Consistent Prebiotic Effects of Carrot RG-I on the Gut Microbiota of Four Human Adult Donors in the SHIME® Model despite Baseline Individual Variability
Abstract
:1. Introduction
2. Materials and Methods
2.1. Chemicals
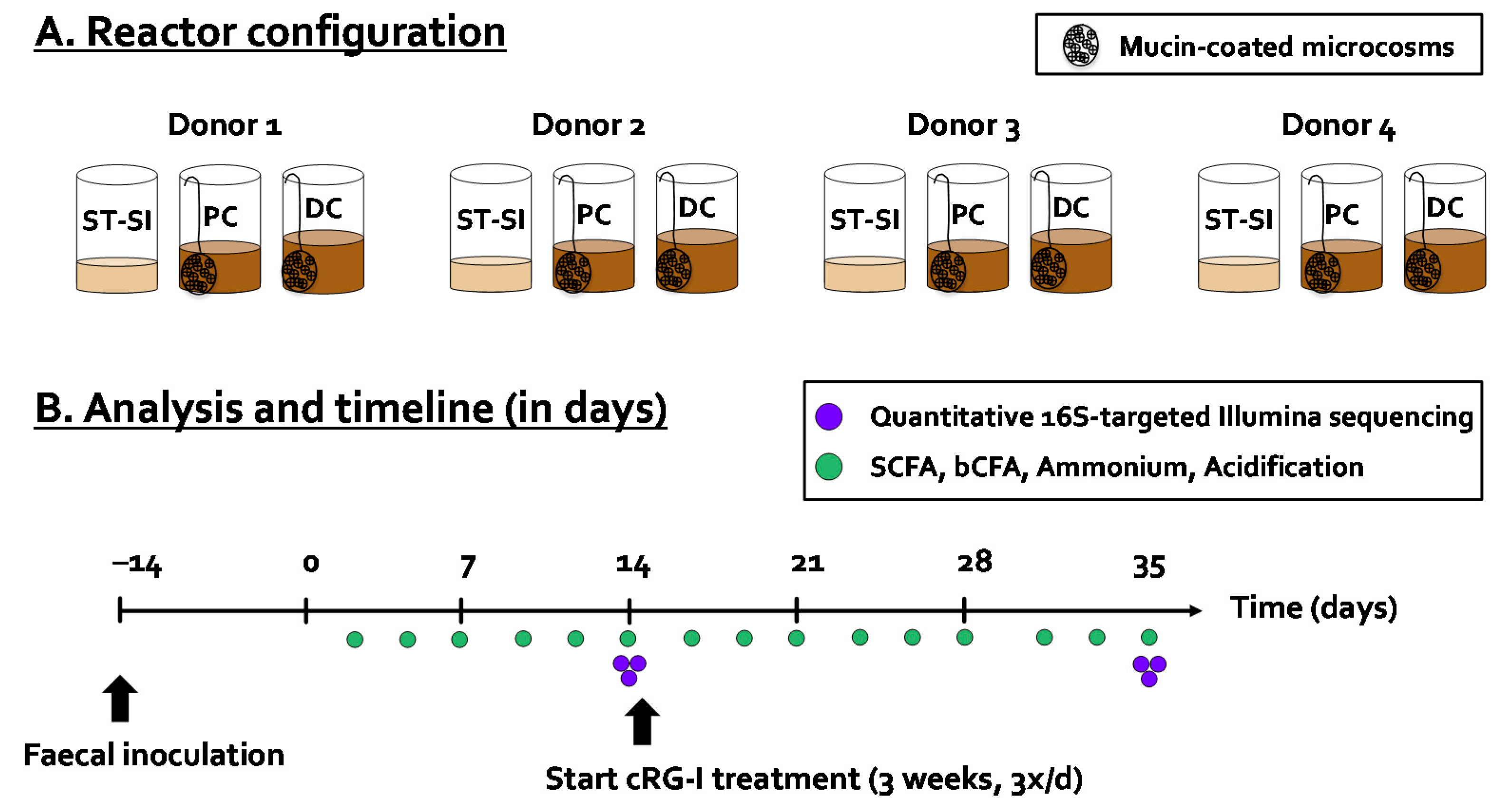
2.2. Mucosal Simulator of the Human Intestinal Microbial Ecosystem (M-SHIME®)
2.3. Microbial Activity Analysis
2.4. Microbial Composition Analysis
2.5. Bifidobacteria Growth Experiments on cRG-I
2.6. Statistics
3. Results
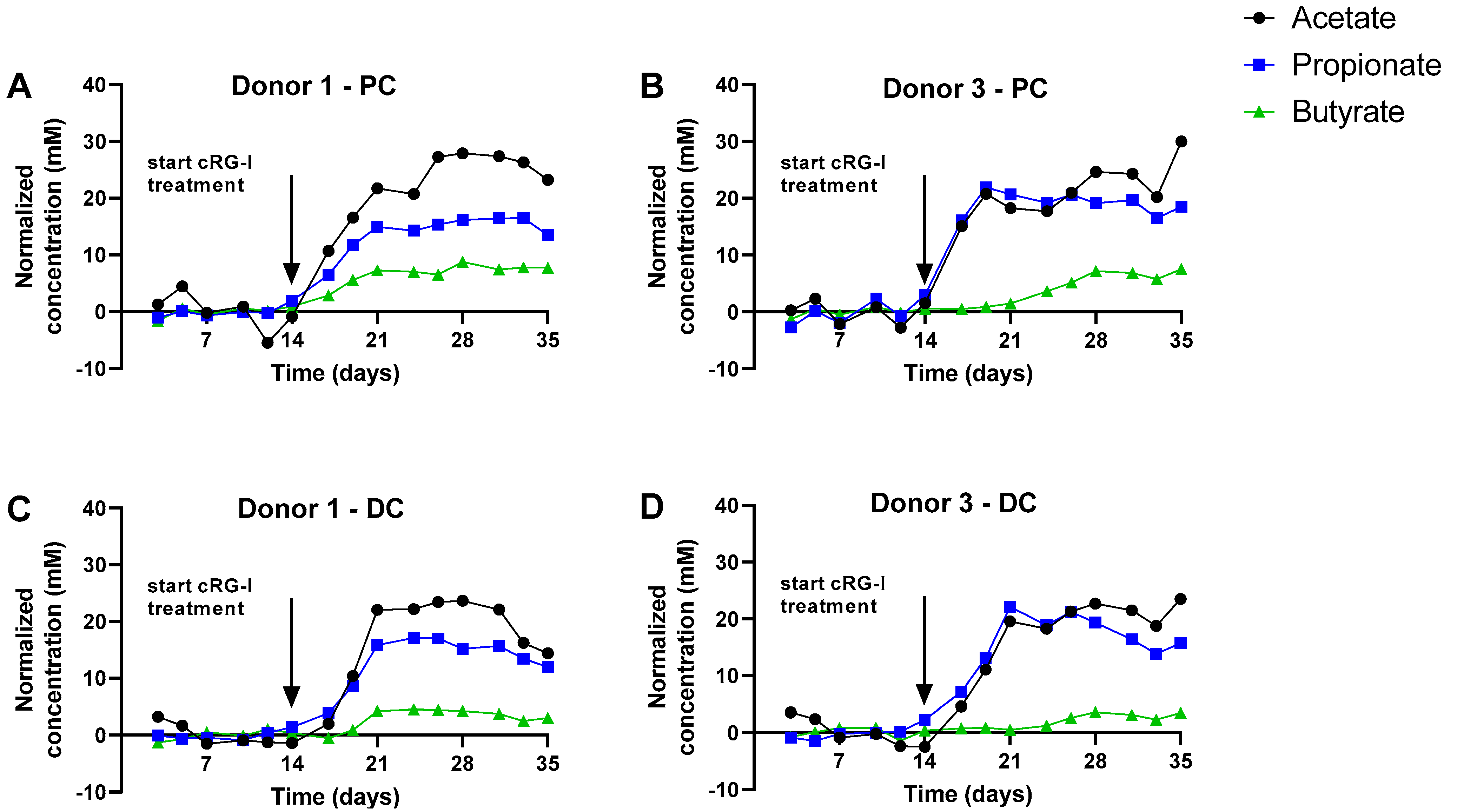
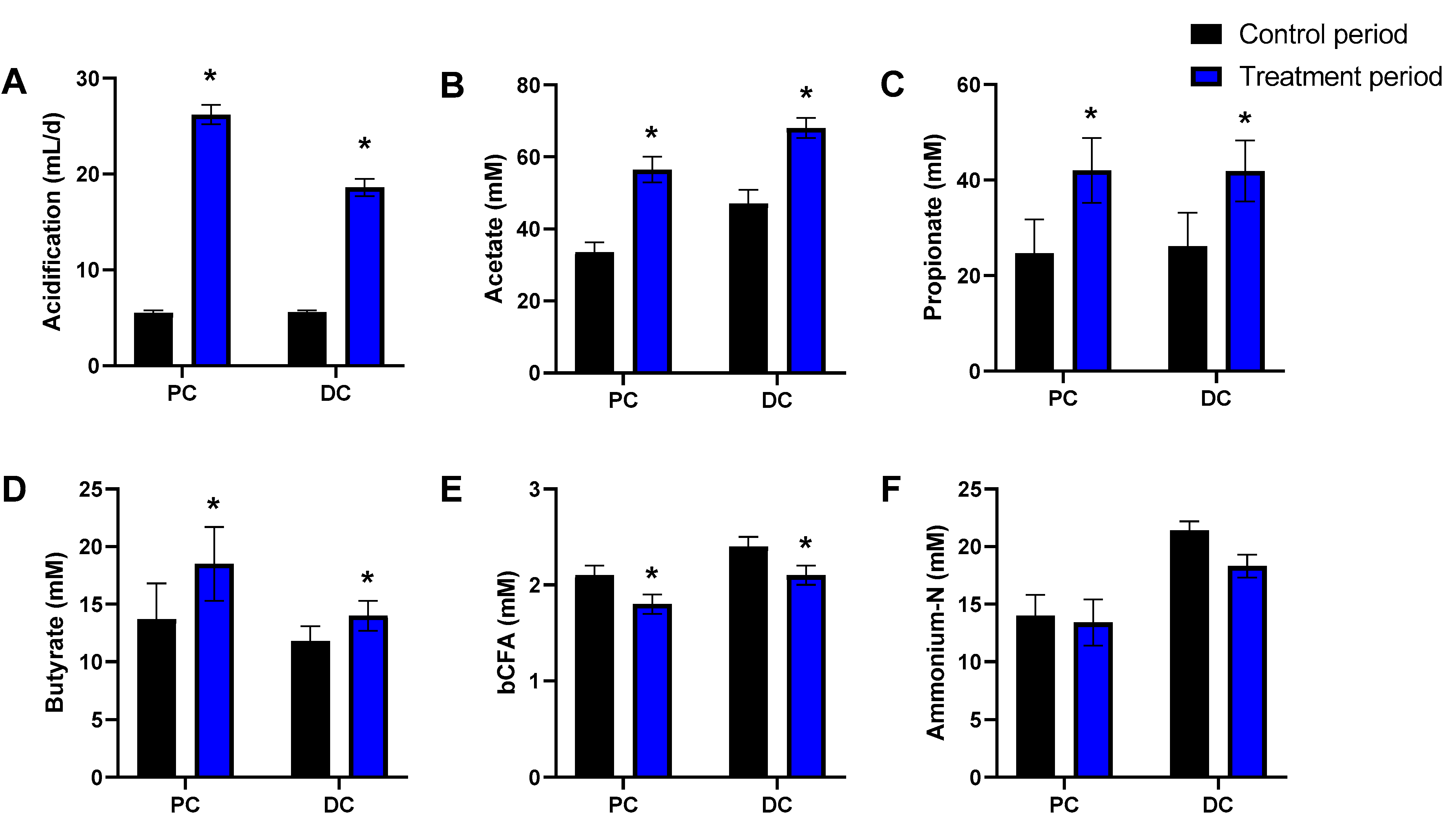
3.1. cRG-I Consistently Stimulated Microbial Activity in the Simulated Proximal and Distal Colon of Four Human Adults Tested in the Quad-M-SHIME
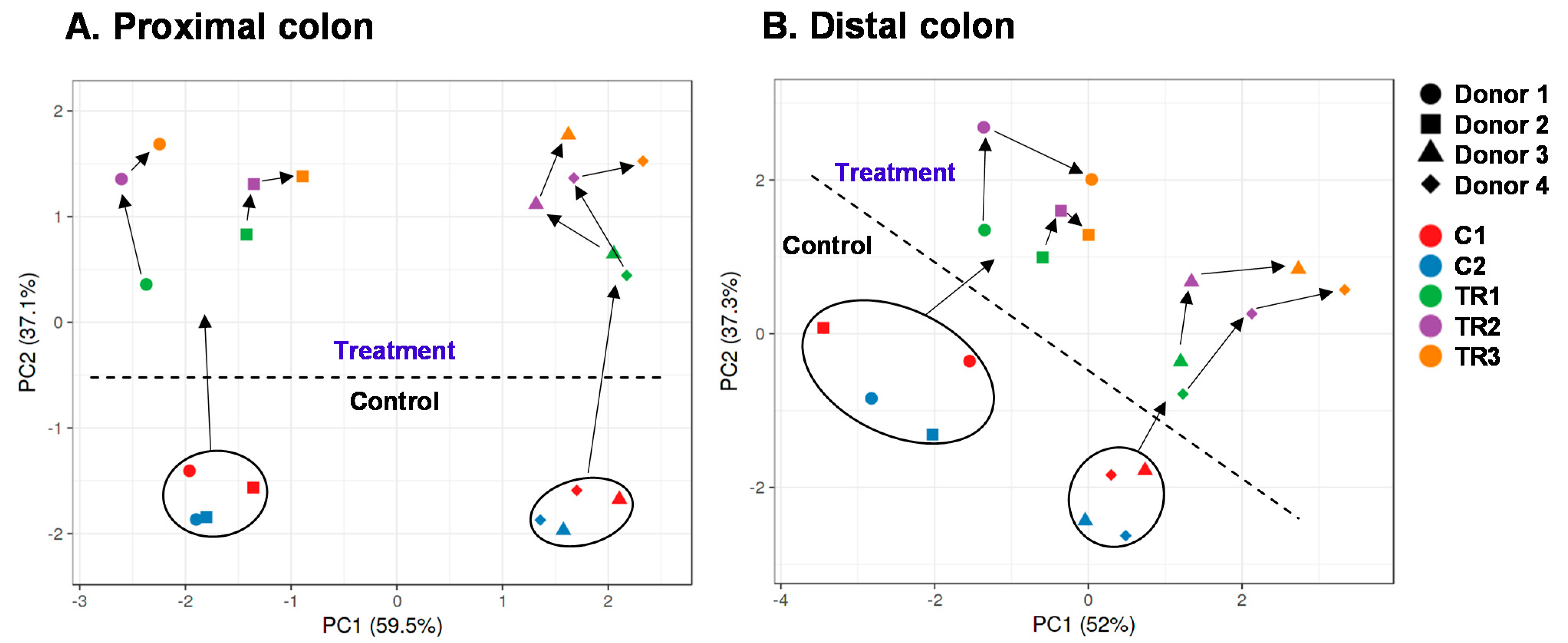
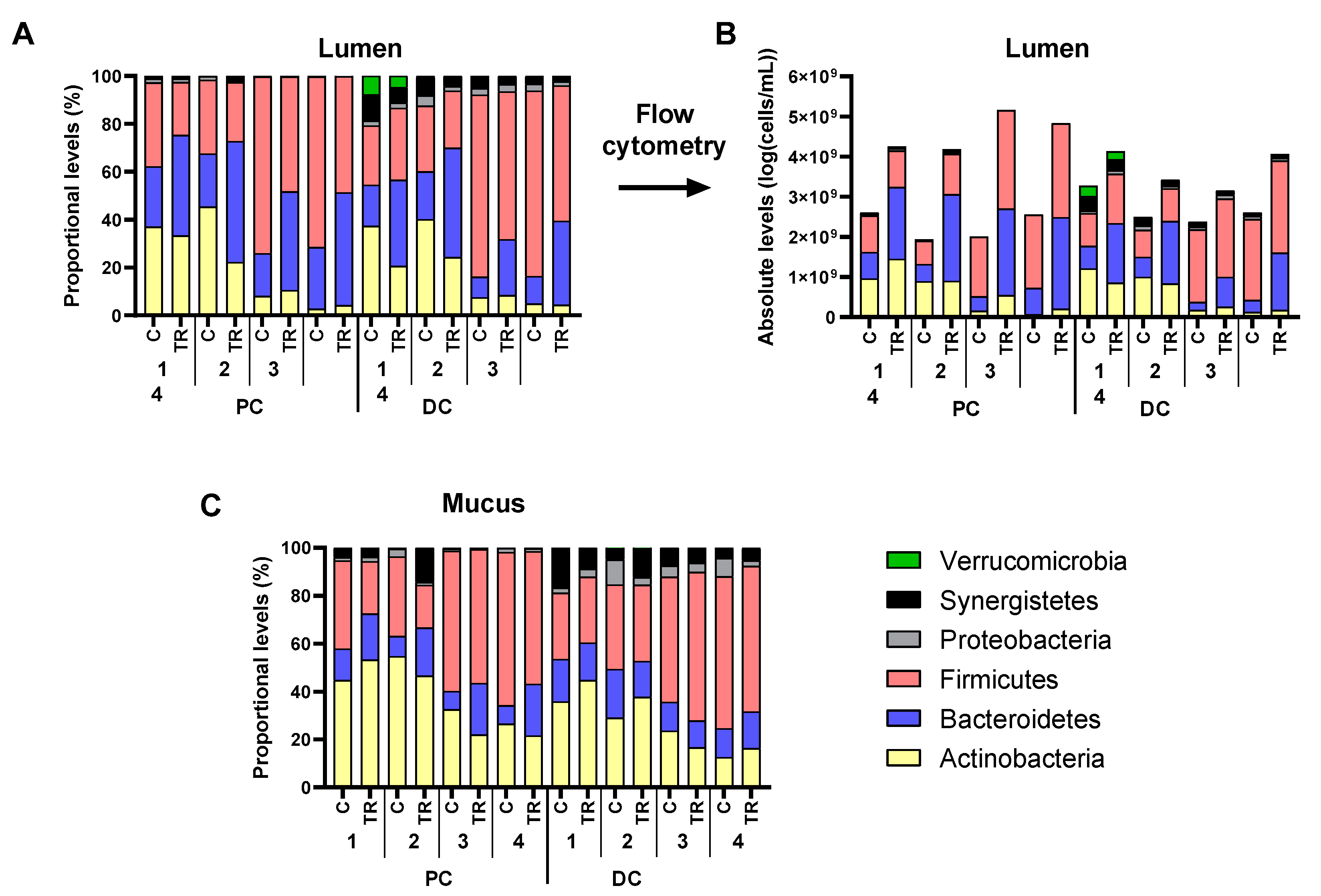
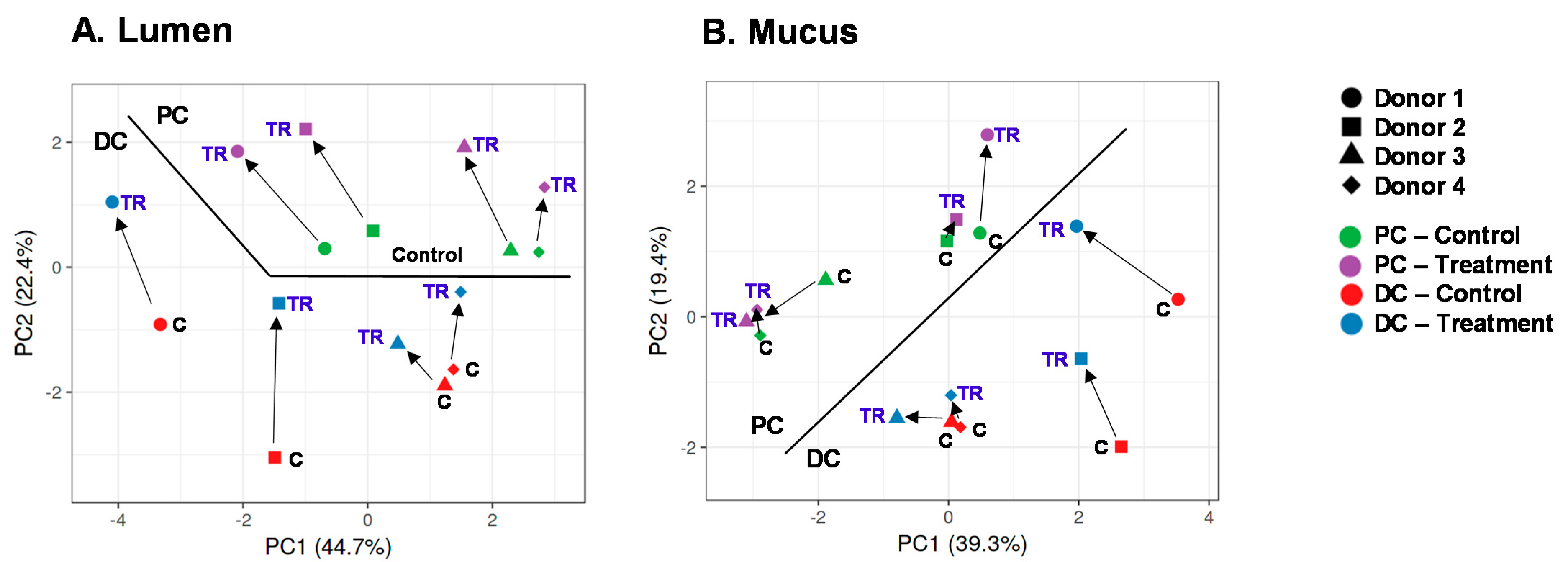
3.2. cRG-I Consistently Modulated Luminal and Mucosal Microbiota Composition in the Simulated Proximal and Distal Colon of Four Human Adults as Tested in the Quad-M-SHIME
3.3. Bifidobacterium Strains Are Unable to Ferment cRG-I as Such
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Marchesi, J.R.; Adams, D.H.; Fava, F.; Hermes, G.D.A.; Hirschfield, G.M.; Hold, G.; Quraishi, M.N.; Kinross, J.; Smidt, H.; Tuohy, K.M.; et al. The gut microbiota and host health: A new clinical frontier. Gut 2016, 65, 330–339. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boulangé, C.L.; Neves, A.L.; Chilloux, J.; Nicholson, J.K.; Dumas, M.-E. Impact of the gut microbiota on inflammation, obesity, and metabolic disease. Genome Med. 2016, 8, 42. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tilg, H.; Cani, P.D.; Mayer, E.A. Gut microbiome and liver diseases. Gut 2016, 65, 2035–2044. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.; Wu, G.D.; Albenberg, L.; Tomov, V.T. Gut microbiota and IBD: Causation or correlation? Nat. Rev. Gastroenterol. Hepatol. 2017, 14, 573–584. [Google Scholar] [CrossRef] [Green Version]
- Ternes, D.; Karta, J.; Tsenkova, M.; Wilmes, P.; Haan, S.; Letellier, E. Microbiome in Colorectal Cancer: How to Get from Meta-omics to Mechanism? Trends Microbiol. 2020, 28, 401–423. [Google Scholar] [CrossRef]
- Flint, H.J.; Scott, K.P.; Louis, P.; Duncan, S.H. The role of the gut microbiota in nutrition and health. Nat. Rev. Gastroenterol. Hepatol. 2012, 9, 577–589. [Google Scholar] [CrossRef]
- Rivière, A.; Selak, M.; Lantin, D.; Leroy, F.; De Vuyst, L. Bifidobacteria and Butyrate-Producing Colon Bacteria: Importance and Strategies for Their Stimulation in the Human Gut. Front. Microbiol. 2016, 7, 979. [Google Scholar] [CrossRef] [Green Version]
- Koh, A.; De Vadder, F.; Kovatcheva-Datchary, P.; Bäckhed, F. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 2016, 165, 1332–1345. [Google Scholar] [CrossRef] [Green Version]
- Gonçalves, P.; Araújo, J.R.; Di Santo, J.P. A Cross-Talk Between Microbiota-Derived Short-Chain Fatty Acids and the Host Mucosal Immune System Regulates Intestinal Homeostasis and Inflammatory Bowel Disease. Inflamm. Bowel Dis. 2018, 24, 558–572. [Google Scholar] [CrossRef] [Green Version]
