Molecular Detection of Avian Pathogenic Escherichia coli (APEC) for the First Time in Layer Farms in Bangladesh and Their Antibiotic Resistance Patterns
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Sample Collection and Processing
2.3. Isolation and Identification of E. coli
2.4. Molecular Detection of APEC-Associated Virulence Genes
2.5. Antimicrobial Susceptibility Test
2.6. Statistical Analysis
3. Results
3.1. Prevalence of E. coli Isolates
3.2. APEC-Associated Virulence Genes
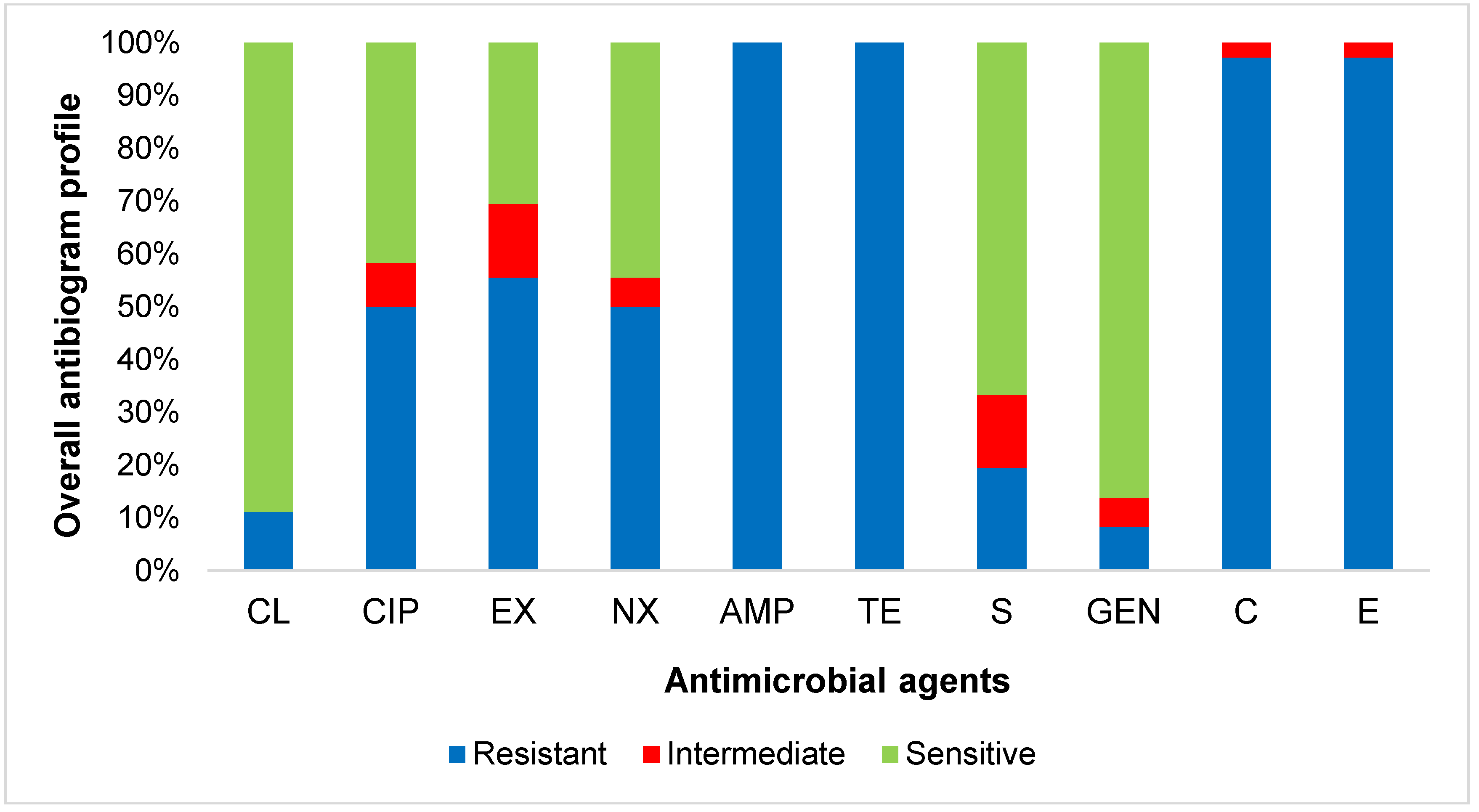
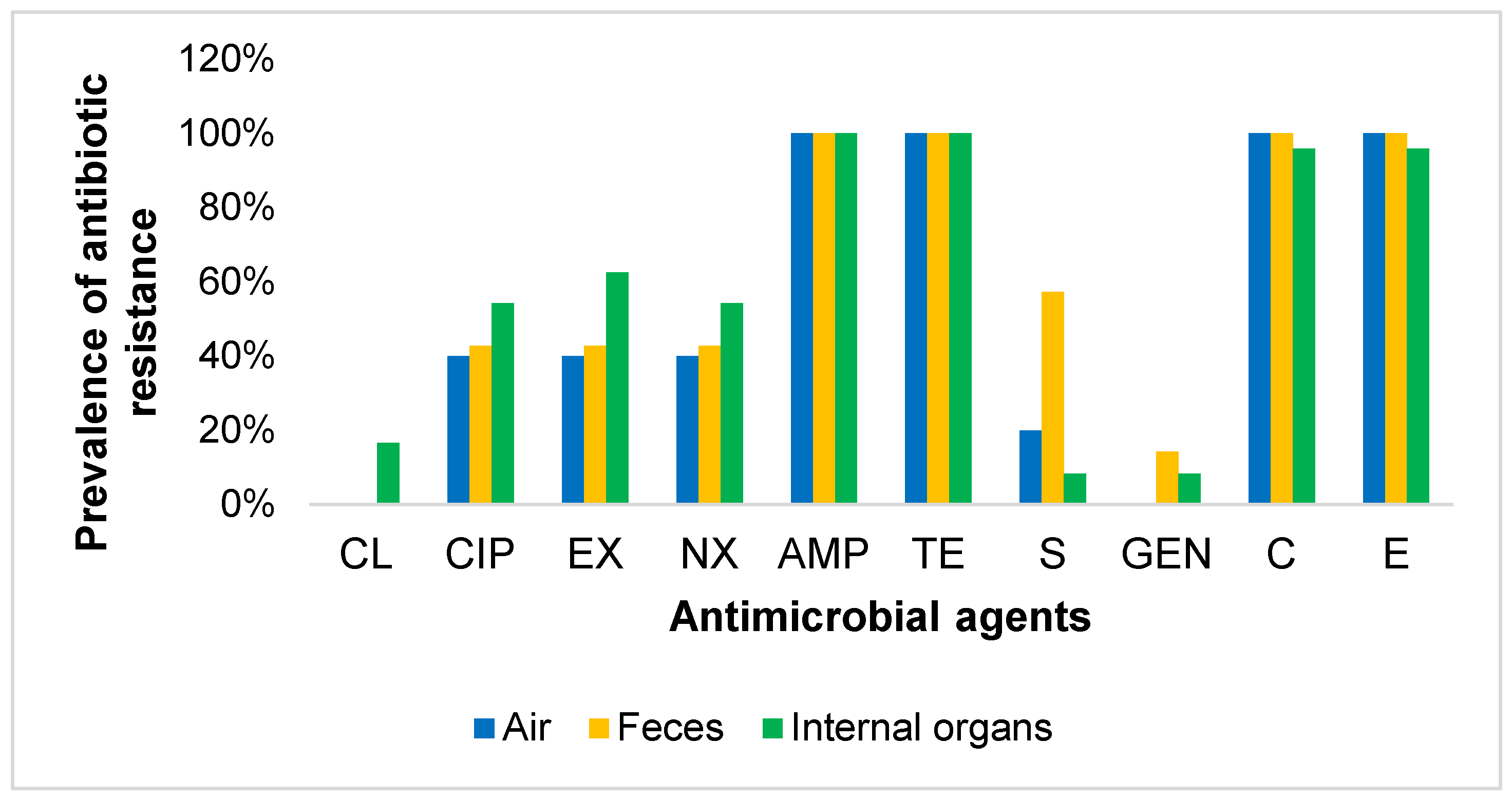
3.3. Antibiogram Profile of E. coli Isolates Carrying APEC-Associated Virulence Genes
3.4. Pairwise Correlation between Resistance to Antimicrobials
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- MOFL, “Annul Report 2018–2019”, Ministry of Fisheries and Livestock, Bangladesh 2019. Available online: http://bbs.gov.bd/site/page/dc2bc6ce-7080-48b3-9a04-73cec782d0df/- (accessed on 28 April 2020).
- Alam, S.B.; Mahmud, M.; Akter, R.; Hasan, M.; Sobur, A.; Nazir, K.H.M.; Noreddin, A.; Rahman, T.; El Zowalaty, M.E.; Rahman, M. Molecular detection of multidrug resistant salmonella species isolated from broiler farm in bangladesh. Pathogens 2020, 9, 201. [Google Scholar] [CrossRef] [PubMed]
- Dho-Moulin, M.; Fairbrother, J.M. Avian pathogenic Escherichia coli (APEC). Vet. Res. 1999, 30, 299–316. [Google Scholar]
- Kim, Y.B.; Yoon, M.Y.; Ha, J.S.; Seo, K.W.; Noh, E.B.; Son, S.H.; Lee, Y.J. Molecular characterization of avian pathogenic Escherichia coli from broiler chickens with colibacillosis. Poult. Sci. 2020, 99, 1088–1095. [Google Scholar] [CrossRef]
- Schouler, C.; Schaeffer, B.; Brée, A.; Mora, A.; Dahbi, G.; Biet, F.; Oswald, E.; Mainil, J.; Blanco, J.; Moulin-Schouleur, M. Diagnostic strategy for identifying avian pathogenic Escherichia coli based on four patterns of virulence genes. J. Clin. Microbiol. 2012, 50, 1673–1678. [Google Scholar] [CrossRef]
- Ewers, C.; Janßen, T.; Kießling, S.; Philipp, H.C.; Wieler, L.H. Molecular epidemiology of avian pathogenic Escherichia coli (APEC) isolated from colisepticemia in poultry. Vet. Microbiol. 2004, 104, 91–101. [Google Scholar] [CrossRef]
- Oh, J.Y.; Kang, M.S.; Kim, J.M.; An, B.K.; Song, E.A.; Kim, J.Y.; Shin, E.G.; Kim, M.J.; Kwon, J.H.; Kwon, Y.K. Characterization of Escherichia coli isolates from laying hens with colibacillosis on 2 commercial egg-producing farms in Korea. Poult. Sci. 2011, 90, 1948–1954. [Google Scholar] [CrossRef] [PubMed]
