Simultaneous Oxidation of Atmospheric Methane, Carbon Monoxide and Hydrogen for Bacterial Growth
Abstract
1. Introduction
2. Materials and Methods
2.1. Cultivation
2.2. Temperature Selection
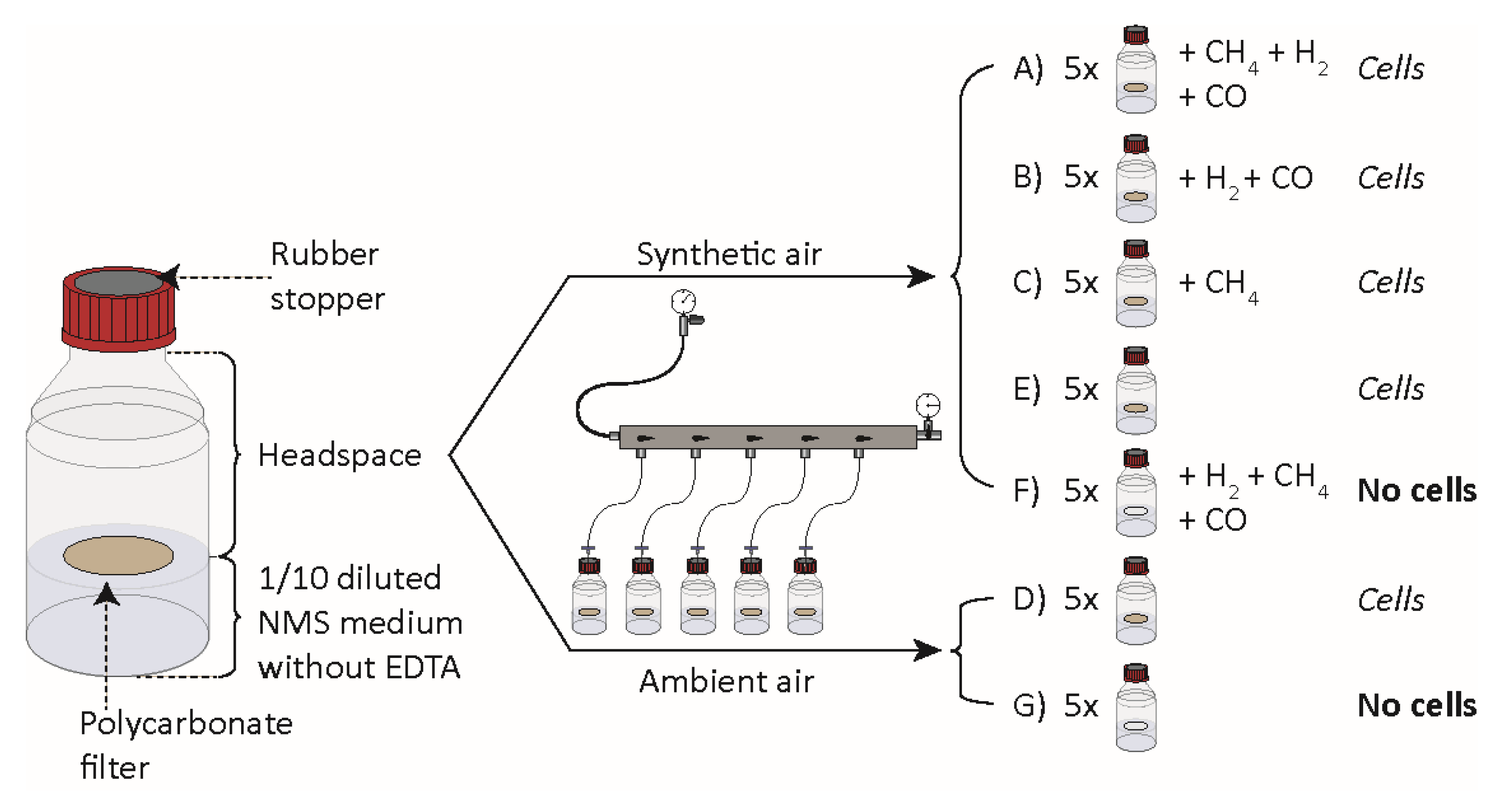
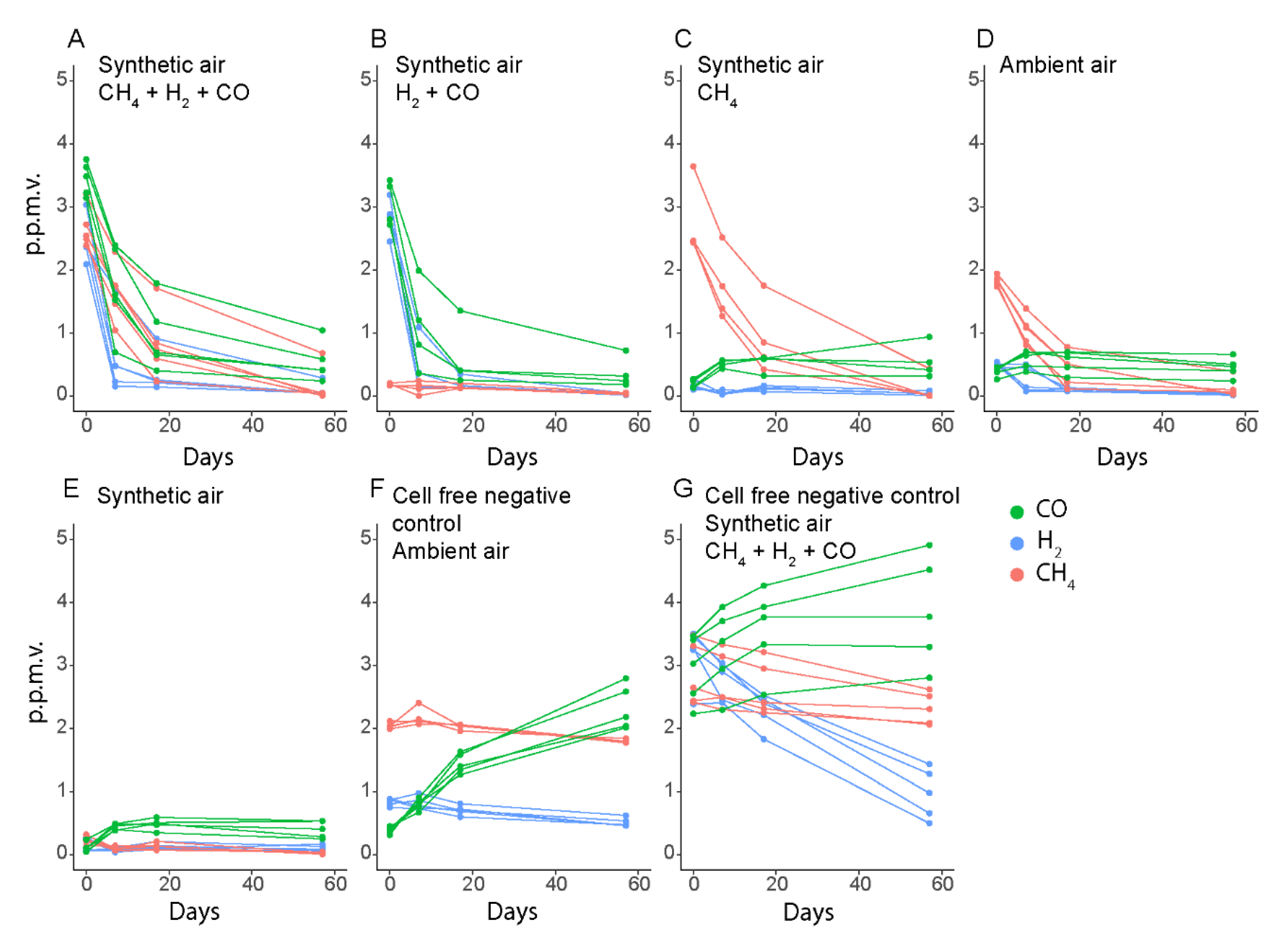
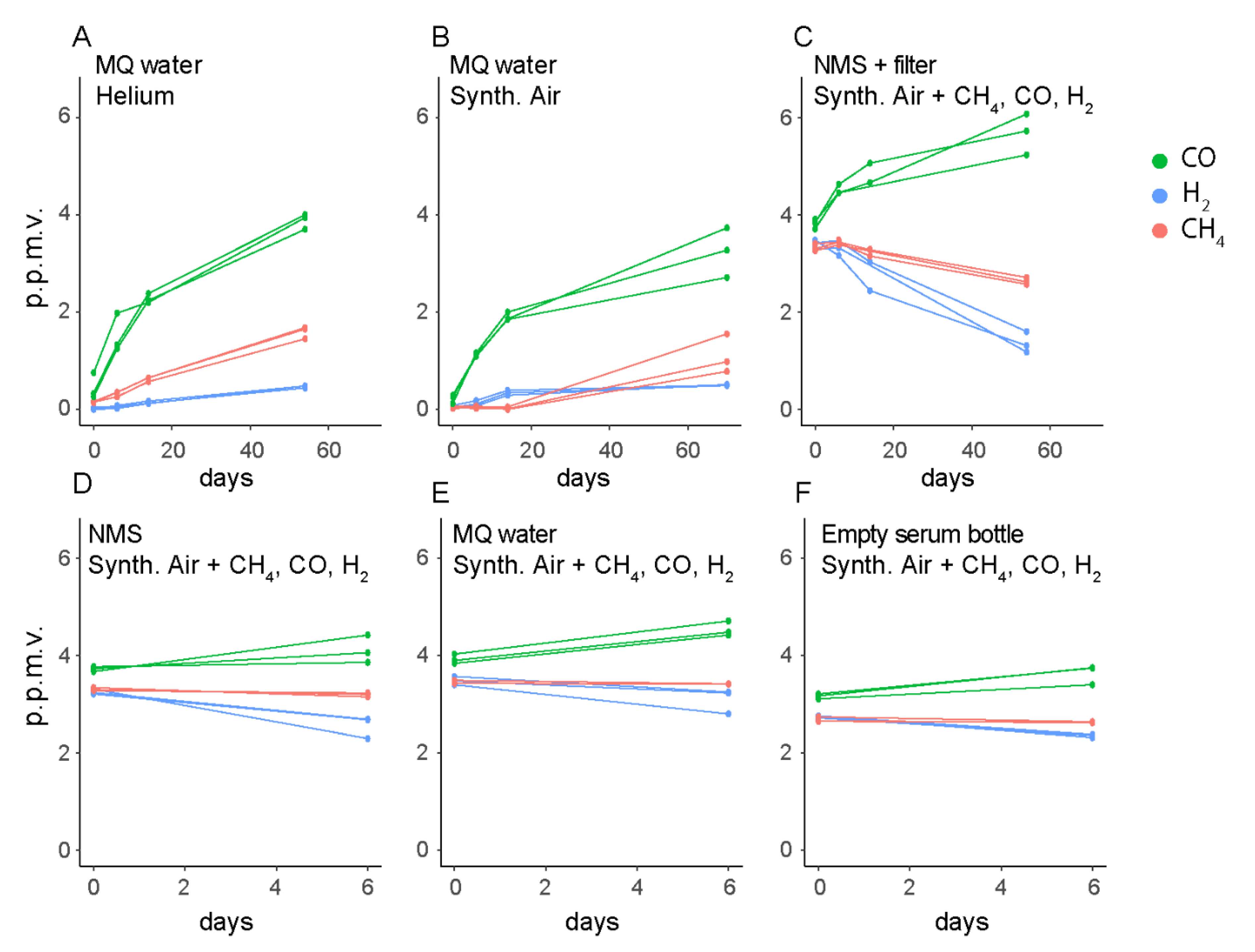
2.3. Gas Uptake and Leakage Experiments
2.4. Cell Quantification
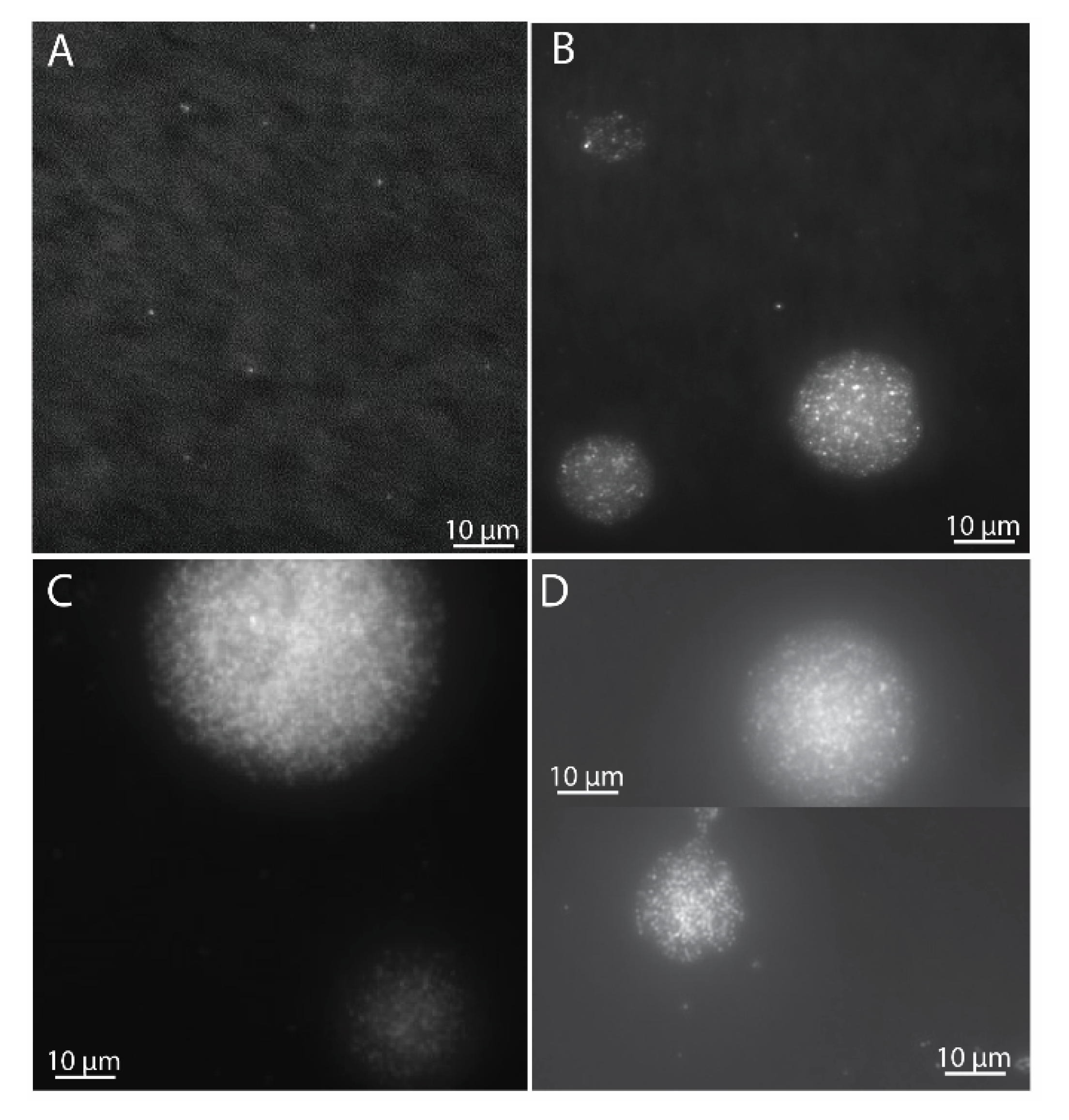
2.5. Contamination Tests and Microscopy
2.6. Plotting and Statistics
2.7. Cell-Specific Oxidation Rates and Free Energy Yield Calculations
2.8. Estimating Cell Dry Weight
3. Results and Discussion
Cell-Specific Oxidation Rates and Energy Yield
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Van Amstel, A. Methane. A review. J. Integr. Environ. Sci. 2012, 9, 5–30. [Google Scholar] [CrossRef]
- Knief, C.; Dunfield, P.F. Response and adaptation of different methanotrophic bacteria to low methane mixing ratios. Environ. Microbiol. 2005, 7, 1307–1317. [Google Scholar] [CrossRef] [PubMed]
- Dunfield, P.F. The Soil Methane Sink. Greenhouse Gas. Sinks; Reay, D.S., Hewitt, C.N., Smith, K.A., Grace, J., Eds.; CABI: Wallingford, UK, 2007; pp. 152–170. [Google Scholar]
- Tveit, A.T.; Hestnes, A.G.; Robinson, S.L.; Schintlmeister, A.; Dedysh, S.N.; Jehmlich, N.; Von Bergen, M.; Herbold, C.; Wagner, M.; Richter, A.; et al. Widespread soil bacterium that oxidizes atmospheric methane. Proc. Natl. Acad. Sci. USA 2019, 116, 8515–8524. [Google Scholar] [CrossRef] [PubMed]
