Norwegian Soils and Waters Contain Mesophilic, Plastic-Degrading Bacteria
Abstract
1. Introduction
2. Materials and Methods
2.1. Samples and Isolation and Enumeration of Bacteria
2.2. Plastic-Containing Agar
2.3. PCR Amplification and Sequencing Studies
2.4. Broth-Based Investigation of PET-Degrading Activity
2.5. API®ZYM (bioMérieux Inc.)
2.6. API®50CH (bioMérieux Inc.)
2.7. Assays for Additional Enzyme Activities
3. Results and Discussion
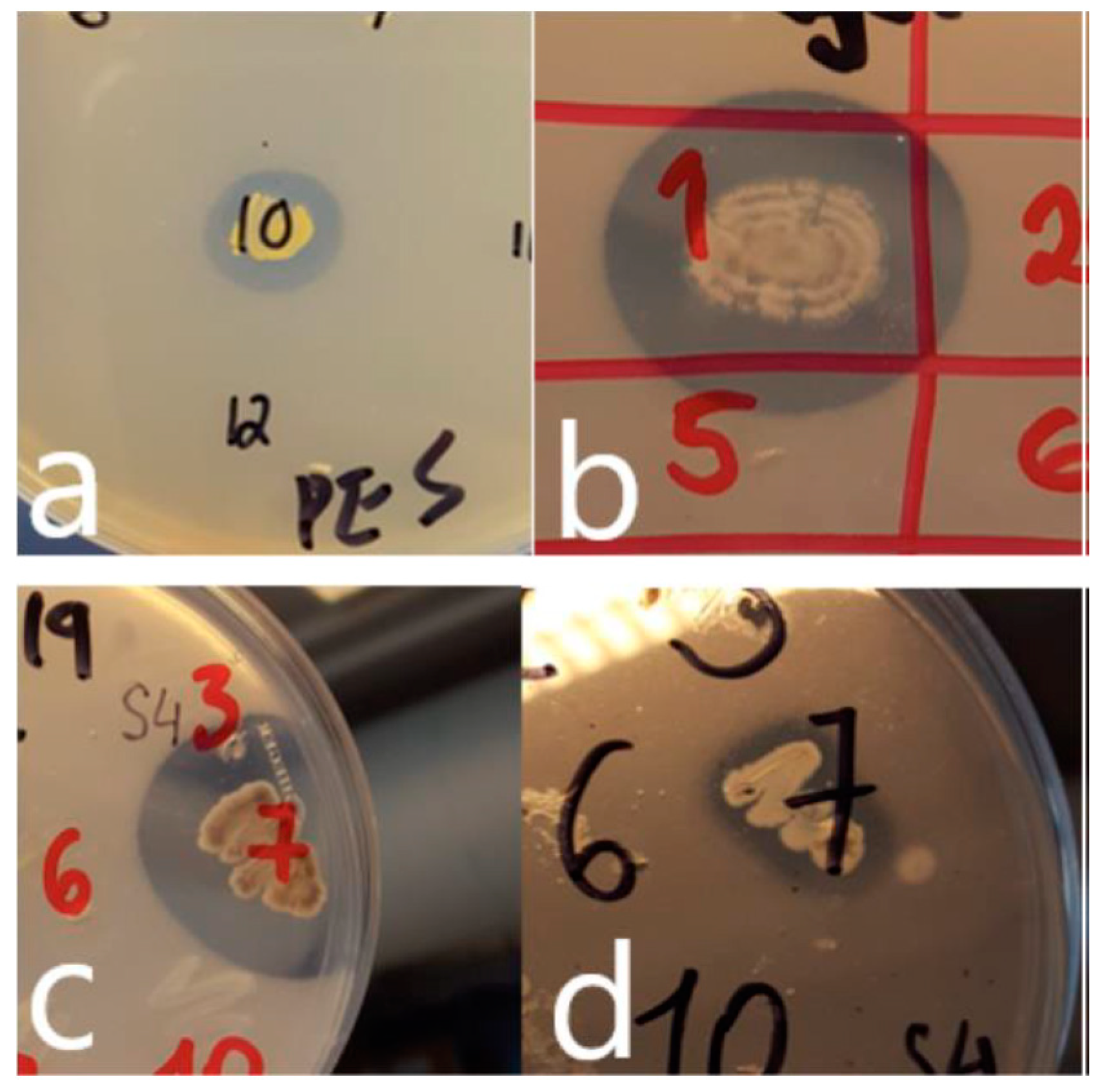
3.1. Numbers, Identities and Range of Activities of Plastic Degrading Bacteria
3.2. Additional Characterizations of Multi-Polymer Degrading Rhodococcus and Streptomyces
3.2.1. API® ZYM and Correlations to Observed Lipid/Polysaccharide Hydrolysis
3.2.2. API®50CH
3.2.3. Additional Enzyme Tests
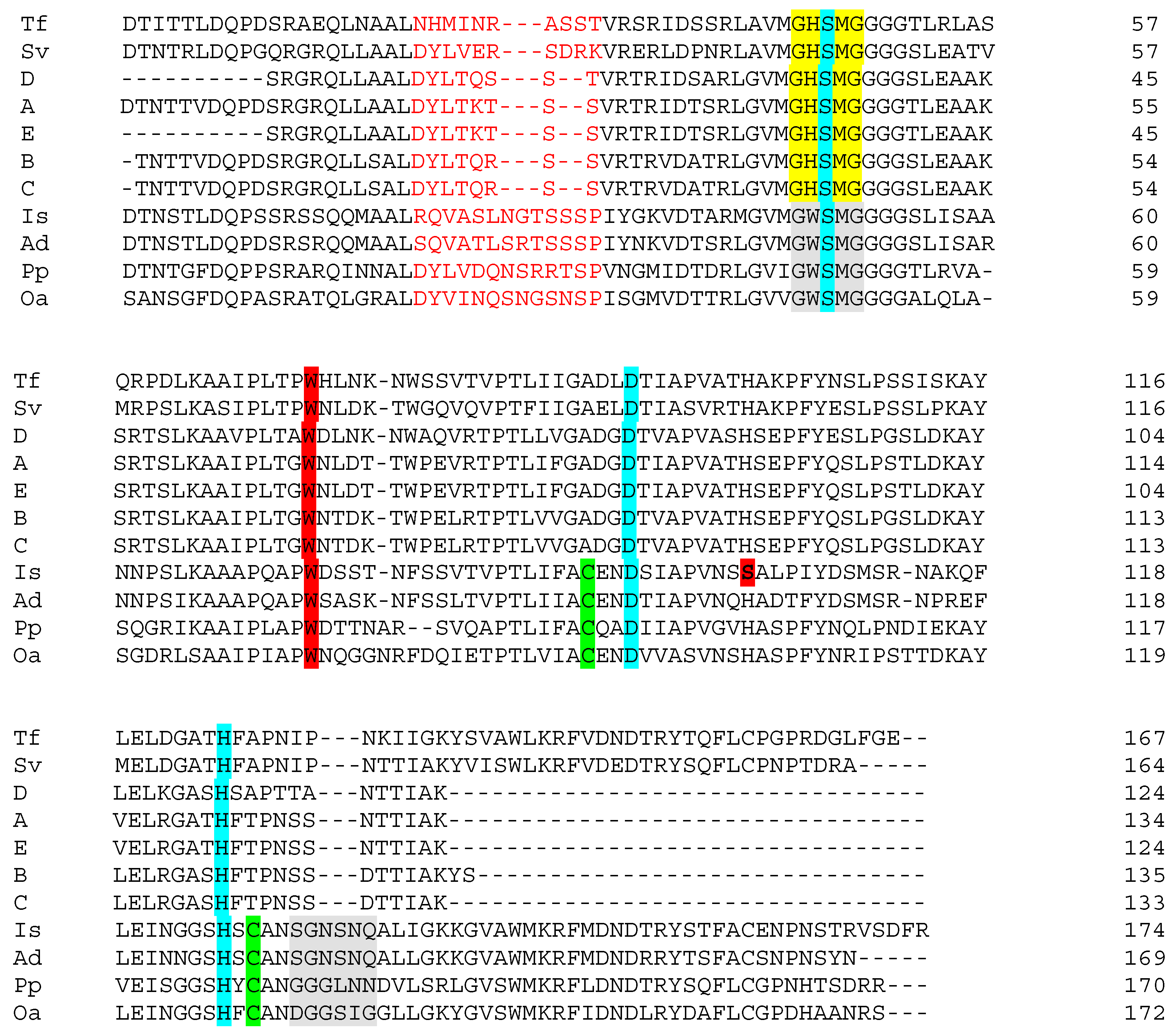
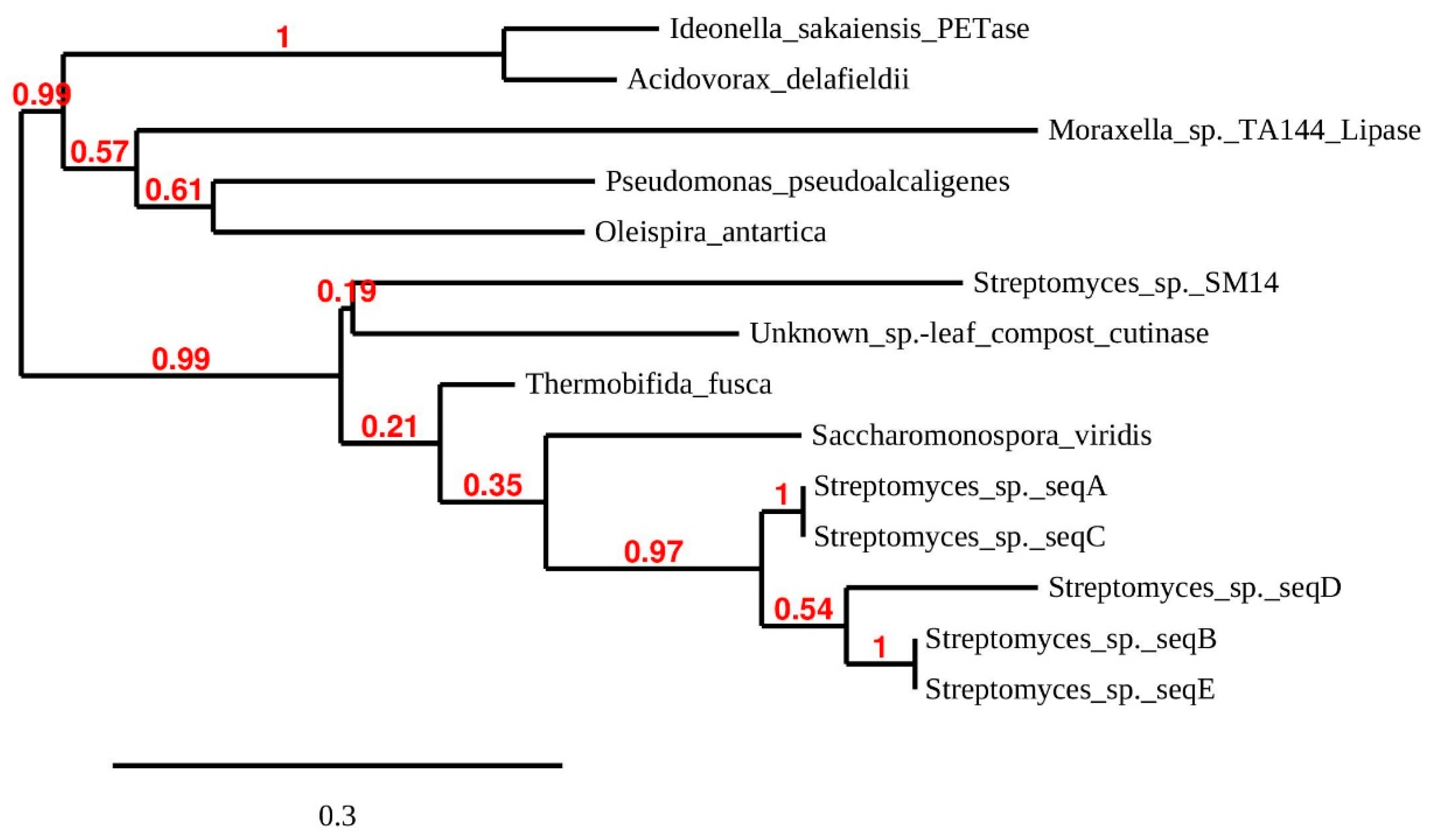
3.3. Presence of Putative PET-Hydrolase Genes
4. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Urbanek, A.K.; Rymowicz, W.; Strzelecki, M.C.; Kociuba, W.; Franczak, Ł.; Mirończuk, A.M. Isolation and characterization of Arctic microorganisms decomposing bioplastics. AMB Express 2017, 7, 1–10. [Google Scholar] [CrossRef]
- Wilkes, R.A.; Aristilde, L. Degradation and metabolism of synthetic plastics and associated products by Pseudomonas sp.: capabilities and challenges. J. Appl. Microbiol. 2017, 123, 1582–1593. [Google Scholar] [CrossRef] [PubMed]
- Tokiwa, Y.; Calabia, B.P.; Ugwu, C.U.; Aiba, S. Biodegradability of plastics. Int. J. Mol. Sci. 2009, 10, 3722–3742. [Google Scholar] [CrossRef] [PubMed]
- Anjana, K.; Hinduja, M.; Sujitha, K.; Dharani, G. Review on plastic wastes in marine environment–Biodegradation and biotechnological solutions. Mar. Pollutt. Bull. 2020, 150, 110733. [Google Scholar]
- Waring, R.H.; Harris, R.M.; Mitchell, S.C. Plastic contamination of the food chain: A threat to human health? Maturitas 2018, 115, 64–68. [Google Scholar] [CrossRef]
- Lehner, R.; Weder, C.; Petri-Fink, A.; Rothen-Rutishauser, B. Emergence of nanoplastic in the environment and possible impact on human health. Environ. Sci. Technol. 2019, 53, 1748–1765. [Google Scholar] [CrossRef]
