Gut–Skin Axis: Current Knowledge of the Interrelationship between Microbial Dysbiosis and Skin Conditions
Abstract
1. Introduction
2. Skin Versus Gut Barrier
3. Skin and Gut Microbiome Involvements
4. Acne Vulgaris
4.1. Acne Vulgaris Pathophysiology
4.2. Acne Vulgaris Skin Microbiome
4.3. Gut Microbiome and Diet Implications in Acne Vulgaris
5. Atopic Dermatitis (AD)
5.1. AD Pathophysiology
5.2. Skin Microbiome in AD
5.3. Gut Microbiome and Diet Implications in AD
6. Psoriasis
6.1. Psoriasis Pathophysiology
6.2. Skin Microbiome in Psoriasis
6.3. Gut Microbiome and Diet Implications in Psoriasis
7. Hidradenitis Suppurativa (HS)
7.1. HS Pathophysiology
7.2. Skin Microbiome in HS
7.3. Gut Microbiome and Diet Implications in HS
8. Rosacea
8.1. Rosacea Pathophysiology
8.2. Skin Microbiome in Rosacea
8.3. Gut Microbiome and Diet Implications in Rosacea
9. Dandruff and Seborrheic Dermatitis
9.1. Dandruff and Seborrheic Dermatitis Pathophysiology
9.2. Skin Microbiome in Dandruff and Seborrheic Dermatitis
9.3. Gut Microbiome and Diet Implications in Dandruff and Seborrheic Dermatitis
10. Alopecia
10.1. Alopecia Pathophysiology
10.2. Skin Microbiome in Alopecia
10.3. Gut Microbiome and Diet Implications in Alopecia
11. Skin Cancer
11.1. Skin Cancer Pathophysiology
11.2. Skin Microbiome in Skin Cancer
11.3. Gut Microbiome and Diet Implications in Skin Cancer
12. Wound Healing
12.1. Wound Pathophysiology
12.2. Wound Skin Microbiome
12.3. Gut Microbiome and Diet Implications in Wound Healing
13. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AD | Atopic Dermatitis |
| AMP | Antimicrobial Peptide |
| ASCA | Anti-Saccharomyces cerevisiae Antibody |
| BCC | Basal Cell Carcinoma |
| CD | Crohn’s Disease |
| CRC | Colorectal Cancer |
| DC | Dendritic Cell |
| EC | Epithelial Cell |
| EDTA | Ethylenediaminetetraacetic Acid |
| FMT | Fecal Microbiota Transplant |
| GABA | Gamma-Aminobutyric Acid |
| Gamma Delta | |
| GI | Gastrointestinal |
| HFD | High-Fat Diet |
| HPI | Helicobacter pylori Infection |
| HS | Hidradenitis Suppurativa |
| IBD | Inflammatory Bowel Disease |
| IBS | Irritable Bowel Syndrome |
| IEC | Intestinal Epithelial Cell |
| I-FABP | Intestinal Fatty Acid Binding Protein |
| IFN | Interferon |
| IFN-ISRE | IFN-Stimulated Response Element |
| IgE | Immunoglobulin E |
| IGF | Insulin-like Growth Factor |
| IL | Interleukin |
| ILC | Innate Lymphoid Cell |
| IMID | Immune-Mediated Inflammatory Disease |
| JAK | Janus Kinase |
| KLK5 | Kallikrein 5 |
| LC | Langerhans Cell |
| LL-37 | Cathelicidin AMP |
| M | Membranous |
| MAMP | Microbial-Associated Molecular Pattern |
| MCPyV | Merkel Cell Polyomavirus |
| MCC | Merkel Cell Carcinoma |
| MOA | Mechanism of Action |
| MRSA | Methicillin-resistant S. aureus |
| NMSC | Non-Melanoma Skin Cancer |
| P-2 | Perforin-2 |
| PUFA | Polyunsaturated Fatty Acid |
| ROS | Reactive Oxygen Species |
| SCC | Squamous Cell Carcinoma |
| SCFA | Short Chain Fatty Acid |
| SIBO | Small Intestinal Bacterial Overgrowth |
| SREBP-1 | Sterol Regulatory Element-Binding Protein 1 |
| STAT | Signal Transducer and Activator of Transcription protein |
| TLR2 | Toll-like Receptor 2 |
| Th | Helper T cell |
| Treg | Regulatory T cell |
| TRP | Transient Receptor Potential |
| TRPV | Transient Receptor Potential Ion Channels of Vanilloid Type |
| UC | Ulcerative Colitis |
| UVB | Ultraviolet B |
| VEGF | Vascularendothelial Growth Factor |
References
- Gallo, R.L. Human skin is the largest epithelial surface for interaction with microbes. J. Investig. Dermatol. 2017, 137, 1213–1214. [Google Scholar] [CrossRef] [PubMed]
- Helander, H.F.; Fändriks, L. Surface area of the digestive tract–revisited. Scand. J. Gastroenterol. 2014, 49, 681–689. [Google Scholar] [CrossRef] [PubMed]
- Thursby, E.; Juge, N. Introduction to the human gut microbiota. Biochem. J. 2017, 474, 1823–1836. [Google Scholar] [CrossRef] [PubMed]
- Williams, R. Benefit and mischief from commensal bacteria. J. Clin. Pathol. 1973, 26, 811. [Google Scholar] [CrossRef]
- Savage, D.C. Microbial ecology of the gastrointestinal tract. Annu. Rev. Microbiol. 1977, 31, 107–133. [Google Scholar] [CrossRef]
- Grice, E.A.; Segre, J.A. The skin microbiome. Nat. Rev. Microbiol. 2011, 9, 244–253. [Google Scholar] [CrossRef]
- Arck, P.; Handjiski, B.; Hagen, E.; Pincus, M.; Bruenahl, C.; Bienenstock, J.; Paus, R. Is there a ‘gut–brain–skin axis’? Exp. Dermatol. 2010, 19, 401–405. [Google Scholar] [CrossRef] [PubMed]
- Shah, K.R.; Boland, C.R.; Patel, M.; Thrash, B.; Menter, A. Cutaneous manifestations of gastrointestinal disease: Part I. J. Am. Acad. Dermatol. 2013, 68, 189.e1–189.e21. [Google Scholar] [CrossRef] [PubMed]
- Thrash, B.; Patel, M.; Shah, K.R.; Boland, C.R.; Menter, A. Cutaneous manifestations of gastrointestinal disease: Part II. J. Am. Acad. Dermatol. 2013, 68, 211.e1–211.e33. [Google Scholar] [CrossRef] [PubMed]
- Gloster, H.M.; Gebauer, L.E.; Mistur, R.L. Cutaneous manifestations of gastrointestinal disease. In Absolute Dermatology Review; Springer: Berlin/Heidelberg, Germany, 2016; pp. 171–179. [Google Scholar]
- O’Neill, C.A.; Monteleone, G.; McLaughlin, J.T.; Paus, R. The gut-skin axis in health and disease: A paradigm with therapeutic implications. BioEssays 2016, 38, 1167–1176. [Google Scholar] [CrossRef]
- Shaykhiev, R.; Bals, R. Interactions between epithelial cells and leukocytes in immunity and tissue homeostasis. J. Leukoc. Biol. 2007, 82, 1–15. [Google Scholar] [CrossRef]
- Bach, J.F. The effect of infections on susceptibility to autoimmune and allergic diseases. N. Engl. J. Med. 2002, 347, 911–920. [Google Scholar] [CrossRef]
- Madison, K.C. Barrier function of the skin:“la raison d’etre” of the epidermis. J. Investig. Dermatol. 2003, 121, 231–241. [Google Scholar] [CrossRef] [PubMed]
- Lange, L.; Huang, Y.; Busk, P.K. Microbial decomposition of keratin in nature—a new hypothesis of industrial relevance. Appl. Microbiol. Biotechnol. 2016, 100, 2083–2096. [Google Scholar] [CrossRef] [PubMed]
- Pelaseyed, T.; Bergström, J.H.; Gustafsson, J.K.; Ermund, A.; Birchenough, G.M.; Schütte, A.; van der Post, S.; Svensson, F.; Rodríguez-Piñeiro, A.M.; Nyström, E.E.; et al. The mucus and mucins of the goblet cells and enterocytes provide the first defense line of the gastrointestinal tract and interact with the immune system. Immunol. Rev. 2014, 260, 8–20. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.S.; Ho, S.B. Intestinal goblet cells and mucins in health and disease: Recent insights and progress. Curr. Gastroenterol. Rep. 2010, 12, 319–330. [Google Scholar] [CrossRef]
- Janeway, C.A., Jr.; Travers, P.; Walport, M.; Shlomchik, M.J. The front line of host defense. In Immunobiology: The Immune System in Health and Disease, 5th ed.; Garland Science: New York, NY, USA, 2001. [Google Scholar]
- Schmid-Wendtner, M.H.; Korting, H.C. The pH of the skin surface and its impact on the barrier function. Skin Pharmacol. Physiol. 2006, 19, 296–302. [Google Scholar] [CrossRef]
- Nakatsuji, T.; Kao, M.C.; Zhang, L.; Zouboulis, C.C.; Gallo, R.L.; Huang, C.M. Sebum free fatty acids enhance the innate immune defense of human sebocytes by upregulating β-defensin-2 expression. J. Investig. Dermatol. 2010, 130, 985–994. [Google Scholar] [CrossRef]
- Dahlhoff, M.; Zouboulis, C.C.; Schneider, M.R. Expression of dermcidin in sebocytes supports a role for sebum in the constitutive innate defense of human skin. J. Dermatol. Sci. 2016, 81, 124–126. [Google Scholar] [CrossRef] [PubMed]
- Patricia, J.J.; Dhamoon, A.S. Physiology, Digestion. 2019. Available online: https://europepmc.org/books/nbk544242 (accessed on 2 November 2020).
- Brown, E.M.; Sadarangani, M.; Finlay, B.B. The role of the immune system in governing host-microbe interactions in the intestine. Nat. Immunol. 2013, 14, 660–667. [Google Scholar] [CrossRef]
- Spits, H.; Cupedo, T. Innate lymphoid cells: Emerging insights in development, lineage relationships, and function. Annu. Rev. Immunol. 2012, 30, 647–675. [Google Scholar] [CrossRef]
- Braff, M.H.; Zaiou, M.; Fierer, J.; Nizet, V.; Gallo, R.L. Keratinocyte production of cathelicidin provides direct activity against bacterial skin pathogens. Infect. Immun. 2005, 73, 6771–6781. [Google Scholar] [CrossRef] [PubMed]
- Gläser, R.; Harder, J.; Lange, H.; Bartels, J.; Christophers, E.; Schröder, J.M. Antimicrobial psoriasin (S100A7) protects human skin from Escherichia coli infection. Nat. Immunol. 2005, 6, 57–64. [Google Scholar] [CrossRef]
- Yamasaki, K.; Schauber, J.; Coda, A.; Lin, H.; Dorschner, R.A.; Schechter, N.M.; Bonnart, C.; Descargues, P.; Hovnanian, A.; Gallo, R.L. Kallikrein-mediated proteolysis regulates the antimicrobial effects of cathelicidins in skin. FASEB J. 2006, 20, 2068–2080. [Google Scholar] [CrossRef]
- Johansson, M.E.; Sjövall, H.; Hansson, G.C. The gastrointestinal mucus system in health and disease. Nat. Rev. Gastroenterol. Hepatol. 2013, 10, 352. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Leblond, C. Origin, differentiation and renewal of the four main epithelial cell types in the mouse small intestine I. Columnar cell. Am. J. Anat. 1974, 141, 461–479. [Google Scholar] [CrossRef] [PubMed]
- Podolsky, D.K.; Lynch-Devaney, K.; Stow, J.L.; Oates, P.; Murgue, B.; DeBeaumont, M.; Sands, B.E.; Mahida, Y.R. Identification of human intestinal trefoil factor. Goblet cell-specific expression of a peptide targeted for apical secretion. J. Biol. Chem. 1993, 268, 6694–6702. [Google Scholar] [CrossRef]
- Johansson, M.E.; Larsson, J.M.H.; Hansson, G.C. The two mucus layers of colon are organized by the MUC2 mucin, whereas the outer layer is a legislator of host–microbial interactions. Proc. Natl. Acad. Sci. USA 2011, 108, 4659–4665. [Google Scholar] [CrossRef]
- Vaishnava, S.; Behrendt, C.L.; Ismail, A.S.; Eckmann, L.; Hooper, L.V. Paneth cells directly sense gut commensals and maintain homeostasis at the intestinal host-microbial interface. Proc. Natl. Acad. Sci. USA 2008, 105, 20858–20863. [Google Scholar] [CrossRef]
- Qiu, J.; Heller, J.J.; Guo, X.; Zong-ming, E.C.; Fish, K.; Fu, Y.X.; Zhou, L. The aryl hydrocarbon receptor regulates gut immunity through modulation of innate lymphoid cells. Immunity 2012, 36, 92–104. [Google Scholar] [CrossRef]
- Sonnenberg, G.F.; Monticelli, L.A.; Elloso, M.M.; Fouser, L.A.; Artis, D. CD4+ lymphoid tissue-inducer cells promote innate immunity in the gut. Immunity 2011, 34, 122–134. [Google Scholar] [CrossRef] [PubMed]
- Niess, J.H.; Brand, S.; Gu, X.; Landsman, L.; Jung, S.; McCormick, B.A.; Vyas, J.M.; Boes, M.; Ploegh, H.L.; Fox, J.G.; et al. CX3CR1-mediated dendritic cell access to the intestinal lumen and bacterial clearance. Science 2005, 307, 254–258. [Google Scholar] [CrossRef] [PubMed]
- Macpherson, A.J.; Uhr, T. Induction of protective IgA by intestinal dendritic cells carrying commensal bacteria. Science 2004, 303, 1662–1665. [Google Scholar] [CrossRef] [PubMed]
- Neutra, M.R.; Pringault, E.; Kraehenbuhl, J.P. Antigen sampling across epithelial barriers and induction of mucosal immune responses. Annu. Rev. Immunol. 1996, 14, 275–300. [Google Scholar] [CrossRef]
- Barker, N. Adult intestinal stem cells: Critical drivers of epithelial homeostasis and regeneration. Nat. Rev. Mol. Cell Biol. 2014, 15, 19–33. [Google Scholar] [CrossRef]
- Hsieh, E.A.; Chai, C.M.; Benito, O.; Neese, R.A.; Hellerstein, M.K. Dynamics of keratinocytes in vivo using 2H2O labeling: A sensitive marker of epidermal proliferation state. J. Investig. Dermatol. 2004, 123, 530–536. [Google Scholar] [CrossRef]
- Sekirov, I.; Russell, S.L.; Antunes, L.C.M.; Finlay, B.B. Gut microbiota in health and disease. Physiol. Rev. 2010, 90, 859–904. [Google Scholar] [CrossRef]