- van der Hee, B.; Wells, J.M. Microbial Regulation of Host Physiology by Short-chain Fatty Acids. Trends Microbiol. 2021, 29, 700–712. [Google Scholar] [CrossRef]
- Ndeh, D.; Gilbert, H.J. Biochemistry of complex glycan depolymerisation by the human gut microbiota. FEMS Microbiol. Rev. 2018, 42, 146–164. [Google Scholar] [CrossRef] [Green Version]
- Martens, E.C.; Lowe, E.C.; Chiang, H.; Pudlo, N.A.; Wu, M.; McNulty, N.P.; Abbott, D.W.; Henrissat, B.; Gilbert, H.J.; Bolam, D.N.; et al. Recognition and Degradation of Plant Cell Wall Polysaccharides by Two Human Gut Symbionts. PLoS Biol. 2011, 9, e1001221. [Google Scholar] [CrossRef]
- Cuskin, F.; Lowe, E.C.; Temple, M.J.; Zhu, Y.; Cameron, E.; Pudlo, N.A.; Porter, N.T.; Urs, K.; Thompson, A.J.; Cartmell, A.; et al. Human gut Bacteroidetes can utilize yeast mannan through a selfish mechanism. Nature 2015, 517, 165–169. [Google Scholar] [CrossRef] [Green Version]
- Larsbrink, J.; Rogers, T.E.; Hemsworth, G.R.; McKee, L.S.; Tauzin, A.S.; Spadiut, O.; Klinter, S.; Pudlo, N.A.; Urs, K.; Koropatkin, N.M.; et al. A discrete genetic locus confers xyloglucan metabolism in select human gut Bacteroidetes. Nature 2014, 506, 498–502. [Google Scholar] [CrossRef]
- Kelly, S.M.; Munoz-Munoz, J.; van Sinderen, D. Plant Glycan Metabolism by Bifidobacteria. Front. Microbiol. 2021, 12, 25. [Google Scholar] [CrossRef] [PubMed]
- Windey, K.; De Preter, V.; Verbeke, K. Relevance of protein fermentation to gut health. Mol. Nutr. Food Res. 2012, 56, 184–196. [Google Scholar] [CrossRef] [PubMed]
- Gibson, G.R.; Hutkins, R.; Sanders, M.E.; Prescott, S.L.; Reimer, R.A.; Salminen, S.J.; Scott, K.; Stanton, C.; Swanson, K.S.; Cani, P.D.; et al. Expert consensus document: The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat. Rev. Gastroenterol. Hepatol. 2017, 14, 491–502. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sabater-Molina, M.; Larqué, E.; Torrella, F.; Zamora, S. Dietary fructooligosaccharides and potential benefits on health. J. Physiol. Biochem. 2009, 65, 315–328. [Google Scholar] [CrossRef] [PubMed]
- Le Bastard, Q.; Chapelet, G.; Javaudin, F.; Lepelletier, D.; Batard, E.; Montassier, E. The effects of inulin on gut microbial composition: A systematic review of evidence from human studies. Eur. J. Clin. Microbiol. Infect. Dis. 2020, 39, 403–413. [Google Scholar] [CrossRef]
- Bode, L. Human milk oligosaccharides: Prebiotics and beyond. Nutr. Rev. 2009, 67, S183–S191. [Google Scholar] [CrossRef]
- Broekaert, W.F.; Courtin, C.M.; Verbeke, K.; Van de Wiele, T.; Verstraete, W.; Delcour, J.A. Prebiotic and other health-related effects of cereal-derived arabinoxylans, arabinoxylan-oligosaccharides, and xylooligosaccharides. Crit. Rev. Food Sci. Nutr. 2011, 51, 178–194. [Google Scholar] [CrossRef] [PubMed]