- Paixao, A.C.; Ferreira, A.C.; Fontes, M.; Themudo, P.; Albuquerque, T.; Soares, M.C.; Fevereiro, M.; Martins, L.; de Sá, M.I.C. Detection of virulence-associated genes in pathogenic and commensal avian Escherichia coli isolates. Poult. Sci. 2016, 95, 1646–1652. [Google Scholar] [CrossRef]
- Redweik, G.A.; Stromberg, Z.R.; Van Goor, A.; Mellata, M. Protection against avian pathogenic Escherichia coli and Salmonella Kentucky exhibited in chickens given both probiotics and live Salmonella vaccine. Poult. Sci. 2020, 99, 752–762. [Google Scholar] [CrossRef] [PubMed]
- Yogaratnam, V. Analysis of the causes of high rates of carcase rejection at a poultry processing plant. Vet. Rec. 1995, 137, 215–217. [Google Scholar] [CrossRef] [PubMed]
- Al-Kandari, F.; Woodward, M.J. Genotypic and phenotypic diversity differences of presumptive commensal and avian pathogenic E. coli. Br. Poult. Sci. 2019, 60, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Janßen, T.; Schwarz, C.; Preikschat, P.; Voss, M.; Philipp, H.C.; Wieler, L.H. Virulence-associated genes in avian pathogenic Escherichia coli (APEC) isolated from internal organs of poultry having died from colibacillosis. Int. J. Med. Microbiol. 2001, 291, 371–378. [Google Scholar] [CrossRef]
- Ghunaim, H.; Abu-Madi, M.A.; Kariyawasam, S. Advances in vaccination against avian pathogenic Escherichia coli respiratory disease: Potentials and limitations. Vet. Microbiol. 2014, 172, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Huja, S.; Oren, Y.; Trost, E.; Brzuszkiewicz, E.; Biran, D.; Blom, J.; Goesmann, A.; Gottschalk, G.; Hacker, J.; Ron, E.Z.; et al. Genomic avenue to avian colisepticemia. MBio 2015, 6, e01681-14. [Google Scholar] [CrossRef] [PubMed]
- Zakariazadeh, N.; Shayegh, J.; Ghorbani, A. Phylogenetic typing and virulence gene profile of pathogenic and commensal avian Escherichia coli in Iran: A notable finding. Comp. Clin. Pathol. 2019, 28, 525–530. [Google Scholar] [CrossRef]
- Mohamed, L.; Ge, Z.; Yuehua, L.; Yubin, G.; Rachid, K.; Mustapha, O.; Junwei, W.; Karine, O. Virulence traits of avian pathogenic (APEC) and fecal (AFEC) E. coli isolated from broiler chickens in Algeria. Trop. Anim. Health Prod. 2018, 50, 547–553. [Google Scholar] [CrossRef] [PubMed]
- Subedi, M.; Luitel, H.; Devkota, B.; Bhattarai, R.K.; Phuyal, S.; Panthi, P.; Shrestha, A.; Chaudhary, D.K. Antibiotic resistance pattern and virulence genes content in avian pathogenic Escherichia coli (APEC) from broiler chickens in Chitwan, Nepal. BMC Vet. Res. 2018, 14, 113. [Google Scholar] [CrossRef]