- Capone, K.A.; Dowd, S.E.; Stamatas, G.N.; Nikolovski, J. Diversity of the human skin microbiome early in life. J. Investig. Dermatol. 2011, 131, 2026–2032. [Google Scholar] [CrossRef] [PubMed]
- Dzutsev, A.; Goldszmid, R.S.; Viaud, S.; Zitvogel, L.; Trinchieri, G. The role of the microbiota in inflammation, carcinogenesis, and cancer therapy. Eur. J. Immunol. 2015, 45, 17–31. [Google Scholar] [CrossRef]
- Vlachos, C.; Gaitanis, G.; Katsanos, K.H.; Christodoulou, D.K.; Tsianos, E.; Bassukas, I.D. Psoriasis and inflammatory bowel disease: Links and risks. Psoriasis 2016, 6, 73. [Google Scholar]
- Clarke, G.; Stilling, R.M.; Kennedy, P.J.; Stanton, C.; Cryan, J.F.; Dinan, T.G. Minireview: Gut microbiota: The neglected endocrine organ. Mol. Endocrinol. 2014, 28, 1221–1238. [Google Scholar] [CrossRef]
- Chapat, L.; Chemin, K.; Dubois, B.; Bourdet-Sicard, R.; Kaiserlian, D. Lactobacillus casei reduces CD8+ T cell-mediated skin inflammation. Eur. J. Immunol. 2004, 34, 2520–2528. [Google Scholar] [CrossRef]
- Guéniche, A.; Benyacoub, J.; Buetler, T.M.; Smola, H.; Blum, S. Supplementation with oral probiotic bacteria maintains cutaneous immune homeostasis after UV exposure. Eur. J. Dermatol. 2006, 16, 511–517. [Google Scholar]
- Benyacoub, J.; Bosco, N.; Blanchard, C.; Demont, A.; Philippe, D.; Castiel-Higounenc, I.; Guéniche, A. Immune modulation property of Lactobacillus paracasei NCC2461 (ST11) strain and impact on skin defences. Benef. Microbes 2014, 5, 129–136. [Google Scholar] [CrossRef]
- Belkaid, Y.; Tamoutounour, S. The influence of skin microorganisms on cutaneous immunity. Nat. Rev. Immunol. 2016, 16, 353–366. [Google Scholar] [CrossRef] [PubMed]
- Johnson, L.R.; Christensen, J.; Jackson, M.J. Physiology of the Gastrointestinal Tract, 2nd ed.; Raven: New York, NY, USA, 1987; pp. 665–693. [Google Scholar]
- Ipci, K.; Altıntoprak, N.; Muluk, N.B.; Senturk, M.; Cingi, C. The possible mechanisms of the human microbiome in allergic diseases. Eur. Arch. Oto-Rhino 2017, 274, 617–626. [Google Scholar] [CrossRef]
- LeBlanc, J.G.; Milani, C.; De Giori, G.S.; Sesma, F.; Van Sinderen, D.; Ventura, M. Bacteria as vitamin suppliers to their host: A gut microbiota perspective. Curr. Opin. Biotechnol. 2013, 24, 160–168. [Google Scholar] [CrossRef]
- Scott, K.P.; Gratz, S.W.; Sheridan, P.O.; Flint, H.J.; Duncan, S.H. The influence of diet on the gut microbiota. Pharmacol. Res. 2013, 69, 52–60. [Google Scholar] [CrossRef] [PubMed]
- Yazdanbakhsh, M.; Kremsner, P.G.; Van Ree, R. Allergy, parasites, and hygiene hypothesis. Science 2002, 296, 490–494. [Google Scholar] [CrossRef]
- Mccall, L.I.; Callewaert, C.; Zhu, Q.; Song, S.J.; Bouslimani, A.; Minich, J.J.; Ernst, M.; Ruiz-Calderon, J.F.; Cavallin, H.; Pereira, H.S.; et al. Home chemical and microbial transitions across urbanization. Nat. Microbiol. 2020, 5, 108–115. [Google Scholar] [CrossRef] [PubMed]
- Callewaert, C.; Helffer, K.R.; Lebaron, P. Skin Microbiome and its Interplay with the Environment. Am. J. Clin. Dermatol. 2020, 21, 4–11. [Google Scholar] [CrossRef]
- Guo, X.; Li, J.; Tang, R.; Zhang, G.; Zeng, H.; Wood, R.J.; Liu, Z. High fat diet alters gut microbiota and the expression of paneth cell-antimicrobial peptides preceding changes of circulating inflammatory cytokines. Mediat. Inflamm. 2017, 2017, 9474896. [Google Scholar] [CrossRef] [PubMed]
- Gee, J.; Wortley, G.; Johnson, I.; Price, K.; Rutten, A.; Houben, G.; Penninks, A. Effects of saponins and glycoalkaloids on the permeability and viability of mammalian intestinal cells and on the integrity of tissue preparations in vitro. Toxicol. Vitr. 1996, 10, 117–128. [Google Scholar] [CrossRef]
- Humbert, P.; Pelletier, F.; Dreno, B.; Puzenat, E.; Aubin, F. Gluten intolerance and skin diseases. Eur. J. Dermatol. 2006, 16, 4–11. [Google Scholar] [PubMed]
- Fry, L.; Riches, D.; Seah, P.; Hoffbrand, A. Clearance of skin lesions in dermatitis herpetiformis after gluten withdrawal. Lancet 1973, 301, 288–291. [Google Scholar] [CrossRef]
- Grossi, E.; Cazzaniga, S.; Crotti, S.; Naldi, L.; Di Landro, A.; Ingordo, V.; Cusano, F.; Atzori, L.; Tripodi Cutrì, F.; Musumeci, M.; et al. The constellation of dietary factors in adolescent acne: A semantic connectivity map approach. J. Eur. Acad. Dermatol. Venereol. 2016, 30, 96–100. [Google Scholar] [CrossRef]
- Bosman, E.S.; Albert, A.Y.; Lui, H.; DUTZ, J.P.; Vallance, B.A. Skin exposure to Narrow Band Ultraviolet (UV) B light modulates the human intestinal microbiome. Front. Microbiol. 2019, 10, 2410. [Google Scholar] [CrossRef]
- Brough, H.A.; Liu, A.H.; Sicherer, S.; Makinson, K.; Douiri, A.; Brown, S.J.; Stephens, A.C.; McLean, W.I.; Turcanu, V.; Wood, R.A.; et al. Atopic dermatitis increases the effect of exposure to peanut antigen in dust on peanut sensitization and likely peanut allergy. J. Allergy Clin. Immunol. 2015, 135, 164–170. [Google Scholar] [CrossRef]
- Bartnikas, L.M.; Gurish, M.F.; Burton, O.T.; Leisten, S.; Janssen, E.; Oettgen, H.C.; Beaupré, J.; Lewis, C.N.; Austen, K.F.; Schulte, S.; et al. Epicutaneous sensitization results in IgE-dependent intestinal mast cell expansion and food-induced anaphylaxis. J. Allergy Clin. Immunol. 2013, 131, 451–460. [Google Scholar] [CrossRef] [PubMed]
- Hoh, R.A.; Joshi, S.A.; Lee, J.Y.; Martin, B.A.; Varma, S.; Kwok, S.; Nielsen, S.C.; Nejad, P.; Haraguchi, E.; Dixit, P.S.; et al. Origins and clonal convergence of gastrointestinal IgE+ B cells in human peanut allergy. Sci. Immunol. 2020, 5, eaay4209. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.L.; Chandra, S.; Shih, D.Q. Skin manifestations of inflammatory bowel disease. Front. Physiol. 2012, 3, 13. [Google Scholar] [CrossRef] [PubMed]
- Bowe, W.P.; Logan, A.C. Acne vulgaris, probiotics and the gut-brain-skin axis-back to the future? Gut Pathog. 2011, 3, 1–11. [Google Scholar] [CrossRef]
- Lyte, M. Microbial endocrinology and the microbiota-gut-brain axis. In Microbial Endocrinology: The Microbiota-Gut-Brain Axis in Health and Disease; Springer: Berlin/Heidelberg, Germany, 2014; pp. 3–24. [Google Scholar]
- Rea, K.; Dinan, T.G.; Cryan, J.F. The microbiome: A key regulator of stress and neuroinflammation. Neurobiol. Stress 2016, 4, 23–33. [Google Scholar] [CrossRef]
- Cummings, J.H.; Macfarlane, G.T. Role of intestinal bacteria in nutrient metabolism. Clin. Nutr. 1997, 16, 3–11. [Google Scholar] [CrossRef]
- Mariadason, J.; Catto-Smith, A.; Gibson, P. Modulation of distal colonic epithelial barrier function by dietary fibre in normal rats. Gut 1999, 44, 394–399. [Google Scholar] [CrossRef]
- Lomholt, H.B.; Kilian, M. Population genetic analysis of Propionibacterium acnes identifies a subpopulation and epidemic clones associated with acne. PLoS ONE 2010, 5, e12277. [Google Scholar] [CrossRef]
- Lomholt, H.; Scholz, C.; Brüggemann, H.; Tettelin, H.; Kilian, M. A comparative study of Cutibacterium (Propionibacterium) acnes clones from acne patients and healthy controls. Anaerobe 2017, 47, 57–63. [Google Scholar] [CrossRef] [PubMed]
- McDowell, A.; Gao, A.; Barnard, E.; Fink, C.; Murray, P.I.; Dowson, C.G.; Nagy, I.; Lambert, P.A.; Patrick, S. A novel multilocus sequence typing scheme for the opportunistic pathogen Propionibacterium acnes and characterization of type I cell surface-associated antigens. Microbiology 2011, 157, 1990–2003. [Google Scholar] [CrossRef] [PubMed]
- Paugam, C.; Corvec, S.; Saint-Jean, M.; Le Moigne, M.; Khammari, A.; Boisrobert, A.; Nguyen, J.; Gaultier, A.; Dréno, B. Propionibacterium acnes phylotypes and acne severity: An observational prospective study. J. Eur. Acad. Dermatol. Venereol. 2017, 31, e398–e399. [Google Scholar] [CrossRef]
- Fitz-Gibbon, S.; Tomida, S.; Chiu, B.H.; Nguyen, L.; Du, C.; Liu, M.; Elashoff, D.; Erfe, M.C.; Loncaric, A.; Kim, J.; et al. Propionibacterium acnes strain populations in the human skin microbiome associated with acne. J. Investig. Dermatol. 2013, 133, 2152–2160. [Google Scholar] [CrossRef] [PubMed]
- Karoglan, A.; Paetzold, B.; De Lima, J.P.; Brüggemann, H.; Tüting, T.; Schanze, D.; Güell, M.; Gollnick, H. Safety and efficacy of topically applied selected cutibacterium acnes strains over five weeks in patients with acne vulgaris: An open-label, pilot study. Acta Derm. Venereol. 2019, 99, 1253–1257. [Google Scholar] [CrossRef]
- Johnson, T.; Kang, D.; Barnard, E.; Li, H. Strain-level differences in porphyrin production and regulation in Propionibacterium acnes elucidate disease associations. Msphere 2016, 1. [Google Scholar] [CrossRef] [PubMed]
- Barnard, E.; Shi, B.; Kang, D.; Craft, N.; Li, H. The balance of metagenomic elements shapes the skin microbiome in acne and health. Sci. Rep. 2016, 6, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Lheure, C.; Grange, P.A.; Ollagnier, G.; Morand, P.; Désiré, N.; Sayon, S.; Corvec, S.; Raingeaud, J.; Marcelin, A.G.; Calvez, V.; et al. TLR-2 recognizes Propionibacterium acnes CAMP factor 1 from highly inflammatory strains. PLoS ONE 2016, 11, e0167237. [Google Scholar] [CrossRef]
- Kong, H.H.; Oh, J.; Deming, C.; Conlan, S.; Grice, E.A.; Beatson, M.A.; Nomicos, E.; Polley, E.C.; Komarow, H.D.; Murray, P.R.; et al. Temporal shifts in the skin microbiome associated with disease flares and treatment in children with atopic dermatitis. Genome Res. 2012, 22, 850–859. [Google Scholar] [CrossRef] [PubMed]
- Shi, B.; Bangayan, N.J.; Curd, E.; Taylor, P.A.; Gallo, R.L.; Leung, D.Y.; Li, H. The skin microbiome is different in pediatric versus adult atopic dermatitis. J. Allergy Clin. Immunol. 2016, 138, 1233–1236. [Google Scholar] [CrossRef]
- Oh, J.; Freeman, A.F.; Park, M.; Sokolic, R.; Candotti, F.; Holland, S.M.; Segre, J.A.; Kong, H.H.; NISC Comparative Sequencing Program. The altered landscape of the human skin microbiome in patients with primary immunodeficiencies. Genome Res. 2013, 23, 2103–2114. [Google Scholar] [CrossRef]
- Chng, K.R.; Tay, A.S.L.; Li, C.; Ng, A.H.Q.; Wang, J.; Suri, B.K.; Matta, S.A.; McGovern, N.; Janela, B.; Wong, X.F.C.C.; et al. Whole metagenome profiling reveals skin microbiome-dependent susceptibility to atopic dermatitis flare. Nat. Microbiol. 2016, 1, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Alekseyenko, A.V.; Perez-Perez, G.I.; De Souza, A.; Strober, B.; Gao, Z.; Bihan, M.; Li, K.; Methé, B.A.; Blaser, M.J. Community differentiation of the cutaneous microbiota in psoriasis. Microbiome 2013, 1, 31. [Google Scholar] [CrossRef] [PubMed]
- Statnikov, A.; Alekseyenko, A.V.; Li, Z.; Henaff, M.; Perez-Perez, G.I.; Blaser, M.J.; Aliferis, C.F. Microbiomic signatures of psoriasis: Feasibility and methodology comparison. Sci. Rep. 2013, 3, 2620. [Google Scholar] [CrossRef] [PubMed]
- Takemoto, A.; Cho, O.; Morohoshi, Y.; Sugita, T.; Muto, M. Molecular characterization of the skin fungal microbiome in patients with psoriasis. J. Dermatol. 2015, 42, 166–170. [Google Scholar] [CrossRef]
- Chang, H.W.; Yan, D.; Singh, R.; Liu, J.; Lu, X.; Ucmak, D.; Lee, K.; Afifi, L.; Fadrosh, D.; Leech, J.; et al. Alteration of the cutaneous microbiome in psoriasis and potential role in Th17 polarization. Microbiome 2018, 6, 154. [Google Scholar] [CrossRef]
- Yerushalmi, M.; Elalouf, O.; Anderson, M.; Chandran, V. The skin microbiome in psoriatic disease: A systematic review and critical appraisal. J. Transl. Autoimmun. 2019, 2, 100009. [Google Scholar] [CrossRef]
- Guet-Revillet, H.; Jais, J.P.; Ungeheuer, M.N.; Coignard-Biehler, H.; Duchatelet, S.; Delage, M.; Lam, T.; Hovnanian, A.; Lortholary, O.; Nassif, X.; et al. The microbiological landscape of anaerobic infections in hidradenitis suppurativa: A prospective metagenomic study. Clin. Infect. Dis. 2017, 65, 282–291. [Google Scholar] [CrossRef]
- Assan, F.; Gottlieb, J.; Tubach, F.; Lebbah, S.; Guigue, N.; Hickman, G.; Pape, E.; Madrange, M.; Delaporte, E.; Sendid, B.; et al. Anti-Saccharomyces cerevisiae IgG and IgA antibodies are associated with systemic inflammation and advanced disease in hidradenitis suppurativa. J. Allergy Clin. Immunol. 2020, 146, 452–455. [Google Scholar] [CrossRef] [PubMed]
