The Toothbrush Microbiome: Impact of User Age, Period of Use and Bristle Material on the Microbial Communities of Toothbrushes
Abstract
1. Introduction
2. Material and Methods
2.1. Sampling
2.2. Sample Preparation
2.3. DNA Extraction
2.4. Detection of Bla, Mcr, and IntI1 Genes
2.5. Microbiome Profiling
2.6. In-Vitro Test of Different Toothbrushes
2.7. Statistical Analysis
3. Results
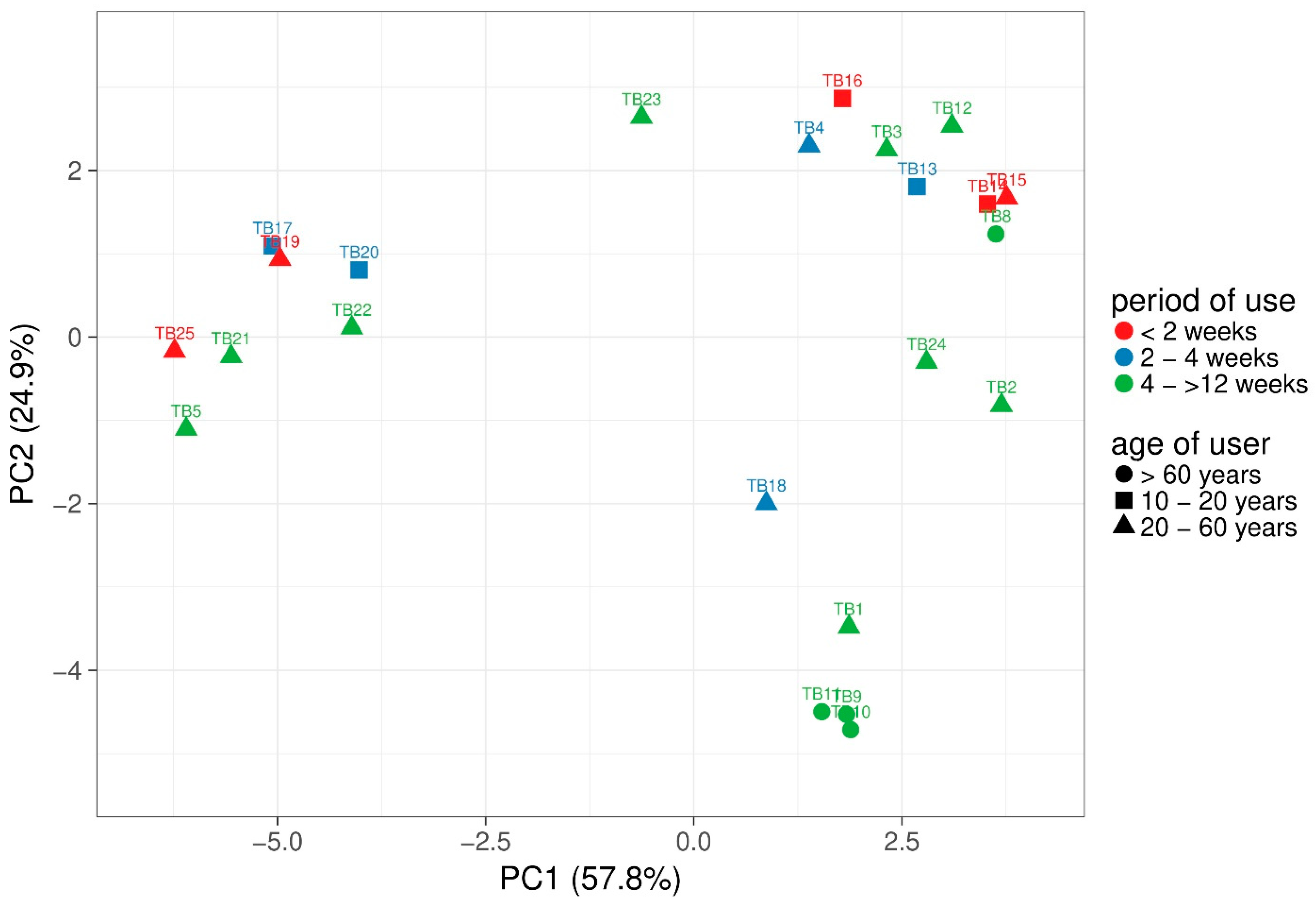
3.1. Taxonomic Distribution
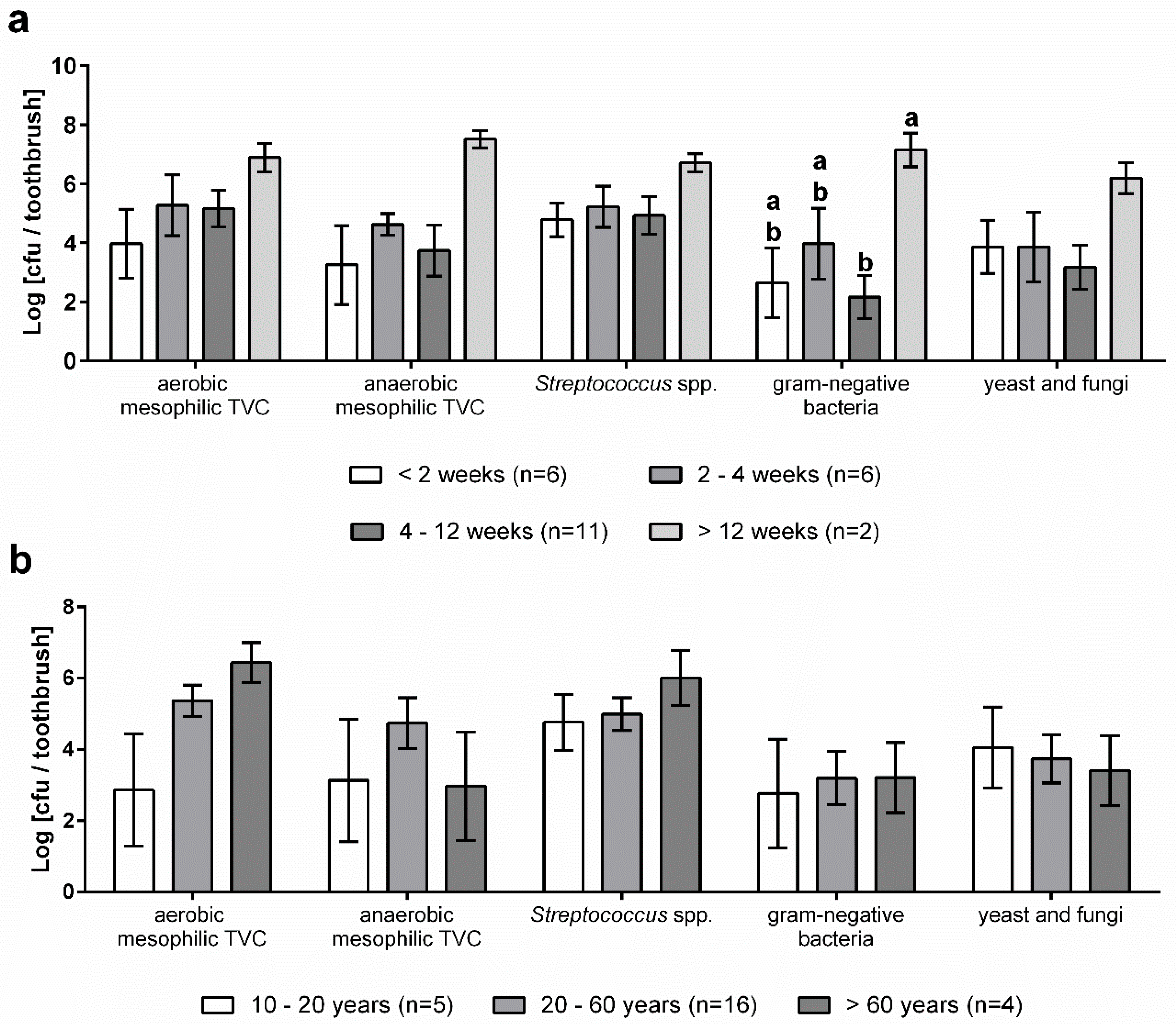
3.2. Microbial Contamination of Toothbrushes
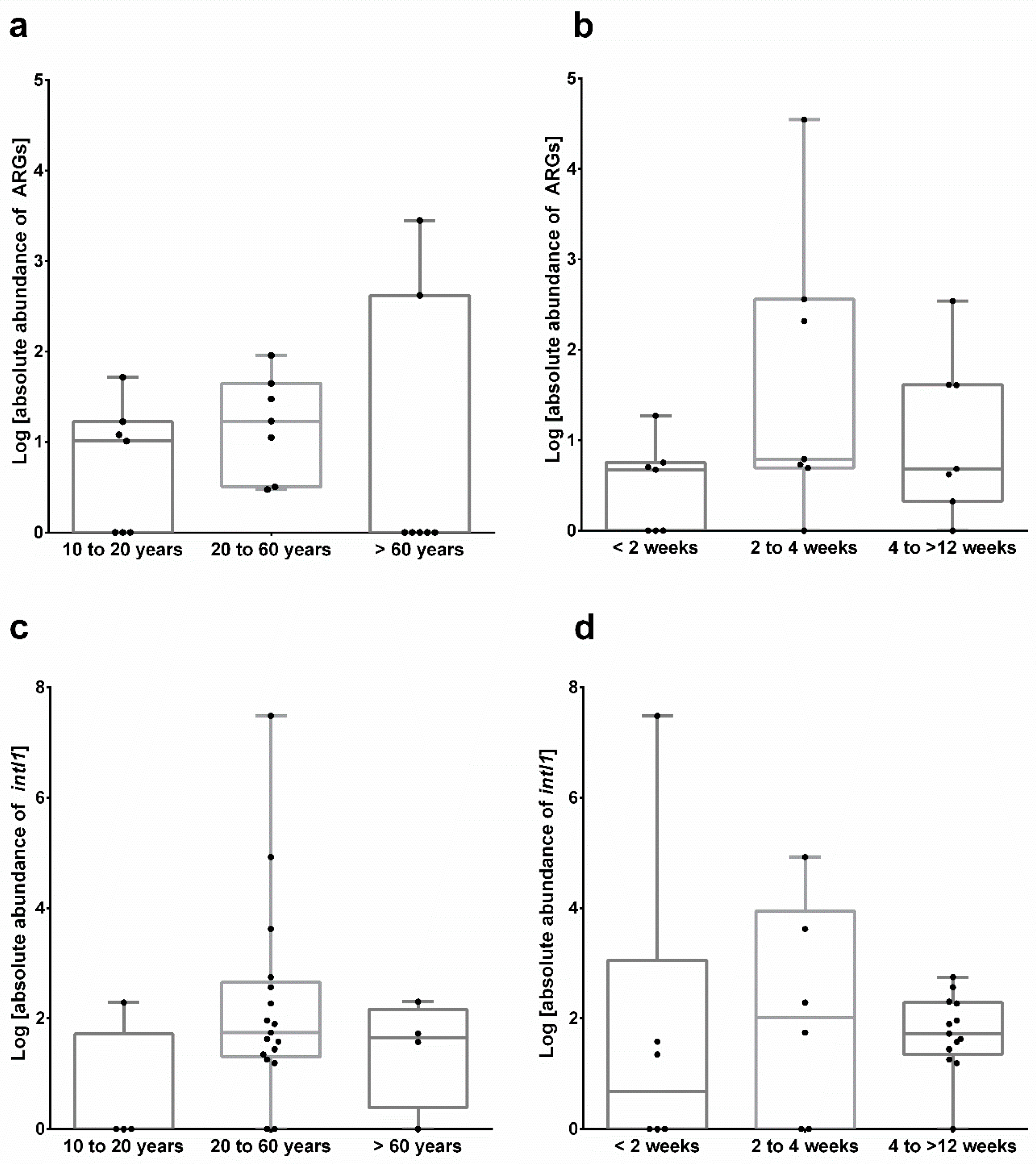
3.3. Occurrence of Antibiotic Resistance Genes
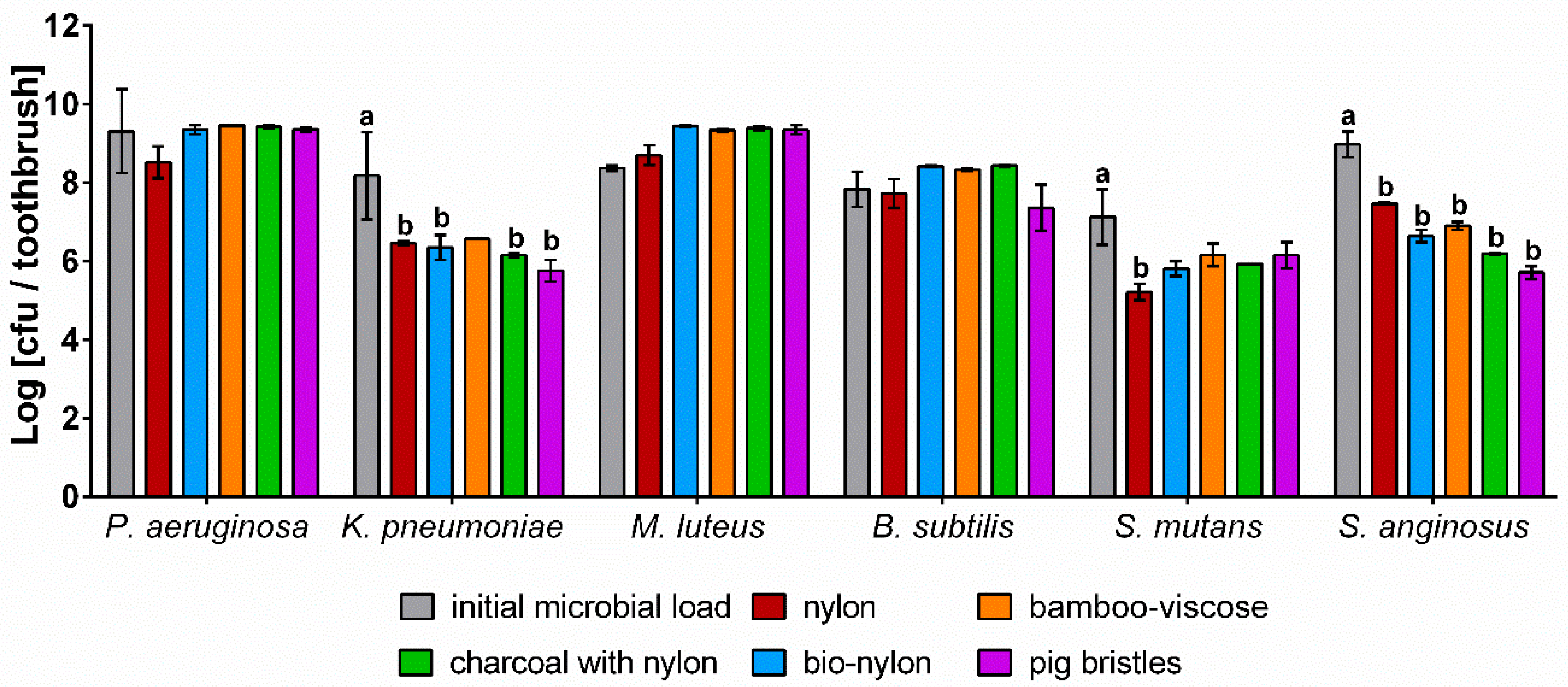
3.4. In-Vitro Study of Different Types of Toothbrush Bristles
4. Discussion
4.1. Bacterial Diversity of Toothbrushes
4.2. Microbial Contamination of Toothbrushes
4.3. Occurrence of Bla and IntI1 Genes in Toothbrush Samples
4.4. In-Vitro Study of Different Bristle Materials
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Glass, R.T. The infected toothbrush, the infected denture, and transmission of disease: A review. Compendium 1992, 13, 592–598. [Google Scholar] [PubMed]
- Naik, R.; Mujib, B.R.A.; Telagi, N.; Anil, B.S.; Spoorthi, B.R. Contaminated tooth brushes–potential threat to oral and general health. J. Fam. Med. Prim. Care 2015, 4, 444–448. [Google Scholar] [CrossRef] [PubMed]
- Frazelle, M.R.; Munro, C.L. Toothbrush Contamination: A Review of the Literature. Nurs. Res. Pract. 2012, 2012, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Thamke, M.V.; Beldar, A.; Thakkar, P.; Murkute, S.; Ranmare, V.; Hudwekar, A. Comparison of bacterial contamination and antibacterial efficacy in bristles of charcoal toothbrushes versus noncharcoal toothbrushes: A microbiological study. Contemp. Clin. Dent. 2018, 3, 463–467. [Google Scholar]
- Contreras, A.; Arce, R.; Botero, J.E.; Jaramillo, A.; Betancourt, M. Toothbrush contamination in family members. Revista Clínica de Periodoncia, Implantología y Rehabilitación Oral 2010, 3, 24–26. [Google Scholar] [CrossRef]
- Caudry, S.; Klitorinos, A.; Chan, E. Contaminated toothbrushes and their disinfection. Can. Dent. Assoc. 1995, 61, 511–516. [Google Scholar]
- Newburn, E. Preventing dental caries: Breaking the chain of transmission. J. Am. Dent. Assoc. 1992, 123, 55–59. [Google Scholar] [CrossRef]