- Knief, C. Diversity and habitat preferences of cultivated and uncultivated aerobic methanotrophic bacteria evaluated based on pmoA as molecular marker. Front. Microbiol. 2015, 6. [Google Scholar] [CrossRef]
- Kolb, S.; Knief, C.; Dunfield, P.F.; Conrad, R. Abundance and activity of uncultured methanotrophic bacteria involved in the consumption of atmospheric methane in two forest soils. Environ. Microbiol. 2005, 7, 1150–1161. [Google Scholar] [CrossRef]
- Bender, M.; Conrad, R. Kinetics of CH4 oxidation in oxic soils exposed to ambient air or high CH4 mixing ratios. FEMS Microbiol. Lett. 1992, 101, 261–270. [Google Scholar] [CrossRef]
- Tijhuis, L.; Van Loosdrecht, M.C.M.; Heijnen, J.J. A thermodynamically based correlation for maintenance gibbs energy requirements in aerobic and anaerobic chemotrophic growth. Biotechnol. Bioeng. 1993, 42, 509–519. [Google Scholar] [CrossRef]
- Cordero, P.R.F.; Bayly, K.; Leung, P.M.; Huang, C.; Islam, Z.F.; Schittenhelm, R.B.; King, G.M.; Greening, C. Atmospheric carbon monoxide oxidation is a widespread mechanism supporting microbial survival. ISME J. 2019, 13, 2868–2881. [Google Scholar] [CrossRef]
- Ji, M.; Greening, C.; VanWonterghem, I.; Carere, C.R.; Bay, S.K.; Steen, J.A.; Montgomery, K.; Lines, T.; Beardall, J.; Van Dorst, J.; et al. Atmospheric trace gases support primary production in Antarctic desert surface soil. Nature 2017, 552, 400–403. [Google Scholar] [CrossRef]
