitis and septicemia. Pediatr. Drugs 2013, 15, 49–58. [Google Scholar] [CrossRef]
- Song, J.Y.; Lim, J.H.; Lim, S.; Yong, Z.; Seo, S.H. Progress towards a group B streptococcal vaccine. Hum. Vaccines Immunother. 2018, 14, 2669–2681. [Google Scholar] [CrossRef]
- Nanduri, S.A.; Petit, S.; Smelser, C.; Apostol, M.; Alden, N.B.; Harrison, L.H.; Lynfield, R.; Vagnone, P.S.; Burzlaff, K.; Spina, N.L.; et al. Epidemiology of invasive early-onset and late-onset group B streptococcal disease in the United States, 2006 to 2015: Multistate laboratory and population-based surveillance. JAMA Pediatr. 2019, 173, 224–233. [Google Scholar] [CrossRef] [PubMed]
- Watkins, L.K.F.; McGee, L.; Schrag, S.J.; Beall, B.; Jain, J.H.; Ponda, T.; Farley, M.M.; Harrison, L.H.; Zansky, S.M.; Baumbach, J.; et al. Epidemiology of invasive group B streptococcal infections among non-pregnant adults in the United States, 2008–2016, JAMA Intern. Med. 2019, 179, 479–488. [Google Scholar]
- World Health Organization. Defeating meningitis by 2030: A Global Road Map. 2018. Available online: https://www.who.int/immunization/research/development/DefeatingMeningitisRoadmap.pdf (accessed on 25 December 2020).
- Black, S.; Shinefield, H.; Fireman, B.; Lewis, E.; Ray, P.; Hansen, J.R.; Elvin, L.; Ensor, K.M.; Hackell, J.; Siber, G.; et al. Efficacy, safety and immunogenicity of heptavalent pneumococcal conjugate vaccine in children. Northern California Kaiser Permanente Vaccine Study Center Group. Pediatr. Infect. Dis. J. 2000, 19, 187–195. [Google Scholar] [CrossRef] [PubMed]
- Kelly, D.F.; Moxon, E.R.; Pollard, A.J. Haemophilus influenzae type b conjugate vaccines. Immunology 2004, 113, 163–174. [Google Scholar] [CrossRef] [PubMed]
- Harrison, L.H. A multivalent conjugate vaccine for prevention of meningococcal disease in infants. JAMA 2009, 299, 217–219. [Google Scholar] [CrossRef] [PubMed]
- McIntyre, P.B.; O’Brien, K.L.; Greenwood, B.; van de Beek, D. Bacterial meningitis: Effect of vaccines on bacterial meningitis worldwide. Lancet 2012, 380, 1703–1711. [Google Scholar] [CrossRef]
- Lin, S.M.; Zhi, Y.; Ahn, K.B.; Lim, S.; Seo, H.S. Status of group B streptococcal vaccine development. Clin. Exp. Vaccine Res. 2018, 7, 76–81. [Google Scholar] [CrossRef] [PubMed]
- Jacups, S.P. The continuing role of Haemophilus influenzae type b carriage surveillance as a mechanism for early detection of invasive disease activity. Human Vaccines 2011, 7, 1254–1260. [Google Scholar] [CrossRef] [PubMed]
- Bogaert, D.; de Groot, R.; Hermans, P.W.M. Streptococcus pneumoniae colonisation: The key to pneumococcal disease. Lancet Infect. Dis 2004, 4, 144–154. [Google Scholar] [CrossRef]
- Caugant, D.A.; Maiden, M.C.J. Meningococcal carriage and disease—Population biology and evolution. Vaccine 2009, 27, B64–B70. [Google Scholar] [CrossRef]
- Carlin, A.F.; Chang, A.C.; Areschoug, T.; Lindahl, G.; Hurtado-Ziola, N.; King, C.C.; Varki, A.; Nizet, V. Group B Streptococcus suppression of phagocyte functions by protein-mediated engagement of human Siglec-5. J. Exp. Med. 2009, 206, 1691–1699. [Google Scholar] [CrossRef]
- Hallstrom, T.; Riesbeck, K. Haemophilus influenzae and the complement system. Trends Microbiol. 2010, 18, 258–265. [Google Scholar] [CrossRef]
- Hogg, J.S.; Hu, F.Z.; Janto, B.; Boissy, R.; Hayes, J.; Keefe, R.; Post, J.C.; Ehrlich, G.D. Characterization and modeling of the Haemophilus influenzae core and supragenomes based on the complete genomic sequences of Rd and 12 clinical nontypeable strains. Genome Biol. 2007, 8, R103. [Google Scholar] [CrossRef]
- Schoen, C.; Tettelin, H.; Parkhill, J.; Frosch, M. Genome flexibility in Neisseria meningitidis. Vaccine 2009, 27S, B103–B111. [Google Scholar] [CrossRef]
- Chaguza, C.; Andam, C.P.; Harris, S.R.; Cornick, J.E.; Yang, M.; Bricio-Moreno, L.; Kamng’ona, A.W.; Parkhill, J.; French, N.; Heyderman, R.S.; et al. Recombination in Streptococcus pneumoniae lineages increase with carriage duration and size of the polysaccharide capsule. mBio 2016, 7, e01053. [Google Scholar] [CrossRef]
- Keefe, G.P. Streptococcus agalactiae mastitis: A review. Can. Vet. J. 1997, 38, 429–437. [Google Scholar] [PubMed]
- Carvalho-Castro, G.A.; Silvaa, J.R.; Paivab, L.V.; Custódio, D.A.C.; Moreirab, R.O.; Miana, G.F.; Ingrid, A.; Prado, I.A.; Chalfun-Junior, A.C.; Costa, G.M. Molecular epidemiology of Streptococcus agalactiae isolated from mastitis in Brazilian dairy herds. Braz. J. Microbiol. 2017, 48, 551–559. [Google Scholar] [CrossRef] [PubMed]
- Manning, S.D.; Springman, C.; Million, A.D.; Milton, N.R.; McNamara, S.E.; Somsel, P.A.; Bartlett, P.; Davies, H.D. Association of group B Streptococcus colonization and bovine exposure: A prospective cross-sectional cohort study. PLoS ONE 2010, 5, e8795. [Google Scholar] [CrossRef] [PubMed]
- Neemuchwala, A.; Teatero, S.; Athey, T.B.T.; McGeer, A.; Fittipaldi, N. Capsular switching and other large-scale recombination events in invasive sequence type 1 group B Streptococcus. Emerg. Infect. Dis. 2016, 22, 1941–1944. [Google Scholar] [CrossRef] [PubMed]
- Pittman, M. Variation and type specificity in the bacterial species Haemophilus influenzae. J. Exp. Med. 1931, 53, 471–492. [Google Scholar] [CrossRef] [PubMed]