- Tingirikari, J.M.R. In-Vitro Prebiotic Analysis of Microbiota Accessible Pectic Polysaccharides. Curr. Microbiol. 2019, 76, 1452–1460. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Zheng, J.; Mao, G.; Hu, W.; Ye, X.; Linhardt, R.J.; Chen, S. Rethinking the impact of RG-I mainly from fruits and vegetables on dietary health. Crit. Rev. Food Sci. Nutr. 2020, 60, 2938–2960. [Google Scholar] [CrossRef]
- Wu, D.; Ye, X.; Linhardt, R.J.; Liu, X.; Zhu, K.; Yu, C.; Ding, T.; Liu, D.; He, Q.; Chen, S. Dietary pectic substances enhance gut health by its polycomponent: A review. Compr. Rev. Food Sci. Food Saf. 2021, 20, 2015–2039. [Google Scholar] [CrossRef] [PubMed]
- Luis, A.S.; Briggs, J.; Zhang, X.; Farnell, B.; Ndeh, D.; Labourel, A.; Baslé, A.; Cartmell, A.; Terrapon, N.; Stott, K.; et al. Dietary pectic glycans are degraded by coordinated enzyme pathways in human colonic Bacteroides. Nat. Microbiol. 2018, 3, 210–219. [Google Scholar] [CrossRef] [PubMed]
- McKay, S.; Oranje, P.; Helin, J.; Koek, J.H.; Kreijveld, E.; van den Abbeele, P.; Pohl, U.; Bothe, G.; Tzoumaki, M.; Aparicio-Vergara, M.; et al. Development of an Affordable, Sustainable and Efficacious Plant-Based Immunomodulatory Food Ingredient Based on Bell Pepper or Carrot RG-I Pectic Polysaccharides. Nutrients 2021, 13, 963. [Google Scholar] [CrossRef]
- Van den Abbeele, P.; Verstrepen, L.; Ghyselinck, J.; Albers, R.; Marzorati, M.; Mercenier, A. A Novel Non-Digestible, Carrot-Derived Polysaccharide (cRG-I) Selectively Modulates the Human Gut Microbiota while Promoting Gut Barrier Integrity: An Integrated In Vitro Approach. Nutrients 2020, 12, 1917. [Google Scholar] [CrossRef] [PubMed]
- Ruppin, H.; Bar-Meir, S.; Soergel, K.H.; Wood, C.M.; Schmitt, M.G. Absorption of Short-Chain Fatty Acids by the Colon. Gastroenterology 1980, 78, 1500–1507. [Google Scholar] [CrossRef]
- Boets, E.; Deroover, L.; Houben, E.; Vermeulen, K.; Gomand, S.V.; Delcour, J.A.; Verbeke, K. Quantification of in Vivo Colonic Short Chain Fatty Acid Production from Inulin. Nutrients 2015, 7, 8916–8929. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sakata, T. Pitfalls in short-chain fatty acid research: A methodological review. Anim. Sci. J. 2019, 90, 3–13. [Google Scholar] [CrossRef] [Green Version]
- Chen, T.; Long, W.; Zhang, C.; Liu, S.; Zhao, L.; Hamaker, B.R. Fiber-utilizing capacity varies in Prevotella—versus Bacteroides—dominated gut microbiota. Sci. Rep. 2017, 7, 2594. [Google Scholar] [CrossRef]
- Molly, K.; Woestyne, M.V.; Smet, I.D.; Verstraete, W. Validation of the Simulator of the Human Intestinal Microbial Ecosystem (SHIME) Reactor Using Microorganism-associated Activities. Microb. Ecol. Health Dis. 1994, 7, 191–200. [Google Scholar] [CrossRef]
- Minekus, M.; Smeets-Peeters, M.; Bernalier, A.; Marol-Bonnin, S.; Havenaar, R.; Marteau, P.; Alric, M.; Fonty, G.; Huis in’t Veld, J.H.J. A computer-controlled system to simulate conditions of the large intestine with peristaltic mixing, water absorption and absorption of fermentation products. Appl. Microbiol. Biotechnol. 1999, 53, 108–114. [Google Scholar] [CrossRef] [PubMed]