- Wang, J.; Tang, P.; Tan, D.; Wang, L.; Zhang, S.; Qiu, Y.; Dong, R.; Liu, W.; Huang, J.; Chen, T.; et al. The pathogenicity of chicken pathogenic Escherichia coli is associated with the numbers and combination patterns of virulence-associated genes. Open J. Vet. Med. 2015, 5, 243. [Google Scholar] [CrossRef]
- Vandekerchove, D.; Vandemaele, F.; Adriaensen, C.; Zaleska, M.; Hernalsteens, J.P.; De Baets, L.; Butaye, P.; Van Immerseel, F.; Wattiau, P.; Laevens, H.; et al. Virulence-associated traits in avian Escherichia coli: Comparison between isolates from colibacillosis-affected and clinically healthy layer flocks. Vet. Microbiol. 2005, 108, 75–87. [Google Scholar] [CrossRef] [PubMed]
- Nie, W.; Wang, J.; Xu, J.; Yao, L.; Qiao, D.; Xue, F.; Tang, F.; Chen, W. A molecule capturer analysis system for visual determination of avian pathogenic Escherichia coli serotype O78 using a lateral flow assay. Microchim. Acta 2020, 187, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Mellata, M. Human and avian extraintestinal pathogenic Escherichia coli: Infections, zoonotic risks, and antibiotic resistance trends. Foodborne Pathog. Dis. 2013, 10, 916–932. [Google Scholar] [CrossRef]
- Manges, A.R.; Johnson, J.R. Food-borne origins of Escherichia coli causing extraintestinal infections. Clin. Infect. Dis. 2012, 55, 712–719. [Google Scholar] [CrossRef] [PubMed]
- Tivendale, K.A.; Logue, C.M.; Kariyawasam, S.; Jordan, D.; Hussein, A.; Li, G.; Wannemuehler, Y.; Nolan, L.K. Avian-pathogenic Escherichia coli strains are similar to neonatal meningitis E. coli strains and are able to cause meningitis in the rat model of human disease. Infect. Immun. 2010, 78, 3412–3419. [Google Scholar] [CrossRef] [PubMed]
- Clifford, K.; Darash, D.; da Costa, C.P.; Meyer, H.; Islam, M.T.; Meyer, H.; Klohe, K.; Winklerc, A.; Rahman, M.T.; Islam, M.T.; et al. The threat of antimicrobial resistance opportunities for a technology-integrated One Health approach. Bull. World Health Organ. 2018, 96, 662–664. [Google Scholar] [CrossRef]
- Jakobsen, L.; Spangholm, D.J.; Pedersen, K.; Jensen, L.B.; Emborg, H.D.; Agersø, Y.; Aarestrup, F.M.; Hammerum, A.M.; Frimodt-Møller, N. Broiler chickens, broiler chicken meat, pigs and pork as sources of ExPEC related virulence genes and resistance in Escherichia coli isolates from community-dwelling humans and UTI patients. Int. J. Food Microbiol. 2010, 142, 264–272. [Google Scholar] [CrossRef] [PubMed]
- WHO. Antimicrobial Resistance in the Food Chain, Retrieved 2019. Available online: https://www.who.int/foodsafety/areas_work/antimicrobial-resistance/amrfoodchain/en/ (accessed on 28 April 2020).