- Harrison, O.B.; Claus, H.; Jiang, Y.; Bennett, J.S.; Bratcher, H.B.; Jolley, K.A.; Corton, C.; Care, R.; Poolman, J.T.; Zollinger, W.D. Description and nomenclature of Neisseria meningitidis capsule locus. Emerg. Infect. Dis. 2013, 19, 566–573. [Google Scholar] [CrossRef] [PubMed]
- Geno, K.A.; Gilbert, G.L.; Song, J.Y.; Skovsted, I.C.; Klugman, K.P.; Jones, C.; Konradsen, H.B.; Nahm, M.H. Pneumococcal capsules and their types: Past, present, and future. Clin. Microbiol. Rev. 2015, 28, 871–899. [Google Scholar] [CrossRef]
- Ganaie, F.; Saad, J.S.; McGee, L.; van Tonder, A.J.; Bentley, S.D.; Lo, S.W.; Gladstone, R.A.; Turner, P.; Keenan, J.D.; Breiman, R.F.; et al. A new pneumococcal capsule type, 10D, is the 100th serotype and has a large cps fragment from an oral Streptococcus. mBio 2020, 11, e00937. [Google Scholar] [CrossRef] [PubMed]
- Madhi, S.A.; Dangor, Z.; Heath, P.T.; Schrag, S.; Izu, A.; Meulen, A.S.T.; Dull, P.M. Considerations for a phase-III trial to evaluate a group B Streptococcus polysaccharide-protein conjugate vaccine in pregnant women for the prevention of early- and late-onset invasive disease in young-infants. Vaccine 2013, 31 (Suppl. 40), D52–D57. [Google Scholar] [CrossRef]
- Buurman, E.T.; Timofeyeva, Y.; Gu, J.; Kim, J.H.; Kodali, S.; Liu, Y.; Mininni, T.; Moghazeh, S.; Pavliakova, D.; Singer, C.; et al. A novel hexavalent capsular polysaccharide conjugate vaccine (GBS6) for the prevention of neonatal group B streptococcal infections by maternal immunization. J. Infect. Dis 2019, 220, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Vos, Q.; Lees, A.; Wu, Z.Q.; Snapper, C.M.; Mond, J.J. B-cell activation by T-cell-independent type 2 antigens as an integral part of the humoral immune response to pathogenic microorganisms. Immunol. Rev. 2000, 176, 154–170. [Google Scholar]
- Mason, E.O.; Kaplan, S.L.; Lamberthm, L.B.; Hinds, D.B.; Kvernland, S.J.; Loiselle, E.M.; Feigin, R.D. Serotype and ampicillin susceptibility of Haemophilus influenzae causing systemic infections in children: 3 years of experience. J. Clin. Microbiol. 1982, 15, 543–546. [Google Scholar] [CrossRef]
- Falla, T.J.; Dobson, S.R.; Crook, D.W.; Kraak, W.A.; Nichols, W.W.; Anderson, E.C.; Jordens, L.J.; Slack, M.P.; Mayon-White, D.; Moxon, E.R. Population-based study of non-typable Haemophilus influenzae invasive disease in children and neonates. Lancet 1993, 341, 851–864. [Google Scholar] [CrossRef]
- Barbour, M.L.; Mayon-White, R.T.; Coles, C.; Crook, D.W.; Moxon, E.R. The impact of conjugate vaccine on carriage of Haemophilus influenzae type b. J. Infect. Dis 1995, 171, 93–98. [Google Scholar] [CrossRef]
- Huang, S.S.; Platt, R.; Rifas-Shiman, S.L.; Pelton, S.I.; Goldmann, D.; Finkelstein, J.A. Post-PCV7 changes in colonizing pneumococcal serotypes in 16 Massachusetts communities, 2001 and 2004. Pediatrics 2005, 116, e408–e413. [Google Scholar] [CrossRef] [PubMed]
- Pizza, M.; Bekkat-Berkani, R.; Rappuoli, R. Vaccines against meningococcal diseases. Microorganism 2020, 8, 1521. [Google Scholar] [CrossRef]
- Zwahlen, A.; Kroll, J.S.; Rubin, L.G.; Moxon, E.R. The molecular basis of pathogenicity in Haemophilus influenzae: Comparative virulence of genetically-related capsular transformants and correlation with changes at the capsulation locus cap. Microb. Pathog. 1989, 7, 225–235. [Google Scholar] [CrossRef]
- Jafri, R.Z.; Ali, A.; Messonnier, N.E.; Tevi-Benissan, C.; Durrheim, D.; Eskola, J.; Fermon, F.; Klugman, K.P.; Ramsay, M.; Sow, S.; et al. Global epidemiology of invasive meningococcal disease. Popul. Health Metr. 2013, 11, 17. [Google Scholar] [CrossRef]
- Pelton, S.I. The global evolution of meningococcal epidemiology following the introduction of meningococcal vaccines. J. Adolesc. Health 2016, 59, S3–S11. [Google Scholar] [CrossRef]
- Lo, S.W.; Gladstone, R.A.; van Tonder, A.J.; Lees, J.A.; du Plessis, M.; Benisty, R.; Givon-Lavi, N.; Hawkins, P.A.; Cornick, J.E.; Kwambana-Adams, B.; et al. The Global Pneumococcal Sequencing Consortium. Pneumococcal lineages associated with serotype replacement and antibiotic resistance in childhood invasive pneumococcal disease in the post-PCV13 era: An international whole-genome sequencing study. Lancet Infect. Dis. 2019, 19, 759–769. [Google Scholar] [CrossRef]
- Farrell, D.J.; Jenkins, S.G.; Reinert, R.R. Global distribution of Streptococcus pneumoniae serotypes isolated from paediatric patients during 1999–2000 and the in vitro efficacy of telithromycin and comparators. J. Med. Microbiol. 2004, 53, 1109–1117. [Google Scholar] [CrossRef] [PubMed]
- Yahiaoui, R.Y.; Bootsma, H.J.; den Heijer, C.D.J.; Pluister, G.N.; Paget, W.J.; Spreeuwenberg, P.; Trzcinski, K.; Stobberingh, E.E. Distribution of serotypes and patterns of antimicrobial resistance among commensal Streptococcus pneumoniae in nine European countries. BMC Infect. Dis. 2018, 18, 440. [Google Scholar] [CrossRef] [PubMed]
- Harrison, L.H.; Shutt, K.A.; Schmink, S.E.; Marsh, J.W.; Harcourt, B.H.; Wang, X.; Whitney, A.M.; Stephens, D.S.; Cohn, A.A.; Messonnier, N.E.; et al. Population structure and capsular switching of invasive Neisseria meningitidis isolates in the pre-meningococcal conjugate vaccine era, United States, 2000–2005. J. Infect. Dis 2010, 201, 1208–1224. [Google Scholar] [CrossRef]