- Glass, R.; Jensen, H. More on the contaminated toothbrushes: The viral story. Quintessence Int. 1988, 19, 713–716. [Google Scholar]
- Ayşegül, O.; Elgin, I.; Gulcin, A.; Nedim, S. The efficacy of chlorhexidine spray vs outhwash in the microbial contamination of child toothbrushes. J. Dent. Child. 2007, 74, 177–181. [Google Scholar]
- Efstratiou, M.; Papaionnou, W.; Nakou, M.; Ktenas, E.; Vrotsos, I.; Panis, V. Contamination of a toothbrush with antibacterial properties by oral microorganisms. J. Dent. 2007, 35, 331–337. [Google Scholar] [CrossRef]
- Nelson-Filho, P.; Macari, S.; Faria, G.; Assed, S.; Ito, I. Microbial contamination of toothbrushes and their decontamination. Pediatr. Dent. 2000, 22, 381–384. [Google Scholar] [PubMed]
- Karibasappa, G.; Nagesh, L.; Sujatha, B. Assessment of microbial contamination of toothbrush head: An in vitro study. Indian J. Dent. Res. 2011, 22, 2–5. [Google Scholar] [CrossRef] [PubMed]
- Taji, S.S.; Rogers, A.H. The microbial contamination of toothbrushes. A pilot study. Aust. Dent. J. 1998, 43, 128–130. [Google Scholar] [CrossRef] [PubMed]
- Bunetel, L.; Tricot-Doulex, S.; Agnani, G.; Bonnaure-Malletm, M. Invitroevaluation of the retention of three species of pathogenic microorganisms by three different types of toothbrush. Oral. Microbiol. Immunol. 2000, 15, 313–316. [Google Scholar] [CrossRef]
- Nascimento, A.; Watanabe, E.; Ito, I.Y. Toothbrush contamination by Candida spp. and efficacy of mouthrinse spray for their disinfection. Mycopathologia 2010, 169, 133–138. [Google Scholar] [CrossRef]
- Svanberg, M. Contamination of toothpaste and toothbrush by Streptococcus mutans. Scand. J. Dent. Res. 1978, 86, 412–414. [Google Scholar] [CrossRef]
- Ferreira, C.A.; Savi, G.D.; Panatto, A.P.; Generoso, J.D.; Barichello, T. Microbiological evaluation of bristles of frequently used toothbrushes. Dent. Press J. Orthod. 2012, 17, 72–76. [Google Scholar] [CrossRef]
- Sammons, R.L.; Kaur, D.; Neal, P. Bacterial survival and biofilm formation on conventional and antibacterial toothbrushes. Biofilms 2004, 1, 123–130. [Google Scholar] [CrossRef]
