Antiviral Effects of Curcumin on Adenovirus Replication
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell lines, Viruses and Reagents
2.2. Infection and Drug Treatments
2.3. Immunoblot Analysis
2.4. Quantitative Real-Time PCR (qPCR)
2.5. Plaque Assay for Virus Yield
2.6. MTS Metabolic Activity Assays
2.7. Statistical Analysis
3. Results
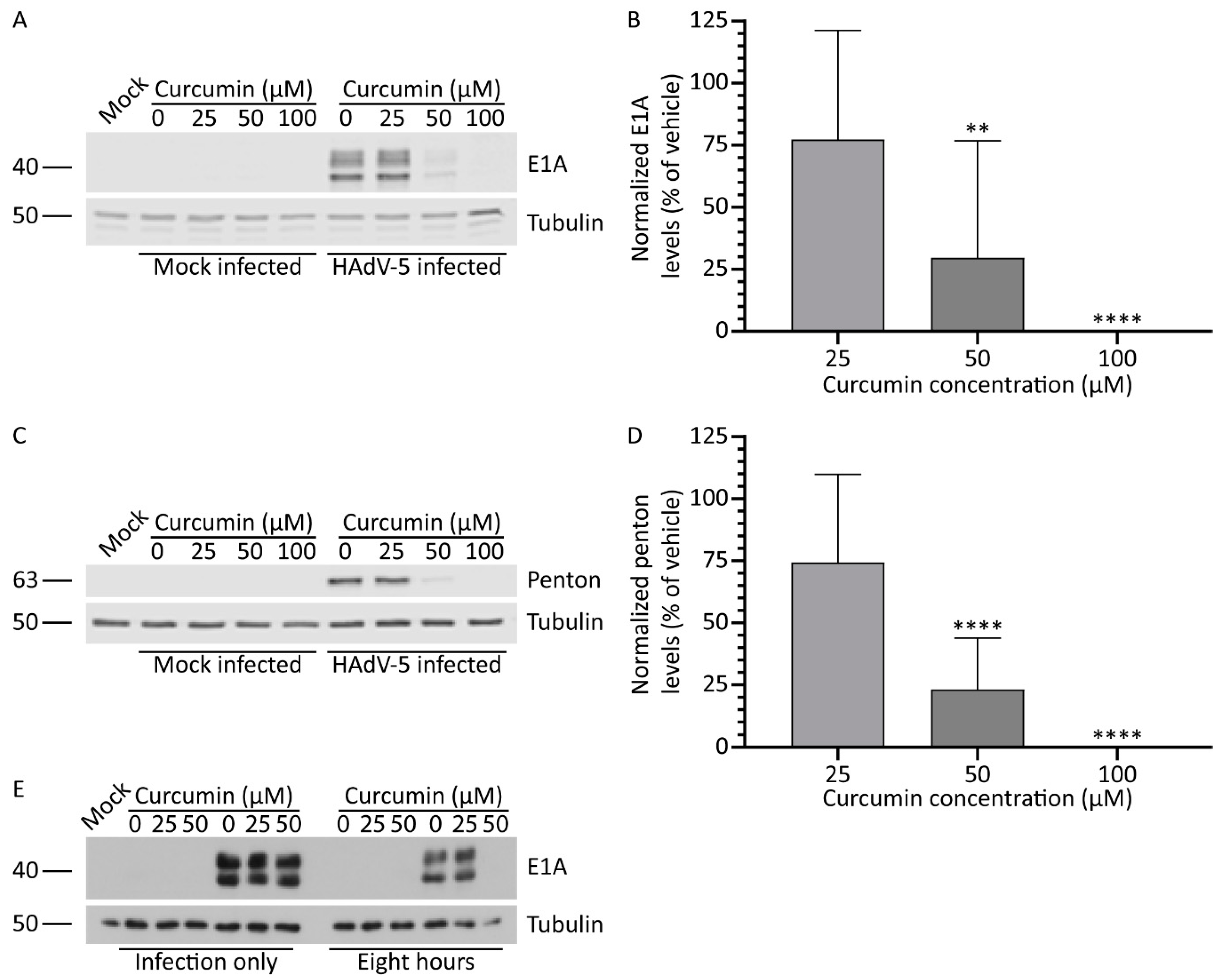
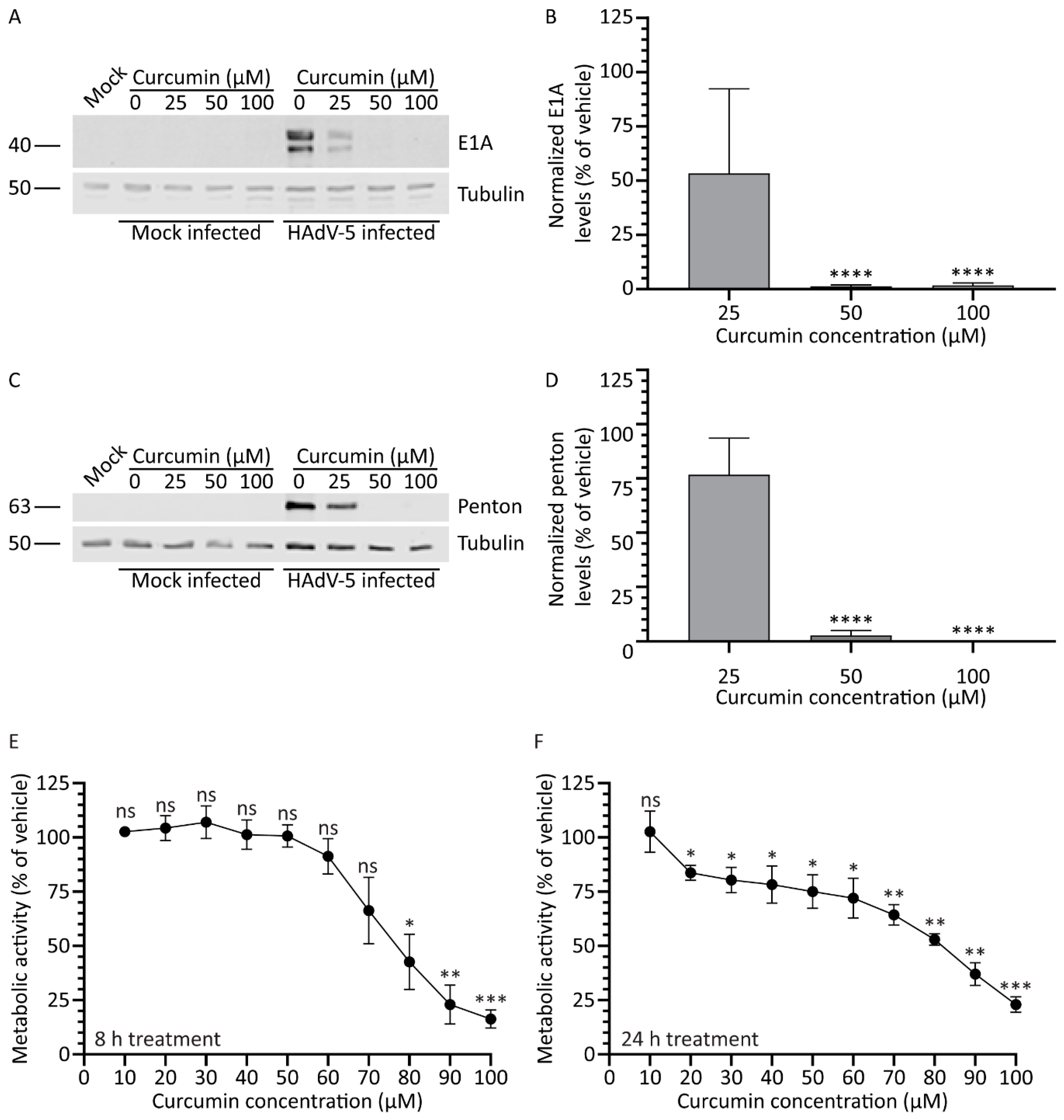
3.1. Treatment with Curcumin Reduces HAdV-5 Protein Expression