- Hu, F.Z.; Eutsey, R.; Ahmed, A.; Frazao, N.; Powell, E.; Hiller, N.L.; Hillman, T.; Buchinsky, F.J.; Boissy, R.; Benjamin Janto, B.; et al. In vivo capsular switch in Streptococcus pneumoniae – analysis by whole genome sequencing. PLoS ONE 2012, 7, e47983. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Maiden, M.C.; Bygraves, J.A.; Feil, E.; Morelli, G.; Russell, J.E.; Urwin, R.; Zhang, Q.; Zurth, K.; Caugant, D.A.; Feavers, I.M.; et al. Multilocus sequence typing: A portable approach to the identification of clones within population of pathogenic microorganisms. Proc. Natl. Acad. Sci. USA 1998, 95, 3140–3145. [Google Scholar] [CrossRef]
- Lucidarme, J.; Lekshmi, A.; Parikh, S.R.; Bray, J.E.; Hill, D.M.; Bratcher, H.B.; Gray, S.J.; Carr, A.D.; Jolley, K.A.; Findlow, J.; et al. Frequent capsule switching in “ultra-virulent” meningococci—Are we ready for a serogroup B ST-11 complex outbreak? J. Infect. 2017, 75, 95–103. [Google Scholar] [CrossRef]
- Swartley, J.S.; Marfin, A.A.; Edupuganti, S.; Liu, L.J.; Cieslar, P.; Perkins, B.; Wenger, J.D.; Stephens, D.S. Capsule switching of Neisseria meningitidis. Proc. Natl. Acad. Sci. USA 1997, 94, 271–276. [Google Scholar] [CrossRef] [PubMed]
- Lipstich, M. Bacterial vaccines and serotype replacement: Lessons from Haemophilus influenzae and prospects for Streptococcus pneumoniae. Emerg. Infect. Dis. 1999, 5, 336–345. [Google Scholar] [CrossRef]
- Weinbergera, D.M.; Malley, R.; Lipsitcha, M. Serotype replacement in disease following pneumococcal vaccination: A discussion of the evidence. Lancet 2011, 378, 1962–1973. [Google Scholar] [CrossRef]
- Mitchell, P.R.; Azarian, T.; Croucher, N.J.; Callendrello, A.; Thompson, C.M.; Pelton, S.I.; Lipsitch, M.; Hanage1, W.P. Population genomics of pneumococcal carriage in Massachusetts children following introduction of PCV-13. Microb. Genom. 2019, 5, e000252. [Google Scholar] [CrossRef]
- Brueggemann, A.B.; Pai, R.; Crook, D.W.; Beall, B. Vaccine escape recombinants emerge after pneumococcal vaccination in the United States. PLoS Pathog. 2007, 3, e168. [Google Scholar] [CrossRef] [PubMed]
- Makarewicz, O.; Lucas, M.; Brandt, C.; Herrmann, L.; Albersmeier, A.; Ruckert, C.; Blom, J.; Goesmann, A.; van der Linden, M.; Kalinowski, J.; et al. Whole genome sequencing of 39 invasive Streptococcus pneumoniae sequence type 199 isolates revealed switches from serotype 19A to 15B. PLoS ONE 2017, 12, e0169370. [Google Scholar] [CrossRef] [PubMed]
- Mostowy, R.; Croucher, N.J.; De Maio, N.; Chewapreecha, C.; Salter, S.J.; Turner, P.; Aanensen, D.M.; Bentley, S.D.; Didelot, X.; Fraser, C. Pneumococcal capsule synthesis locus cps as evolutionary hotspot with potential to generate novel serotypes by recombination. Mol. Biol. Evol. 2017, 34, 2537–2554. [Google Scholar] [CrossRef]
- Croucher, N.J.; Kagedan, L.; Thompson, C.M.; Parkhill, J.; Bentley, S.D.; Finkelstein, J.A.; Lipsitch, M.; Hanage, W.P. Selective and genetic constraints on pneumococcal serotype switching. PLoS Genet. 2015, 11, e1005095. [Google Scholar] [CrossRef] [PubMed]
- Wyres, K.L.; Lambertsen, L.M.; Croucher, M.J.; McGee, L.; von Gottberg, A.; Liñares, J.; Jacobs, M.R.; Kristinsson, K.G.; Beall, B.W.; Klugman, K.P.; et al. Pneumococcal capsular switching: A historical perspective. J. Infect. Dis. 2013, 207, 439–449. [Google Scholar] [CrossRef]
- Groves, N.; Sheppard, C.L.; Litt, D.; Rose, S.; Silva, A.; Njoku, N.; Rodrigues, S.; Amin-Chowdhury, Z.; Andrews, N.; Ladhani, S.; et al. Evolution of Streptococcus pneumoniae serotype 3 in England and Wales: A major vaccine evader. Genes 2019, 10, 845. [Google Scholar] [CrossRef] [PubMed]
- Kandasamy, R.; Voysey, M.; Collins, S.; Berbers, G.; Robinson, H.; Noel, I.; Hughes, H.; Ndimah, S.; Gould, K.; Fry, N.; et al. Persistent circulation of vaccine serotypes and serotype replacement after 5 years of infant immunization with 13-valent pneumococcal conjugate vaccine in the United Kingdom. J. Infect. Dis. 2020, 221, 1361–1370. [Google Scholar] [CrossRef] [PubMed]
- Balsells, E.; Guillot, L.; Nair, H.; Kyaw, M.H. Serotype distribution of Streptococcus pneumoniae causing invasive disease in children in the post-PCV era: A systematic review and meta-analysis. PLoS ONE 2017, 12, e0177113. [Google Scholar] [CrossRef]
- Dagan, R.; Ben-Shimol, S.; Benisty, R.; Regev-Yochay, G.; Ron, M.; Givon-Lavi, N.; Rokney, A. 1888. A nationwide outbreak of invasive pneumococcal disease (IPD) caused by a novel Streptococcus pneumoniae serotype 2 (SP2) clone in the PCV13 era, in Israel. Open Forum Infect. Dis. 2019, 6, S54. [Google Scholar] [CrossRef]
- Levy, C.; Ouldali, N.; Caeymaex, L.; Angoulvant, F.; Varon, E.; Cohen, R. Diversity of serotype replacement after pneumococcal conjugate vaccine implementation in Europe. J. Pediatr. 2019, 213, 252–256. [Google Scholar] [CrossRef] [PubMed]
- Neves, F.P.G.; Cardoso, N.T.; Cardoso, C.A.C.; Teixeira, L.M.; Riley, L.W. Direct effect of the 13-valent pneumococcal conjugate vaccine use on pneumococcal colonization among children in Brazil. Vaccine 2019, 37, 5265–5269. [Google Scholar] [CrossRef] [PubMed]
- Pick, H.; Daniel, P.l.; Rodrigo, C.; Bewick, T.; Ashton, D.; Lawrence, H.; Baskaran, V.; Edwards-Pritchard, R.C.; Sheppard, C.; Seyi, D.; et al. Pneumococcal serotype trends, surveillance and risk factors in UK adult pneumonia, 2013–2018. Thorax 2020, 75, 38–49. [Google Scholar] [CrossRef] [PubMed]