- Forton, F.; Seys, B. Density of Demodex folliculorum in rosacea: A case-control study using standardized skin-surface biopsy. Br. J. Dermatol. 1993, 128, 650–659. [Google Scholar] [CrossRef] [PubMed]
- Woo, Y.R.; Lim, J.H.; Cho, D.H.; Park, H.J. Rosacea: Molecular mechanisms and management of a chronic cutaneous inflammatory condition. Int. J. Mol. Sci. 2016, 17, 1562. [Google Scholar] [CrossRef]
- Dawson, T.L., Jr. Malassezia globosa and restricta: Breakthrough understanding of the etiology and treatment of dandruff and seborrheic dermatitis through whole-genome analysis. J. Investig. Dermatol. Symp. Proc. 2007, 12, 15–19. [Google Scholar] [CrossRef]
- MacKee, G.M.; Lewis, G.M.; WTTA of Martha; Spence, J.; WTTA of Mary; Hopper, E. Dandruff and seborrhea: I. flora of “normal” and diseased scalps. J. Investig. Dermatol. 1938, 1, 131–139. [Google Scholar] [CrossRef]
- Xu, Z.; Wang, Z.; Yuan, C.; Liu, X.; Yang, F.; Wang, T.; Wang, J.; Manabe, K.; Qin, O.; Wang, X.; et al. Dandruff is associated with the conjoined interactions between host and microorganisms. Sci. Rep. 2016, 6, 1–9. [Google Scholar] [CrossRef]
- Clavaud, C.; Jourdain, R.; Bar-Hen, A.; Tichit, M.; Bouchier, C.; Pouradier, F.; El Rawadi, C.; Guillot, J.; Ménard-Szczebara, F.; Breton, L.; et al. Dandruff is associated with disequilibrium in the proportion of the major bacterial and fungal populations colonizing the scalp. PLoS ONE 2013, 8, e58203. [Google Scholar] [CrossRef]
- Skinner, R.B., Jr.; Light, W.H.; Leonardi, C.; Bale, G.F.; Rosenberg, E.W. A molecular approach to alopecia areata. J. Investig. Dermatol. 1995, 104, 3S. [Google Scholar] [CrossRef]
- Rudnicka, L.; Lukomska, M. Alternaria scalp infection in a patient with alopecia areata. Coexistence or causative relationship? J. Dermatol. Case Rep. 2012, 6, 120. [Google Scholar] [CrossRef][Green Version]
- Pinto, D.; Sorbellini, E.; Marzani, B.; Rucco, M.; Giuliani, G.; Rinaldi, F. Scalp bacterial shift in Alopecia areata. PLoS ONE 2019, 14, e0215206. [Google Scholar] [CrossRef] [PubMed]
- Wood, D.L.; Lachner, N.; Tan, J.M.; Tang, S.; Angel, N.; Laino, A.; Linedale, R.; Lê Cao, K.A.; Morrison, M.; Frazer, I.H.; et al. A natural history of actinic keratosis and cutaneous squamous cell carcinoma microbiomes. MBio 2018, 9. [Google Scholar] [CrossRef]
- Cheng, J.; Zens, M.S.; Duell, E.; Perry, A.E.; Chapman, M.S.; Karagas, M.R. History of allergy and atopic dermatitis in relation to squamous cell and basal cell carcinoma of the skin. Cancer Epidemiol. Prev. Biomarkers 2015, 24, 749–754. [Google Scholar] [CrossRef]
- Mrázek, J.; Mekadim, C.; Kučerová, P.; Švejstil, R.; Salmonová, H.; Vlasáková, J.; Tarasová, R.; Čížková, J.; Červinková, M. Melanoma-related changes in skin microbiome. Folia Microbiol. 2019, 64, 435–442. [Google Scholar] [CrossRef] [PubMed]
- Sherwani, M.A.; Tufail, S.; Muzaffar, A.F.; Yusuf, N. The skin microbiome and immune system: Potential target for chemoprevention? Photodermatol. Photoimmunol. Photomed. 2018, 34, 25–34. [Google Scholar] [CrossRef] [PubMed]
- Sikorska, H.; Smoragiewicz, W. Role of probiotics in the prevention and treatment of meticillin-resistant Staphylococcus aureus infections. Int. J. Antimicrob. Agents 2013, 42, 475–481. [Google Scholar] [CrossRef]
- Guo, H.; Zheng, Y.; Wang, B.; Li, Z. A note on an improved self-healing group key distribution scheme. Sensors 2015, 15, 25033–25038. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.N.; Mann, N.J.; Braue, A.; Mäkeläinen, H.; Varigos, G.A. A low-glycemic-load diet improves symptoms in acne vulgaris patients: A randomized controlled trial. Am. J. Clin. Nutr. 2007, 86, 107–115. [Google Scholar] [CrossRef] [PubMed]
- Song, H.; Yoo, Y.; Hwang, J.; Na, Y.C.; Kim, H.S. Faecalibacterium prausnitzii subspecies–level dysbiosis in the human gut microbiome underlying atopic dermatitis. J. Allergy Clin. Immunol. 2016, 137, 852–860. [Google Scholar] [CrossRef] [PubMed]
- Kalliomäki, M.; Kirjavainen, P.; Eerola, E.; Kero, P.; Salminen, S.; Isolauri, E. Distinct patterns of neonatal gut microflora in infants in whom atopy was and was not developing. J. Allergy Clin. Immunol. 2001, 107, 129–134. [Google Scholar] [CrossRef]
- Penders, J.; Thijs, C.; van den Brandt, P.A.; Kummeling, I.; Snijders, B.; Stelma, F.; Adams, H.; van Ree, R.; Stobberingh, E.E. Gut microbiota composition and development of atopic manifestations in infancy: The KOALA Birth Cohort Study. Gut 2007, 56, 661–667. [Google Scholar] [CrossRef]
- Lee, E.; Lee, S.Y.; Kang, M.J.; Kim, K.; Won, S.; Kim, B.J.; Choi, K.Y.; Kim, B.S.; Cho, H.J.; Kim, Y.; et al. Clostridia in the gut and onset of atopic dermatitis via eosinophilic inflammation. Ann. Allergy Asthma Immunol. 2016, 117, 91–92. [Google Scholar] [CrossRef]
- Kirjavainen, P.; Arvola, T.; Salminen, S.; Isolauri, E. Aberrant composition of gut microbiota of allergic infants: A target of bifidobacterial therapy at weaning? Gut 2002, 51, 51–55. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, S.; Narisawa, Y.; Arase, S.; Okamatsu, H.; Ikenaga, T.; Tajiri, Y.; Kumemura, M. Differences in fecal microflora between patients with atopic dermatitis and healthy control subjects. J. Allergy Clin. Immunol. 2003, 111, 587–591. [Google Scholar] [CrossRef]
- Fujimura, K.E.; Sitarik, A.R.; Havstad, S.; Lin, D.L.; Levan, S.; Fadrosh, D.; Panzer, A.R.; LaMere, B.; Rackaityte, E.; Lukacs, N.W.; et al. Neonatal gut microbiota associates with childhood multisensitized atopy and T cell differentiation. Nat. Med. 2016, 22, 1187–1191. [Google Scholar] [CrossRef]
- Fiocchi, A.; Pawankar, R.; Cuello-Garcia, C.; Ahn, K.; Al-Hammadi, S.; Agarwal, A.; Beyer, K.; Burks, W.; Canonica, G.W.; Ebisawa, M.; et al. World allergy organization-McMaster university guidelines for allergic disease prevention (GLAD-P): Probiotics. World Allergy Organ. J. 2015, 8, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Sikora, M.; Stec, A.; Chrabaszcz, M.; Knot, A.; Waskiel-Burnat, A.; Rakowska, A.; Olszewska, M.; Rudnicka, L. Gut microbiome in psoriasis: An updated review. Pathogens 2020, 9, 463. [Google Scholar] [CrossRef]
- Grine, L.; Steeland, S.; Van Ryckeghem, S.; Ballegeer, M.; Lienenklaus, S.; Weiss, S.; Sanders, N.N.; Vandenbroucke, R.E.; Libert, C. Topical imiquimod yields systemic effects due to unintended oral uptake. Sci. Rep. 2016, 6, 20134. [Google Scholar] [CrossRef] [PubMed]
- Jung, G.W.; Tse, J.E.; Guiha, I.; Rao, J. Prospective, randomized, open-label trial comparing the safety, efficacy, and tolerability of an acne treatment regimen with and without a probiotic supplement and minocycline in subjects with mild to moderate acne. J. Cutan. Med. Surg. 2013, 17, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.J.; Nguyen, T.U.; Poon, K.Y.T.; Herrinton, L.J. The association of psoriasis with autoimmune diseases. J. Am. Acad. Dermatol. 2012, 67, 924–930. [Google Scholar] [CrossRef]
- Pietrzak, D.; Pietrzak, A.; Krasowska, D.; Borzęcki, A.; Franciszkiewicz-Pietrzak, K.; Polkowska-Pruszyńska, B.; Baranowska, M.; Reich, K. Digestive system in psoriasis: An update. Arch. Dermatol. Res. 2017, 309, 679–693. [Google Scholar] [CrossRef] [PubMed]
- Yeh, N.L.; Hsu, C.Y.; Tsai, T.F.; Chiu, H.Y. Gut microbiome in psoriasis is perturbed differently during secukinumab and ustekinumab therapy and associated with response to treatment. Clin. Drug Investig. 2019, 39, 1195–1203. [Google Scholar] [CrossRef]
- Wark, K.J.; Cains, G.D. The Microbiome in Hidradenitis Suppurativa: A Review. In Dermatology and Therapy; Springer: Cham, Switzerland, 2020; Volume 11, pp. 39–52. [Google Scholar]
- Brooks, M. Gut microbe curbs systemic inflammation in psoriasis. In Proceedings of the 29th European Academy of Dermatology and Venereology Congress (EADV), Vienna, Austria, 29–31 October 2020. [Google Scholar]
- Rebora, A.; Drago, F.; Parodi, A. May Helicohacter pylori be important for dermatologists. Dermatology 1995, 191, 6–8. [Google Scholar] [CrossRef] [PubMed]
- Parodi, A.; Paolino, S.; Greco, A.; Drago, F.; Mansi, C.; Rebora, A.; Parodi, A.; Savarino, V. Small intestinal bacterial overgrowth in rosacea: Clinical effectiveness of its eradication. Clin. Gastroenterol. Hepatol. 2008, 6, 759–764. [Google Scholar] [CrossRef]
- Nam, J.H.; Yun, Y.; Kim, H.S.; Kim, H.N.; Jung, H.J.; Chang, Y.; Ryu, S.; Shin, H.; Kim, H.L.; Kim, W.S. Rosacea and its association with enteral microbiota in Korean females. Exp. Dermatol. 2018, 27, 37–42. [Google Scholar] [CrossRef]
- Reygagne, P.; Bastien, P.; Couavoux, M.; Philippe, D.; Renouf, M.; Castiel-Higounenc, I.; Gueniche, A. The positive benefit of Lactobacillus paracasei NCC2461 ST11 in healthy volunteers with moderate to severe dandruff. Benef. Microbes 2017, 8, 671–680. [Google Scholar] [CrossRef]
- Moreno-Arrones, O.; Serrano-Villar, S.; Perez-Brocal, V.; Saceda-Corralo, D.; Morales-Raya, C.; Rodrigues-Barata, R.; Moya, A.; Jaen-Olasolo, P.; Vano-Galvan, S. Analysis of the gut microbiota in alopecia areata: Identification of bacterial biomarkers. J. Eur. Acad. Dermatol. Venereol. 2020, 34, 400–405. [Google Scholar] [CrossRef]
- Rebello, D.; Wang, E.; Yen, E.; Lio, P.A.; Kelly, C.R. Hair growth in two alopecia patients after fecal microbiota transplant. ACG Case Rep. J. 2017, 4, e107. [Google Scholar] [CrossRef]
- Chen, J.; Domingue, J.C.; Sears, C.L. Microbiota dysbiosis in select human cancers: Evidence of association and causality. In Seminars in Immunology; Elsevier: Amsterdam, The Netherlands, 2017; Volume 32, pp. 25–34. [Google Scholar]
- Guo, Y.; Liu, W.; Wu, J. Helicobacter pylori infection and pancreatic cancer risk: A meta-analysis. J. Cancer Res. Ther. 2016, 12, 229. [Google Scholar]
- Pichon, M.; Burucoa, C. Impact of the gastro-intestinal bacterial microbiome on Helicobacter-associated diseases. Healthcare 2019, 7, 34. [Google Scholar] [CrossRef] [PubMed]
- Silva, Y.P.; Bernardi, A.; Frozza, R.L. The role of short-chain fatty acids from gut microbiota in gut-brain communication. Front. Endocrinol. 2020, 11, 25. [Google Scholar] [CrossRef]
- Torki, M.; Gholamrezaei, A.; Mirbagher, L.; Danesh, M.; Kheiri, S.; Emami, M.H. Vitamin D deficiency associated with disease activity in patients with inflammatory bowel diseases. Dig. Dis. Sci. 2015, 60, 3085–3091. [Google Scholar] [CrossRef] [PubMed]
- Kammeyer, A.; Peters, C.P.; Meijer, S.L.; te Velde, A.A. Anti-inflammatory effects of urocanic acid derivatives in models ex vivo and in vivo of inflammatory bowel disease. ISRN Inflamm. 2012, 2012, 898153. [Google Scholar] [CrossRef]
- Strandwitz, P. Neurotransmitter modulation by the gut microbiota. Brain Res. 2018, 1693, 128–133. [Google Scholar] [CrossRef]
- Akiyama, T.; Carstens, M.I.; Carstens, E. Transmitters and pathways mediating inhibition of spinal itch-signaling neurons by scratching and other counterstimuli. PLoS ONE 2011, 6, e22665. [Google Scholar] [CrossRef] [PubMed]
- Langan, E.; Lisztes, E.; Bíró, T.; Funk, W.; Kloepper, J.; Griffiths, C.; Paus, R. Dopamine is a novel, direct inducer of catagen in human scalp hair follicles in vitro. Br. J. Dermatol. 2013, 168, 520–525. [Google Scholar] [CrossRef]
- Lee, H.; Park, M.; Kim, S.; Park Choo, H.; Lee, A.; Lee, C. Serotonin induces melanogenesis via serotonin receptor 2A. Br. J. Dermatol. 2011, 165, 1344–1348. [Google Scholar] [CrossRef]
- Yokoyama, S.; Hiramoto, K.; Koyama, M.; Ooi, K. Impairment of skin barrier function via cholinergic signal transduction in a dextran sulphate sodium-induced colitis mouse model. Exp. Dermatol. 2015, 24, 779–784. [Google Scholar] [CrossRef]