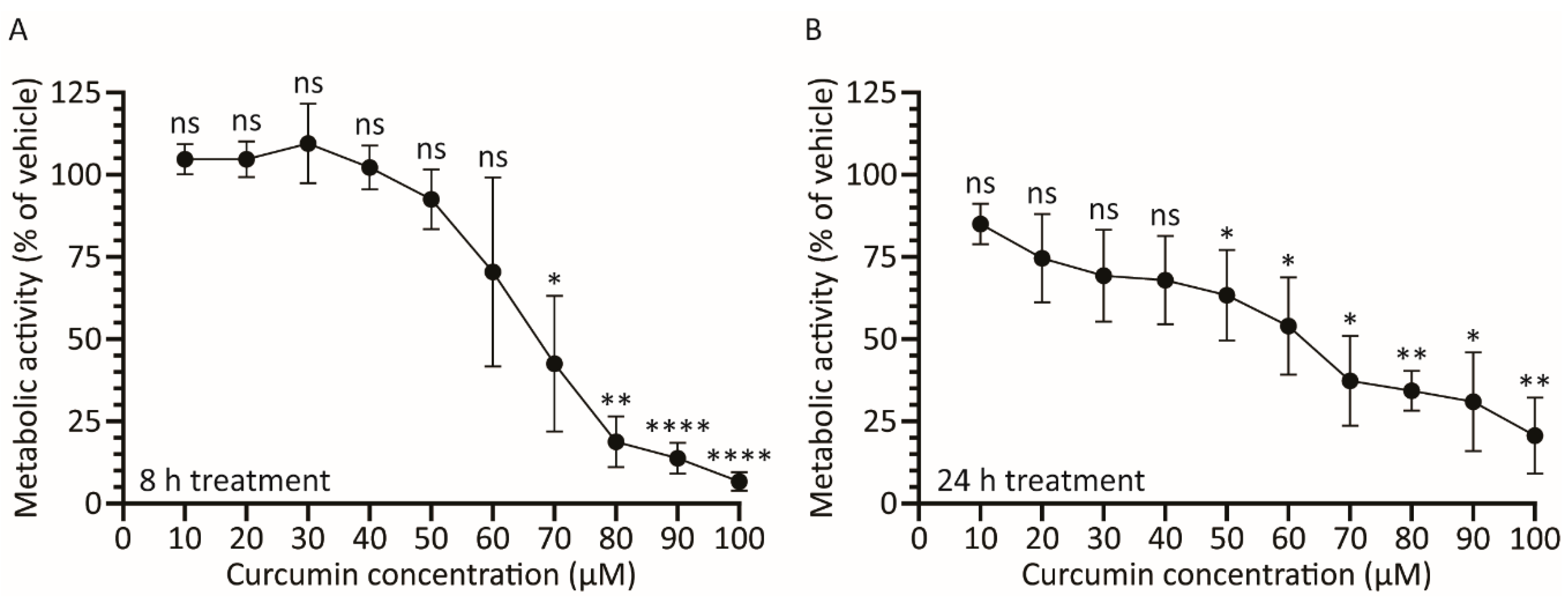
3.2. Treatment with Curcumin Causes a Dose-Dependent Decrease in A459 Cellular Metabolic Activity
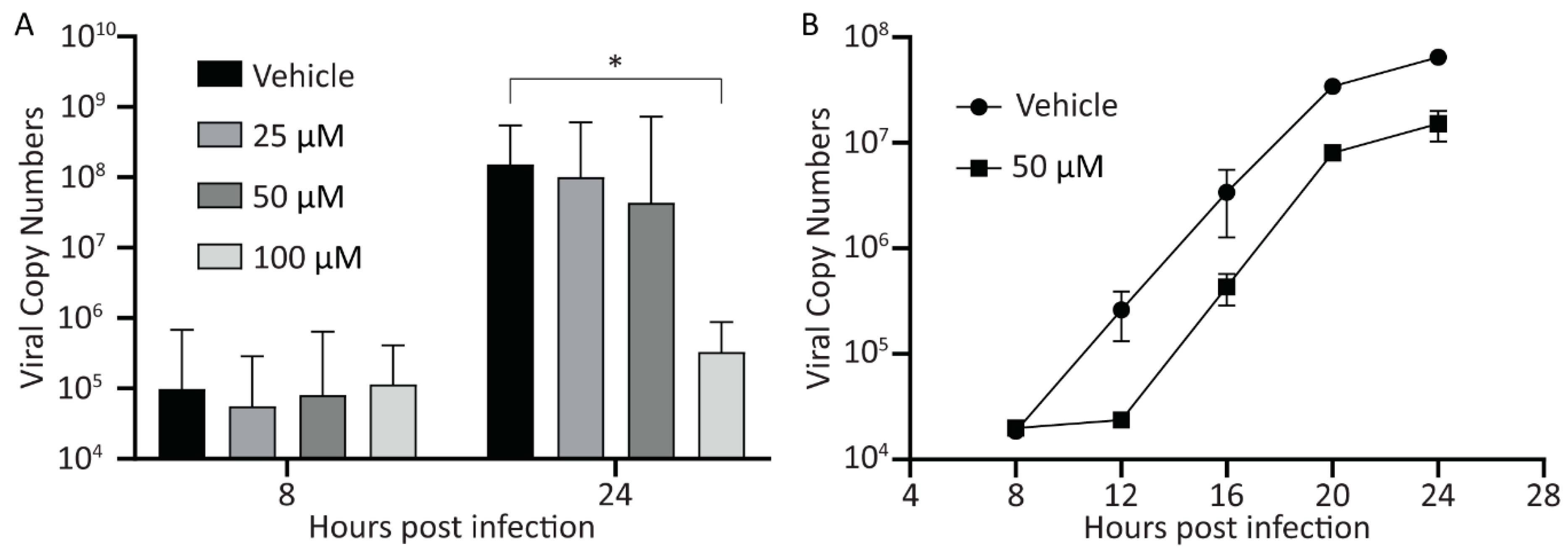
3.3. Treatment with Curcumin Reduces