- Lucassen, R.; Rehberg, L.; Heyden, M.; Bockmühl, D.P. Strong correlation of total phenotypic resistance of samples from household environments and the prevalence of class 1 integrons suggests for the use of the relative prevalence of intI1 as a screening tool for multi-resistance. PLoS ONE 2019, 14, e0218277. [Google Scholar] [CrossRef]
- Rehberg, L.; Frontzek, A.; Melhus, Å.; Bockmühl, D.P. Prevalence of β-lactamase genes in domestic washing machines and dishwashers and the impact of laundering processes on antibiotic-resistant bacteria. J. Appl. Microbiol. 2017, 123, 3218–3221. [Google Scholar] [CrossRef]
- Marshall, B.M.; Robleto, E.; Dumont, T.; Levy, S.B. The frequency of antibiotic-resistant bacteria in homes differing in their use of surface antibacterial agents. Curr. Microbiol. 2012, 65, 407–415. [Google Scholar] [CrossRef] [PubMed]
- Penders, J.; Stobberingh, E.E.; Savelkoul, P.H.M.; Wolffs, P.F.G. The human microbiome as a reservoir of antimicrobial resistance. Front. Microbiol. 2013, 4, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Baron, S.A.; Diene, S.M.; Rolain, J.M. Human microbiomes and antibiotic resistance. Hum. Microbiome J. 2018, 10, 43–52. [Google Scholar] [CrossRef]
- Sommer, M.O.A.; Dantas, G.; Church, G.M. Functional characterization of the antibiotic resistance reservoir in the human microflora. Science 2009, 325, 1128–1131. [Google Scholar] [CrossRef] [PubMed]
- Nelson-Filho, P.; Pereira, M.S.S.; de Rossi, A.; da Silva, R.A.B.; de Mesquita, K.S.F.; de Queiroz, A.M.; da Silva, L.A.B. Children’s toothbrush contamination in day-care centers: How to solve this problem? Clin. Oral. Investig. 2014, 18, 1969–1974. [Google Scholar] [CrossRef]
- Malmberg, E.; Birkhed, D.; Norvenius, G.; Norén, J.G.; Dahlén, G. Microorganisms on toothbrushes at day-care centers. Acta Odontol. Scand. 1994, 52, 93–98. [Google Scholar] [CrossRef]
- Raiyani, C.M.; Arora, R.; Bhayya, D.P.; Dogra, S.; Katageri, A.A.; Singh, V. Assessment of microbial contamination on twice a day used toothbrush head after 1-month and 3 months: An in vitro study. J. Nat. Sci. Biol. Med. 2015, 6, S44–S48. [Google Scholar] [CrossRef]
- Schages, L.; Wichern, F.; Kalscheuer, R.; Bockmühl, D. Winter is coming—Impact of temperature on the variation of beta-lactamase and mcr genes in a wastewater treatment plant. Sci. Total Environ. 2020, 712, 136499. [Google Scholar] [CrossRef]