- Miyazaki, K.; Masuoka, N.; Kano, M.; Iizuka, R. Bifidobacterium fermented milk and galacto-oligosaccharides lead to improved skin health by decreasing phenols production by gut microbiota. Benef. Microbes 2014, 5, 121–128. [Google Scholar] [CrossRef]
- Rhodes, L.E.; Darby, G.; Massey, K.A.; Clarke, K.A.; Dew, T.P.; Farrar, M.D.; Bennett, S.; Watson, R.E.; Williamson, G.; Nicolaou, A. Oral green tea catechin metabolites are incorporated into human skin and protect against UV radiation-induced cutaneous inflammation in association with reduced production of pro-inflammatory eicosanoid 12-hydroxyeicosatetraenoic acid. Br. J. Nutr. 2013, 110, 891–900. [Google Scholar] [CrossRef]
- Giampieri, F.; Alvarez-Suarez, J.M.; Mazzoni, L.; Forbes-Hernandez, T.Y.; Gasparrini, M.; Gonzàlez-Paramàs, A.M.; Santos-Buelga, C.; Quiles, J.L.; Bompadre, S.; Mezzetti, B.; et al. Polyphenol-rich strawberry extract protects human dermal fibroblasts against hydrogen peroxide oxidative damage and improves mitochondrial functionality. Molecules 2014, 19, 7798–7816. [Google Scholar] [CrossRef]
- Gibson, G.R.; Hutkins, R.; Sanders, M.E.; Prescott, S.L.; Reimer, R.A.; Salminen, S.J.; Scott, K.; Stanton, C.; Swanson, K.S.; Cani, P.D.; et al. Expert consensus document: The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat. Rev. Gastroenterol. Hepatol. 2017, 14, 491. [Google Scholar] [CrossRef] [PubMed]
- Rizwan, M.; Rodriguez-Blanco, I.; Harbottle, A.; Birch-Machin, M.; Watson, R.; Rhodes, L. Tomato paste rich in lycopene protects against cutaneous photodamage in humans in vivo: A randomized controlled trial. Br. J. Dermatol. 2011, 164, 154–162. [Google Scholar] [CrossRef] [PubMed]
- Yoon, H.J.; Jang, M.S.; Kim, H.W.; Song, D.U.; Nam, K.I.; Bae, C.S.; Kim, S.J.; Lee, S.R.; Ku, C.S.; Jang, D.I.; et al. Protective effect of diet supplemented with rice prolamin extract against DNCB-induced atopic dermatitis in BALB/c mice. BMC Complement. Altern. Med. 2015, 15, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Tundis, R.; Loizzo, M.; Bonesi, M.; Menichini, F. Potential role of natural compounds against skin aging. Curr. Med. Chem. 2015, 22, 1515–1538. [Google Scholar] [CrossRef]
- Cordain, L.; Lindeberg, S.; Hurtado, M.; Hill, K.; Eaton, S.B.; Brand-Miller, J. Acne vulgaris: A disease of Western civilization. Arch. Dermatol. 2002, 138, 1584–1590. [Google Scholar] [CrossRef] [PubMed]
- Zouboulis, C.C.; Jourdan, E.; Picardo, M. Acne is an inflammatory disease and alterations of sebum composition initiate acne lesions. J. Eur. Acad. Dermatol. Venereol. JEADV 2014, 28, 527–532. [Google Scholar] [CrossRef] [PubMed]
- Agak, G.W.; Qin, M.; Nobe, J.; Kim, M.H.; Krutzik, S.R.; Tristan, G.R.; Elashoff, D.; Garbán, H.J.; Kim, J. Propionibacterium acnes induces an IL-17 response in acne vulgaris that is regulated by vitamin A and vitamin D. J. Investig. Dermatol. 2014, 134, 366–373. [Google Scholar] [CrossRef] [PubMed]
- Thiboutot, D.M.; Layton, A.M.; Eady, E.A. IL-17: A key player in the P. acnes inflammatory cascade? J. Investig. Dermatol. 2014, 134, 307–310. [Google Scholar] [CrossRef] [PubMed]
- Mattii, M.; Lovászi, M.; Garzorz, N.; Atenhan, A.; Quaranta, M.; Lauffer, F.; Konstantinow, A.; Küpper, M.; Zouboulis, C.; Kemeny, L.; et al. Sebocytes contribute to skin inflammation by promoting the differentiation of T helper 17 cells. Br. J. Dermatol. 2018, 178, 722–730. [Google Scholar] [CrossRef] [PubMed]
- Ben-Amitai, D.; Laron, Z. Effect of insulin-like growth factor-1 deficiency or administration on the occurrence of acne. J. Eur. Acad. Dermatol. Venereol. 2011, 25, 950–954. [Google Scholar] [CrossRef]
- Melnik, B.C.; Schmitz, G. Role of insulin, insulin-like growth factor-1, hyperglycaemic food and milk consumption in the pathogenesis of acne vulgaris. Exp. Dermatol. 2009, 18, 833–841. [Google Scholar] [CrossRef]
- Çerman, A.A.; Aktaş, E.; Altunay, İ.K.; Arıcı, J.E.; Tulunay, A.; Ozturk, F.Y. Dietary glycemic factors, insulin resistance, and adiponectin levels in acne vulgaris. J. Am. Acad. Dermatol. 2016, 75, 155–162. [Google Scholar] [CrossRef]
- Nast, A.; Dréno, B.; Bettoli, V.; Bukvic Mokos, Z.; Degitz, K.; Dressler, C.; Finlay, A.Y.; Haedersdal, M.; Lambert, J.; Layton, A.; et al. European evidence-based (S3) guideline for the treatment of acne–update 2016–short version. J. Eur. Acad. Dermatol. Venereol. 2016, 30, 1261–1268. [Google Scholar] [CrossRef] [PubMed]
- Iinuma, K.; Sato, T.; Akimoto, N.; Noguchi, N.; Sasatsu, M.; Nishijima, S.; Kurokawa, I.; Ito, A. Involvement of Propionibacterium acnes in the augmentation of lipogenesis in hamster sebaceous glands in vivo and in vitro. J. Investig. Dermatol. 2009, 129, 2113–2119. [Google Scholar] [CrossRef]
- Yu, Y.; Champer, J.; Agak, G.W.; Kao, S.; Modlin, R.L.; Kim, J. Different Propionibacterium acnes phylotypes induce distinct immune responses and express unique surface and secreted proteomes. J. Investig. Dermatol. 2016, 136, 2221–2228. [Google Scholar] [CrossRef] [PubMed]
- Dagnelie, M.A.; Corvec, S.; Saint-Jean, M.; Bourdès, V.; Nguyen, J.M.; Khammari, A.; Dréno, B. Decrease in diversity of Propionibacterium acnes phylotypes in patients with severe acne on the back. Acta Derm. Venereol. 2018, 98, 262–267. [Google Scholar] [CrossRef] [PubMed]
- Borelli, C.; Merk, K.; Schaller, M.; Jacob, K.; Vogeser, M.; Weindl, G.; Berger, U.; Plewig, G. In vivo porphyrin production by P. acnes in untreated acne patients and its modulation by acne treatment. Acta Derm. Venereol. 2006, 86, 316–319. [Google Scholar] [CrossRef] [PubMed]
- Kasimatis, G.; Fitz-Gibbon, S.; Tomida, S.; Wong, M.; Li, H. Analysis of complete genomes of Propionibacterium acnes reveals a novel plasmid and increased pseudogenes in an acne associated strain. BioMed. Res. Int. 2013, 2013. [Google Scholar] [CrossRef] [PubMed]
- Brüggemann, H.; Lomholt, H.B.; Tettelin, H.; Kilian, M. CRISPR/cas loci of type II Propionibacterium acnes confer immunity against acquisition of mobile elements present in type I P. acnes. PLoS ONE 2012, 7, e34171. [Google Scholar] [CrossRef]
- Sanford, J.A.; O’Neill, A.M.; Zouboulis, C.C.; Gallo, R.L. Short-chain fatty acids from Cutibacterium acnes activate both a canonical and epigenetic inflammatory response in human sebocytes. J. Immunol. 2019, 202, 1767–1776. [Google Scholar] [CrossRef] [PubMed]
- Brüggemann, H. Skin: Cutibacterium (formerly Propionibacterium) acnes and Acne Vulgaris. In Health Consequences of Microbial Interactions with Hydrocarbons, Oils, and Lipids. Handbook of Hydrocarbon and Lipid Microbiology; Springer: Cham, Switzerland, 2020; pp. 225–243. [Google Scholar]
- O’Neill, A.M.; Gallo, R.L. Host-microbiome interactions and recent progress into understanding the biology of acne vulgaris. Microbiome 2018, 6, 177. [Google Scholar] [CrossRef] [PubMed]
- Andersson, T.; Bergdahl, G.E.; Saleh, K.; Magnúsdóttir, H.; Stødkilde, K.; Andersen, C.B.F.; Lundqvist, K.; Jensen, A.; Brüggemann, H.; Lood, R. Common skin bacteria protect their host from oxidative stress through secreted antioxidant RoxP. Sci. Rep. 2019, 9, 1–10. [Google Scholar] [CrossRef]
- Dréno, B.; Pécastaings, S.; Corvec, S.; Veraldi, S.; Khammari, A.; Roques, C. Cutibacterium acnes (Propionibacterium acnes) and acne vulgaris: A brief look at the latest updates. J. Eur. Acad. Dermatol. Venereol. 2018, 32, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Callewaert, C.; Knödlseder, N.; Karoglan, A.; Güell, M.; Paetzold, B. Skin microbiome transplantation and manipulation: Current state of the art. Comput. Struct. Biotechnol. J. 2021, 19, 624–631. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Wang, H.; Zhou, J.; Mou, Y.; Wang, G.; Xiong, X. Patients with acne vulgaris have a distinct gut microbiota in comparison with healthy controls. Acta Derm. Venereol. 2018, 98, 783–790. [Google Scholar] [CrossRef]
- Fabbrocini, G.; Bertona, M.; Picazo, O.; Pareja-Galeano, H.; Monfrecola, G.; Emanuele, E. Supplementation with Lactobacillus rhamnosus SP1 normalises skin expression of genes implicated in insulin signalling and improves adult acne. Benef. Microbes 2016, 7, 625–630. [Google Scholar] [CrossRef]
- Kim, J.; Ko, Y.; Park, Y.K.; Kim, N.I.; Ha, W.K.; Cho, Y. Dietary effect of lactoferrin-enriched fermented milk on skin surface lipid and clinical improvement of acne vulgaris. Nutrition 2010, 26, 902–909. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. Linking diet to acne metabolomics, inflammation, and comedogenesis: An update. Clin. Cosmetic Investig. dermatol. 2015, 8, 371–388. [Google Scholar] [CrossRef]
- Bieber, T. Mechanisms of disease. N. Engl. J. Med. 2008, 358, 1483–1494. [Google Scholar] [CrossRef]
- Williams, H.C. Epidemiology of atopic dermatitis. Clin. Exp. Dermatol. 2000, 25, 522–529. [Google Scholar] [CrossRef] [PubMed]
- Brunner, P.M.; Guttman-Yassky, E.; Leung, D.Y. The immunology of atopic dermatitis and its reversibility with broad-spectrum and targeted therapies. J. Allergy Clin. Immunol. 2017, 139, S65–S76. [Google Scholar] [CrossRef] [PubMed]
- Leung, D.Y.; Guttman-Yassky, E. Deciphering the complexities of atopic dermatitis: Shifting paradigms in treatment approaches. J. Allergy Clin. Immunol. 2014, 134, 769–779. [Google Scholar] [CrossRef]
- Cho, S.H.; Strickland, I.; Tomkinson, A.; Fehringer, A.P.; Gelfand, E.W.; Leung, D.Y. Preferential binding of Staphylococcus aureus to skin sites of Th2-mediated inflammation in a murine model. J. Investig. Dermatol. 2001, 116, 658–663. [Google Scholar] [CrossRef] [PubMed]
- Morar, N.; Cookson, W.O.; Harper, J.I.; Moffatt, M.F. Filaggrin mutations in children with severe atopic dermatitis. J. Investig. Dermatol. 2007, 127, 1667–1672. [Google Scholar] [CrossRef]
- Tanei, R. Atopic Dermatitis in Older Adults: A Review of Treatment Options. In Drugs & Aging; Springer: Cham, Switzerland, 2020; pp. 1–12. [Google Scholar]
- Paller, A.S.; Kabashima, K.; Bieber, T. Therapeutic pipeline for atopic dermatitis: End of the drought? J. Allergy Clin. Immunol. 2017, 140, 633–643. [Google Scholar] [CrossRef]
- Guttman-Yassky, E.; Pavel, A.B.; Zhou, L.; Estrada, Y.D.; Zhang, N.; Xu, H.; Peng, X.; Wen, H.C.; Govas, P.; Gudi, G.; et al. GBR 830, an anti-OX40, improves skin gene signatures and clinical scores in patients with atopic dermatitis. J. Allergy Clin. Immunol. 2019, 144, 482–493. [Google Scholar] [CrossRef]
- Shirley, M. Dupilumab: First global approval. Drugs 2017, 77, 1115–1121. [Google Scholar] [CrossRef]
- Bjerre, R.; Bandier, J.; Skov, L.; Engstrand, L.; Johansen, J. The role of the skin microbiome in atopic dermatitis: A systematic review. Br. J. Dermatol. 2017, 177, 1272–1278. [Google Scholar] [CrossRef]
- Leyden, J.J.; Marples, R.R.; Kligman, A.M. Staphylococcus aureus in the lesions of atopic dermatitis. Br. J. Dermatol. 1974, 90, 525. [Google Scholar] [CrossRef]
- Callewaert, C.; Nakatsuji, T.; Knight, R.; Kosciolek, T.; Vrbanac, A.; Kotol, P.; Ardeleanu, M.; Hultsch, T.; Guttman-Yassky, E.; Bissonnette, R.; et al. IL-4Rα blockade by dupilumab decreases Staphylococcus aureus colonization and increases microbial diversity in atopic dermatitis. J. Investig. Dermatol. 2020, 140, 191–202. [Google Scholar] [CrossRef]
- Matsui, K.; Nishikawa, A. Peptidoglycan-induced T helper 2 immune response in mice involves interleukin-10 secretion from Langerhans cells. Microbiol. Immunol. 2013, 57, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Ruíz-González, V.; Cancino-Diaz, J.C.; Rodríguez-Martínez, S.; Cancino-Diaz, M.E. Keratinocytes treated with peptidoglycan from Staphylococcus aureus produce vascular endothelial growth factor, and its expression is amplified by the subsequent production of interleukin-13. Int. J. Dermatol. 2009, 48, 846–854. [Google Scholar] [CrossRef]
- Travers, J.B. Toxic interaction between Th2 cytokines and Staphylococcus aureus in atopic dermatitis. J. Investig. Dermatol. 2014, 134, 2069–2071. [Google Scholar] [CrossRef] [PubMed]