- Greening, C.; Carere, C.R.; Rushton-Green, R.; Harold, L.K.; Hards, K.; Taylor, M.C.; Morales, S.E.; Stott, M.B.; Cook, G.M. Persistence of the dominant soil phylum Acidobacteria by trace gas scavenging. Proc. Natl. Acad. Sci. USA 2015, 112, 10497. [Google Scholar] [CrossRef]
- Greening, C.; Berney, M.; Hards, K.; Cook, G.M.; Conrad, R. A soil actinobacterium scavenges atmospheric H2 using two membrane-associated, oxygen-dependent [NiFe] hydrogenases. Proc. Natl. Acad. Sci. USA 2014, 111, 4257. [Google Scholar] [CrossRef] [PubMed]
- Greening, C.; Biswas, A.; Carere, C.R.; Jackson, C.J.; Taylor, M.C.; Stott, M.B.; Cook, G.M.; Morales, S.E. Genomic and metagenomic surveys of hydrogenase distribution indicate H2 is a widely utilised energy source for microbial growth and survival. ISME J. 2016, 10, 761–777. [Google Scholar] [CrossRef] [PubMed]
- Ray, A.E.; Zhang, E.; Terauds, A.; Ji, M.; Kong, W.; Ferrari, B.C. Soil Microbiomes With the Genetic Capacity for Atmospheric Chemosynthesis Are Widespread Across the Poles and Are Associated With Moisture, Carbon, and Nitrogen Limitation. Front. Microbiol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- World Meteorological Organization. The State of Greenhouse Gases in the Atmosphere Using Global Observations through 2018. WMO GHG Bullet. 2019, 15, 1–8, ISSN 2078-0796. [Google Scholar]