- Gillings, M.R.; Gaze, W.H.; Pruden, A.; Smalla, K.; Tiedje, J.M.; Zhu, Y.G. Using the class 1 integron-integrase gene as a proxy for anthropogenic pollution. ISME J. 2015, 9, 1269–1279. [Google Scholar] [CrossRef]
- Lebuhn, M.; Hanreich, A.; Klocke, M.; Schlüter, A.; Bauer, C.; Pérez, C.M. Towards molecular biomarkers for biogas production from lignocellulose-rich substrates. Anaerobe 2014, 29, 10–21. [Google Scholar] [CrossRef]
- Metsalu, T.; Vilo, J. ClustVis: A web tool for visualizing clustering of multivariate data using Principal Component Analysis and heatmap. Nucleic Acids Res. 2015, 43, W566–W570. [Google Scholar] [CrossRef] [PubMed]
- Malta, A.L.; Carvalho, J.; Barroso, H. The effect of toothbrush covers on microbial contamination. Ann. Med. 2019, 51, 112. [Google Scholar] [CrossRef]
- Meyer, D.H.; Fives-Taylor, P.M. Oral pathogens: From dental plaque to cardiac disease. Curr. Opin. Microbiol. 1998, 1, 88–95. [Google Scholar] [CrossRef]
- Saravia, M.E.; da Silva, R.A.B.; Rossi, M.A.; Nelson-filho, P.; Faria, G.; Rossi, M.A.; Ito, I.Y. Viability of Streptococcus mutans toothbrush bristles. J. Dent. Child. 2008, 75, 29–32. [Google Scholar]
- Gendron, R.; Grenier, D.; Maheu-Robert, L.F. The oral cavity as a reservoir of bacterial pathogens for focal infections. Microbes Infect. 2000, 2, 897–906. [Google Scholar] [CrossRef]
- Jenkinson, H.F. Beyond the oral microbiome. Environ. Microbiol. 2011, 13, 3077–3087. [Google Scholar] [CrossRef]
- Dayoub, M.B.; Rusilko, D.; Gross, A. Microbial Contamination of Toothbrushes. J. Dent. Res. 1977, 56, 706. [Google Scholar] [CrossRef]
- Gattlen, J.; Amberg, C.; Zinn, M.; Mauclaire, L. Biofilms isolated from washing machines from three continents and their tolerance to a standard detergent. Biofouling 2010, 26, 873–882. [Google Scholar] [CrossRef]
- Kelley, S.T.; Theisen, U.; Angenent, L.T.; Amand, A.S.; Pace, N.R. Molecular Analysis of Shower Curtain Biofilm Microbes. Appl. Environ. Microbiol. 2004, 70, 4187–4192. [Google Scholar] [CrossRef]
- Dotis, J.; Printza, N.; Stabouli, S.; Papachristou, F. Kocuria species peritonitis: Although rare, we have to care. Perit. Dial. Int. 2015, 35, 26–30. [Google Scholar] [CrossRef]