- Ahmed, A.M.; Shimamoto, T.; Shimamoto, T. Molecular characterization of multidrug-resistant avian pathogenic Escherichia Coli isolated from septicemic broilers. Int. J. Med. Microbiol. 2013, 303, 475–483. [Google Scholar] [CrossRef]
- Ozaki, H.; Matsuoka, Y.; Nakagawa, E.; Murase, T. Characteristics of Escherichia coli isolated from broiler chickens with colibacillosis in commercial farms from a common hatchery. Poult. Sci. 2017, 96, 3717–3724. [Google Scholar] [CrossRef]
- Hossain, M.T.; Siddique, M.P.; Hossain, F.M.A.; Zinnah, M.A.; Hossain, M.M.; Alam, M.K.; Rahman, M.T.; Choudury, K.A. Isolation, identification, toxin profile and antibiogram of Escherichia coli isolated from broilers and layers in Mymensingh district of Bangladesh. Bangladesh J. Vet. Med. 2008, 6, 1–5. [Google Scholar] [CrossRef]
- Akond, M.A.; Alam, S.; Hassan, S.M.R.; Shirin, M. Antibiotic resistance of Escherichia coli isolated from poultry and poultry environment of Bangladesh. Int. J. Food Saf. 2009, 11, 19–23. [Google Scholar]
- Parvej, M.S.; Alam, M.A.; Shono, M.; Zahan, M.N.; Parvez, M.M.M.; Ansari, W.K.; Jowel, M.S.; Uddin, M.S.; Kage-Nakadai, E.; Rahman, M.T.; et al. Prevalence of virulence genes of diarrheagenic Escherichia coli in fecal samples obtained from cattle, poultry and diarrheic patients in Bangladesh. Jpn. J. Infect. Dis. 2020, 73, 76–82. [Google Scholar] [CrossRef]
- Sobur, M.A.; Ievy, S.; Haque, Z.F.; Nahar, A.; Zaman, S.B.; Rahman, M.T. Emergence of colistin-resistant Escherichia coli in poultry, house flies, and pond water in Mymensingh, Bangladesh. J. Adv. Vet. Anim. Res. 2019, 6, 50–53. [Google Scholar]
- Mbamalu, O.; Uebel, R.; Meki, B. Control of airborne microbes in a poultry setting using Dioxy MP 14. Braz. J. Poult. Sci. 2015, 17, 77–86. [Google Scholar] [CrossRef][Green Version]
- Bergey, D.H.; Buchanan, R.E.; Gibbons, N.E. Bergey’s Manual of Determinative Bacteriology, 8th ed.; American Society for Microbiology: Washington, DC, USA, 1974; pp. 966–1097. [Google Scholar]
- Sobur, A.; Haque, Z.F.; Sabuj, A.A.; Ievy, S.; Rahman, A.T.; El Zowalaty, M.E.; Rahman, T. Molecular detection of multidrug and colistin-resistant Escherichia coli isolated from house flies in various environmental settings. Future Microbiol. 2019, 14, 847–858. [Google Scholar] [CrossRef]
- Mahmud, S.; Nazir, K.N.H.; Rahman, M.T. Prevalence and molecular detection of fluoroquinolone-resistant genes (qnrA and qnrS) in Escherichia coli isolated from healthy broiler chickens. Vet. World 2018, 11, 1720–1724. [Google Scholar] [CrossRef] [PubMed]
- Bayer, A.W.; Kirby, W.M.; Sherris, J.C.; Turck, M. Antibiotic susceptibility testing by a standardized single disc method. Am. J. Clin. Pathol. 1966, 45, 493–496. [Google Scholar] [CrossRef]
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 26th ed.; CLSI Supplement M100s; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2016. [Google Scholar]
- Sweeney, M.T.; Lubbers, B.V.; Schwarz, S.; Watts, J.L. Applying definitions for multidrug resistance, extensive drug resistance and pandrug resistance to clinically significant livestock and companion animal bacterial pathogens. J. Antimicrob. Chemother. 2018, 73, 1460–1463. [Google Scholar] [CrossRef]
- Varga, C.; Brash, M.L.; Slavic, D.; Boerlin, P.; Ouckama, R.; Weis, A.; Petrik, M.; Philippe, C.; Barham, M.; Guerin, M.T. Evaluating virulence-associated genes and antimicrobial resistance of avian pathogenic Escherichia coli isolates from broiler and broiler breeder chickens in Ontario, Canada. Avian Dis. 2018, 62, 291–299. [Google Scholar] [CrossRef]
- Logue, C.; Newman, D.; Nolan, L.; Barbieri, N. Characterizing avian pathogenic Escherichia coli from diagnostic cases in Georgia, USA–comparison of gene profiles with tissue of isolation. Acc. Microbiol. 2019, 1, 716. [Google Scholar] [CrossRef]
- Hadiujjaman, M.; Rahman, M.M.; Ahasan, M.D.; Banu, M.A.; Khatun, M.M.; Islam, M.A. Isolation and identification of Escherichia coli from apparently healthy chicken of selected areas of Bangladesh. Int. J. Nat. Soc. Sci. 2016, 3, 15–23. [Google Scholar]