- Danso, D.; Chow, J.; Streit, W.R. Plastics: environmental and biotechnological perspectives on microbial degradation. Appl. Environ. Microbiol. 2019, 85, e01095-19. [Google Scholar] [CrossRef]
- Shah, A.A.; Hasan, F.; Hameed, A.; Ahmed, S. Biological degradation of plastics: a comprehensive review. Biotechnol. Adv. 2008, 26, 246–265. [Google Scholar] [CrossRef]
- Chen, C.C.; Han, X.; Ko, T.P.; Liu, W.; Guo, R.T. Structural studies reveal the molecular mechanism of PETase. FEBS J. 2018, 285, 3717–3723. [Google Scholar] [CrossRef]
- Nakajima-Kambe, T.; Ichihashi, F.; Matsuzoe, R.; Kato, S.; Shintani, N. Degradation of aliphatic–aromatic copolyesters by bacteria that can degrade aliphatic polyesters. Polym. Degrad. Stab. 2009, 94, 1901–1905. [Google Scholar] [CrossRef]
- Müller, R.J.; Kleeberg, I.; Deckwer, W.D. Biodegradation of polyesters containing aromatic constituents. J. Biotechnol. 2001, 86, 87–95. [Google Scholar] [CrossRef]
- Mouriño, V.; Boccaccini, A.R. Bone tissue engineering therapeutics: controlled drug delivery in three-dimensional scaffolds. J. R. Soc. Interface 2010, 7, 209–2027. [Google Scholar] [CrossRef] [PubMed]
- Müller, R.J.; Schrader, H.; Profe, J.; Dresler, K.; Deckwer, W.D. Enzymatic degradation of poly (ethylene terephthalate): rapid hydrolyse using a hydrolase from T. fusca. Macromol. Rapid Commun. 2005, 26, 1400–1405. [Google Scholar] [CrossRef]
- Joo, S.; Cho, I.J.; Seo, H.; Son, H.F.; Sagong, H.Y.; Shin, T.; Choi, S.Y.; Lee, S.Y.; Kim, K.J. Structural insight into molecular mechanism of poly (ethylene terephthalate) degradation. Nat. Commun. 2018, 9, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, S.; Hiraga, K.; Takehana, T.; Taniguchi, I.; Yamaji, H.; Maeda, Y.; Toyohara, K.; Miyamoto, K.; Kimura, Y.; Oda, K. A bacterium that degrades and assimilates poly (ethylene terephthalate). Science 2016, 351, 1196–1199. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; He, L.; Wang, L.; Li, T.; Li, C.; Liu, H.; Bao, R. Protein crystallography and site-direct mutagenesis analysis of the poly (ethylene terephthalate) hydrolase PETase from Ideonella sakaiensis. ChemBioChem 2018, 19, 1471–1475. [Google Scholar] [CrossRef]