- Von Graevenitz, A. Rothia dentocariosa: Taxonomy and differential diagnosis. Clin. Microbiol. Infect. 2004, 10, 399–402. [Google Scholar] [CrossRef] [PubMed]
- Lira-Junior, R.; Åkerman, S.; Klinge, B.; Boström, E.A.; Gustafsson, A. Salivary microbial profiles in relation to age, periodontal, and systemic diseases. PLoS ONE 2018, 13, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Takeshita, T.; Kageyama, S.; Furuta, M.; Tsuboi, H.; Takeuchi, K.; Shibata, Y.; Shimazaki, Y.; Akifusa, S.; Ninomiya, T.; Kiyohara, Y.; et al. Bacterial diversity in saliva and oral health-related conditions: The Hisayama Study. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef]
- Xu, X.; He, J.; Xue, J.; Wang, Y.; Li, K.; Zhang, K.; Guo, Q.; Liu, X.; Zhou, Y.; Cheng, L.; et al. Oral cavity contains distinct niches with dynamic microbial communities. Environ. Microbiol. 2015, 17, 699–710. [Google Scholar] [CrossRef] [PubMed]
- VuMA Touchpoints, Umfrage in Deutschland zur Häufigkeit des Wechselns der Zahnbürste bis 2018. 2019. Available online: https://de.statista.com/statistik/daten/studie/181217/umfrage/haeufigkeit-wechsel-der-zahnbuerste/ (accessed on 16 August 2020).
- Ziebolz, D.; van Küss, K.; Hornecker, E.; Mausberg, R. Eine Untersuchung gebrauchter Handzahnbürsten—Ergebnisse einer Umtauschaktion. Oralprophylaxe und Kinderzahnheilkunde 2006, 28, 54–59. [Google Scholar]
- Wetzel, W.E.; Schaumburg, C.; Ansari, F.; Kroeger, T.; Sziegoleit, A. Microbial contamination of toothbrushes with different principles of filament anchoring. J. Am. Dent. Assoc. 2005, 136, 758–765. [Google Scholar] [CrossRef]
- Quirynen, M.; de Soete, M.; Pauwels, M.; Gizani, S.; van Meerbeek, B.; van Steenberghe, D. Can Toothpaste or a Toothbrush with Antibacterial Tufts Prevent Toothbrush Contamination? J. Periodontol. 2003, 74, 312–322. [Google Scholar] [CrossRef]
- Schwartz, T.; Kohnen, W.; Jansen, B.; Obst, U. Detection of antibiotic resistant bacteria and their resistance genes in wastewater, surface water, and drinking water biofilms. FEMS Microbiol. Ecol. 2003, 43, 325–335. [Google Scholar] [CrossRef]
- Jacoby, G.A. AmpC Β-Lactamases. Clin. Microbiol. Rev. 2009, 22, 161–182. [Google Scholar] [CrossRef]
- Deng, Y.; Bao, X.; Ji, L.; Chen, L.; Liu, J.; Miao, J.; Chen, D.; Bian, H.; Li, Y.; Yu, G. Resistance integrons: Class 1, 2 and 3 integrons. Ann. Clin. Microbiol. Antimicrob. 2015, 14, 45. [Google Scholar] [CrossRef]