- Wollenberg, A.; Zoch, C.; Wetzel, S.; Plewig, G.; Przybilla, B. Predisposing factors and clinical features of eczema herpeticum: A retrospective analysis of 100 cases. J. Am. Acad. Dermatol. 2003, 49, 198–205. [Google Scholar] [CrossRef]
- Mathes, E.F.; Oza, V.; Frieden, I.J.; Cordoro, K.M.; Yagi, S.; Howard, R.; Kristal, L.; Ginocchio, C.C.; Schaffer, J.; Maguiness, S.; et al. “Eczema coxsackium” and unusual cutaneous findings in an enterovirus outbreak. Pediatrics 2013, 132, e149–e157. [Google Scholar] [CrossRef]
- Eichenfield, L.F.; Tom, W.L.; Berger, T.G.; Krol, A.; Paller, A.S.; Schwarzenberger, K.; Bergman, J.N.; Chamlin, S.L.; Cohen, D.E.; Cooper, K.D.; et al. Guidelines of care for the management of atopic dermatitis: Section 2. Management and treatment of atopic dermatitis with topical therapies. J. Am. Acad. Dermatol. 2014, 71, 116–132. [Google Scholar] [CrossRef]
- Nakatsuji, T.; Chen, T.H.; Narala, S.; Chun, K.A.; Two, A.M.; Yun, T.; Shafiq, F.; Kotol, P.F.; Bouslimani, A.; Melnik, A.V.; et al. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci. Transl. Med. 2017, 9, eaah4680. [Google Scholar] [CrossRef]
- Nylund, L.; Nermes, M.; Isolauri, E.; Salminen, S.; De Vos, W.; Satokari, R. Severity of atopic disease inversely correlates with intestinal microbiota diversity and butyrate-producing bacteria. Allergy 2015, 70, 241–244. [Google Scholar] [CrossRef]
- Rather, I.A.; Bajpai, V.K.; Kumar, S.; Lim, J.; Paek, W.K.; Park, Y.H. Probiotics and atopic dermatitis: An overview. Front. Microbiol. 2016, 7, 507. [Google Scholar] [CrossRef] [PubMed]
- West, C.E.; Dzidic, M.; Prescott, S.L.; Jenmalm, M.C. Bugging allergy; role of pre-, pro-and synbiotics in allergy prevention. Allergol. Int. 2017, 66, 529–538. [Google Scholar] [CrossRef]
- Piqué, N.; Berlanga, M.; Miñana-Galbis, D. Health benefits of heat-killed (Tyndallized) probiotics: An overview. Int. J. Mol. Sci. 2019, 20, 2534. [Google Scholar] [CrossRef]
- Chang, Y.S.; Trivedi, M.K.; Jha, A.; Lin, Y.F.; Dimaano, L.; Garcia-Romero, M.T. Synbiotics for prevention and treatment of atopic dermatitis: A meta-analysis of randomized clinical trials. JAMA Pediatr. 2016, 170, 236–242. [Google Scholar] [CrossRef]
- Zuccotti, G.; Meneghin, F.; Aceti, A.; Barone, G.; Callegari, M.L.; Di Mauro, A.; Fantini, M.; Gori, D.; Indrio, F.; Maggio, L.; et al. Probiotics for prevention of atopic diseases in infants: Systematic review and meta-analysis. Allergy 2015, 70, 1356–1371. [Google Scholar] [CrossRef]
- Iemoli, E.; Trabattoni, D.; Parisotto, S.; Borgonovo, L.; Toscano, M.; Rizzardini, G.; Clerici, M.; Ricci, E.; Fusi, A.; De Vecchi, E.; et al. Probiotics reduce gut microbial translocation and improve adult atopic dermatitis. J. Clin. Gastroenterol. 2012, 46, S33–S40. [Google Scholar] [CrossRef] [PubMed]
- Kim, N.Y.; Ji, G.E. Effects of probiotics on the prevention of atopic dermatitis. Korean J. Pediatr. 2012, 55, 193. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.E.; Kim, H.S. Microbiome of the skin and gut in atopic dermatitis (AD): Understanding the pathophysiology and finding novel management strategies. J. Clin. Med. 2019, 8, 444. [Google Scholar] [CrossRef]
- Black, P.; Sharpe, S. Dietary fat and asthma: Is there a connection? Eur. Respir. J. 1997, 10, 6–12. [Google Scholar] [CrossRef] [PubMed]
- Devereux, G.; Seaton, A. Diet as a risk factor for atopy and asthma. J. Allergy Clin. Immunol. 2005, 115, 1109–1117. [Google Scholar] [CrossRef] [PubMed]
- Mabin, D.; Sykes, A.; David, T. Controlled trial of a few foods diet in severe atopic dermatitis. Arch. Dis. Child. 1995, 73, 202–207. [Google Scholar] [CrossRef]
- Caputo, V.; Strafella, C.; Termine, A.; Dattola, A.; Mazzilli, S.; Lanna, C.; Cosio, T.; Campione, E.; Novelli, G.; Giardina, E.; et al. Overview of the molecular determinants contributing to the expression of Psoriasis and Psoriatic Arthritis phenotypes. J. Cell. Mol. Med. 2020, 24, 13554–13563. [Google Scholar] [CrossRef] [PubMed]
- Nestle, F.; Kaplan, D.; Barker, J. Mechanisms of Disease: Psoriasis. N. Engl. J. Med. 2009, 361, 496–509. [Google Scholar] [CrossRef]
- Grine, L.; Lambert, J. Psoriasis: Burning Down the Host. 2016. Available online: https://www.tandfonline.com/doi/pdf/10.3109/09546634.2015.1117567 (accessed on 15 November 2020).
- Li, Q.; Chandran, V.; Tsoi, L.; O’Rielly, D.; Nair, R.P.; Gladman, D.; Elder, J.T.; Rahman, P. Quantifying differences in heritability among psoriatic arthritis (PsA), cutaneous psoriasis (PsC) and psoriasis vulgaris (PsV). Sci. Rep. 2020, 10, 1–6. [Google Scholar] [CrossRef]
- Dand, N.; Mahil, S.K.; Capon, F.; Smith, C.H.; Simpson, M.A.; Barker, J.N. Psoriasis and genetics. Acta Derm Venereol 2020, 100, adv00030. [Google Scholar] [CrossRef]
- Ovejero-Benito, M.C.; Muñoz-Aceituno, E.; Sabador, D.; Almoguera, B.; Prieto-Pérez, R.; Hakonarson, H.; Coto-Segura, P.; Carretero, G.; Reolid, A.; Llamas-Velasco, M.; et al. Genome-wide association analysis of psoriasis patients treated with anti-TNF drugs. Exp. Dermatol. 2020, 29, 1225–1232. [Google Scholar] [CrossRef] [PubMed]
- Grozdev, I.; Korman, N.; Tsankov, N. Psoriasis as a systemic disease. Clin. Dermatol. 2014, 32, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Takeshita, J.; Grewal, S.; Langan, S.M.; Mehta, N.N.; Ogdie, A.; Van Voorhees, A.S.; Gelfand, J.M. Psoriasis and comorbid diseases: Implications for management. J. Am. Acad. Dermatol. 2017, 76, 393–403. [Google Scholar] [CrossRef] [PubMed]
- Remröd, C.; Sjöström, K.; Svensson, Å. Subjective stress reactivity in psoriasis–a cross sectional study of associated psychological traits. BMC Dermatol. 2015, 15, 6. [Google Scholar] [CrossRef]
- Peters, E.M. Stressed skin?–a molecular psychosomatic update on stress-causes and effects in dermatologic diseases. J. Der Dtsch. Dermatol. Ges. 2016, 14, 233–252. [Google Scholar] [CrossRef] [PubMed]
- Grine, L.; Dejager, L.; Libert, C.; Vandenbroucke, R.E. An inflammatory triangle in psoriasis: TNF, type I IFNs and IL-17. Cytokine Growth Factor Rev. 2015, 26, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Song, Y.; Zhu, L.; Wang, X.; Yang, B.; Lu, P.; Chen, Q.; Bin, L.; Deng, L. Interferon Kappa Is Up-Regulated in Psoriasis and It Up-Regulates Psoriasis-Associated Cytokines in vivo. Clin. Cosmet. Investig. Dermatol. 2019, 12, 865. [Google Scholar] [CrossRef]
- Zhang, L.j. Type1 interferons: Potential initiating factors linking skin wounds with psoriasis pathogenesis. Front. Immunol. 2019, 10, 1440. [Google Scholar] [CrossRef] [PubMed]
- Conrad, C.; Di Domizio, J.; Mylonas, A.; Belkhodja, C.; Demaria, O.; Navarini, A.A.; Lapointe, A.K.; French, L.E.; Vernez, M.; Gilliet, M. TNF blockade induces a dysregulated type I interferon response without autoimmunity in paradoxical psoriasis. Nat. Commun. 2018, 9, 1–11. [Google Scholar] [CrossRef]
- Frasca, L.; Palazzo, R.; Chimenti, M.S.; Alivernini, S.; Tolusso, B.; Bui, L.; Botti, E.; Giunta, A.; Bianchi, L.; Petricca, L.; et al. Anti-LL37 antibodies are present in psoriatic arthritis (PsA) patients: New biomarkers in PsA. Front. Immunol. 2018, 9, 1936. [Google Scholar] [CrossRef]
- Kong, H.H.; Andersson, B.; Clavel, T.; Common, J.E.; Jackson, S.A.; Olson, N.D.; Segre, J.A.; Traidl-Hoffmann, C. Performing skin microbiome research: A method to the madness. J. Investig. Dermatol. 2017, 137, 561–568. [Google Scholar] [CrossRef] [PubMed]
- Thorleifsdottir, R.H.; Sigurdardottir, S.L.; Sigurgeirsson, B.; Olafsson, J.H.; Sigurdsson, M.I.; Petersen, H.; Gudjonsson, J.E.; Johnston, A.; Valdimarsson, H. Patient-reported outcomes and clinical response in patients with moderate-to-severe plaque psoriasis treated with tonsillectomy: A randomized controlled trial. Acta Derm. Venereol. 2017, 97, 340–345. [Google Scholar] [CrossRef] [PubMed]
- Cohn, J.E.; Pfeiffer, M.; Vernose, G. Complete resolution of guttate psoriasis after tonsillectomy. Ear, Nose Throat J. 2018, 97, 62–63. [Google Scholar] [CrossRef]
- Haapasalo, K.; Koskinen, L.L.; Suvilehto, J.; Jousilahti, P.; Wolin, A.; Suomela, S.; Trembath, R.; Barker, J.; Vuopio, J.; Kere, J.; et al. The psoriasis risk allele HLA-C* 06: 02 shows evidence of association with chronic or recurrent streptococcal tonsillitis. Infect. Immun. 2018, 86. [Google Scholar] [CrossRef]
- Assarsson, M.; Duvetorp, A.; Dienus, O.; Söderman, J.; Seifert, O. Significant changes in the skin microbiome in patients with chronic plaque psoriasis after treatment with narrowband ultraviolet B. Acta Derm. Venereol. 2018, 98, 428–436. [Google Scholar] [CrossRef]
- Langan, E.; Künstner, A.; Miodovnik, M.; Zillikens, D.; Thaçi, D.; Baines, J.F.; Ibrahim, S.; Solbach, W.; Knobloch, J. Combined culture and metagenomic analyses reveal significant shifts in the composition of the cutaneous microbiome in psoriasis. Br. J. Dermatol. 2019, 181, 1254–1264. [Google Scholar] [CrossRef] [PubMed]
- Paniz Mondolfi, A.; Hernandez Perez, M.; Blohm, G.; Marquez, M.; Mogollon Mendoza, A.; Hernandez-Pereira, C.; Escalona, M.; Lodeiro Colatosti, A.; Rothe DeArocha, J.; Rodriguez Morales, A. Generalized pustular psoriasis triggered by Zika virus infection. Clin. Exp. Dermatol. 2018, 43, 171–174. [Google Scholar] [CrossRef]
- Sbidian, E.; Madrange, M.; Viguier, M.; Salmona, M.; Duchatelet, S.; Hovnanian, A.; Smahi, A.; Le Goff, J.; Bachelez, H. Respiratory virus infection triggers acute psoriasis flares across different clinical subtypes and genetic backgrounds. Br. J. Dermatol. 2019, 181, 1304–1306. [Google Scholar] [CrossRef]
- Sanchez, I.M.; Jiang, W.; Yang, E.J.; Singh, R.K.; Beck, K.; Liu, C.; Afifi, L.; Liao, W. Enteropathy in psoriasis: A systematic review of gastrointestinal disease epidemiology and subclinical inflammatory and functional gut alterations. Curr. Dermatol. Rep. 2018, 7, 59–74. [Google Scholar] [CrossRef]
- Hueber, W.; Sands, B.E.; Lewitzky, S.; Vandemeulebroecke, M.; Reinisch, W.; Higgins, P.D.; Wehkamp, J.; Feagan, B.G.; Yao, M.D.; Karczewski, M.; et al. Secukinumab, a human anti-IL-17A monoclonal antibody, for moderate to severe Crohn’s disease: Unexpected results of a randomised, double-blind placebo-controlled trial. Gut 2012, 61, 1693–1700. [Google Scholar] [CrossRef]
- Fobelo Lozano, M.J.; Serrano Giménez, R.; Castro Fernández, M. Emergence of inflammatory bowel disease during treatment with secukinumab. J. Crohn’s Colitis 2018, 12, 1131–1133. [Google Scholar] [CrossRef] [PubMed]
- Barry, R.; Salmon, P.; Read, A.; Warin, R. Mucosal architecture of the small bowel in cases of psoriasis. Gut 1971, 12, 873–877. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Humbert, P.; Bidet, A.; Treffel, P.; Drobacheff, C.; Agache, P. Intestinal permeability in patients with psoriasis. J. Dermatol. Sci. 1991, 2, 324–326. [Google Scholar] [CrossRef]
- Sikora, M.; Stec, A.; Chrabaszcz, M.; Waskiel-Burnat, A.; Zaremba, M.; Olszewska, M.; Rudnicka, L. Intestinal fatty acid binding protein, a biomarker of intestinal barrier, is associated with severity of psoriasis. J. Clin. Med. 2019, 8, 1021. [Google Scholar] [CrossRef]
- Adarsh, M.; Dogra, S.; Vaiphei, K.; Vaishnavi, C.; Sinha, S.; Sharma, A. Evaluation of subclinical gut inflammation using faecal calprotectin levels and colonic mucosal biopsy in patients with psoriasis and psoriatic arthritis. Br. J. Dermatol. 2019, 181, 401–402. [Google Scholar] [CrossRef] [PubMed]
- Munz, O.H.; Sela, S.; Baker, B.S.; Griffiths, C.E.; Powles, A.V.; Fry, L. Evidence for the presence of bacteria in the blood of psoriasis patients. Arch. Dermatol. Res. 2010, 302, 495–498. [Google Scholar] [CrossRef] [PubMed]
- Ramírez-Boscá, A.; Navarro-López, V.; Martínez-Andrés, A.; Such, J.; Francés, R.; de la Parte, J.H.; Asín-Llorca, M. Identification of bacterial DNA in the peripheral blood of patients with active psoriasis. JAMA Dermatol. 2015, 151, 670–671. [Google Scholar] [CrossRef]