- Khalil, M.A.K.; Rasmussen, R.A. The global cycle of carbon monoxide: Trends and mass balance. Chemosphere 1990, 20, 227–242. [Google Scholar] [CrossRef]
- Novelli, P.C.; Masarie, K.A.; Lang, P.M. Distributions and recent changes of carbon monoxide in the lower troposphere. J. Geophys. Res. Atmos. 1998, 103, 19015–19033. [Google Scholar] [CrossRef]
- Chi, X.; Winderlich, J.; Mayer, J.-C.; Panov, A.V.; Heimann, M.; Birmili, W.; Heintzenberg, J.; Cheng, Y.; Andreae, M.O. Long-term measurements of aerosol and carbon monoxide at the ZOTTO tall tower to characterize polluted and pristine air in the Siberian taiga. Atmos. Chem. Phys. 2013, 13, 12271–12298. [Google Scholar] [CrossRef]
- Petrenko, V.V.; Martinerie, P.; Novelli, P.; Etheridge, D.M.; Levin, I.; Wang, Z.; Blunier, T.; Chappellaz, J.; KaiseriD, J.; Lang, P.; et al. A 60 yr record of atmospheric carbon monoxide reconstructed from Greenland firn air. Atmos. Chem. Phys. 2013, 13, 7567–7585. [Google Scholar] [CrossRef]
- Novelli, P.C.; Lang, P.M.; Masarie, K.A.; Hurst, D.F.; Myers, R.; Elkins, J.W. Molecular hydrogen in the troposphere: Global distribution and budget. J. Geophys. Res. Atmos. 1999, 104, 30427–30444. [Google Scholar] [CrossRef]
- Schmitz, R.A.; Pol, A.; Mohammadi, S.S.; Hogendoorn, C.; Van Gelder, A.H.; Jetten, M.S.M.; Daumann, L.J.; Camp, H.J.M.O.D. The thermoacidophilic methanotroph Methylacidiphilum fumariolicum SolV oxidizes subatmospheric H2 with a high-affinity, membrane-associated [NiFe] hydrogenase. ISME J. 2020, 14, 1223–1232. [Google Scholar] [CrossRef]