HAdV-5 Genome Copy Number within Cells
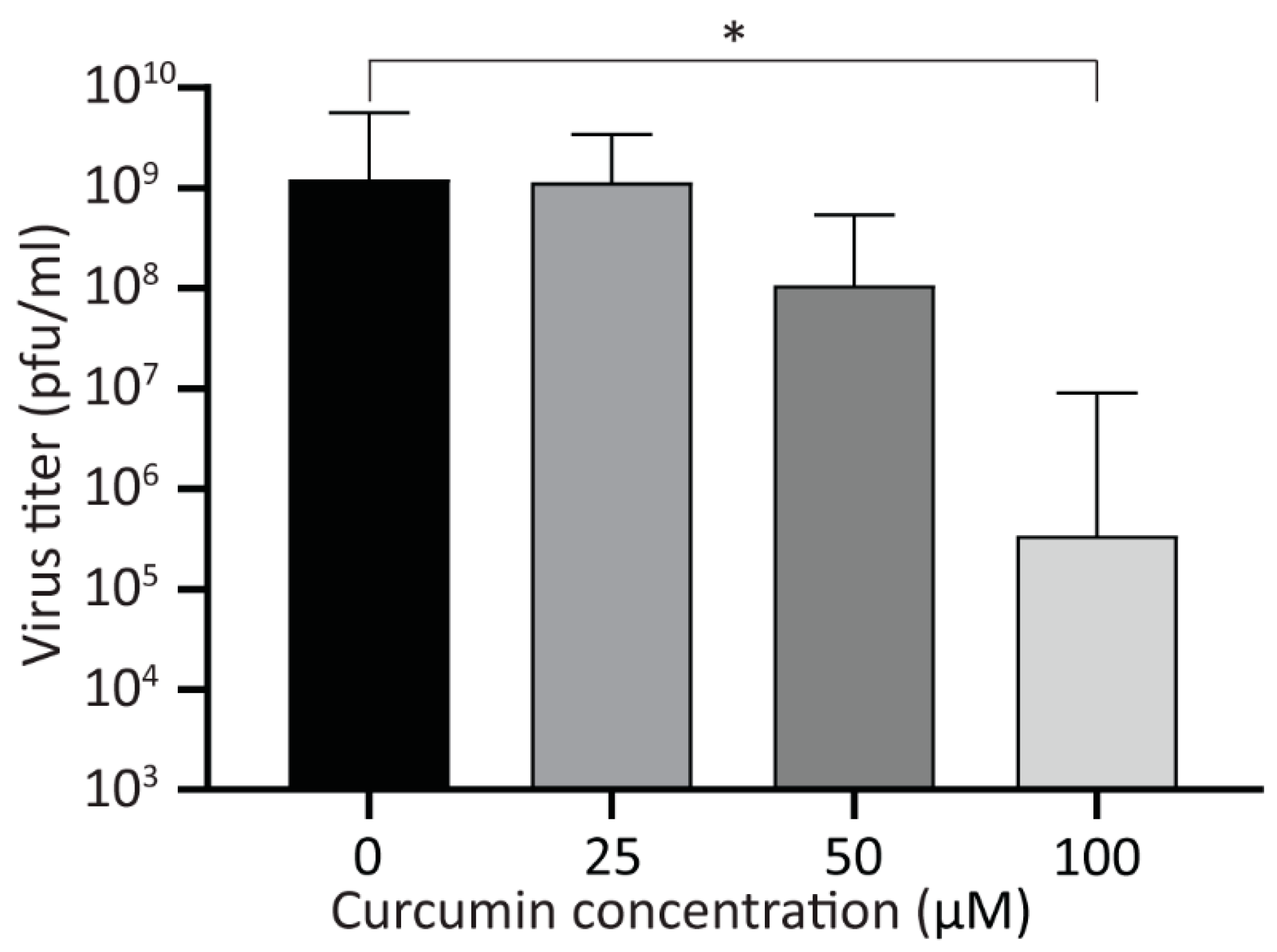
3.4. Treatment with Curcumin Reduces Viral Yield
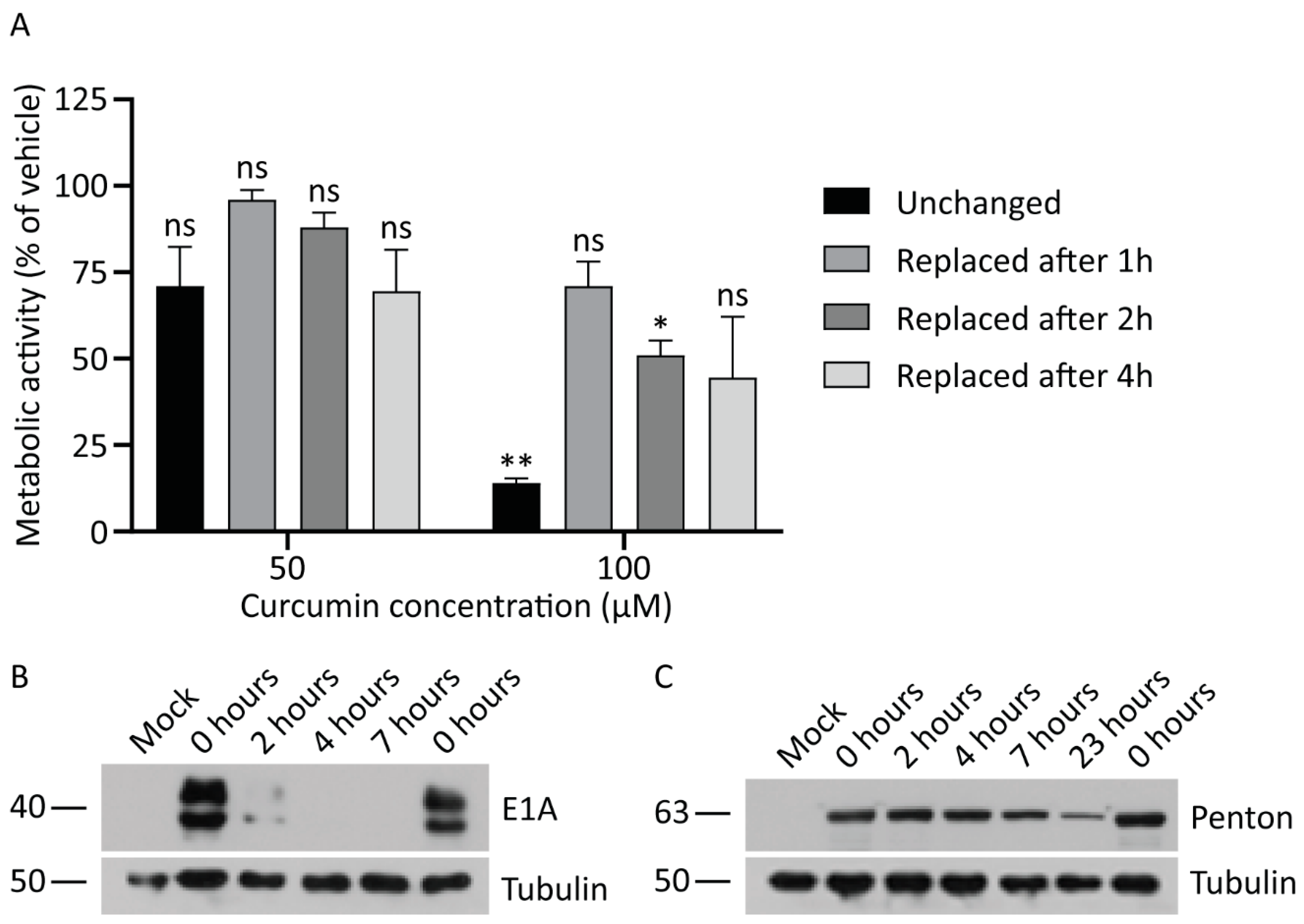
3.5. Continued Exposure to Curcumin is Required to Inhibit HAdV Protein Expression