- Islam, K.; Kabir, S.L.; Haque, A.K.M.Z.; Sarker, Y.A.; Sikder, M.H. Molecular detection and characterization of Escherichia coli, Salmonella spp. and Campylobacter spp. isolated from broiler meat in Jamalpur, Tangail, Netrokona and Kishoreganj districts of Bangladesh. Afr. J. Microbiol. Res. 2018, 12, 761–770. [Google Scholar]
- Al Azad, M.; Rahman, A.; Rahman, M.; Amin, R.; Begum, M.; Ara, I.; Fries, R.; Husna, A.; Khairalla, A.S.; Badruzzaman, A.T.M.; et al. Susceptibility and multidrug resistance patterns of Escherichia coli isolated from cloacal swabs of live broiler chickens in Bangladesh. Pathogens 2019, 8, 118. [Google Scholar] [CrossRef]
- Tonu, N.S.; Sufian, M.A.; Sarker, S.; Kamal, M.M.; Rahman, M.H.; Hossain, M.M. Pathological study on colibacillosis in chickens and detection of Escherichia coli by PCR. Bangladesh J. Vet. Med. 2011, 9, 17–25. [Google Scholar] [CrossRef]
- Khaton, R.; Haider, M.G.; Paul, P.K.; Das, P.M.; Hossain, M.M. Colibacillosis in commercial chickens in Bangladesh. Bangladesh Vet. 2008, 25, 17–24. [Google Scholar] [CrossRef][Green Version]
- Ibrahim, R.A.; Cryer, T.L.; Lafi, S.Q.; Basha, E.A.; Good, L.; Tarazi, Y.H. Identification of Escherichia coli from broiler chickens in Jordan, their antimicrobial resistance, gene characterization and the associated risk factors. BMC Vet. Res. 2019, 15, 159. [Google Scholar] [CrossRef]
- Duan, H.; Chai, T.; Cai, Y.; Zhong, Z.; Yao, M.; Zhang, X. Transmission identification of Escherichia coli aerosol in chicken houses to their environments using ERIC-PCR. Sci. China Life Sci. 2008, 51, 164–173. [Google Scholar] [CrossRef]
- Stromberg, Z.R.; Johnson, J.R.; Fairbrother, J.M.; Kilbourne, J.; Van Goor, A.; Curtiss, R., III; Mellata, M. Evaluation of Escherichia coli isolates from healthy chickens to determine their potential risk to poultry and human health. PLoS ONE 2017, 12, e0180599. [Google Scholar] [CrossRef] [PubMed]
- Azam, M.; Mohsin, M.; Saleemi, M.K. Virulence-associated genes and antimicrobial resistance among avian pathogenic Escherichia coli from colibacillosis affected broilers in Pakistan. Trop. Anim. Health Prod. 2019, 51, 1259–1265. [Google Scholar] [CrossRef]
- Handrova, L.; Kmet, V. Antibiotic resistance and virulence factors of Escherichia coli from eagles and goshawks. J. Environ. Sci. Health B 2019, 54, 605–614. [Google Scholar] [CrossRef] [PubMed]
- Chui, L.; Couturier, M.R.; Chiu, T.; Wang, G.; Olson, A.B.; McDonald, R.R.; Antonishyn, N.A.; Horsman, G.; Gilmour, M.W. Comparison of Shiga toxin-producing Escherichia coli detection methods using clinical stool samples. J. Mol. Diagn. 2010, 12, 469–475. [Google Scholar] [CrossRef] [PubMed]
- Sgariglia, E.; Mandolini, N.A.; Napoleoni, M.; Medici, L.; Fraticelli, R.; Conquista, M.; Gianfelici, P.; Staffolani, M.; Fisichella, S.; Capuccella, M.; et al. Antibiotic resistance pattern and virulence genes in avian pathogenic Escherichia coli (APEC) from different breeding systems. Vet. Ital. 2019, 55, 27–33. [Google Scholar]
- Kemmett, K.; Williams, N.J.; Chaloner, G.; Humphrey, S.; Wigley, P.; Humphrey, T. The contribution of systemic Escherichia coli infection to the early mortalities of commercial broiler chickens. Avian Pathol. 2014, 43, 37–42. [Google Scholar] [CrossRef]
- Dho, M.; Lafont, J.P. Escherichia coli colonization of the trachea in poultry: Comparison of virulent and avirulent strains in gnotoxenic chickens. Avian Dis. 1982, 26, 787–797. [Google Scholar] [CrossRef] [PubMed]
- Mol, N.; Peng, L.; Esnault, E.; Quéré, P.; Haagsman, H.P.; Veldhuizen, E.J. Avian pathogenic Escherichia coli infection of a chicken lung epithelial cell line. Vet. Immunol. Immunopathol. 2019, 210, 55–59. [Google Scholar] [CrossRef] [PubMed]
- Ewers, C.; Antão, E.M.; Diehl, I.; Philipp, H.C.; Wieler, L.H. Intestine and environment of the chicken as reservoirs for extraintestinal pathogenic Escherichia coli strains with zoonotic potential. Appl. Environ. Microbiol. 2009, 75, 184–192. [Google Scholar] [CrossRef] [PubMed]
- Obeng, A.S.; Rickard, H.; Ndi, O.; Sexton, M.; Barton, M. Antibiotic resistance, phylogenetic grouping and virulence potential of Escherichia coli isolated from the faeces of intensively farmed and free range poultry. Vet. Microbiol. 2012, 154, 305–315. [Google Scholar] [CrossRef]