- Rupp, R.; Hurley, D.; Grayson, S.; Li, J.; Nolan, K.; McFetridge, R.D. A dose ranging study of 2 different formulations of 15-valent pneumococcal conjugate vaccine (PCV15) in healthy infants. Hum. Vaccines Immunother. 2019, 15, 549–559. [Google Scholar] [CrossRef] [PubMed]
- Thompson, A.; Lamberth, E.; Severs, J.; Scully, I.; Tarabar, S.; Ginis, J. Phase 1 trial of a 20 valent pneumococcal conjugate vaccine in healthy adults. Vaccine 2019, 37, 6201–6207. [Google Scholar] [CrossRef]
- Løchen, A.; Croucher, N.J.; Anderson, R.M. Divergent serotype replacement trends and increasing diversity in pneumococcal disease in high income settings reduce the benefit of expanding vaccine valency. Sci. Rep. 2020, 10, 18977. [Google Scholar] [CrossRef] [PubMed]
- Peters, T.R.; Poehling, K.A. Invasive pneumococcal disease, the target is moving. JAMA 2017, 297, 1825–1926. [Google Scholar] [CrossRef] [PubMed]
- Hanage, W.P. Serotype replacement in invasive pneumococcal disease: Where do we go from here? (editorial commentary). J. Infect. Dis. 2007, 196, 1282–1284. [Google Scholar] [CrossRef]
- Knol, M.J.; van der Ende, A. Continuous surveillance of invasive pneumococcal disease is key. Lancet Infect. Dis. 2021, 21, 13–14. [Google Scholar] [CrossRef]
- Klugman, K.P.; Rodgers, G.L. Time for a third-generation pneumococcal vaccine. Lancet Infect. Dis. 2021, 21, 14–16. [Google Scholar] [CrossRef]
- Advisory Committee on Immunization Practices, Centers for Disease Control and Prevention (CDC). Revised recommendations of the Advisory Committee on Immunization Practices to vaccinate all persons aged 11–18 years with meningococcal conjugate vaccine. Morb. Mortal. Wkly. Rep. 2007, 56, 794–795. [Google Scholar]
- Wang, X.; Shutt, K.A.; Vuong, J.T.; Cohn, A.; MacNeil, J.; Schmink, S.; Plikaytis, B.; Messonnoer, N.E.; Harrison, L.H.; Clark, T.A.; et al. Changes in the population structure of invasive Neisseria meningitidis in the United States after quadrivalent meningococcal conjugate vaccine licensure. J. Infect. Dis. 2015, 211, 1887–1894. [Google Scholar] [CrossRef]
- Law, D.K.S.; Lorange, M.; Ringuette, L.; Dion, R.; Giguère, M.; Henderson, A.M.; Stoltz, J.; Zollinger, W.D.; De Wals, P.; Tsang, R.S.W. Invasive meningococcal disease in Quebec, Canada, due to an emerging clone of ST-269 serogroup B meningococci with serotype antigen 17 and serosubtype antigen P1.19 (B:17:P1.19). J. Clin. Microbiol. 2006, 44, 2743–2749. [Google Scholar] [CrossRef]
- Wals, P.; Dionne, M.; Douville-Fradet, M.; Boulianne, N.; Drapeau, J.; De Serres, D. Impact of a mass immunization campaign against serogroup C meningococcus in the Province of Quebec, Canada. Bull. WHO 1996, 74, 407–411. [Google Scholar] [PubMed]
- Wals, P.; Deceuninck, G.; Boulianne, N.; De Serres, G. Effectiveness of a mass immunization campaign using serogroup C meningococcal conjugate vaccine. JAMA 2004, 292, 2491–2494. [Google Scholar] [CrossRef]
- Krone, M.; Gray, S.; Abad, R.; Skoczyńska, A.; Stefanelli, P.; van der Ende, A.; Tzanakaki, G.; Mölling, P.; Simões, M.J.; Křížová, P.; et al. Increase of invasive meningococcal serogroup W disease in Europe, 2013 to 2017. Eurosurveill 2019, 24, 1800245. [Google Scholar] [CrossRef]
- Taha, M.K.; Deghmane, A.E.; Knol, M.; van der Ende, A. Whole genome sequencing reveals Trans-European spread of an epidemic Neisseria meningitidis serogroup W clone. Clin. Microbiol. Infect. 2019, 25, 765–767. [Google Scholar] [CrossRef] [PubMed]
- Martin, N.V.; Ong, K.S.; Howden, B.P.; Lahra, M.M.; Lambert, S.B.; Beard, F.H.; Dowse, G.K.; Saul, N. Communicable Diseases Network Australia MenW Working Group. Rise in invasive serogroup W meningococcal disease in Australia 2013–2015. Commun. Dis. Intell. Q. Rep. 2016, 40, E454–E459. [Google Scholar] [PubMed]
- Tsang, R.S.W.; Hoang, L.; Tyrrell, G.J.; Horsman, G.; Caeseele, P.; Jamieson, F.B.; Lefebvre, B.; Haldane, D.; Gad, R.R.; German, G.J.; et al. Increase in Neisseria meningitidis serogroup W invasive disease in Canada: 2009-2016. Can. Commun. Dis. Rep. 2017, 43, 144–149. [Google Scholar] [CrossRef] [PubMed]
- Mowlaboccus, S.; Jolley, K.A.; Bray, J.E.; Pang, S.; Lee, Y.T.; Bew, J.D.; Speers, D.S.; Keil, A.D.; Coombs, G.W.; Kahler, C.M. Clonal expansion of new penicillin-resistant clade of Neisseria meningitidis serogroup W clonal complex 11, Australia. Emerg. Infect. Dis. 2017, 23, 1364–1367. [Google Scholar] [CrossRef]
- Trotter, C.L.; Lingani, C.; Fernandez, K.; Cooper, L.V.; Bita, A.; Tevi-Benissan, C.; Ronveaux, O.; Préziosi, M.P.; Stuart, J.M. Impact of MenAfriVac in nine countries of the African meningitis belt, 2010–2015: An analysis of surveillance data. Lancet Infect. Dis. 2017, 17, 867–872. [Google Scholar] [CrossRef]
- Boisier, P.; Nicolas, P.; Djibo, S.; Taha, M.K.; Jeanne, J.; Maïnassara, H.B.; Tenebray, B.; Kairo, K.K.; Giorgini, D.; Chanteau, S. Meningococcal meningitis: Unprecedented incidence of serogroup X-related cases in 2006 in Niger. Clin. Infect. Dis. 2007, 44, 657–663. [Google Scholar] [CrossRef]
- Xie, O.; Pollard, A.J.; Mueller, J.E.; Norheim, G. Emergence of serogroup X meningococcal disease in Africa: Need for a vaccine. Vaccine 2013, 31, 2852–2861. [Google Scholar] [CrossRef]
- Agnememel, A.; Hong, E.; Giorgini, D.; Nuñez-Samudio, V.; Deghmane, A.E.; Taha, M.K. Neisseria meningitidis serogroup X in Sub-Saharan Africa. Emerg. Infect. Dis. 2016, 22, 698–702. [Google Scholar] [CrossRef]