- Berner, A.Z.; Fuentes, S.; Dostal, A.; Payne, A.N.; Gutierrez, P.V.; Chassard, C.; Grattepanche, F.; de Vos, W.M.; Lacroix, C. Novel Polyfermentor Intestinal Model (PolyFermS) for Controlled Ecological Studies: Validation and Effect of pH. PLoS ONE 2013, 8, e77772. [Google Scholar] [CrossRef] [Green Version]
- Chung, W.S.F.; Walker, A.W.; Louis, P.; Parkhill, J.; Vermeiren, J.; Bosscher, D.; Duncan, S.H.; Flint, H.J. Modulation of the human gut microbiota by dietary fibres occurs at the species level. BMC Biol. 2016, 14, 3. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Salonen, A.; Lahti, L.; Salojärvi, J.; Holtrop, G.; Korpela, K.; Duncan, S.H.; Date, P.; Farquharson, F.; Johnstone, A.M.; Lobley, G.E.; et al. Impact of diet and individual variation on intestinal microbiota composition and fermentation products in obese men. ISME J. 2014, 8, 2218–2230. [Google Scholar] [CrossRef] [PubMed]
- Healey, G.R.; Murphy, R.; Brough, L.; Butts, C.A.; Coad, J. Interindividual variability in gut microbiota and host response to dietary interventions. Nutr. Rev. 2017, 75, 1059–1080. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van den Abbeele, P.; Belzer, C.; Goossens, M.; Kleerebezem, M.; De Vos, W.M.; Thas, O.; De Weirdt, R.; Kerckhof, F.-M.; Van de Wiele, T. Butyrate-producing Clostridium cluster XIVa species specifically colonize mucins in an in vitro gut model. ISME J. 2013, 7, 949–961. [Google Scholar] [CrossRef] [Green Version]
- Van den Abbeele, P.; Marzorati, M.; Derde, M.; De Weirdt, R.; Joan, V.; Possemiers, S.; Van de Wiele, T. Arabinoxylans, inulin and Lactobacillus reuteri 1063 repress the adherent-invasive Escherichia coli from mucus in a mucosa-comprising gut model. NPJ Biofilms Microbiomes 2016, 2, 16016. [Google Scholar] [CrossRef]
- Moens, F.; Duysburgh, C.; van den Abbeele, P.; Morera, M.; Marzorati, M. Lactobacillus rhamnosus GG and Saccharomyces cerevisiae boulardii exert synergistic antipathogenic activity in vitro against enterotoxigenic Escherichia coli. Benef. Microbes 2019, 10, 923–935. [Google Scholar] [CrossRef] [Green Version]
- De Weirdt, R.; Possemiers, S.; Vermeulen, G.; Moerdijk-Poortvliet, T.C.W.; Boschker, H.T.S.; Verstraete, W.; Van de Wiele, T. Human faecal microbiota display variable patterns of glycerol metabolism. FEMS Microbiol. Ecol. 2010, 74, 601–611. [Google Scholar] [CrossRef] [Green Version]
- Van de Wiele, T.; Boon, N.; Possemiers, S.; Jacobs, H.; Verstraete, W. Prebiotic effects of chicory inulin in the simulator of the human intestinal microbial ecosystem. FEMS Microbiol. Ecol. 2004, 51, 143–153. [Google Scholar] [CrossRef] [PubMed]
- Boon, N.; Top, E.M.; Verstraete, W.; Siciliano, S.D. Bioaugmentation as a Tool To Protect the Structure and Function of an Activated-Sludge Microbial Community against a 3-Chloroaniline Shock Load. Appl. Environ. Microbiol. 2003, 69, 1511–1520. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duysburgh, C.; Van den Abbeele, P.; Krishnan, K.; Bayne, T.F.; Marzorati, M. A synbiotic concept containing spore-forming Bacillus strains and a prebiotic fiber blend consistently enhanced metabolic activity by modulation of the gut microbiome in vitro. Int. J. Pharm. X 2019, 1, 100021. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L. Assessing and Improving Methods Used in Operational Taxonomic Unit-Based Approaches for 16S rRNA Gene Sequence Analysis. Appl. Environ. Microbiol. 2011, 77, 3219–3226. [Google Scholar] [CrossRef] [Green Version]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a Dual-Index Sequencing Strategy and Curation Pipeline for Analyzing Amplicon Sequence Data on the MiSeq Illumina Sequencing Platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef] [Green Version]
- Cole, J.R.; Wang, Q.; Cardenas, E.; Fish, J.; Chai, B.; Farris, R.J.; Kulam-Syed-Mohideen, A.S.; McGarrell, D.M.; Marsh, T.; Garrity, G.M.; et al. The Ribosomal Database Project: Improved alignments and new tools for rRNA analysis. Nucleic Acids Res. 2009, 37, D141–D145. [Google Scholar] [CrossRef] [Green Version]
- Hammer, O.; Harper, D.A.T.; Ryan, P.D. PAST: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontol. Electron. 2001, 4, 9. [Google Scholar]
- Vandeputte, D.; Kathagen, G.; D’hoe, K.; Vieira-Silva, S.; Valles-Colomer, M.; Sabino, J.; Wang, J.; Tito, R.Y.; De Commer, L.; Darzi, Y.; et al. Quantitative microbiome profiling links gut community variation to microbial load. Nature 2017, 551, 507–511. [Google Scholar] [CrossRef]
- Metsalu, T.; Vilo, J. ClustVis: A web tool for visualizing clustering of multivariate data using Principal Component Analysis and heatmap. Nucleic Acids Res. 2015, 43, W566–W570. [Google Scholar] [CrossRef]
- Lee, S.; Lee, D.K. What is the proper way to apply the multiple comparison test? Korean J. Anesthesiol. 2018, 71, 353–360. [Google Scholar] [CrossRef] [Green Version]
- Falony, G.; Vlachou, A.; Verbrugghe, K.; Vuyst, L.D. Cross-Feeding between Bifidobacterium longum BB536 and Acetate-Converting, Butyrate-Producing Colon Bacteria during Growth on Oligofructose. Appl. Environ. Microbiol. 2006, 72, 7835–7841. [Google Scholar] [CrossRef] [Green Version]
- Rios-Covian, D.; Gueimonde, M.; Duncan, S.H.; Flint, H.J.; de los Reyes-Gavilan, C.G. Enhanced butyrate formation by cross-feeding between Faecalibacterium prausnitzii and Bifidobacterium adolescentis. FEMS Microbiol. Lett. 2015, 362, fnv176. [Google Scholar] [CrossRef] [Green Version]
- Ng, S.K.C.; Hamilton, I.R. Lactate Metabolism by Veillonella parvula. J. Bacteriol. 1971, 105, 999–1005. [Google Scholar] [CrossRef] [Green Version]
- Ndeh, D.; Rogowski, A.; Cartmell, A.; Luis, A.S.; Baslé, A.; Gray, J.; Venditto, I.; Briggs, J.; Zhang, X.; Labourel, A.; et al. Complex pectin metabolism by gut bacteria reveals novel catalytic functions. Nature 2017, 544, 65–70. [Google Scholar] [CrossRef]
- Chung, W.S.F.; Meijerink, M.; Zeuner, B.; Holck, J.; Louis, P.; Meyer, A.S.; Wells, J.M.; Flint, H.J.; Duncan, S.H. Prebiotic potential of pectin and pectic oligosaccharides to promote anti-inflammatory commensal bacteria in the human colon. FEMS Microbiol. Ecol. 2017, 93, fix127. [Google Scholar] [CrossRef] [PubMed]