- Kogovšek, P.; Ambrožič-Avguštin, J.; Dovč, A.; Dreo, T.; Hristov, H.; Krapež, U.; Ravnikar, M.; Slavec, B.; Lotrič, M.; Žel, J.; et al. Loop-mediated isothermal amplification: Rapid molecular detection of virulence genes associated with avian pathogenic Escherichia coli in poultry. Poult. Sci. 2019, 98, 1500–1510. [Google Scholar] [CrossRef]
- Jin, W.J.; Zheng, Z.M.; Zhang, Y.Z.; Qin, A.J.; Shao, H.X.; Liu, Y.L.; Jiao, W.A.N.G.; Wang, Q.Q. Distribution of virulence-associated genes of avian pathogenic Escherichia coli isolates in China. Agr. Sci. China 2008, 7, 1511–1515. [Google Scholar] [CrossRef]
- Kawano, M.; Yaguchi, K.; Osawa, R. Genotypic analyses of Escherichia coli isolated from chickens with colibacillosis and apparently healthy chickens in Japan. Microbiol. Immunol. 2006, 50, 961–966. [Google Scholar] [CrossRef]
- Camarda, A.; Circella, E.; Giovanardi, D.; Pennelli, D.; Battista, P.; Campagnari, E.; Bruni, G.; Tagliabue, S. Avian Pathogenic Escherichia coli in Audouin gulls (Larus audouinii) Could they affect the surviving of the bird colonies? Ital. J. Anim. 2007, 6, 317–320. [Google Scholar] [CrossRef]
- Jan, A.W.; Javed, M.T.; Lone, S.Q.; Aslam, M.S.; Javed, A. Association of six selected pathogenicity genes of Escherichia coli with gross and histopathological lesions in broiler chickens from field cases. Pak. J. Agri. Sci. 2018, 55, 433–440. [Google Scholar]
- Gao, Q.; Wang, X.; Xu, H.; Xu, Y.; Ling, J.; Zhang, D.; Gao, S.; Liu, X. Roles of iron acquisition systems in virulence of extraintestinal pathogenic Escherichia coli: Salmochelin and aerobactin contribute more to virulence than heme in a chicken infection model. BMC Microbiol. 2012, 12, 143. [Google Scholar] [CrossRef]
- Wright, G.D. The antibiotic resistome: The nexus of chemical and genetic diversity. Nat. Rev. Microbiol. 2007, 5, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Miles, T.D.; McLaughlin, W.; Brown, P.D. Antimicrobial resistance of Escherichia coli isolates from broiler chickens and humans. BMC Vet. Res. 2006, 2, 7. [Google Scholar] [CrossRef] [PubMed]
- Awad, A.M.; El-Shall, N.A.; Khalil, D.S.; El-Hack, A.; Mohamed, E.; Swelum, A.A.; Mahmoud, A.H.; Ebaid, H.; Komany, A.; Sammour, R.H.; et al. Incidence, pathotyping, and antibiotic susceptibility of avian pathogenic Escherichia coli among diseased broiler chicks. Pathogens 2020, 9, 114. [Google Scholar] [CrossRef] [PubMed]
- Ozawa, M.; Harada, K.; Kojima, A.; Asai, T.; Sameshima, T. Antimicrobial susceptibilities, serogroups, and molecular characterization of avian pathogenic Escherichia coli isolates in Japan. Avian Dis. 2008, 52, 392–397. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, M.O.; Ribeiro, A.R.; Santos, L.R.D.; Pilotto, F.; de Moraes, H.L.; Salle, C.T.P.; Rocha, S.L.D.S.; Nascimento, V.P.D. Antibiotic resistance in Salmonella enteritidis isolated from broiler carcasses. Braz. J. Microbiol. 2006, 37, 368–371. [Google Scholar] [CrossRef]
- Sarker, M.S.; Mannan, M.S.; Ali, M.Y.; Bayzid, M.; Ahad, A.; Bupasha, Z.B. Antibiotic resistance of Escherichia coli isolated from broilers sold at live bird markets in Chattogram, Bangladesh. J. Adv. Vet. Anim. Res. 2019, 6, 272–277. [Google Scholar] [CrossRef]
- Azad, M.A.R.A.; Amin, R.; Begum, M.I.A.; Fries, R.; Lampang, K.N.; Hafez, H.M. Prevalence of antimicrobial resistance of Escherichia coli isolated from broiler at Rajshahi region, Bangladesh. Br. J. Biomed. Multidisc. Res. 2017, 1, 6–12. [Google Scholar]
- Bashar, T.; Rahman, M.; Rabbi, F.A.; Noor, R.; Rahman, M.M. Enterotoxin profiling and antibiogram of Escherichia coli isolated from poultry feces in Dhaka district of Bangladesh. Stamford J. Microbiol. 2011, 1, 51–57. [Google Scholar] [CrossRef]
- Kim, T.E.; Jeong, Y.W.; Cho, S.H.; Kim, S.J.; Kwon, H.J. Chronological study of antibiotic resistances and their relevant genes in Korean avian pathogenic Escherichia Coli isolates. J. Clin. Microbiol. 2007, 45, 3309–3315. [Google Scholar] [CrossRef]
- Dou, X.; Gong, J.; Han, X.; Xu, M.; Shen, H.; Zhang, D.; Zhuang, L.; Liu, J.; Zou, J. Characterization of avian pathogenic Escherichia coli isolated in eastern China. Gene 2016, 576, 244–248. [Google Scholar] [CrossRef]