- Kim, M.; Han, K.D.; Lee, J.H. Bodyweight variability and the risk of psoriasis: A nationwide population-based cohort study. J. Eur. Acad. Dermatol. Venereol. 2020, 34, 1019–1025. [Google Scholar] [CrossRef]
- Jensen, P.; Christensen, R.; Zachariae, C.; Geiker, N.R.; Schaadt, B.K.; Stender, S.; Hansen, P.R.; Astrup, A.; Skov, L. Long-term effects of weight reduction on the severity of psoriasis in a cohort derived from a randomized trial: A prospective observational follow-up study. Am. J. Clin. Nutr. 2016, 104, 259–265. [Google Scholar] [CrossRef]
- Nakamizo, S.; Honda, T.; Adachi, A.; Nagatake, T.; Kunisawa, J.; Kitoh, A.; Otsuka, A.; Dainichi, T.; Nomura, T.; Ginhoux, F.; et al. High fat diet exacerbates murine psoriatic dermatitis by increasing the number of IL-17-producing γδ T cells. Sci. Rep. 2017, 7, 1–13. [Google Scholar]
- Herbert, D.; Franz, S.; Popkova, Y.; Anderegg, U.; Schiller, J.; Schwede, K.; Lorz, A.; Simon, J.C.; Saalbach, A. High-fat diet exacerbates early psoriatic skin inflammation independent of obesity: Saturated fatty acids as key players. J. Investig. Dermatol. 2018, 138, 1999–2009. [Google Scholar] [CrossRef] [PubMed]
- Castaldo, G.; Galdo, G.; Aufiero, F.R.; Cereda, E. Very low-calorie ketogenic diet may allow restoring response to systemic therapy in relapsing plaque psoriasis. Obes. Res. Clin. Pract. 2016, 10, 348–352. [Google Scholar] [CrossRef]
- Damiani, G.; Watad, A.; Bridgewood, C.; Pigatto, P.D.M.; Pacifico, A.; Malagoli, P.; Bragazzi, N.L.; Adawi, M. The impact of ramadan fasting on the reduction of PASI score, in moderate-to-severe psoriatic patients: A real-life multicenter study. Nutrients 2019, 11, 277. [Google Scholar] [CrossRef]
- Codoñer, F.M.; Ramírez-Bosca, A.; Climent, E.; Carrión-Gutierrez, M.; Guerrero, M.; Pérez-Orquín, J.M.; De La Parte, J.H.; Genovés, S.; Ramón, D.; Navarro-López, V.; et al. Gut microbial composition in patients with psoriasis. Sci. Rep. 2018, 8, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Navarro-López, V.; Martínez-Andrés, A.; Ramírez-Boscá, A.; Ruzafa-Costas, B.; Núñez-Delegido, E.; Carrión-Gutiérrez, M.A.; Prieto-Merino, D.; Codoñer-Cortés, F.; Ramón-Vidal, D.; Genovés-Martínez, S.; et al. Efficacy and safety of oral administration of a mixture of probiotic strains in patients with psoriasis: A randomized controlled clinical trial. Acta Derm. Venereol. 2019, 99, 1078–1084. [Google Scholar] [CrossRef]
- Itano, A.; Cormack, T.; Ramani, K.; Barth, K.; Wang, I.; Mukherjee, A.; Ponichtera, H.; McKenna, C.; Jahic, M.; Bodmer, M. Orally-administered EDP1815, a single strain of Prevotella histicola, has potent systemic anti-inflammatory effects in Type 1, Type 2, and Type 3 inflammatory models. In Proceedings of the 29th European Academy of Dermatology and Venereology Congress (EADV), Vienna, Austria, 29–31 October 2020. [Google Scholar]
- Phan, K.; Charlton, O.; Smith, S.D. Global prevalence of hidradenitis suppurativa and geographical variation—systematic review and meta-analysis. Biomed. Dermatol. 2020, 4, 1–6. [Google Scholar] [CrossRef]
- Jørgensen, A.H.R.; Thomsen, S.F.; Karmisholt, K.E.; Ring, H.C. Clinical, microbiological, immunological and imaging characteristics of tunnels and fistulas in hidradenitis suppurativa and Crohn’s disease. Exp. Dermatol. 2020, 29, 118–123. [Google Scholar] [CrossRef]
- Moran, B.; Sweeney, C.M.; Hughes, R.; Malara, A.; Kirthi, S.; Tobin, A.M.; Kirby, B.; Fletcher, J.M. Hidradenitis suppurativa is characterized by dysregulation of the Th17: Treg cell axis, which is corrected by anti-TNF therapy. J. Investig. Dermatol. 2017, 137, 2389–2395. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.T.; Chi, C.C. Association of hidradenitis suppurativa with inflammatory bowel disease: A systematic review and meta-analysis. JAMA Dermatol. 2019, 155, 1022–1027. [Google Scholar] [CrossRef]
- Giudici, F.; Maggi, L.; Santi, R.; Cosmi, L.; Scaletti, C.; Annunziato, F.; Nesi, G.; Barra, G.; Bassotti, G.; De Palma, R.; et al. Perianal Crohn’s disease and hidradenitis suppurativa: A possible common immunological scenario. Clin. Mol. Allergy 2015, 13, 12. [Google Scholar] [CrossRef]
- Barta, Z.; Zöld, É.; Csípõ, I.; Zeher, M. ASCAs in (auto-) Immune Small Bowel Diseases. 2020. Available online: https://gut.bmj.com/content/ascas-auto-immune-small-bowel-diseases (accessed on 20 November 2020).
- Denny, G.; Anadkat, M.J. The effect of smoking and age on the response to first-line therapy of hidradenitis suppurativa: An institutional retrospective cohort study. J. Am. Acad. Dermatol. 2017, 76, 54–59. [Google Scholar] [CrossRef]
- König, A.; Lehmann, C.; Rompel, R.; Happle, R. Cigarette smoking as a triggering factor of hidradenitis suppurativa. Dermatology 1999, 198, 261–264. [Google Scholar] [CrossRef]
- Kromann, C.B.; Ibler, K.S.; Kristiansen, V.B.; Jemec, G.B. The influence of body weight on the prevalence and severity of hidradenitis suppurativa. Acta Derm. Venereol. 2014, 94, 553–557. [Google Scholar] [CrossRef] [PubMed]
- Aboud, C.; Zamaria, N.; Cannistrà, C. Treatment of hidradenitis suppurativa: Surgery and yeast (Saccharomyces cerevisiae)–exclusion diet. Results after 6 years. Surgery 2020, 167, 1012–1015. [Google Scholar] [CrossRef] [PubMed]
- Silfvast-Kaiser, A.; Youssef, R.; Paek, S.Y. Diet in hidradenitis suppurativa: A review of published and lay literature. Int. J. Dermatol. 2019, 58, 1225–1230. [Google Scholar] [CrossRef]
- Buechner, S.A. Rosacea: An update. Dermatology 2005, 210, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Rainer, B.M.; Fischer, A.H.; Da Silva, D.L.F.; Kang, S.; Chien, A.L. Rosacea is associated with chronic systemic diseases in a skin severity–dependent manner: Results of a case-control study. J. Am. Acad. Dermatol. 2015, 73, 604–608. [Google Scholar] [CrossRef] [PubMed]
- Tan, J.; Berg, M. Rosacea: Current state of epidemiology. J. Am. Acad. Dermatol. 2013, 69, S27–S35. [Google Scholar] [CrossRef] [PubMed]
- Duman, N.; Ersoy Evans, S.; Atakan, N. Rosacea and cardiovascular risk factors: A case control study. J. Eur. Acad. Dermatol. Venereol. 2014, 28, 1165–1169. [Google Scholar] [CrossRef]
- Yamasaki, K.; Di Nardo, A.; Bardan, A.; Murakami, M.; Ohtake, T.; Coda, A.; Dorschner, R.A.; Bonnart, C.; Descargues, P.; Hovnanian, A.; et al. Increased serine protease activity and cathelicidin promotes skin inflammation in rosacea. Nat. Med. 2007, 13, 975–980. [Google Scholar] [CrossRef]
- Nizet, V.; Ohtake, T.; Lauth, X.; Trowbridge, J.; Rudisill, J.; Dorschner, R.A.; Pestonjamasp, V.; Piraino, J.; Huttner, K.; Gallo, R.L. Innate antimicrobial peptide protects the skin from invasive bacterial infection. Nature 2001, 414, 454–457. [Google Scholar] [CrossRef] [PubMed]
- Two, A.M.; Wu, W.; Gallo, R.L.; Hata, T.R. Rosacea: Part I. Introduction, categorization, histology, pathogenesis, and risk factors. J. Am. Acad. Dermatol. 2015, 72, 749–758. [Google Scholar] [CrossRef]
- Gallo, R.L.; Ono, M.; Povsic, T.; Page, C.; Eriksson, E.; Klagsbrun, M.; Bernfield, M. Syndecans, cell surface heparan sulfate proteoglycans, are induced by a proline-rich antimicrobial peptide from wounds. Proc. Natl. Acad. Sci. USA 1994, 91, 11035–11039. [Google Scholar] [CrossRef]
- Yang, D.; Chen, Q.; Schmidt, A.P.; Anderson, G.M.; Wang, J.M.; Wooters, J.; Oppenheim, J.J.; Chertov, O. LL-37, the neutrophil granule–and epithelial cell–derived cathelicidin, utilizes formyl peptide receptor–like 1 (FPRL1) as a receptor to chemoattract human peripheral blood neutrophils, monocytes, and T cells. J. Exp. Med. 2000, 192, 1069–1074. [Google Scholar] [CrossRef] [PubMed]
- Koczulla, R.; Von Degenfeld, G.; Kupatt, C.; Krötz, F.; Zahler, S.; Gloe, T.; Issbrücker, K.; Unterberger, P.; Zaiou, M.; Lebherz, C.; et al. An angiogenic role for the human peptide antibiotic LL-37/hCAP-18. J. Clin. Investig. 2003, 111, 1665–1672. [Google Scholar] [CrossRef]
- Sulk, M.; Seeliger, S.; Aubert, J.; Schwab, V.D.; Cevikbas, F.; Rivier, M.; Nowak, P.; Voegel, J.J.; Buddenkotte, J.; Steinhoff, M. Distribution and expression of non-neuronal transient receptor potential (TRPV) ion channels in rosacea. J. Investig. Dermatol. 2012, 132, 1253–1262. [Google Scholar] [CrossRef]
- Ferrer, L.; Ravera, I.; Silbermayr, K. Immunology and pathogenesis of canine demodicosis. Vet. Dermatol. 2014, 25, 427–e65. [Google Scholar] [CrossRef] [PubMed]
- Kocak, M.; Yagli, S.; Vahapoğlu, G.; Ekşioğlu, M. Permethrin 5% cream versus metronidazole 0.75% gel for the treatment of papulopustular rosacea. Dermatology 2002, 205, 265–270. [Google Scholar] [CrossRef]
- O’Reilly, N.; Menezes, N.; Kavanagh, K. Positive correlation between serum immunoreactivity to Demodex-associated Bacillus proteins and erythematotelangiectatic rosacea. Br. J. Dermatol. 2012, 167, 1032–1036. [Google Scholar] [CrossRef]
- Yamasaki, K.; Gallo, R.L. The molecular pathology of rosacea. J. Dermatol. Sci. 2009, 55, 77–81. [Google Scholar] [CrossRef]
- Woo, Y.R.; Lee, S.H.; Cho, S.H.; Lee, J.D.; Kim, H.S. Characterization and Analysis of the Skin Microbiota in Rosacea: Impact of Systemic Antibiotics. J. Clin. Med. 2020, 9, 185. [Google Scholar] [CrossRef]
- Eriksson, G.; Nord, C. Impact of topical metronidazole on the skin and colon microflora in patients with rosacea. Infection 1987, 15, 8–10. [Google Scholar] [CrossRef] [PubMed]
- Zaidi, A.K.; Spaunhurst, K.; Sprockett, D.; Thomason, Y.; Mann, M.W.; Fu, P.; Ammons, C.; Gerstenblith, M.; Tuttle, M.S.; Popkin, D.L. Characterization of the facial microbiome in twins discordant for rosacea. Exp. Dermatol. 2018, 27, 295–298. [Google Scholar] [CrossRef] [PubMed]
- Weiss, E.; Katta, R. Diet and rosacea: The role of dietary change in the management of rosacea. Dermatol. Pract. Concept. 2017, 7, 31. [Google Scholar] [CrossRef] [PubMed]
- Egeberg, A.; Weinstock, L.; Thyssen, E.; Gislason, G.; Thyssen, J. Rosacea and gastrointestinal disorders: A population-based cohort study. Br. J. Dermatol. 2017, 176, 100–106. [Google Scholar] [CrossRef]
- Chen, Y.J.; Lee, W.H.; Ho, H.J.; Tseng, C.H.; Wu, C.Y. An altered fecal microbial profiling in rosacea patients compared to matched controls. J. Formos. Med. Assoc. 2020, 120, 256–264. [Google Scholar] [CrossRef]
- Gupta, V.K.; Paul, S.; Dutta, C. Geography, ethnicity or subsistence-specific variations in human microbiome composition and diversity. Front. Microbiol. 2017, 8, 1162. [Google Scholar] [CrossRef]
- Scheman, A.; Rakowski, E.M.; Chou, V.; Chhatriwala, A.; Ross, J.; Jacob, S.E. Balsam of Peru: Past and future. Dermatitis 2013, 24, 153–160. [Google Scholar] [CrossRef]
- Aubdool, A.A.; Brain, S.D. Neurovascular aspects of skin neurogenic inflammation. J. Investig. Dermatol. Symp. Proc. 2011, 15, 33–39. [Google Scholar] [CrossRef]
- Kamamoto, C.; Nishikaku, A.; Gompertz, O.; Melo, A.; Hassun, K.; Bagatin, E. Cutaneous fungal microbiome: Malassezia yeasts in seborrheic dermatitis scalp in a randomized, comparative and therapeutic trial. Dermato-endocrinology 2017, 9, e1361573. [Google Scholar] [CrossRef]
- Tucker, D.; Masood, S. Seborrheic Dermatitis. In StatPearls [Internet]; StatPearls Publishing: Treasure Island, FL, USA, 2019. [Google Scholar]
- Borda, L.J.; Wikramanayake, T.C. Seborrheic dermatitis and dandruff: A comprehensive review. J. Clin. Investig. Dermatol. 2015, 3. [Google Scholar] [CrossRef]
- Mokos, Z.B.; Kralj, M.; Basta-Juzbasic, A.; Jukic, I.L. Seborrheic dermatitis: An update. Acta Dermatovenerol Croat 2012, 20, 98–104. [Google Scholar]
- Rudramurthy, S.M.; Honnavar, P.; Chakrabarti, A.; Dogra, S.; Singh, P.; Handa, S. Association of Malassezia species with psoriatic lesions. Mycoses 2014, 57, 483–488. [Google Scholar] [CrossRef] [PubMed]