- Ghosh, T.S.; Gupta, S.S.; Nair, G.B.; Mande, S.S. In silico analysis of antibiotic resistance genes in the gut microflora of individuals from diverse geographies and age-groups. PLoS ONE 2013, 8, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Bonomo, R.A.; Burd, E.M.; Conly, J.; Limbago, B.M.; Poirel, L.; Segre, J.A.; Westblade, L.F. Carbapenemase-Producing Organisms: A Global Scourge. Clin. Infect. Dis. 2018, 66, 1290–1297. [Google Scholar] [CrossRef] [PubMed]


| Primer | 5′–3′ |
|---|---|
| KPC-1 | GCCGTGCAATACAGTGATAAC |
| KPC-2 | GAACGTGGTATCGCCGATAG |
| KPC-probe | TexRed-CCGCCGCCAATTTGTTGCTGAAGG-BHQ2 |
| OXA48-1 | GTTGGAATGCTCACTTTACTGAA |
| OXA48-2 | TTCGCCCGTTTAAGATTATTGG |
| OXA48-probe | FAM-ATTCTCATTCCAGAGCACAACTACGCC-BHQ1 |
| GES-1 | CTCTGTGAGTCGGCTAGACC |
| GES-2 | CGATCAGCCACCTCTCAATG |
| GES-probe | HEX-ACACCTGGCGACCTCAGAGATACAACT-BHQ1 |
| VIM-1 | GTTTGGTYGCATATCGCAAC |
| VIM-2 | CTTYTCAATCTCCGCGAGAAG |
| VIM-probe1 | FAM-AGCAACTCATCRCCATCACGGACAATG-BHQ1 |
| VIM-probe2 | FAM-AACTCGGTGACACGGTGTACTCGTCT-BHQ1 |
| NDM-1 | CGACTTATGCCAATGCGTTG |
| NDM-2 | CGGGGTAAAATACCTTGAGC |
| NDM-probe | HEX-AGCCTGACTTTCGCCGCCAATG-BHQ1 |
| OXA23-1 | CCTGATCGGATTGGAGAACC |
| OXA23-2 | GTTCCTGATAGACTGGGACTG |
| OXA23-probe | FAM-TGGCTTCTCCTAGTGTCATGTCTT-BHQ1 |
| OXA58-1 | GTTGGTATGTGGGTTTTGTTG |
| OXA58-2 | CGTAGAGCAATATCATCACCAG |
| OXA58-probe | Cy5-TGCCACCACTTGCCCATCTGCC-BBQ®-650 |
| CTXM-1-1 | AATCTGACGCTGGGTAAAGC |
| CTXM-1-2 | GATATCGTTGGTGGTGCCATA |
| CTXM-1-Probe | FAM-CCTGAATGCTCGCTGCACCGG-BHQ1 |
| CTXM-9-1 | CCGATCTGGTTAACTACAATCC |
| CTXM-9-2 | GCTGGGCAATCAATTTGTTC |
| CTXM-9-probe | TexRed-CAACGGCACAATGACGCTGGC-BHQ2 |
| CMY2-1 | GATGCAGGAGCAGGCTATTC |
| CMY2-2 | AACACGCCGTTAAACGTCTTAC |
| CMY2-probe | FAM-CCAATAACCACCCAGTCACGCAG-BHQ1 |
| FOX-1 | GTTCGAGATTGGCTCGGTCA |
| FOX-2 | CACTGTAGGTGGCAAGCTCG |
| FOX-probe | HEX-TGGCTCACCTTGTCATCCAGC-BHQ1 |
| ACT_MIR-1 | ACTGGCAGCCGCAGTGGAAG |
| ACT_MIR-2 | ACGTTAATCCASGTATGGTCCAGC |
| ACT_MIR-probe | FAM-AGACCCGCGTCGTYATGGCCTG-BHQ1 |
| DHA-1 | GGCGATATGCGTCTGTATGC |
| DHA-2 | GTCAGCAACTGCTCATACGG |
| DHA-probe | TexRed-CCTGTTTGGTGCTCTGACCGC-BHQ2 |
| mcr-1-1 | ATGGCACGGTCTATGATACG |
| mcr-1-2 | CACACCCAAACCAATGATACG |