- Soeters, H.M.; Diallo, A.O.; Bicaba, B.W.; Kadadé, G.; Dembélé, A.Y.; Acyl, M.A.; Nikiema, C.; Sadji, A.Y.; Poy, A.N.; Lingani, C.; et al. Bacterial meningitis epidemiology in 5 countries in the meningitis belt of sub-Saharan Africa, 2015–2017. J. Infect. Dis. 2019, 220 (Suppl. 4), S165–S174. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, K.; Lingani, C.; Aderinola, O.M.; Goumbi, K.; Bicaba, B.; Edea, Z.A.; Glèlè, C.; Sarkodie, B.; Tamekloe, A.; Ngomba, A.; et al. Meningococcal meningitis outbreaks in the African meningitis belt after meningococcal serogroup A conjugate vaccine introduction, 2011–2017. J. Infect. Dis. 2019, 220 (Suppl. 4), S225–S232. [Google Scholar] [CrossRef]
- Brynildsruda, O.B.; Eldholma, V.; Bohlina, J.; Uadialeb, K.; Obaro, S.; Cauganta, D.A. Acquisition of virulence genes by a carrier strain gave rise to the ongoing epidemics of meningococcal disease in West Africa. Proc. Natl. Acad. Sci. USA 2018, 115, 5510–5515. [Google Scholar] [CrossRef]
- Retchless, A.C.; Hu, F.; Ouédraogo, A.; Diarra, S.; Knipe, K.; Sheth, M.; Rowe, L.A.; Sangaré, L.; Ba, A.K.; Ouangraoua, S.; et al. The establishment and diversification of epidemic-associated serogroup W meningococcus in the African meningitis belt, 1994 to 2012. mSphere 2016, 1, e00201–e00216. [Google Scholar] [CrossRef] [PubMed]
- Leimkugel, J.; Hodgson, A.; Forgor, A.A.; Pfluger, V.; Dangy, J.P.; Smith, T.; Achtman, M.; Sebastien Gagneux, S.; Gerd Pluschke, G. Clonal waves of Neisseria colonisation and disease in the African meningitis belt: Eight year longitudinal study in Northern Ghana. PLoS Med. 2007, 4, e101. [Google Scholar] [CrossRef]
- Mbaeyi, S.; Sampo, E.; Dinanibè, K.; Yaméogo, I.; Congo-Ouédraogo, M.; Tamboura, M.; Sawadogo, G.; Ouattara, K.; Sanou, M.; Kiemtoré, K.; et al. Meningococcal carriage 7 years after introduction of a serogroup A meningococcal conjugate vaccine in Burkina Faso: Results from four cross-sectional carriage surveys. Lancet Infect. Dis. 2020, 20, 1418–1425. [Google Scholar] [CrossRef]
- Langereis, J.D.; de Jonge, M.I. Invasive disease caused by non-typeable Haemophilus influenzae. Emerg. Infect. Dis. 2015, 21, 1711–1718. [Google Scholar] [CrossRef] [PubMed]
- Whittaker, R.; Economopoulou, A.; Dias, J.G.; Bancroft, E.; Ramliden, M.; Celentano, L.P. European Centre for Disease Prevention and Control Country Experts for Invasive Haemophilus influenzae Disease. Epidemiology of invasive Haemophilus influenzae disease, Europe, 2007–2014. Emerg. Infect. Dis. 2017, 23, 396–404. [Google Scholar] [CrossRef] [PubMed]
- Soeters, H.M.; Blain, A.; Pondo, T.; Doman, B.; Farley, M.M.; Harrison, L.H.; Lynfield, R.; Miller, L.; Petit, S.; Reinglod, A.; et al. Current epidemiology and trends in invasive Haemophilus influenzae disease—United States, 2009-2015. Clin. Infect. Dis. 2018, 67, 881–889. [Google Scholar] [CrossRef] [PubMed]
- Ulanova, M. Global epidemiology of invasive Haemophilus influenzae type a disease: Do we need a new vaccine? J. Vaccine 2013, 14, 941461. [Google Scholar] [CrossRef]
- Tsang, R.S.W.; Ulanova, M. The changing epidemiology of invasive Haemophilus influenzae disease: Emergence and global presence of serotype a strains that may require a new vaccine for control. Vaccine 2017, 35, 4270–4275. [Google Scholar] [CrossRef]
- Boisvert, A.A.; Moore, D. Invasive disease due to Haemophilus influenzae type a in Canada’s north: A priority for prevention. Can. J. Infect. Dis. Med. Microbiol. 2015, 26, 291–292. [Google Scholar] [CrossRef] [PubMed]
- Soeters, H.M.; Oliver, S.E.; Plumb, I.D.; Blain, A.E.; Zulz, T.; Simons, B.C.; Barnes, M.; Farley, M.M.; Harrison, L.H.; Lynfield, R.; et al. Epidemiology of invasive Haemophilus influenzae serotype a disease—United States, 2008–2017. Clin. Infect. Dis. 2020, ciaa875. [Google Scholar] [CrossRef] [PubMed]
- Bozio, C.H.; Blain, A.; Edge, K.; Farley, M.M.; Harrison, L.H.; Poissant, T.; Schaffner, W.; Scheuer, T.; Torres, S.; Triden, L.; et al. Clinical characteristics and adverse clinical outcomes of invasive Haemophilus influenzae serotype a cases—United States, 2011–2015. Clin. Infect. Dis. 2020, ciaa990. [Google Scholar] [CrossRef]
- Cox, A.D.; Williams, D.; Cairns, C.; St. Michael, F.; Fleming, P.; Vinogradov, E.; Arbour, M.; Masson, L.; Zou, W. Investigating the candidacy of a capsular polysaccharide-based glycoconjugate as a vaccine to combat Haemophilus influenzae type a disease: A solution for an unmet public health need. Vaccine 2017, 35, 6129–6136. [Google Scholar] [CrossRef] [PubMed]
- Bruce, M.G.; Zulz, T.; DeByle, C.; Singleton, R.; Hurlburt, D.; Bruden, D.; Rudolph, K.; Hennessy, T.; Klejka, J.; Wenger, J.D. Haemophilus influenzae serotype a invasive disease, Alaska, USA, 1983–2011. Emerg. Infect. Dis. 2013, 19, 932–937. [Google Scholar] [CrossRef] [PubMed]
- Tsang, R.S.W.; Shuel, M.; Ahmad, T.; Hayden, K.; Knox, N.; Van Domselaar, G.; Hoang, L.; Tyrrell, G.; Minion, J.; Van Caeseele, P.; et al. Whole genome sequencing to study the phylogenetic structure of serotype a Haemophilus influenzae recovered from patients in Canada. Can. J. Microbiol. 2019, 66, 99–110. [Google Scholar] [CrossRef]