- Han, X.; Liu, W.; Huang, J.W.; Ma, J.; Zheng, Y.; Ko, T.P.; Xu, L.; Cheng, Y.S.; Chen, C.C.; Guo, R. T Structural insight into catalytic mechanism of PET hydrolase. Nat. Commun. 2017, 8, 1–6. [Google Scholar] [CrossRef]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Söding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Sayers, E.W.; Agarwala, R.; Bolton, E.E.; Brister, J.R.; Canese, K.; Clark, K.; Connor, R.; Fiorini, N.; Funk, K.; Hefferon, T.; et al. Database resources of the national center for biotechnology information. Nucleic Acids Res. 2019, 47, D23. [Google Scholar] [CrossRef] [PubMed]
- Pearson, W.R. Rapid and sensitive sequence comparison with FASTP and FASTA. Methods Enzymol. 1990, 183, 63–98. [Google Scholar] [PubMed]
- UniProt Consortium. UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. 2019, 47, D506-15. [Google Scholar]
- Dereeper, A.; Guignon, V.; Blanc, G.; Audic, S.; Buffet, S.; Chevenet, F.; Dufayard, J.F.; Guindon, S.; Lefort, V.; Lescot, M.; et al. Phylogeny. fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008, 1, W465–W469. [Google Scholar] [CrossRef]
- Dereeper, A.; Audic, S.; Claverie, J.M.; Blanc, G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol. Biol. 2010, 10, 8. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 2004, 5, 113–119. [Google Scholar] [CrossRef]
- Castresana, J. Selection of conserved blocks for multiple alignments for their use in phylogenetic alignments. Mol. Biol. Evol. 2000, 17, 540–552. [Google Scholar] [CrossRef]
- Guindon, S.; Gascuel, O. A simple. Fast and accurate alogorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef]
- Anisimova, M.; Gascuel, O. Approximate likelihood ratio test for branches: a fast accurate and powerful alternative. Syst. Biol. 2006, 55, 539–552. [Google Scholar] [CrossRef]
- Whelan, S.; Goldman, N. A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol. Biol. Evol. 2001, 18, 691–699. [Google Scholar] [CrossRef]