| mcr-1-probe | FAM-ACCGACCAAGCCGAGACCAAGGA-BHQ1 |
| mcr-2-1 | GCCAACAGACACCATCTATC |
| mcr-2-2 | TAGCCATTGAACTGCACATG |
| mcr-2-probe | HEX-ACCACCAAGCCGAGCGAGCG-BHQ1 |
| Primer | 5′–3′ |
|---|---|
| intI1 F165 | CGAACGAGTGGCGGAGGGTG [29] |
| intI1 R476 | TACCCGAGAGCTTGGCACCCA [29] |
| 16S rDNA F919 | GAATTGACGGGGGCCCGCACAAG [30] |
| 16S rDNA R1378 | CGGTGTG TACAAGGCCCGGGAACG [30] |
| Bacterial Order | <2 Weeks (n = 6) | 2 to 4 Weeks (n = 6) | 4 to >12 Weeks (n = 13) | 10 to 20 Years (n = 5) | 20 to 60 Years (n = 16) | >60 Years (n = 4) |
|---|---|---|---|---|---|---|
| Micrococcales | 4.9% | 10.8% | 27.6% * | 5.6% | 12.5% | 56.9% |
| Enterobacterales | 29.7% * | 22.6% | 7.0% | 17.0% | 19.0% | 0.0% |
| Lactobacillales | 22.6% | 14.0% | 12.6% | 25.0% | 14.8% | 6.2% |
| Actinomycetales | 15.8% | 11.3% | 10.1% | 16.8% | 11.1% | 8.2% |
| Neisserales | 6.5% | 2.7% | 1.6% | 7.7% | 1.8% | 2.4% |
| Pseudomonadales | 0.0% | 1.2% | 2.0% | 0.0% | 2.1% | 0.25% |
| Sphingomonadales | 0.0% | 0.2% | 1.0% | 0.3% | 0.8% | 0.4% |
| Sample | Period of Use | User Age | Storage | Sample | Period of Use | User Age | Storage |
|---|---|---|---|---|---|---|---|
| TB1 | 4–12 weeks | 20–60 | upright | TB14 | <2 weeks | 10–20 | upright |
| TB2 | 4–12 weeks | 20–60 | upright | TB15 | <2 weeks | 20–60 | upright |
| TB3 | 4–12 weeks | 20–60 | upright | TB16 | <2 weeks | 10–20 | upright |
| TB4 | 2–4 weeks | 20–60 | upright | TB17 | 2–4 weeks | 10–20 | horizontal |
| TB5 | 4–12 weeks | 20–60 | upright | TB18 | 2–4 weeks | 20–60 | horizontal |
| TB6 | <2 weeks | 20–60 | upright | TB19 | <2 weeks | 20–60 | upright |
| TB7 | 2–4 weeks | 20–60 | upright | TB20 | 2–4 weeks | 10–20 | upright |
| TB8 | 4–12 weeks | >60 | upright | TB21 | >12 weeks | 20–60 | horizontal |
| TB9 | 4–12 weeks | >60 | horizontal | TB22 | >12 weeks | 20–60 | horizontal |
| TB10 | 4–12 weeks | >60 | upright | TB23 | 4–12 weeks | 20–60 | upright |
| TB11 | 4–12 weeks | >60 | upright | TB24 | 4–12 weeks | 20–60 | upright |
| TB12 | 4–12 weeks | 20–60 | upright | TB25 | <2 weeks | 20–60 | upright |
| TB13 | 2–4 weeks | 10–20 | upright |
| Culture Medium | Microorganisms | Microbial Count (cfu/toothbrush) | Standard Error |
|---|---|---|---|
| TSA (aerobic), 37 °C | aerobic mesophilic bacterial count | 7.05 × 106 | 5.10 × 105 |