- Hoelzer, K.; Wong, N.; Thomas, J.; Talkington, K.; Jungman, E.; Coukell, A. Antimicrobial drug use in food-producing animals and associated human health risks: What, and how strong, is the evidence? BMC Vet. Res. 2017, 13, 211. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Liu, Z.; Zhang, Y.; Yuan, X.; Hu, M.; Liu, Y. Prevalence and molecular characteristics of avian-origin mcr-1-harboring Escherichia coli in Shandong province, China. Front. Microbiol. 2020, 11, 255. [Google Scholar] [CrossRef] [PubMed]
- van den Bogaard, A.E.; Stobberingh, E.E. Epidemiology of resistance to antibiotics: Links between animals and humans. Int. J. Antimicrob. Agents 2000, 14, 327–335. [Google Scholar] [CrossRef]
| Target Genes | Primer Sequence (5′–3′) | Amplicon Size (bp) | Annealing Temperature (°C) | References |
|---|---|---|---|---|
| 16S rRNA | F: GACCTCGGTTTAGTTCACAGA R: CACACGCTGACGCTGACCA | 585 | 55 | [35] |
| fimC | F: GGGTAGAAAATGCCGATGGTG R: CGTCATTTTGGGGGTAAGTGC | 496 | 59 | [12] |
| iucD | F: ACAAAAAGTTCTATCGCTTCC R: CCTGATCCAGCTGATGCTC | 692 | 55 | |
| papC | F: TGATATCACGCAGTCAGTAGC R: CCGGCCATATTCACATAA | 483 | 59 |
| Sample Source/Nature of Sample | No. of Samples Analyzed | No. Overall Analyzed | No. E. coli Positive Samples | Overall Positive for E. coli | Prevalence (%) | p-Value | |
|---|---|---|---|---|---|---|---|
| Air | 31 | 31 | 21 | 21 | 67.74 | 0.003 | |
| Feces | 32 | 32 | 32 | 32 | 100 | ||
| Internal organs | Trachea | 5 | 36 | 3 | 29 | 80.56 | |
| Intestine | 8 | 7 | |||||
| Liver | 14 | 11 | |||||
| Lung | 7 | 6 | |||||
| Egg Yolk | 2 | 2 | |||||
| Total | 99 | 99 | 82 | 82 | 82.83 | ||
| Samples | Name of Positive Isolate | APEC-Associated Virulence Genes | No. of Positive Isolates (%) | p-Value | |||
|---|---|---|---|---|---|---|---|
| fimC | iucD | papC | |||||
| Air (n = 31) | A1 | + | + | + | 5 (16.13) | ˂0.001 | |
| A2 | + | - | + | ||||
| A3 | + | + | + | ||||
| A4 | + | + | + | ||||
| A5 | + | + | + | ||||
| Feces (n = 32) | F2 | + | + | - | 7 (21.87) | ||
| F3 | + | + | + | ||||
| F4 | + | - | + | ||||
| F5 | + | - | + | ||||
| F6 | + | + | + | ||||
| F7 | + | - | - | ||||
| F9 | - | - | + | ||||
| Internal organs (n = 36) | Trachea (n = 5) | Io-T2 | + | + | - | 24 (66.67) | |
| Io-T3 | + | + | + | ||||
| Intestine (n = 8) | Io-I2 | + | + | - | |||
| Io-I3 | + | + | - | ||||
| Io-I4 | + | - | - | ||||
| Io-I5 | + | + | - | ||||
| Io-I6 | + | + | - | ||||
| Io-I7 | + | - | + | ||||
| Io-I8 | + | - | - | ||||
| Liver (n = 14) | Io-L1 | + | + | - | |||
| Io-L2 | + | + | - | ||||
| Io-L3 | + | - | - | ||||
| Io-L4 | + | - | - | ||||
| Io-L6 | + | - | - | ||||
| Io-L7 | + | - | - | ||||
| Io-L9 | + | + | - | ||||
| Io-L10 | + | + | - | ||||
| Io-L13 | + | + | - | ||||
| Lung (n = 7) | Io-Lu1 | + | - | - | |||
| Io-Lu2 | + | - | - | ||||
| Io-Lu5 | + | - | - | ||||
| Io-Lu7 | + | + | - | ||||
| Egg Yolk (n = 2) | Io-Y1 | + | + | - | |||
| Io-Y2 | + | + | - | ||||
| Total (n = 99) | 36 | 35 (97.22%) | 21 (58.33%) | 12 (33.33%) | 36 (36.36%) | ||
| p-value | 0.003 | ||||||
| SL No. | Isolate Name | CL | CIP | EX | NX | AMP | TE | S | GEN | C | E |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | A1 | S | S | S | S | R | R | S | S | R | R |
| 2 | A2 | S | S | S | S | R | R | I | I | R | R |
| 3 | A3 | S | R | R | R | R | R | R | S | R | R |
| 4 | A4 | S | R | R | R | R | R | R | S | R | R |
| 5 | A5 | S | S | I | S | R | R | S | S | R | R |
| 6 | F2 | S | R | R | R | R | R | S | R | R | R |
| 7 | F3 | S | R | R | R | R | R | R | S | R | R |
| 8 | F4 | S | R | R | R | R | R | I | S | R | R |
| 9 | F5 | S | S | I | S | R | R | R | S | R | R |
| 10 | F6 | S | S | I | S | R | R | R | S | R | R |
| 11 | F7 | S | S | S | S | R | R | S | S | R | R |
| 12 | F9 | S | S | S | R | R | R | S | S | R | R |
| 13 | Io-T2 | S | S | S | S | R | R | S | I | R | R |
| 14 | Io-T3 | S | R | R | R | R | R | R | S | R | R |
| 15 | Io-I2 | S | R | R | R | R | R | S | S | R | R |
| 16 | Io-I3 | S | R | R | R | R | R | S | S | R | R |
| 17 | Io-I4 | S | I | R | I | R | R | S | S | R | R |
| 18 | Io-I5 | R | R | R | I | R | R | S | R | R | R |
| 19 | Io-I6 | S | S | R | S | R | R | S | S | R | R |
| 20 | Io-I7 | S | I | R | R | R | R | S | R | R | R |