- Chevenet, F.; Brun, C.; Bañuls, A.L.; Jacq, B.; Christen, R. TreeDyn: Towards dynamic graphics and annotations for analyses of trees. BMC Bioinform. 2006, 7, 439. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.K.; Pal, S.; Ray, S. Study of microbes having potentiality for biodegradation of plastics. Environ. Sci. Pollut. Res. 2013, 20, 4339–4355. [Google Scholar] [CrossRef] [PubMed]
- Penkhrue, W.; Khanongnuch, C.; Masaki, K.; Pathom-aree, W.; Punyodom, W.; Lumyong, S. Isolation and screening of biopolymer-degrading microorganisms from northern Thailand. World J. Microbiol. Biotechnol. 2015, 31, 1431–1442. [Google Scholar] [CrossRef] [PubMed]
- Nishida, H.; Yutaka, T. Distribution of poly (β-hydroxybutyrate) and poly (ε-caprolactone) aerobic degrading microorganisms in different environments. J. Environ. Polym Degrad. 1993a, 1, 227–233. [Google Scholar] [CrossRef]
- Chen, Q.; Wang, Q.; Wei, G.; Liang, G.; Qi, Q. Production in Escherichia coli of poly (3-hydroxybutyrate-co-3-hydroxyvalerate) with differing monomer compositions from unrelated carbon sources. Appl. Environ. Microbiol. 2011, 77, 4886–4893. [Google Scholar] [CrossRef] [PubMed]
- Weng, Y.X.; Wang, X.L.; Wang, Y.Z. Biodegradation behavior of PHAs with different chemical structures under controlled composting conditions. Polym. Test. 2011, 30, 372–380. [Google Scholar] [CrossRef]
- Boyandin, A.N.; Rudnev, V.P.; Ivonin, V.N.; Prudnikova, S.V.; Korobikhina, K.I.; Filipenko, M.L.; Volova, T.G.; Sinskey, A.J. Biodegradation of polyhydroxyalkanoate films in natural environments. Macromol. Symp. 2012, 320, 38–42. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, R.; Cai, J.; Liu, Z.; Zheng, Y. Biosynthesis and thermal properties of PHBV produced from levulinic acid by Ralstonia eutropha. PLoS ONE 2013, 8, e60318. [Google Scholar] [CrossRef]
- Shang, L.; Fei, Q.; Zhang, Y.H.; Wang, X.Z.; Fan, D.D.; Chang, H.N. Thermal properties and biodegradability studies of poly (3-hydroxybutyrate-co-3-hydroxyvalerate). J. Polym. Environ. 2012, 20, 23–28. [Google Scholar] [CrossRef]
- Iram, D.; Riaz, R.A.; Iqbal, R.K. Usage of Potential Micro-organisms for Degradation of Plastics. Open J. Environ. Biol 2019, 4, 007–0015. [Google Scholar]