- Lima, J.B.; Ribeiro, G.S.; Cordeiro, S.M.; Gouveia, E.L.; Salgado, K.; Spratt, B.G.; Godoy, D.; Reis, M.G.; Ko, A.I.; Reis, J.N. Poor clinical outcome for meningitis caused by Haemophilus influenzae serotype a strains containing the IS1016-bexA deletion. J. Infect. Dis. 2010, 202, 1577–1584. [Google Scholar] [CrossRef] [PubMed]
- Crandall, H.; Christiansen, J.; Varghese, A.A.; Russon, A.; Korgenski, E.K.; Bengtson, E.K.; Dickey, M.; Killpack, J.; Knackstedt, E.D.; Daly, J.A.; et al. Clinical and molecular epidemiology of invasive Haemophilus influenzae serotype a infections in Utah children. J. Pediatr. Infect. Dis. Soc. 2019, piz088. [Google Scholar] [CrossRef] [PubMed]
- LaCross, N.C.; Marrs, C.F.; Patel, M.; Sandstedt, S.A.; Gilsdorf, J.R. High genetic diversity of nontypeable Haemophilus influenzae isolates from two children attending a day care center. J. Clin. Microbiol. 2008, 46, 3817–3821. [Google Scholar] [CrossRef]
- Pinto, M.; González-Díaz, A.; Machado, M.P.; Duarte, S.; Vieira, L.; Carriço, J.A.; Marti, S.; Bajanca-Lavado, M.P.; Gomes, J.P. Insights into the population structure and pan-genome of Haemophilus influenzae. Infect. Gene Evol. 2019, 67, 126–135. [Google Scholar] [CrossRef]
- Chewapreecha, C.; Harris, S.R.; Croucher, N.J.; Turner, C.; Marttinen, P.; Cheng, L.; Pessia, A.; Aanensen, D.M.; Mather, A.E.; Page, A.J.; et al. Dense genomic sampling identifies highways of pneumococcal recombination. Nat. Genet. 2014, 46, 305–309. [Google Scholar] [CrossRef]
- Harrison, O.B.; Brueggemann, A.B.; Caugant, D.A.; van der Ende, A.; Frosch, M.; Gray, S.; Heuberger, S.; Krizova, P.; Olcen, P.; Slack, M.; et al. Molecular typing methods for outbreak detection and surveillance of invasive disease caused by Neisseria meningitidis, Haemophilus influenzae and Streptococcus pneumoniae, a review. Microbiology 2011, 157, 2181–2195. [Google Scholar] [CrossRef] [PubMed]
- LaClaire, L.L.; Tondella, M.L.C.; Beall, D.S.; Noble, C.A.; Raghunathan, P.I.; Rosenstein, N.E.; Popovic, T. The Active Bacterial Core Surveillance Team Members. Identification of Haemophilus influenzae serotypes by standard slide agglutination serotyping and PCR-based capsule typing. J. Clin. Microbiol. 2003, 41, 393–396. [Google Scholar] [CrossRef]
- Falla, T.J.; Crook, D.W.M.; Brophy, L.N.; Maskell, D.; Kroll, J.S.; Moxon, E.R. PCR for capsular typing of Haemophilus influenzae. J. Clin. Microbiol. 1994, 32, 2382–2386. [Google Scholar] [CrossRef] [PubMed]
- Kong, F.; Gowan, S.; Martin, D.; James, G.; Gilbert, G.L. Serotype identification of group B streptococci by PCR and sequencing. J. Clin. Microbiol. 2002, 40, 216–226. [Google Scholar] [CrossRef]
- Brito, D.A.; Ramirez, M.; de Lencastre, H. Serotyping Streptococcus pneumoniae by multiplex PCR. J. Clin. Microbiol. 2003, 41, 2378–2384. [Google Scholar] [CrossRef]
- Zhu, H.; Wang, Q.; Wen, L.; Xu, J.; Shao, Z.; Chen, M.; Chen, M.; Reeves, P.R.; Cao, B.; Wanga, L. Development of a multiplex PCR assay for detection and genogrouping of Neisseria meningitidis. J. Clin. Microbiol. 2012, 50, 46–51. [Google Scholar] [CrossRef]
- Enright, M.C.; Spratt, B.G. A multilocus sequence typing scheme for Streptococcus pneumoniae: Identification of clones associated with serious invasive disease. Microbiology 1998, 144, 3049–3060. [Google Scholar] [CrossRef]
- Meats, E.; Feil, E.J.; Stringer, S.; Cody, A.J.; Goldstein, R.; Kroll, J.S.; Popovic, T.; Spratt, B.G. Characterization of encapsulated and non-encapsulated Haemophilus influenzae and determination of phylogenetic relationships by multilocus sequence typing. J. Clin. Microbiol. 2003, 41, 1623–1626. [Google Scholar] [CrossRef]
- Serruto, D.; Bottomley, M.J.; Ram, S.; Giuliliani, M.M.; Rappuoli, R. The new multicomponent vaccine against meningococcal serogroup B, 4CMenB: Immunological, functional and structural characterization of the antigens. Vaccine 2012, 30 (Suppl. 2), B87–B97. [Google Scholar] [CrossRef] [PubMed]
- Gandhi, A.; Balmer, P.; York, L.J. Characteristics of a new meningococcal serogroup B vaccine, bivalent rLP2086 (MenB-FHbp; Trumenba®). Postgrad. Med. 2016, 128, 548–556. [Google Scholar] [CrossRef] [PubMed]
- Browall, S.; Norman, M.; Tångrot, J.; Galanis, I.; Sjöström, K.; Dagerhamn, J.; Hellberg, C.; Pathak, A.; Spadafina, T.; Sandgren, A.; et al. Intraclonal variations among Streptococcus pneumoniae isolates influence the likelihood of invasive disease in children. J. Infect. Dis 2014, 209, 377–388. [Google Scholar] [CrossRef] [PubMed]
- Cremers, A.J.H.; Mobegi, F.M.; van der Gaast–de Jongh, C.; van Weert, M.; van Opzeeland, F.J.; Vehkala, M.; Knol, M.J.; Bootsma, H.J.; Välimäki, N.; Croucher, N.J.; et al. The contribution of genetic variation of Streptococcus pneumoniae to the clinical manifestation of invasive pneumococcal disease. Clin. Infect. Dis. 2019, 68, 61–69. [Google Scholar] [CrossRef]
- Pettigrewa, M.M.; Ahearnb, C.P.; Gentd, J.F.; Konge, Y.; Gallob, M.C.; Munroh, J.B.; D’Melloh, A.; Sethic, S.; Tettelinh, H.; Murphy, T.F. Haemophilus influenzae genome evolution during persistence in the human airways in chronic obstructive pulmonary disease. Proc. Natl. Acad. Sci. USA 2018, 115, E3256–E3265. [Google Scholar] [CrossRef] [PubMed]
- The European Committee on Antimicrobial Susceptibility Testing—EUCAST. Breakpoint Tables for Interpretation of MICs and Zone Diameters, Version 10.0. 2020. Available online: http://www.eucast.org/clinical_breakpoints/ (accessed on 15 December 2020).