- Louis, P.; Flint, H.J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 2017, 19, 29–41. [Google Scholar] [CrossRef] [Green Version]
- Yoshida, N.; Emoto, T.; Yamashita, T.; Watanabe, H.; Hayashi, T.; Tabata, T.; Hoshi, N.; Hatano, N.; Ozawa, G.; Sasaki, N.; et al. Bacteroides vulgatus and Bacteroides dorei Reduce Gut Microbial Lipopolysaccharide Production and Inhibit Atherosclerosis. Circulation 2018, 138, 2486–2498. [Google Scholar] [CrossRef] [PubMed]
- O’Connell Motherway, M.; Fitzgerald, G.F.; van Sinderen, D. Metabolism of a plant derived galactose-containing polysaccharide by Bifidobacterium breve UCC2003. Microb. Biotechnol. 2011, 4, 403–416. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Komeno, M.; Hayamizu, H.; Fujita, K.; Ashida, H. Two Novel α-l-Arabinofuranosidases from Bifidobacterium longum subsp. longum Belonging to Glycoside Hydrolase Family 43 Cooperatively Degrade Arabinan. Appl. Environ. Microbiol. 2019, 85, e02582-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franke, T.; Deppenmeier, U. Physiology and central carbon metabolism of the gut bacterium Prevotella copri. Mol. Microbiol. 2018, 109, 528–540. [Google Scholar] [CrossRef] [Green Version]
- Wu, F.; Guo, X.; Zhang, J.; Zhang, M.; Ou, Z.; Peng, Y. Phascolarctobacterium faecium abundant colonization in human gastrointestinal tract. Exp. Ther. Med. 2017, 14, 3122–3126. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dot, T.; Osawa, R.; Stackebrandt, E. Phascolarctobacterium faecium gen. nov, spec. nov., a Novel Taxon of the Sporomusa Group of Bacteria. Syst. Appl. Microbiol. 1993, 16, 380–384. [Google Scholar] [CrossRef]
- Li, L.; Su, Q.; Xie, B.; Duan, L.; Zhao, W.; Hu, D.; Wu, R.; Liu, H. Gut microbes in correlation with mood: Case study in a closed experimental human life support system. Neurogastroenterol. Motil. Off. J. Eur. Gastrointest. Motil. Soc. 2016, 28, 1233–1240. [Google Scholar] [CrossRef] [PubMed]
- Lavelle, A.; Lennon, G.; O’Sullivan, O.; Docherty, N.; Balfe, A.; Maguire, A.; Mulcahy, H.E.; Doherty, G.; O’Donoghue, D.; Hyland, J.; et al. Spatial variation of the colonic microbiota in patients with ulcerative colitis and control volunteers. Gut 2015, 64, 1553–1561. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van den Abbeele, P.; Grootaert, C.; Marzorati, M.; Possemiers, S.; Verstraete, W.; Gérard, P.; Rabot, S.; Bruneau, A.; El Aidy, S.; Derrien, M.; et al. Microbial community development in a dynamic gut model is reproducible, colon region specific, and selective for Bacteroidetes and Clostridium cluster IX. Appl. Environ. Microbiol. 2010, 76, 5237–5246. [Google Scholar] [CrossRef] [Green Version]
- Dupont, D.; Alric, M.; Blanquet-Diot, S.; Bornhorst, G.; Cueva, C.; Deglaire, A.; Denis, S.; Ferrua, M.; Havenaar, R.; Lelieveld, J.; et al. Can dynamic in vitro digestion systems mimic the physiological reality? Crit. Rev. Food Sci. Nutr. 2019, 59, 1546–1562. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Šuligoj, T.; Vigsnæs, L.K.; den Abbeele, P.V.; Apostolou, A.; Karalis, K.; Savva, G.M.; McConnell, B.; Juge, N. Effects of Human Milk Oligosaccharides on the Adult Gut Microbiota and Barrier Function. Nutrients 2020, 12, 2808. [Google Scholar] [CrossRef]