- Suyama, T.; Tokiwa, Y.; Oichanpagdee, P.; Kanagawa, T.; Kamagata, Y. Phylogenetic affiliation of soil bacteria that degrade aliphatic polyesters available commercially as biodegradable plastics. Appl. Environ. Microbiol. 1998a, 64, 5008–5011. [Google Scholar] [CrossRef] [PubMed]
- Pranamuda, H.; Tokiwa, Y.; Tanaka, H. Polylactide degradation by an Amycolatopsis sp. Appl. Environ. Microbiol. 1997, 63, 1637–1640. [Google Scholar] [CrossRef] [PubMed]
- Tansengco, M.L.; Tokiwa, Y. Thermophilic microbial degradation of polyethylene succinate. World J. Microbiol. Biotechnol. 1998a, 14, 133–138. [Google Scholar] [CrossRef]
- Goethals, K.; Vereecke, D.; Jaziri, M.; Van Montagu, M.; Holsters, M. Leafy gall formation by Rhodococcus fascians. Ann. Rev. Phytopathol. 2001, 39, 27–52. [Google Scholar] [CrossRef] [PubMed]
- Almeida, E.L.; Carrillo Rincon, A.F.; Jackson, S.A.; Dobson, A.D. In silico screening and heterologous expression of a Polyethylene Terephthalate hydrolase (PETase)-like enzyme (SM14est) with Polycaprolactone (PCL)-degrading activity, from the marine sponge-derived strain Streptomyces sp. SM14. Front. Microbiol. 2019, 10, 2187. [Google Scholar] [CrossRef]
- Zucchi, T.D.; Kim, B.Y.; Kshetrimayum, J.D.; Weon, H.Y.; Kwon, S.W.; Goodfellow, M. Streptomyces brevispora sp. nov. and Streptomyces laculatispora sp. nov., actinomycetes isolated from soil. Int. J. Syst. Evol. 2012, 62, 478–483. [Google Scholar] [CrossRef] [PubMed]
- Calabia, B.P.; Tokiwa, Y. Microbial degradation of poly (D-3-hydroxybutyrate) by a new thermophilic Streptomyces isolate. Biotechnol. Lett. 2004, 26, 15–19. [Google Scholar] [CrossRef]
- Shao, H.; Chen, M.; Fei, X.; Zhang, R.; Zhong, Y.; Ni, W.; Tao, X.; He, X.Y.; Zhang, E.; Yong, B.; et al. Complete genome sequence and characterization of a polyethylene biodegradation strain, Streptomyces Albogriseolus LBX-2. Microorganisms 2019, 7, 379. [Google Scholar] [CrossRef]
- Kouker, G.; Jaeger, K.E. Specific and sensitive plate assay for bacterial lipases. Appl. Environ. Microbiol. 1987, 53, 211–213. [Google Scholar] [CrossRef]
- Nikolaivits, E.; Kanelli, M.; Dimarogona, M.; Topakas, E. A middle-aged enzyme still in its prime: recent advances in the field of cutinases. Catalysts 2018, 8, 612. [Google Scholar] [CrossRef]