| TSA (anaerobic), 37 °C | anaerobic mesophilic bacterial count | 1.19 × 107 | 1.24 × 106 |
| Columbia Blood Agar, 35 °C | Streptococcus spp. | 3.85 × 106 | 3.04 × 105 |
| MacConkey, 35 °C | gram-neg. bacteria | 5.04 × 106 | 7.27 × 105 |
| MEA, 30 °C | yeast and fungi | 1.42 × 106 | 1.26 × 105 |
| Sample | Resistance Genes | Sample | Resistance Genes |
|---|---|---|---|
| TB1 | intI1, blaACT/MIR, blaOXA-23, blaOXA-58 | TB14 | no genes detected |
| TB2 | intI1, blaOXA-23, blaOXA-58 | TB15 | intI1 |
| TB3 | intI1, blaOXA-48 | TB16 | blaKPC |
| TB4 | intI1, blaCMY-2, blaGES, blaOXA-23, blaOXA-58 | TB17 | blaCMY-2, blaOXA-23 |
| TB5 | intI1, blaGES, blaOXA-58 | TB18 | intI1, blaCMY-2, blaOXA-23 |
| TB6 | no genes detected | TB19 | intI1 |
| TB7 | blaOXA-58 | TB20 | intI1, blaACT/MIR, blaCMY-2, blaOXA-48 |
| TB8 | intI1, blaOXA-23, blaOXA-58 | TB21 | intI1, blaGES |
| TB9 | blaOXA-23, blaOXA-58 | TB22 | intI1, blaKPC, blaGES |
| TB10 | intI1, blaOXA-58 | TB23 | intI1, blaKPC |
| TB11 | intI1 | TB24 | intI1, blaGES |
| TB12 | intI1, blaACT/MIR, blaGES, blaOXA-58 | TB25 | intI1, blaACT/MIR, blaCMY-2, blaOXA-58 |
| TB13 | intI1, blaOXA-58 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zinn, M.-K.; Schages, L.; Bockmühl, D. The Toothbrush Microbiome: Impact of User Age, Period of Use and Bristle Material on the Microbial Communities of Toothbrushes. Microorganisms 2020, 8, 1379. https://doi.org/10.3390/microorganisms8091379
Zinn M-K, Schages L, Bockmühl D. The Toothbrush Microbiome: Impact of User Age, Period of Use and Bristle Material on the Microbial Communities of Toothbrushes. Microorganisms. 2020; 8(9):1379. https://doi.org/10.3390/microorganisms8091379
Chicago/Turabian StyleZinn, Marc-Kevin, Laura Schages, and Dirk Bockmühl. 2020. "The Toothbrush Microbiome: Impact of User Age, Period of Use and Bristle Material on the Microbial Communities of Toothbrushes" Microorganisms 8, no. 9: 1379. https://doi.org/10.3390/microorganisms8091379
APA StyleZinn, M.-K., Schages, L., & Bockmühl, D. (2020). The Toothbrush Microbiome: Impact of User Age, Period of Use and Bristle Material on the Microbial Communities of Toothbrushes. Microorganisms, 8(9), 1379. https://doi.org/10.3390/microorganisms8091379