- Marzorati, M.; Van den Abbeele, P.; Bubeck, S.; Bayne, T.; Krishnan, K.; Young, A. Treatment with a spore-based probiotic containing five strains of Bacillus induced changes in the metabolic activity and community composition of the gut microbiota in a SHIME® model of the human gastrointestinal system. Food Res. Int. 2021, 149, 110676. [Google Scholar] [CrossRef]
- Moens, F.; Van den Abbeele, P.; Basit, A.W.; Dodoo, C.; Chatterjee, R.; Smith, B.; Gaisford, S. A four-strain probiotic exerts positive immunomodulatory effects by enhancing colonic butyrate production in vitro. Int. J. Pharm. 2019, 555, 1–10. [Google Scholar] [CrossRef] [PubMed]
| Phylum | Family | Lumen | Mucus | ||
|---|---|---|---|---|---|
| PC | DC | PC | DC | ||
| Actinobacteria | Atopobiaceae | 0.43 | 0.30 | 0.11% | 0.02% |
| Bifidobacteriaceae | 0.30 | 0.02 | −6.00% | 3.98% | |
| Coriobacteriaceae | 0.23 | −0.02 | 2.16% | −0.41% | |
| Eggerthellaceae | 0.00 | −0.17 | 0.00% | −0.02% | |
| Bacteroidetes | Bacteroidaceae | 0.61 | 0.33 | 3.99% | −2.81% |
| Barnesiellaceae | 0.00 | 0.27 | 0.00% | 0.05% | |
| Marinifilaceae | 0.00 | 0.01 | 0.00% | 0.01% | |
| Muribaculaceae | 0.00 | −0.01 | 0.00% | 0.02% | |
| Prevotellaceae | 0.52 | 0.69 | 7.55% | 1.85% | |
| Rikenellaceae | −0.02 | 0.07 | −0.07% | −0.30% | |
| Tannerellaceae | 0.38 | −0.10 | −0.12% | −0.03% | |
| Firmicutes | Acidaminococcaceae | 0.47 | 0.39 | 0.46% | 0.16% |
| Christensenellaceae | 0.00 | 0.44 | 0.01% | 0.21% | |
| Clostridiaceae | 0.00 | 0.00 | −0.01% | −1.00% | |
| Erysipelotrichaceae | 0.16 | 0.68 | 0.22% | 0.18% | |
| Family XIII | 0.00 | 0.01 | 0.00% | −0.03% | |
| Lachnospiraceae | 0.24 | 0.19 | −3.83% | 0.96% | |
| Lactobacillaceae | 0.00 | 0.03 | −0.15% | 0.02% | |
| Ruminococcaceae | 0.43 | −0.21 | 0.08% | −1.15% | |
| Staphylococcaceae | 0.03 | 0.19 | 0.01% | 0.07% | |
| Veillonellaceae | −0.08 | −0.13 | −7.19% | 1.41% | |
| Proteobacteria | Burkholderiaceae | 0.18 | 0.12 | −0.36% | −0.36% |
| Desulfovibrionaceae | −0.28 | −0.06 | −0.60% | −0.75% | |
| Enterobacteriaceae | 0.06 | 0.25 | 0.32% | −0.27% | |
| Pseudomonadaceae | 0.00 | 0.00 | −0.01% | 0.00% | |
| uncultured | 0.00 | 0.64 | 0.00% | 0.20% | |
| Xanthomonadaceae | 0.39 | 0.42 | 0.04% | 0.00% | |
| Synergistetes | Synergistaceae | 0.54 | −0.09 | 3.36% | −0.22% |
| Verrucomicrobia | Akkermansiaceae | 0.00 | −0.09 | 0.00% | −0.01% |
| Phylum | Family | OTU | Related Species | Lumen | Mucus | ||
|---|---|---|---|---|---|---|---|
| PC | DC | PC | DC | ||||
| Actinobacteria | Bifidobacteriaceae | 1 | Bifidobacterium adolescentis | 0.26 | −0.01 | −1.03% | 6.64% |
| 8 | Bifidobacterium bifidum | −0.56 | −0.84 | −9.28% | −5.96% | ||
| 10 | Bifidobacterium longum | 1.32 | 1.05 | 4.13% | 3.23% | ||
| Coriobacteriaceae | 12 | Collinsella aerofaciens | 0.23 | −0.02 | 2.16% | −0.41% | |