- Sankararaman, S.; Sferra, T.J. Are we going nuts on coconut oil? Curr. Nutr. Rep. 2018, 7, 107–115. [Google Scholar] [CrossRef] [PubMed]
- Nesterenko, O.A.; Nogina, T.M.; Kasumova, S.A.; Kvasnikov, E.I.; Batrakov, S.G. Rhodococcus luteus nom. nov. and Rhodococcus maris nom. nov. Int. J. Syst. Evol. 1982, 32, 1–14. [Google Scholar]
- Poelarends, G.J.; Zandstra, M.; Bosma, T.; Kulakov, L.A.; Larkin, M.J.; Marchesi, J.R.; Weightman, A.J.; Janssen, D.B. Haloalkane-utilizing Rhodococcus strains isolated from geographically distinct locations possess a highly conserved gene cluster encoding haloalkane catabolism. J. Bacteriol. 2000, 182, 2725–2731. [Google Scholar] [CrossRef] [PubMed]
- Rajesh, T.; Jeon, J.M.; Kim, Y.H.; Kim, H.J.; Park, S.H.; Choi, K.Y.; Kim, Y.G.; Kim, J.; Jung, S.; Park, H.Y. Functional analysis of the gene SCO1782 encoding Streptomyces hemolysin (S-hemolysin) in Streptomyces coelicolor M145. Toxicon 2013, 71, 159–165. [Google Scholar] [CrossRef]
- Sulaiman, S.; Yamato, S.; Kanaya, E.; Kim, J.J.; Koga, Y.; Takano, K.; Kanaya, S. Isolation of a novel cutinase homolog with polyethylene terephthalate-degrading activity from leaf-branch compost by using a metagenomic approach. Appl. Environ. Microbiol. 2012, 78, 1556–1562. [Google Scholar] [CrossRef]
- Jabloune, R.; Khalil, M.; Moussa, I.E.B.; Simao-Beaunoir, A.M.; Lerat, S.; Brzezinski, R.; Beaulieu, C. Enzymatic Degradation of p-Nitrophenyl Esters, Polyethylene Terephthalate, Cutin, and Suberin by Sub1, a Suberinase Encoded by the Plant Pathogen Streptomyces scabies. Microbes Environ. 2020, 35, ME19086. [Google Scholar] [CrossRef]


| Sample | Sampling Site | Coordinates | Sample Appearance | Additional Information |
|---|---|---|---|---|
| 1 | Soil from immediate vicinity of a small factory making plastic products to order. | 59°40′12.8″N 10°47′45.0″E | Dry soil, with a dark appearance. | Sample was taken a few cm beneath the top soil. |
| 2 | Bank of a stream passing through an industrial area, close (20 m) to a small factory making plastic products to order | 59°42′33.0″N 10°51′10.7″E | Brown, wet material apparent mix of soil and sand. | Sample was taken a few cm beneath the top layer |