| 21 | Io-I8 | S | R | S | S | R | R | S | S | R | R |
| 22 | Io-L1 | R | R | R | R | R | R | S | S | R | R |
| 23 | Io-L2 | S | R | R | R | R | R | S | S | R | R |
| 24 | Io-L3 | S | R | R | R | R | R | I | S | R | I |
| 25 | Io-L4 | S | S | S | S | R | R | I | S | R | R |
| 26 | Io-L6 | S | S | S | S | R | R | S | S | R | R |
| 27 | Io-L7 | S | S | S | S | R | R | S | S | R | R |
| 28 | Io-L9 | S | S | S | S | R | R | S | S | R | R |
| 29 | Io-L10 | S | R | R | R | R | R | S | S | I | R |
| 30 | Io-L13 | S | R | R | R | R | R | S | S | R | R |
| 31 | Io-Lu1 | S | R | R | R | R | R | I | S | R | R |
| 32 | Io-Lu2 | R | R | R | R | R | R | R | S | R | R |
| 33 | Io-Lu5 | S | S | S | S | R | R | S | S | R | R |
| 34 | Io-Lu7 | S | I | I | S | R | R | S | S | R | R |
| 35 | Io-Y1 | S | R | R | R | R | R | S | S | R | R |
| 36 | Io-Y2 | R | S | I | S | R | R | S | S | R | R |
| Pattern No. | Antibiotic Resistance Pattern | No. of Antibiotics (Classes) | Isolate No. | No. of Isolates (%) |
|---|---|---|---|---|
| 1 | AMP, TE, C, E | 4 (4) | A2, A5, F6, F7, Io-T2, Io-L4, Io-L6, Io-L7, Io-L9, Io-Lu5, Io-Lu7 | 11 (30.55) |
| 2 | AMP, TE, S, C, E | 5 (5) | F5, F6 | 2 (5.55) |
| 3 | CL, AMP, TE, C, E | 5 (5) | Io-Y2 | 1 (2.78) |
| 4 | EX, AMP, TE, C, E | 5 (5) | Io-I4, Io-I6 | 2 (5.55) |
| 5 | NX, AMP, TE, C, E | 5 (5) | F9 | 1 (2.78) |
| 6 | CIP, AMP, TE, C, E | 5 (5) | Io-I8 | 1 (2.78) |
| 7 | CIP, EX, NX, AMP, TE, E | 6 (4) | Io-L10 | 1 (2.78) |
| 8 | CIP, EX, NX, AMP, TE, C | 6 (4) | Io-L3 | 1 (2.78) |
| 9 | CIP, EX, NX, AMP, TE, C, E | 7 (5) | F4, Io-L2, Io-Y1, Io-I2, Io-I3, Io-L13 | 6 (16.67) |
| 10 | EX, NX, AMP, TE, GEN, C, E | 7 (6) | Io-I7 | 1 (2.78) |
| 11 | CIP, EX, NX, AMP, TE, C, E | 7 (5) | Io-Lu1 | 1 (2.78) |
| 12 | CIP, EX, NX, AMP, TE, S, C, E | 8 (6) | A3, A4, F3, Io-T3 | 4 (11.11) |
| 13 | CL, CIP, EX, AMP, TE, GEN, C, E | 8 (7) | Io-I5 | 1 (2.78) |
| 14 | CIP, EX, NX, AMP, TE, GEN, C, E | 8 (6) | F2 | 1 (2.78) |
| 15 | CL, CIP, EX, NX, AMP, TE, C, E | 8 (6) | Io-L1 | 1 (2.78) |
| 16 | CL, CIP, EX, NX, AMP, TE, S, C, E | 9 (7) | Io-Lu2 | 1 (2.78) |
| Total | 36 |
| CL | CIP | EX | NX | AMP | TE | S | C | GEN | E | |
|---|---|---|---|---|---|---|---|---|---|---|
| CL | 1.0000 | - | - | - | - | - | - | - | - | - |
| CIP | 0.1195 | 1.0000 | - | - | - | - | - | - | - | - |
| EX | 0.2345 | 0.6625 * | 1.0000 | - | - | - | - | - | - | - |
| NX | 0.1383 | 0.8315 * | 0.6203 * | 1.0000 | - | - | - | - | - | - |
| AMP | - | - | - | - | - | - | - | - | - | - |
| TE | - | - | - | - | - | - | - | - | - | - |
| S | −0.0625 | 0.1195 | 0.2132 | 0.1581 | - | - | 1.0000 | - | - | - |
| C | - | - | - | - | - | - | - | - | - | - |
| GEN | 0.1136 | 0.0136 | −0.0823 | 0.0359 | - | - | −0.1136 | - | 1.0000 | - |
| E | - | - | - | - | - | - | - | - | - | - |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ievy, S.; Islam, M.S.; Sobur, M.A.; Talukder, M.; Rahman, M.B.; Khan, M.F.R.; Rahman, M.T. Molecular Detection of Avian Pathogenic Escherichia coli (APEC) for the First Time in Layer Farms in Bangladesh and Their Antibiotic Resistance Patterns. Microorganisms 2020, 8, 1021. https://doi.org/10.3390/microorganisms8071021
Ievy S, Islam MS, Sobur MA, Talukder M, Rahman MB, Khan MFR, Rahman MT. Molecular Detection of Avian Pathogenic Escherichia coli (APEC) for the First Time in Layer Farms in Bangladesh and Their Antibiotic Resistance Patterns. Microorganisms. 2020; 8(7):1021. https://doi.org/10.3390/microorganisms8071021
Chicago/Turabian StyleIevy, Samina, Md. Saiful Islam, Md. Abdus Sobur, Mithun Talukder, Md. Bahanur Rahman, Mohammad Ferdousur Rahman Khan, and Md. Tanvir Rahman. 2020. "Molecular Detection of Avian Pathogenic Escherichia coli (APEC) for the First Time in Layer Farms in Bangladesh and Their Antibiotic Resistance Patterns" Microorganisms 8, no. 7: 1021. https://doi.org/10.3390/microorganisms8071021
APA StyleIevy, S., Islam, M. S., Sobur, M. A., Talukder, M., Rahman, M. B., Khan, M. F. R., & Rahman, M. T. (2020). Molecular Detection of Avian Pathogenic Escherichia coli (APEC) for the First Time in Layer Farms in Bangladesh and Their Antibiotic Resistance Patterns. Microorganisms, 8(7), 1021. https://doi.org/10.3390/microorganisms8071021