| Bacteroidetes | Bacteroidaceae | 6 | Bacteroides dorei | 0.83 | 0.73 | 5.22% | −0.14% |
| 16 | Bacteroides ovatus | −0.38 | 0.03 | −0.22% | 0.04% | ||
| 28 | Bacteroides fragilis | −0.58 | −0.80 | −0.29% | −2.09% | ||
| 17 | Bacteroides massiliensis | −0.25 | −0.57 | −1.12% | −1.25% | ||
| 13 | Bacteroides intestinalis | −0.19 | 0.02 | −0.17% | 0.20% | ||
| Prevotellaceae | 2 | Prevotella sp. | 0.52 | 0.69 | 7.55% | 1.85% | |
| Firmicutes | Acidaminococcaceae | 11 | Phascolarctobacterium faecium | 0.47 | 0.39 | 0.46% | 0.16% |
| Clostridiaceae | 32 | Clostridium butyricum | 0.00 | 0.00 | −0.01% | −0.98% | |
| Lachnospiraceae | 5 | Clostridium clostridioforme/bolteae | 0.04 | 0.15 | −3.74% | 2.27% | |
| 9 | Lachnoclostridium sp. | 0.57 | 0.16 | −2.16% | −0.09% | ||
| 14 | Eubacterium contortum | 0.00 | 0.04 | 0.00% | −1.44% | ||
| 25 | Clostridium sp. | 0.34 | 0.32 | 0.10% | 0.11% | ||
| Ruminococcaceae | 19 | Gemmiger formicilis | 0.36 | −0.05 | 0.06% | −0.74% | |
| 42 | Faecalibacterium prausnitzii | 0.00 | 0.00 | 0.00% | −0.10% | ||
| Veillonellaceae | 3 | Megamonas funiformis | 0.05 | −0.01 | −1.70% | 0.52% | |
| 4 | Megamonas funiformis | 0.06 | 0.01 | −0.26% | 1.21% | ||
| 18 | Megasphaera sp. | −0.05 | −0.13 | −1.78% | −0.02% | ||
| 26 | Anaeroglobus geminatus | −0.01 | −0.24 | −1.49% | −0.21% | ||
| Proteobacteria | Pseudomonadaceae | 15 | Pseudomonas aeruginosa | 0.31 | −0.06 | 0.04% | −1.88% |
| Synergistetes | Synergistaceae | 7 | Cloacibacillus sp. | 0.54 | −0.09 | 3.36% | 0.04% |
| Verrucomicrobia | Akkermansiaceae | 20 | Akkermansia muciniphila | 0.00 | −0.09 | 0.00% | −0.01% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Van den Abbeele, P.; Duysburgh, C.; Cleenwerck, I.; Albers, R.; Marzorati, M.; Mercenier, A. Consistent Prebiotic Effects of Carrot RG-I on the Gut Microbiota of Four Human Adult Donors in the SHIME® Model despite Baseline Individual Variability. Microorganisms 2021, 9, 2142. https://doi.org/10.3390/microorganisms9102142
Van den Abbeele P, Duysburgh C, Cleenwerck I, Albers R, Marzorati M, Mercenier A. Consistent Prebiotic Effects of Carrot RG-I on the Gut Microbiota of Four Human Adult Donors in the SHIME® Model despite Baseline Individual Variability. Microorganisms. 2021; 9(10):2142. https://doi.org/10.3390/microorganisms9102142
Chicago/Turabian StyleVan den Abbeele, Pieter, Cindy Duysburgh, Ilse Cleenwerck, Ruud Albers, Massimo Marzorati, and Annick Mercenier. 2021. "Consistent Prebiotic Effects of Carrot RG-I on the Gut Microbiota of Four Human Adult Donors in the SHIME® Model despite Baseline Individual Variability" Microorganisms 9, no. 10: 2142. https://doi.org/10.3390/microorganisms9102142
APA StyleVan den Abbeele, P., Duysburgh, C., Cleenwerck, I., Albers, R., Marzorati, M., & Mercenier, A. (2021). Consistent Prebiotic Effects of Carrot RG-I on the Gut Microbiota of Four Human Adult Donors in the SHIME® Model despite Baseline Individual Variability. Microorganisms, 9(10), 2142. https://doi.org/10.3390/microorganisms9102142