| 3 | Natural marsh, several km from housing/industry. | 59°34′12.1″N 10°40′17.7″E | Dark brown, watery sediment. | Sample taken under shallow water. |
| 4 | Bank of a stream running through housing estate and construction site. | 59°33′51.1″N 10°44′02.6″E | Clay-like material. | Sample was taken a few cm beneath the top layer |
| 5 | Bank of a river passing through agricultural area. | 59°36′34.1″N 10°45′51.2″E | Brown, watery material with plant debris. | Sample was taken a few cm beneath the top layer |
| 6 | Edge of a small ditch below the outlet of a drainage pipe probably leading water from housing area and surroundings. | 59°36′30.6″N 10°45′55.4″E | Mixture of sand and clay-like material. | Sample was taken a few cm beneath the top layer |
| 7 | Bank of a stream about 100 m in the terrain below a sewage-treatment plant. | 59°34′47.2″N 10°39′13.7″E | Light, finely-grained sand mixed with clay-like material. | Sample was taken a few cm beneath the top layer. |
| 8 | River bank in an urban area close to outlet to the Oslo fjord. | 59°54′45.7″N 10°45′26.8″E | Coarse mix of sand and clay-like material. | Sample was taken a few cm beneath the top layer. |
| 9 | River bank in an urban/industrialized area. | 59°54′46.8″N 10°49′40.7″E | Mix of sand and clay-like material. | Sample taken from top layer of sediment |
| Agar | Isolate i–xi (sample of origin 1–9;—Table 1) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| i * | ii * | iii (1) | iv (9) | v (7) | vi (7) | vii (7) | viii (4) | ix (5) | x (5) | xi (3) | |
| PHB | + | + | − | + | + | + | − | + | − | − | + |
| PHBV | + | + | + | + | + | + | − | + | + | − | + |
| PES | − | − | − | − | − | − | + | + | − | − | − |
| PBS | − | − | − | − | − | − | + | [+] | − | − | − |
| PCL | − | [+] | − | − | − | − | [+] | + | − | [+] | − |
| PLLA | − | − | − | − | − | − | − | − | − | − | − |
| RES | − | − | − | − | − | − | + | + | − | + | − |
| Starch (amylase) | − | + | − | + | + | − | − | + | + | − | + |
| CMC (cellulase) | − | + | + | + | + | − | − | + | ‘ | − | + |
| DNA (DNase) | − | − | − | − | − | − | − | − | − | − | − |