- DeAngelis, Y.M.; Gemmer, C.M.; Kaczvinsky, J.R.; Kenneally, D.C.; Schwartz, J.R.; Dawson, T.L., Jr. Three etiologic facets of dandruff and seborrheic dermatitis: Malassezia fungi, sebaceous lipids, and individual sensitivity. J. Investig. Dermatol. Symp. Proc. 2005, 10, 295–297. [Google Scholar] [CrossRef]
- Odintsova, I.; Dyudyun, A. Features of the composition of microorganisms inhabiting the intestinal mucosa in patients with seborrheic dermatitis. Dermatovenerol. Cosmetol. Sexopathol. 2019, 31–34. [Google Scholar] [CrossRef]
- Sakuma, T.H.; Maibach, H.I. Oily skin: An overview. Skin Pharmacol. Physiol. 2012, 25, 227–235. [Google Scholar] [CrossRef]
- Bett, D.; Morland, J.; Yudkin, J. Sugar consumption in acne vulgaris and seborrhoeic dermatitis. Br. Med J. 1967, 3, 153. [Google Scholar] [CrossRef] [PubMed]
- Pochi, P.E.; Downing, D.T.; Strauss, J.S. Sebaceous gland response in man to prolonged total caloric deprivation. J. Investig. Dermatol. 1970, 55, 303–309. [Google Scholar] [CrossRef]
- Boelsma, E.; Van de Vijver, L.P.; Goldbohm, R.A.; Klöpping-Ketelaars, I.A.; Hendriks, H.F.; Roza, L. Human skin condition and its associations with nutrient concentrations in serum and diet. Am. J. Clin. Nutr. 2003, 77, 348–355. [Google Scholar] [CrossRef]
- Tamer, F. Relationship between diet and seborrheic dermatitis. Our Dermatol. Online 2018, 9, 261–264. [Google Scholar] [CrossRef]
- Lee, H.H.; Gwillim, E.; Patel, K.R.; Hua, T.; Rastogi, S.; Ibler, E.; Silverberg, J.I. Epidemiology of alopecia areata, ophiasis, totalis, and universalis: A systematic review and meta-analysis. J. Am. Acad. Dermatol. 2020, 82, 675–682. [Google Scholar] [CrossRef] [PubMed]
- Simakou, T.; Butcher, J.P.; Reid, S.; Henriquez, F.L. Alopecia areata: A multifactorial autoimmune condition. J. Autoimmun. 2019, 98, 74–85. [Google Scholar] [CrossRef]
- Juhasz, M.; Chen, S.; Khosrovi-Eghbal, A.; Ekelem, C.; Landaverde, Y.; Baldi, P.; Mesinkovska, N.A. Characterizing the Skin and Gut Microbiome of Alopecia Areata Patients. SKIN J. Cutan. Med. 2020, 4, 23–30. [Google Scholar] [CrossRef]
- Polak-Witka, K.; Rudnicka, L.; Blume-Peytavi, U.; Vogt, A. The role of the microbiome in scalp hair follicle biology and disease. Exp. Dermatol. 2020, 29, 286–294. [Google Scholar] [CrossRef]
- Migacz-Gruszka, K.; Branicki, W.; Obtulowicz, A.; Pirowska, M.; Gruszka, K.; Wojas-Pelc, A. What’s new in the pathophysiology of alopecia areata? the possible contribution of skin and gut microbiome in the pathogenesis of alopecia–Big opportunities, big challenges, and novel perspectives. Int. J. Trichology 2019, 11, 185. [Google Scholar] [CrossRef] [PubMed]
- Guo, E.L.; Katta, R. Diet and hair loss: Effects of nutrient deficiency and supplement use. Dermatol. Pract. Concept. 2017, 7, 1. [Google Scholar] [CrossRef]
- Pham, C.T.; Romero, K.; Almohanna, H.M.; Griggs, J.; Ahmed, A.; Tosti, A. The Role of Diet as an Adjuvant Treatment in Scarring and Nonscarring Alopecia. Skin Appendage Disord. 2020, 6, 88–96. [Google Scholar] [CrossRef] [PubMed]
- Grosu-Bularda, A.; Lăzărescu, L.; Stoian, A.; Lascăr, I. Immunology and skin cancer. Arch. Clin. Cases 2018, 5. [Google Scholar] [CrossRef]
- Carr, S.; Smith, C.; Wernberg, J. Epidemiology and risk factors of melanoma. Surg. Clin. 2020, 100, 1–12. [Google Scholar] [CrossRef]
- Marks, R. An overview of skin cancers. Cancer 1995, 75, 607–612. [Google Scholar] [CrossRef]
- Rangwala, S.; Tsai, K. Roles of the immune system in skin cancer. Br. J. Dermatol. 2011, 165, 953–965. [Google Scholar] [CrossRef] [PubMed]
- Vergara, D.; Simeone, P.; Damato, M.; Maffia, M.; Lanuti, P.; Trerotola, M. The cancer microbiota: EMT and inflammation as shared molecular mechanisms associated with plasticity and progression. J. Oncol. 2019, 2019, 1253727. [Google Scholar] [CrossRef]
- Nakatsuji, T.; Chen, T.H.; Butcher, A.M.; Trzoss, L.L.; Nam, S.J.; Shirakawa, K.T.; Zhou, W.; Oh, J.; Otto, M.; Fenical, W.; et al. A commensal strain of Staphylococcus epidermidis protects against skin neoplasia. Sci. Adv. 2018, 4, eaao4502. [Google Scholar] [CrossRef] [PubMed]
- Allhorn, M.; Arve, S.; Brüggemann, H.; Lood, R. A novel enzyme with antioxidant capacity produced by the ubiquitous skin colonizer Propionibacterium acnes. Sci. Rep. 2016, 6, 36412. [Google Scholar] [CrossRef] [PubMed]
- Frosali, S.; Pagliari, D.; Gambassi, G.; Landolfi, R.; Pandolfi, F.; Cianci, R. How the intricate interaction among toll-like receptors, microbiota, and intestinal immunity can influence gastrointestinal pathology. J. Immunol. Res. 2015, 2015, 489821. [Google Scholar] [CrossRef]
- Eming, S.A.; Martin, P.; Tomic-Canic, M. Wound repair and regeneration: Mechanisms, signaling, and translation. Sci. Transl. Med. 2014, 6, 265sr6. [Google Scholar] [CrossRef]
- Heyer, K.; Herberger, K.; Protz, K.; Glaeske, G.; Augustin, M. Epidemiology of chronic wounds in Germany: Analysis of statutory health insurance data. Wound Repair Regen. 2016, 24, 434–442. [Google Scholar] [CrossRef] [PubMed]
- Guest, J.F.; Ayoub, N.; McIlwraith, T.; Uchegbu, I.; Gerrish, A.; Weidlich, D.; Vowden, K.; Vowden, P. Health economic burden that wounds impose on the National Health Service in the UK. BMJ Open 2015, 5. [Google Scholar] [CrossRef]
- Gould, L.; Abadir, P.; Brem, H.; Carter, M.; Conner-Kerr, T.; Davidson, J.; DiPietro, L.; Falanga, V.; Fife, C.; Gardner, S.; et al. Chronic wound repair and healing in older adults: Current status and future research. Wound Repair Regen. 2015, 23, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Sawaya, A.P.; Stone, R.C.; Brooks, S.R.; Pastar, I.; Jozic, I.; Hasneen, K.; O’Neill, K.; Mehdizadeh, S.; Head, C.R.; Strbo, N.; et al. Deregulated immune cell recruitment orchestrated by FOXM1 impairs human diabetic wound healing. Nat. Commun. 2020, 11, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Tomic-Canic, M.; Burgess, J.L.; O’Neill, K.E.; Strbo, N.; Pastar, I. Skin Microbiota and its Interplay with Wound Healing. Am. J. Clin. Dermatol. 2020, 21, 36–43. [Google Scholar] [CrossRef]
- Ramirez, H.A.; Pastar, I.; Jozic, I.; Stojadinovic, O.; Stone, R.C.; Ojeh, N.; Gil, J.; Davis, S.C.; Kirsner, R.S.; Tomic-Canic, M. Staphylococcus aureus triggers induction of miR-15B-5P to diminish DNA repair and deregulate inflammatory response in diabetic foot ulcers. J. Investig. Dermatol. 2018, 138, 1187–1196. [Google Scholar] [CrossRef]
- Stone, R.C.; Stojadinovic, O.; Rosa, A.M.; Ramirez, H.A.; Badiavas, E.; Blumenberg, M.; Tomic-Canic, M. A bioengineered living cell construct activates an acute wound healing response in venous leg ulcers. Sci. Transl. Med. 2017, 9, eaaf8611. [Google Scholar] [CrossRef]
- Stone, R.C.; Stojadinovic, O.; Sawaya, A.P.; Glinos, G.D.; Lindley, L.E.; Pastar, I.; Badiavas, E.; Tomic-Canic, M. A bioengineered living cell construct activates metallothionein/zinc/MMP8 and inhibits TGFβ to stimulate remodeling of fibrotic venous leg ulcers. Wound Repair Regen. 2020, 28, 164–176. [Google Scholar] [CrossRef] [PubMed]
- Pastar, I.; Wong, L.L.; Egger, A.N.; Tomic-Canic, M. Descriptive vs mechanistic scientific approach to study wound healing and its inhibition: Is there a value of translational research involving human subjects? Exp. Dermatol. 2018, 27, 551–562. [Google Scholar] [CrossRef] [PubMed]
- Thom, S.R.; Hampton, M.; Troiano, M.A.; Mirza, Z.; Malay, D.S.; Shannon, S.; Jennato, N.B.; Donohue, C.M.; Hoffstad, O.; Woltereck, D.; et al. Measurements of CD34+/CD45-dim stem cells predict healing of diabetic neuropathic wounds. Diabetes 2016, 65, 486–497. [Google Scholar] [CrossRef] [PubMed]
- Zeeuwen, P.L.; Boekhorst, J.; van den Bogaard, E.H.; de Koning, H.D.; van de Kerkhof, P.M.; Saulnier, D.M.; van Swam, I.I.; van Hijum, S.A.; Kleerebezem, M.; Schalkwijk, J.; et al. Microbiome dynamics of human epidermis following skin barrier disruption. Genome Biol. 2012, 13, 1–18. [Google Scholar] [CrossRef]
- Pastar, I.; Nusbaum, A.G.; Gil, J.; Patel, S.B.; Chen, J.; Valdes, J.; Stojadinovic, O.; Plano, L.R.; Tomic-Canic, M.; Davis, S.C. Interactions of methicillin resistant Staphylococcus aureus USA300 and Pseudomonas aeruginosa in polymicrobial wound infection. PLoS ONE 2013, 8, e56846. [Google Scholar] [CrossRef]
- Harrison, O.J.; Linehan, J.L.; Shih, H.Y.; Bouladoux, N.; Han, S.J.; Smelkinson, M.; Sen, S.K.; Byrd, A.L.; Enamorado, M.; Yao, C.; et al. Commensal-specific T cell plasticity promotes rapid tissue adaptation to injury. Science 2019, 363, eaat6280. [Google Scholar] [CrossRef]
- Lai, Y.; Di Nardo, A.; Nakatsuji, T.; Leichtle, A.; Yang, Y.; Cogen, A.L.; Wu, Z.R.; Hooper, L.V.; Schmidt, R.R.; Von Aulock, S.; et al. Commensal bacteria regulate Toll-like receptor 3–dependent inflammation after skin injury. Nat. Med. 2009, 15, 1377. [Google Scholar] [CrossRef]
- McCormack, R.M.; de Armas, L.R.; Shiratsuchi, M.; Fiorentino, D.G.; Olsson, M.L.; Lichtenheld, M.G.; Morales, A.; Lyapichev, K.; Gonzalez, L.E.; Strbo, N.; et al. Perforin-2 is essential for intracellular defense of parenchymal cells and phagocytes against pathogenic bacteria. Elife 2015, 4, e06508. [Google Scholar] [CrossRef]
- Pastar, I.; O’Neill, K.; Padula, L.; Head, C.R.; Burgess, J.L.; Chen, V.; Garcia, D.; Stojadinovic, O.; Hower, S.; Plano, G.V.; et al. Staphylococcus epidermidis boosts innate immune response by activation of Gamma Delta T cells and induction of Perforin-2 in human skin. Front. Immunol. 2020, 11, 2253. [Google Scholar] [CrossRef] [PubMed]
- Luqman, A.; Muttaqin, M.Z.; Yulaipi, S.; Ebner, P.; Matsuo, M.; Zabel, S.; Tribelli, P.M.; Nieselt, K.; Hidayati, D.; Götz, F. Trace amines produced by skin bacteria accelerate wound healing in mice. Commun. Biol. 2020, 3, 1–10. [Google Scholar] [CrossRef]
- Misic, A.M.; Gardner, S.E.; Grice, E.A. The wound microbiome: Modern approaches to examining the role of microorganisms in impaired chronic wound healing. Adv. Wound Care 2014, 3, 502–510. [Google Scholar] [CrossRef]
- Kalan, L.R.; Meisel, J.S.; Loesche, M.A.; Horwinski, J.; Soaita, I.; Chen, X.; Uberoi, A.; Gardner, S.E.; Grice, E.A. Strain-and species-level variation in the microbiome of diabetic wounds is associated with clinical outcomes and therapeutic efficacy. Cell Host Microbe 2019, 25, 641–655. [Google Scholar] [CrossRef] [PubMed]
- Huseini, H.F.; Rahimzadeh, G.; Fazeli, M.R.; Mehrazma, M.; Salehi, M. Evaluation of wound healing activities of kefir products. Burns 2012, 38, 719–723. [Google Scholar] [CrossRef] [PubMed]
- Poutahidis, T.; Kearney, S.M.; Levkovich, T.; Qi, P.; Varian, B.J.; Lakritz, J.R.; Ibrahim, Y.M.; Chatzigiagkos, A.; Alm, E.J.; Erdman, S.E. Microbial symbionts accelerate wound healing via the neuropeptide hormone oxytocin. PLoS ONE 2013, 8, e78898. [Google Scholar] [CrossRef]
- Levkovich, T.; Poutahidis, T.; Smillie, C.; Varian, B.J.; Ibrahim, Y.M.; Lakritz, J.R.; Alm, E.J.; Erdman, S.E. Probiotic bacteria induce a ‘glow of health’. PLoS ONE 2013, 8, e53867. [Google Scholar] [CrossRef] [PubMed]
- Gueniche, A.; Philippe, D.; Bastien, P.; Reuteler, G.; Blum, S.; Castiel-Higounenc, I.; Breton, L.; Benyacoub, J. Randomised double-blind placebo-controlled study of the effect of Lactobacillus paracasei NCC 2461 on skin reactivity. Benef. Microbes 2014, 5, 137–145. [Google Scholar] [CrossRef]
- Krutmann, J. Pre-and probiotics for human skin. J. Dermatol. Sci. 2009, 54, 1–5. [Google Scholar] [CrossRef]
- Hill, C.; Guarner, F.; Reid, G.; Gibson, G.R.; Merenstein, D.J.; Pot, B.; Morelli, L.; Canani, R.B.; Flint, H.J.; Salminen, S.; et al. Expert consensus document: The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 506. [Google Scholar] [CrossRef]
- Farris, P.K. Are skincare products with probiotics worth the hype? DermatologyTimes. 2016. Available online: https://www.dermatologytimes.com/view/are-skincare-products-probiotics-worth-hype (accessed on 20 November 2020).