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 30th ed.; Supplement M100-S30; CLSI: Wayne, PA, USA, 2020. [Google Scholar]
- Yazdankhah, S.P.; Kriz, P.; Tzanakaki, G.; Kremastinou, J.; Kalmusova, J.; Musilek, M.; Alvestad, T.; Jolley, K.A.; Wilson, D.J.; McCarthy, N.D.; et al. Distribution of serogroups and genotypes among disease-associated and carried isolates of Neisseria meningitidis from the Czech Republic, Greece, and Norway. J. Clin. Microbiol. 2004, 42, 5146–5153. [Google Scholar] [CrossRef]
- Sandgren, A.; Sjostrom, K.; Olsson-Liljequist, O.; Christensson, B.; Samuelsson, A.; Kronvall, G.; Normark, B.H. Effect of clonal and serotype-specific properties on the invasive capacity of Streptococcus pneumoniae. J. Infect. Dis. 2004, 189, 784–796. [Google Scholar] [CrossRef]
- Zhou, J.; Jamieson, F.; Dolman, S.; Hoang, L.M.; Rawte, P.; Tsang, R.S.W. Genetic and antigenic analysis of invasive serogroup C Neisseria meningitidis in Canada: A decrease in the electrophoretic type (ET)-15 clonal type and an increase in the proportion of isolates belonging to the ET-37 (but not ET-15) clonal type during the period from 2002 to 2009. Can. J. Infect. Dis Med. Microbiol. 2012, 23, e55–e59. [Google Scholar] [PubMed]
- Fleischmann, R.D.; Adams, M.D.; White, O.; Clayton, R.A.; Kirkness, E.F.; Kerlavage, A.R.; Bult, C.J.; Tomb, J.F.; Dougherty, B.A.; Merrick, J.M.; et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science 1995, 269, 496–512. [Google Scholar] [CrossRef]
- Gladstone, R.A.; Lo, S.W.; Lees, J.A.; Croucher, N.J.; van Tonder, A.J.; Corander, J.; Page, A.J.; Marttinen, P.; Bentley, L.J.; Ochoa, T.J.; et al. International genomic definition of pneumococcal lineages, to contextualize disease, antibiotic resistance and vaccine impact. EBioMedicine 2019, 43, 338–346. [Google Scholar] [CrossRef]
- Caugant, D.A. Population genetics and molecular epidemiology of Neisseria meningitidis. APMIS 1998, 106, 505–525. [Google Scholar] [CrossRef] [PubMed]
- Musser, J.M.; Kroll, J.S.; Moxon, E.R.; Selander, R.K. Clonal population structure of encapsulated Haemophilus influenzae. Infect. Immun. 1988, 56, 1837–1845. [Google Scholar] [CrossRef] [PubMed]
- De Chiaraa, M.; Hood, D.; Muzzia, A.; Pickardd, D.J.; Perkinse, T.; Pizzaa, M.; Dougand, G.; Rappuolia, R.; Moxxon, E.R.; Soriani, M.; et al. Genome sequencing of disease and carriage isolates of nontypeable Haemophilus influenzae identifies discrete population structure. Proc. Natl. Acad. Sci. USA 2014, 111, 5439–5444. [Google Scholar] [CrossRef]
- Epping, L.; Tonder, A.J.; Gladstone, R.A.; The Global Pneumococcal Sequencing Consortium; Bentley, S.D.; Page, A.J.; Keane, J.A. SeroBA: Rapid high-throughput serotyping of Streptococcus pneumoniae from whole genome sequence data. Microb. Genom. 2018, 4, e000204. [Google Scholar]
- Watts, S.C.; Holt, K.E. Hicap: In Silico serotyping of the Haemophilus influenzae capsule locus. J. Clin. Microbiol. 2019, 57, e00190-19. [Google Scholar] [CrossRef] [PubMed]
- Marjuki, H.; Topaz, N.; Rodriguez-Rivera, L.D.; Ramos, E.; Potts, C.A.; Chen, A.; Retchless, A.C.; Doho, G.H.; Wang, X. Whole-genome sequencing for characterization of capsule locus and prediction of serogroup of invasive meningococcal isolates. J. Clin. Microbiol. 2019, 57, e01609–e01618. [Google Scholar] [CrossRef]
- Jolley, K.A.; Maiden, M.C.J. BIGSdb: Scalable analysis of bacterial genome variation at the population level. BMC Bioinform. 2010, 11, 595. [Google Scholar] [CrossRef] [PubMed]
- Davis, J.J.; Boisvert, S.; Brettin, T.; Kenyon, R.W.; Mao, C.; Olson, R.; Overbeek, R.; Santerre, J.; Shukla, M.; Wattam, A.R.; et al. Antimicrobial resistance prediction in PATRIC and RAST. Sci. Rep. 2015, 6, 27930. [Google Scholar] [CrossRef]
- Metcalf, B.J.; Chochua, S.; Gertz, R.E., Jr.; Li, Z.; Walker, H.; Tran, T.; Hawkins, P.A.; Glennen, A.; Lynfield, R.; Li, Y.; et al. Active Bacterial Core surveillance team. Using whole genome sequencing to identify resistance determinants and predict antimicrobial resistance phenotypes for year 2015 invasive pneumococcal disease isolates recovered in the United States. Clin. Microbiol. Infect. 2016, 22, 1002.e1–1002.e8. [Google Scholar] [CrossRef]
- Conteville, L.C.; de Filippis, A.M.B.; Nogueira, R.M.R.; de Mendonça, M.C.L.; Vicente, A.C.P. Metagenomics in undiagnosed cases of fulminant fever. J. Infect. 2018, 77, 160–161. [Google Scholar] [CrossRef] [PubMed]
- Bozio, C.H.; Vuong, J.; Dokubo, E.K.; Fallah, M.P.; McNamara, L.A.; Potts, C.C.; Doedeh, J.; Gbanya, M.; Retchless, A.C.; Jaymin, C.P.; et al. Outbreak of Neisseria meningitidis serogroup C outside the meningitis belt—Liberia, 2017: An epidemiological and laboratory investigation. Lancet Infect. Dis. 2018, 18, 1360–1367. [Google Scholar] [CrossRef]
- Rodgers, E.; Bentley, S.D.; Borrow, R.; Bratcher, H.B.; Brisse, S.; Brueggemann, A.B.; Caugant, D.A.; Findlow, J.; Fox, L.; Glennie, L.; et al. The global meningitis genome partnership. J. Infect. 2020, 81, 510–520. [Google Scholar] [CrossRef] [PubMed]
- Public Health England. Guidance for Public Health Management of Meningococcal Disease in the UK. August 2019. Available online: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/829326/PHE_meningo_disease_guideline.pdf (accessed on 5 February 2021).
- Public Health England. Revised Recommendations for the Prevention of Secondary Haemophilus influenzae type b (Hib) Disease. July 2013. Available online: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/231009/Revised_recommendations_for_the_preventions_of_secondary_Haemophilus_influenzae_type_b_disease.pdf (accessed on 5 February 2021).
- American Academy of Pediatrics. Red Book: 2018 Report of the Committee on Infectious Diseases, 31st ed.; Kimberlin, D.W., Brady, M.T., Jackson, M.A., Long, S.S., Eds.; American Academy of Pediatrics: Elk Grove Village, IL, USA, 2018. [Google Scholar]
- Public Health England. Guidelines for the Public Health Management of Clusters of Severe Pneumococcal Disease in Closed Settings. Updated January 2020. Available online: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/867175/Pneumococccal_cluster_guidelines.pdf (accessed on 5 February 2021).