- Hakobyan, A.; Zhu, J.; Glatter, T.; Paczia, N.; Liesack, W. Hydrogen utilization by Methylocystis sp. strain SC2 expands the known metabolic versatility of type IIa methanotrophs. Metab. Eng. 2020, 61, 181–196. [Google Scholar] [CrossRef] [PubMed]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013; Available online: http://www.R-project.org/ (accessed on 23 January 2020).
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016; Available online: https://ggplot2.tidyverse.org (accessed on 23 January 2020).
- Pieja, A.J.; Sundstrom, E.R.; Criddle, C.S. Poly-3-Hydroxybutyrate Metabolism in the Type II Methanotroph Methylocystis parvus OBBP. Appl. Environ. Microbiol. 2011, 77, 6012–6019. [Google Scholar] [CrossRef] [PubMed]
- Ratcliff, W.C.; Kadam, S.V.; Denison, R.F. Poly-3-hydroxybutyrate (PHB) supports survival and reproduction in starving rhizobia. FEMS Microbiol. Ecol. 2008, 65, 391–399. [Google Scholar] [CrossRef]
- Lange, S.F.; Allaire, S.E.; van Bochove, E. Transfer of CO2, N2O and CH4 to butyl rubber (polyisobutylene) septa during storage. J. Environ. Monit. 2008, 10, 775–777. [Google Scholar] [CrossRef] [PubMed]
- Magnusson, T. A method for equilibration chamber sampling and gas chromatographic analysis of the soil atmosphere. Plant Soil 1989, 120, 39–47. [Google Scholar] [CrossRef]