- Grant, M.C.; Baker, J.S. An overview of the effect of probiotics and exercise on mood and associated health conditions. Crit. Rev. Food Sci. Nutr. 2017, 57, 3887–3893. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, B.; Delgado, S.; Blanco-Míguez, A.; Lourenço, A.; Gueimonde, M.; Margolles, A. Probiotics, gut microbiota, and their influence on host health and disease. Mol. Nutr. Food Res. 2017, 61, 1600240. [Google Scholar] [CrossRef]
- Sarao, L.K.; Arora, M. Probiotics, prebiotics, and microencapsulation: A review. Crit. Rev. Food Sci. Nutr. 2017, 57, 344–371. [Google Scholar] [CrossRef] [PubMed]
- Muizzuddin, N.; Maher, W.; Sullivan, M.; Schnittger, S.; Mammone, T. Physiological effect of a probiotic on skin. J. Cosmet. Sci. 2012, 63, 385–395. [Google Scholar]
- Frei, R.; Akdis, M.; O’Mahony, L. Prebiotics, probiotics, synbiotics, and the immune system: Experimental data and clinical evidence. Curr. Opin. Gastroenterol. 2015, 31, 153–158. [Google Scholar] [CrossRef] [PubMed]
- Longo, V.D.; Cortellino, S. Fasting, dietary restriction, and immunosenescence. J. Allergy Clin. Immunol. 2020, 146, 1002–1004. [Google Scholar] [CrossRef]
- Bronsnick, T.; Murzaku, E.C.; Rao, B.K. Diet in dermatology: Part I. Atopic dermatitis, acne, and nonmelanoma skin cancer. J. Am. Acad. Dermatol. 2014, 71, 1039–e1. [Google Scholar] [CrossRef] [PubMed]
- Parkar, S.G.; Kalsbeek, A.; Cheeseman, J.F. Potential role for the gut microbiota in modulating host circadian rhythms and metabolic health. Microorganisms 2019, 7, 41. [Google Scholar] [CrossRef]
- Voigt, R.; Forsyth, C.; Green, S.; Engen, P.; Keshavarzian, A. Circadian rhythm and the gut microbiome. In International Review of Neurobiology; Elsevier: Amsterdam, The Netherlands, 2016; Volume 131, pp. 193–205. [Google Scholar]
- Deaver, J.A.; Eum, S.Y.; Toborek, M. Circadian disruption changes gut microbiome taxa and functional gene composition. Front. Microbiol. 2018, 9, 737. [Google Scholar] [CrossRef]
- Zeb, F.; Wu, X.; Chen, L.; Fatima, S.; Haq, I.u.; Chen, A.; Majeed, F.; Feng, Q.; Li, M. Effect of time-restricted feeding on metabolic risk and circadian rhythm associated with gut microbiome in healthy males. Br. J. Nutr. 2020, 123, 1216–1226. [Google Scholar] [CrossRef] [PubMed]
- Jakubowicz, D.; Barnea, M.; Wainstein, J.; Froy, O. High caloric intake at breakfast vs. dinner differentially influences weight loss of overweight and obese women. Obesity 2013, 21, 2504–2512. [Google Scholar] [CrossRef]
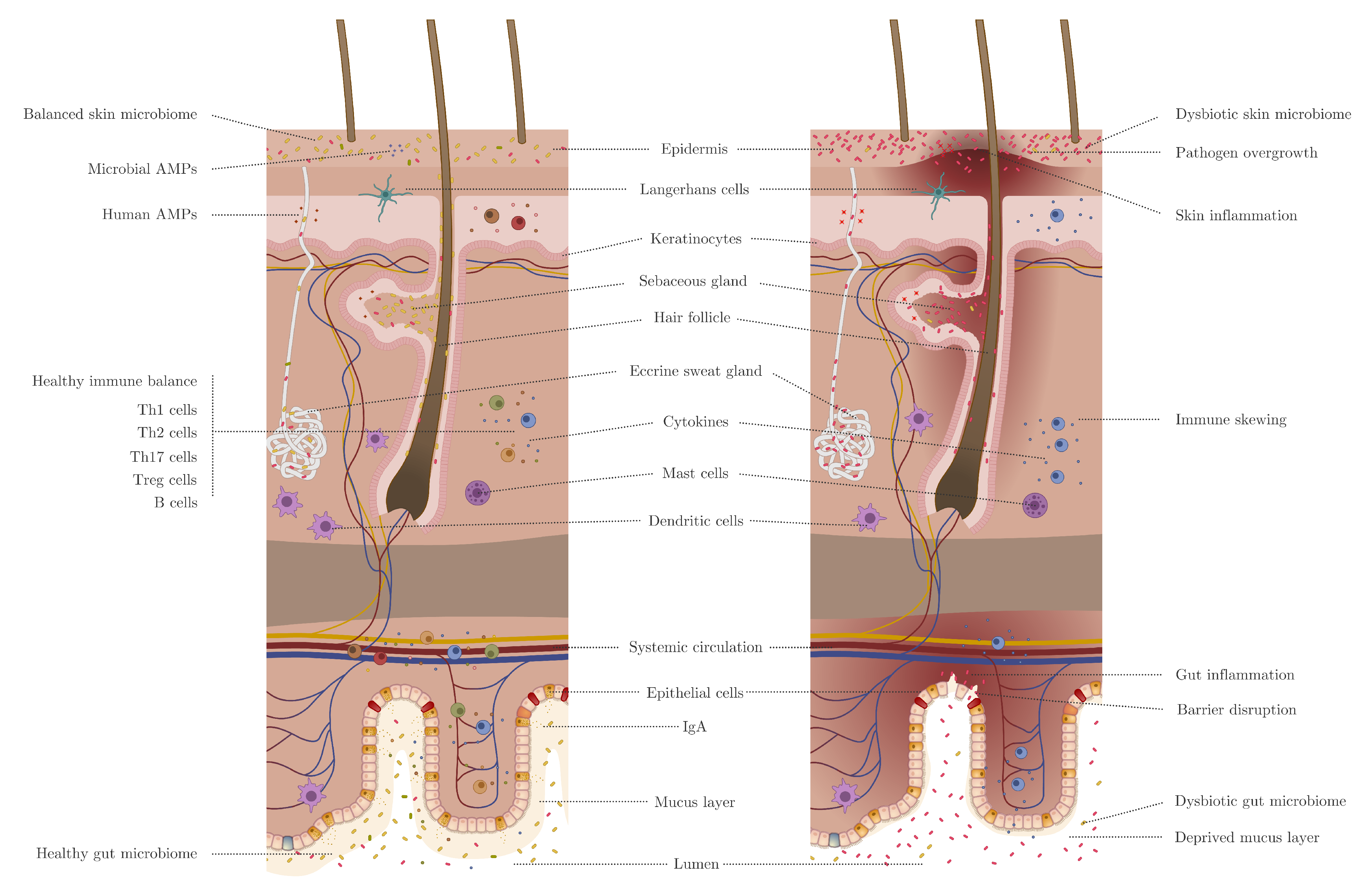
| Disease | Associated Skin Microbiota | Additional Remarks | Reference |
|---|---|---|---|
| 1. Acne vulgaris | Particular C. acnes strains | Administered probiotic bacteria could play a protective role. | [71,72,73,74,75,76,77,78,79] |
| 2. Atopic Dermatitis | Decreased bacterial diversity. Increased abundance of S. aureus. | Herpes simplex virus and coxsackie virus can infect AD * skin. | [55,80,81,82,83] |
| 3. Psoriasis | Higher abundance of Staphylococcus and Streptococcus. | Anti-psoriasis treatments lead to skin microbial changes. | [84,85,86,87,88] |
| 4. Hidradenitis suppurativa | Saccharomyces cerevisiae(yeast), Prevotella, and Porphyromonas (bacteria) | Anaerobic species in lesions. | [89,90] |
| 5. Rosacea | Demodex folliculorum(mites) | C. acnes decreased and Snodgrassella alvi increased. Geobacillus and Gordonia. | [91,92] |
| 6. Dandruff and Seborrheic dermatitis | Malassezia spp. (yeast) | Potential bacterial imbalance. | [93,94,95,96] |
| 7. Alopecia areata | Limited data. Possible imbalance C. acnes/ S. epidermidis. | Potential role of cytomegalovirus and/or Alternaria fungi. | [97,98,99] |
| 8. Skin cancer | Merkel bell Polyomavirus, Fusobacterium, and Trueperella, S. aureus. | Increase in certain strains of S. aureus in combination with a decrease in skin commensals can be associated to SCC * or BCC *, and that in MCPyV * can be associated to MCC *. | [100,101,102,103] |
| 9. Wound healing | S. aureus and biofilm-forming bacteria. | Lactobacilli and fermented products can be beneficial. | [104,105] |
| Disease | Associated Gut Microbiota | Additional Remarks | Reference |
|---|---|---|---|
| 1. Acne vulgaris | Decrease in Firmicutes and increase in Bacteroides. | Distinct gut microbiome composition and decreased diversity. | [106] |
| 2. Atopic Dermatitis | Higher levels of Faecalibacterium prausnitzii, Clostridium, and Escherichia (in infants). Lower levels of Akkermansia, Bacteroidetes, and Bifidobacterium. | Probiotics consumption can prevent AD *. | [107,108,109,110,111,112,113,114] |
| 3. Psoriasis | Changes in -diversity. Gut microbiome changes in reaction to biologicals. | Increased risk of intestinal immune disorders. Diet and gut microbiome can have an impact on inflammation. | [115,116,117,118,119,120] |
| 4. Hidradenitis suppurativa | Unknown | Increased risk in developing CD * and UC *. | [121,122] |
| 5. Rosacea | Can be associated with SIBO *. Acidaminococcus and Megasphaera increase and Peptococcaceae and Methanobrevibacter decrease. | Can be associated with H. pylori infection. | [123,124,125] |
| 6. Dandruff and Seborrheic dermatitis | Unclear | Probiotic consumption can alleviate moderate to severe dandruff | [126] |
| 7. Alopecia areata | No major differences | FMT * in 2 patients showed restoration of hair growth | [127,128] |
| 8. Skin cancer | Not reported | Other cancers are associated with microbial dysbiosis | [129,130,131] |
| 9. Wound healing | Not reported | Not reported |
| Molecule | Documented/Possible effect in gut | Documented/Possible effect on skin | Reference |
|---|---|---|---|
| Bacterial metabolites | |||
| SCFAs * | Anti-inflammatory effects | Anti-inflammatory effects | [132] |
| Vitamin D | Suppress inflammation in IBD* | Not reported | [133] |
| Urocanic Acid | Suppress inflammation in IBD* | Not reported | [134] |
| GABA * | Neurotransmitter modulation | Itch restriction | [135,136] |
| Dopamine | Neurotransmitter modulation | Inhibition of hair growth | [135,137] |
| Serotonin | Neurotransmitter modulation | Melatonin modulation | [135,138] |
| Acetylcholine | Neurotransmitter modulation | Barrier function | [135,139] |
| Phenol and p-cresol | Biomarker of gut dysbiosis | Impaired epidermal barrier function | [140] |
| Dietary components | |||
| Catechins | Anti-inflammatory effects | Anti-inflammatory effects | [141] |
| Polyphenols | Anti-inflammatory effects | Anti-inflammatory effects | [142] |
| Lycopene | Selectively utilized by host microbiota | Protection against photodamage | [143,144] |
| Prolamin | Not reported | Protection against AD * | [145] |
| Phytomolecules | Not reported | Anti-ageing | [146] |
| Gluten | Coeliac disease | Skin Rashes | [58,59] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Pessemier, B.; Grine, L.; Debaere, M.; Maes, A.; Paetzold, B.; Callewaert, C. Gut–Skin Axis: Current Knowledge of the Interrelationship between Microbial Dysbiosis and Skin Conditions. Microorganisms 2021, 9, 353. https://doi.org/10.3390/microorganisms9020353
De Pessemier B, Grine L, Debaere M, Maes A, Paetzold B, Callewaert C. Gut–Skin Axis: Current Knowledge of the Interrelationship between Microbial Dysbiosis and Skin Conditions. Microorganisms. 2021; 9(2):353. https://doi.org/10.3390/microorganisms9020353
Chicago/Turabian StyleDe Pessemier, Britta, Lynda Grine, Melanie Debaere, Aglaya Maes, Bernhard Paetzold, and Chris Callewaert. 2021. "Gut–Skin Axis: Current Knowledge of the Interrelationship between Microbial Dysbiosis and Skin Conditions" Microorganisms 9, no. 2: 353. https://doi.org/10.3390/microorganisms9020353
APA StyleDe Pessemier, B., Grine, L., Debaere, M., Maes, A., Paetzold, B., & Callewaert, C. (2021). Gut–Skin Axis: Current Knowledge of the Interrelationship between Microbial Dysbiosis and Skin Conditions. Microorganisms, 9(2), 353. https://doi.org/10.3390/microorganisms9020353