| Protein (protease) | − | + | − | − | − | − | − | + | − | + | − |
| Tributyrin (esterase) | + | + | + | [+] | + | − | + | + | + | − | + |
| Olive oil (lipase) | − | + | − | − | − | − | − | − | − | − | − |
| Coconut oil (lipase) | − | [+] | − | − | − | − | + | − | − | − | − |
| Number + | 3 | 9 | 3 | 5 | 5 | 2 | 6 | 10 | 3 | 3 | 5 |
| PCR product (PET-primers) | − | − | − | + | − | − | − | + | − | − | + |
| Isolate | Sequence Length (bp) | Phylogenetic Affiliation * | Closest Species (Accession Number) | Identity-% (Query Coverage) |
|---|---|---|---|---|
| i | 625 | Acidovorax | A. facilis LMG2193 (EU024133) | 99.36 (100) |
| ii | 526 | Streptomyces | S. similanensis KC-106 ** (AB773850) | 99.62 (100) |
| iii | 508 | Streptomycetaceae | S. urticae NEAU-PCY-1 (KY788226) | 99.80 (100) |
| iv | 540 | Streptomyces | S. lunaelactis MM109 ** (KM207217) | 100 (100) |
| v | 592 | Streptomyces | S. alboniger DSM40043 ** (AY845349) | 99.49 (100) |
| vi | 510 | Pseudomonas | P. floridensis GEV388 ** (KY614191) | 100 (100) |
| vii | 731 | Rhodococcus | R. fascians CF17 (X79186) | 99.86 (100) |
| viii | 1403 | Streptomyces | S. brevispora BK160 (FR692104) | 99.86 (100) |
| ix | 528 | Oxalobacteraceae | Rugamonas rubra MOM 28/2/79 (HM038005) | 99.43 (99) |
| x | 422 | Nakamurella | N. lactea ** DSM19367 (HE599561) | 98.82 (100) |
| xi | 951 | Streptomyces | S. fulvissimus DSM40593 ** | 100 (100) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Charnock, C. Norwegian Soils and Waters Contain Mesophilic, Plastic-Degrading Bacteria. Microorganisms 2021, 9, 94. https://doi.org/10.3390/microorganisms9010094
Charnock C. Norwegian Soils and Waters Contain Mesophilic, Plastic-Degrading Bacteria. Microorganisms. 2021; 9(1):94. https://doi.org/10.3390/microorganisms9010094
Chicago/Turabian StyleCharnock, Colin. 2021. "Norwegian Soils and Waters Contain Mesophilic, Plastic-Degrading Bacteria" Microorganisms 9, no. 1: 94. https://doi.org/10.3390/microorganisms9010094
APA StyleCharnock, C. (2021). Norwegian Soils and Waters Contain Mesophilic, Plastic-Degrading Bacteria. Microorganisms, 9(1), 94. https://doi.org/10.3390/microorganisms9010094