- Bonk, F.; Popp, D.; Weinrich, S.; Sträuber, H.; Becker, D.; Kleinsteuber, S.; Harms, H.; Centler, F. Determination of Microbial Maintenance in Acetogenesis and Methanogenesis by Experimental and Modeling Techniques. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tveit, A.T.; Schmider, T.; Hestnes, A.G.; Lindgren, M.; Didriksen, A.; Svenning, M.M. Simultaneous Oxidation of Atmospheric Methane, Carbon Monoxide and Hydrogen for Bacterial Growth. Microorganisms 2021, 9, 153. https://doi.org/10.3390/microorganisms9010153
Tveit AT, Schmider T, Hestnes AG, Lindgren M, Didriksen A, Svenning MM. Simultaneous Oxidation of Atmospheric Methane, Carbon Monoxide and Hydrogen for Bacterial Growth. Microorganisms. 2021; 9(1):153. https://doi.org/10.3390/microorganisms9010153
Chicago/Turabian StyleTveit, Alexander Tøsdal, Tilman Schmider, Anne Grethe Hestnes, Matteus Lindgren, Alena Didriksen, and Mette Marianne Svenning. 2021. "Simultaneous Oxidation of Atmospheric Methane, Carbon Monoxide and Hydrogen for Bacterial Growth" Microorganisms 9, no. 1: 153. https://doi.org/10.3390/microorganisms9010153
APA StyleTveit, A. T., Schmider, T., Hestnes, A. G., Lindgren, M., Didriksen, A., & Svenning, M. M. (2021). Simultaneous Oxidation of Atmospheric Methane, Carbon Monoxide and Hydrogen for Bacterial Growth. Microorganisms, 9(1), 153. https://doi.org/10.3390/microorganisms9010153