- Hughes, R.G.; Brocklehurst, P.; Steer, P.J.; Heath, P.; Stenson, B.M.; On behalf of the Royal College of Obstetricians and Gynaecologists. Prevention of early-onset neonatal group B streptococcal disease. Green-top guideline No. 36. BJOG 2017, 124, e280–e305. [Google Scholar] [CrossRef]
- de Gans, J.; van de Beek, D. For the European Dexamethasone in Adulthood Bacterial Meningitis Study Investigators. Dexamethasone in adults with bacterial meningitis. N. Engl. J. Med. 2002, 347, 1549–1556. [Google Scholar] [CrossRef]
- Brouwer, M.C.; McIntyre, P.; Prasad, K.; van de Beek, D. Corticosteroids for acute bacterial meningitis (review). Cochrane Database Syst. Rev. 2015, CD004405. [Google Scholar] [CrossRef]
- van der Flier, M.; Geelen, S.P.M.; Kimpen, J.L.L.; Hoepelman, I.M.; Tuomanen, E.I. Reprogramming the host response in bacterial meningitis: How best to improve outcome? Clin. Microbiol. Rev. 2003, 16, 415–429. [Google Scholar] [CrossRef][Green Version]
- Heide, E.C.; Bindila, L.; Post, J.M.; Malzahn, D.; Lutz, B.; Seele, J.; Nau, R.; Ribes, S. Prophylactic palmitoylethanolamide prolongs survival and decreases detrimental inflammation in aged mice with bacterial meningitidis. Front. Immunol. 2018, 9, 2671. [Google Scholar] [CrossRef]
- Makwana, N.; Riordan, A.I. Bacterial meningitis: The impact of vaccination. CNS Drugs 2007, 21, 355–366. [Google Scholar] [CrossRef]
- Bradshaw, J.L.; McDaniel, L.S. Selective pressure: Rise of the non-encapsulated pneumococcus. PLoS Pathog. 2019, 15, e1007911. [Google Scholar] [CrossRef]
- Tsang, R.S.W.; Tsai, C.M.; Henderson, A.M.; Tyler, S.; Law, D.K.S.; Zollinger, W.; Jamieson, F. Immunochemical studies and genetic background of two Neisseria meningitidis isolates expressing unusual capsule polysaccharide antigens with specificities of both serogroup Y and W135. Can. J. Microbiol. 2008, 54, 229–234. [Google Scholar] [CrossRef]
- Oliver, M.B.; van der Linden, M.P.G.; Küntzel, S.A.; Saad, J.S.; Nahm, M.H. Discovery of Streptococcus pneumoniae serotype 6 variants with glycosyltransferases synthesizing two differing repeating units. J. Biol. Chem. 2013, 288, 25976–25985. [Google Scholar] [CrossRef]
- Lessa, F.C.; Milucky, J.; Rouphael, N.G.; Bennett, N.M.; Talbot, H.K.; Harrison, L.H.; Farley, M.M.; Walston, J.; Pimenta, F.; Gertz, R.E.; et al. Streptococcus mitis expressing pneumococcal serotype 1 capsule. Sci. Rep. 2018, 8, 17959. [Google Scholar] [CrossRef] [PubMed]
- Sette, A.; Rappuoli, R. Reverse vaccinology: Developing vaccines in the era of genomics. Immunity 2010, 33, 530–541. [Google Scholar] [CrossRef]
- Obolski, U.; Gori, A.; Lourenço, J.; Thompson, C.; Thompson, R.; French, N.; Heyderman, R.S.; Gupta, S. Identifying genes associated with invasive disease in S. pneumoniae by applying a machine learning approach to whole genome sequence typing data. Sci. Rep. 2019, 9, 4049. [Google Scholar] [CrossRef]
| Organism | Capsular Serotype/Serogroup Antigens | Reference |
|---|---|---|
| H. influenzae | Serotypes a, b, c, d, e, and f, | Pittman, 1931 [26] |
| N. meningitidis | Serogroups A, B, C, E, H, I, K, L, W, X, Y, and Z | Harrison et al., 2013 [27] |
| S. agalactiae | Serotypes Ia, Ib, II, III, IV, V, VI, VII, VIII, and IX | Song et al., 2018 [5]; Lin et al., 2018 [13] |
| S. pneumonia * | 100 serotypes have been identified and only a few are listed here; Serotypes 1, 2, 3, 4, 5, 6A, 6B, 6C, …… 11E, 20B, … 35D, 7D, 10D (complete list can be found in the references provided) | Geno et al., 2015 [28]; Ganaie et al., 2020 [29] |
| Origanism | Vaccine Type | Serotype/Serogroup Targets | Protein Carrier † | Year First Licensed |
|---|---|---|---|---|
| H. influenzae | Hib conjugate | b | TT, OMP | 1987 |
| N. meningitidis | tetravalent polysaccharide | A, C, Y and W | None | 1974 |
| N. meningitidis | monovalent C conjugate | C | CRM197, TT | 1999 |
| N. meningitidis | monovalent A conjugate | A | TT | 2010 |
| N. meningitidis | tetravalent conjugate | A, C, Y and W | CRM197, DT, TT | 2005 |
| N. meningitidis | 4 component MenB | B | protein base vaccine | 2013 |
| N. meningitidis | factor H binding protein | B | protein base vaccine | 2018 |
| S. pneumoniae | PCV7 conjugate | 4, 6B, 9V, 14, 18C, 19F, 23F | CRM197 | 2000 |
| S. pneumoniae | PCV10 conjugate | 1, 4, 5, 6B, 7F, 9V, 14, 18C, 19F, 23F | CRM197, TT, DT | 2009 |
| S. pneumoniae | PCV13 conjugate | 1, 3, 4, 5, 6A, 6B, 7F, 9V, 14, 18C, 19A, 19F, 23F | CRM197 | 2011 |
| S. pneumoniae | PPV23 plain polysaccharide | 1, 2, 3, 4, 5, 6B, 7F, 8, 9N, 9V, 10A, 11A, 12F, 14, 15B, 17F, 18C, 19A, 19F, 20, 22F, 23F, 33F | None | 1983 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsang, R.S.W. A Narrative Review of the Molecular Epidemiology and Laboratory Surveillance of Vaccine Preventable Bacterial Meningitis Agents: Streptococcus pneumoniae, Neisseria meningitidis, Haemophilus influenzae and Streptococcus agalactiae. Microorganisms 2021, 9, 449. https://doi.org/10.3390/microorganisms9020449
Tsang RSW. A Narrative Review of the Molecular Epidemiology and Laboratory Surveillance of Vaccine Preventable Bacterial Meningitis Agents: Streptococcus pneumoniae, Neisseria meningitidis, Haemophilus influenzae and Streptococcus agalactiae. Microorganisms. 2021; 9(2):449. https://doi.org/10.3390/microorganisms9020449
Chicago/Turabian StyleTsang, Raymond S. W. 2021. "A Narrative Review of the Molecular Epidemiology and Laboratory Surveillance of Vaccine Preventable Bacterial Meningitis Agents: Streptococcus pneumoniae, Neisseria meningitidis, Haemophilus influenzae and Streptococcus agalactiae" Microorganisms 9, no. 2: 449. https://doi.org/10.3390/microorganisms9020449
APA StyleTsang, R. S. W. (2021). A Narrative Review of the Molecular Epidemiology and Laboratory Surveillance of Vaccine Preventable Bacterial Meningitis Agents: Streptococcus pneumoniae, Neisseria meningitidis, Haemophilus influenzae and Streptococcus agalactiae. Microorganisms, 9(2), 449. https://doi.org/10.3390/microorganisms9020449