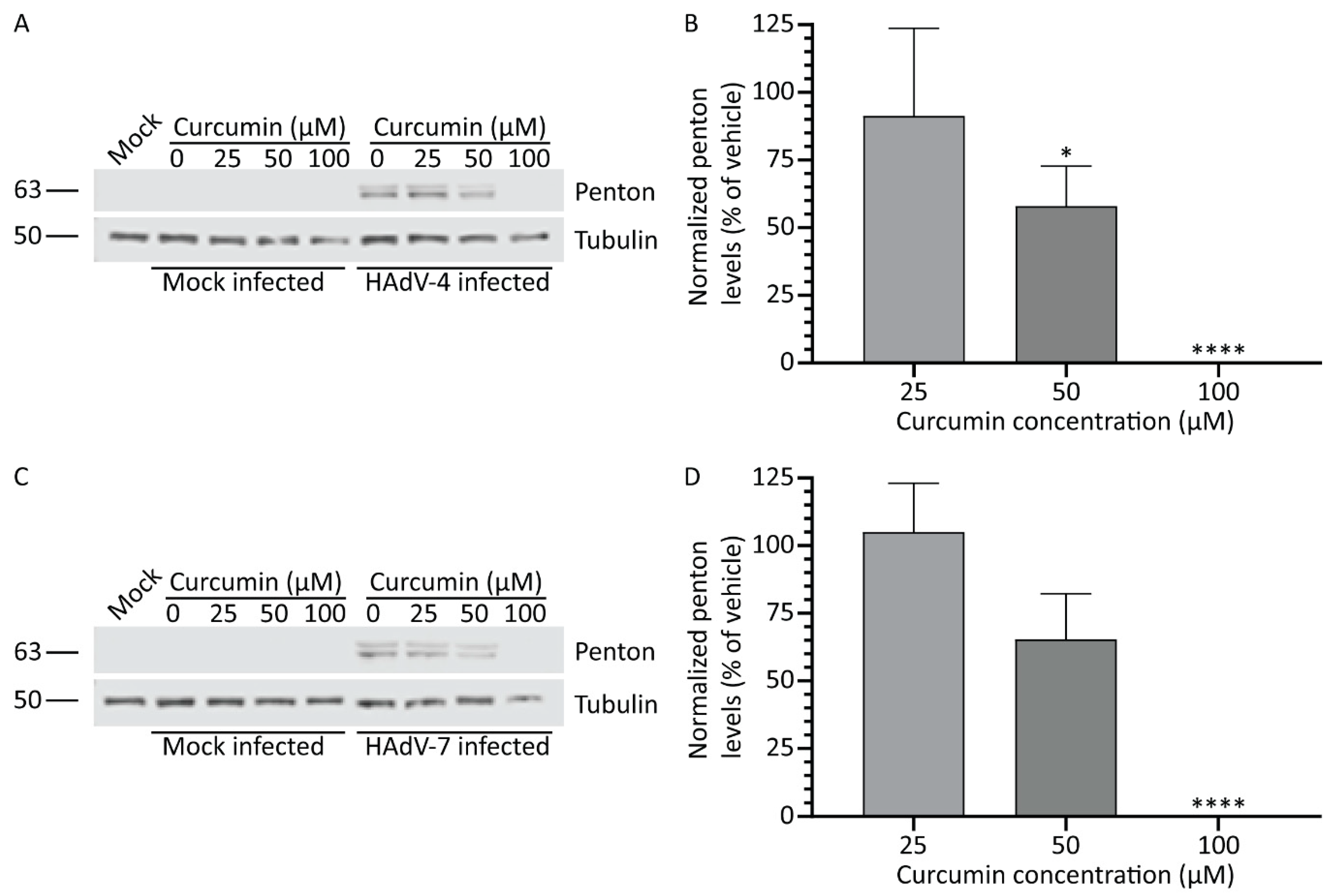
3.6. Treatment with Curcumin Reduces HAdV Types 4 and 7 Protein Levels
3.7. Treatment with Curcumin of Higher Purity Improves Efficacy and Selectivity against HAdV
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Butt, A.L.; Chodosh, J. Adenoviral keratoconjunctivitis in a tertiary care eye clinic. Cornea 2006, 25, 199–202. [Google Scholar] [CrossRef] [PubMed]
- Dhingra, A.; Hage, E.; Ganzenmueller, T.; Böttcher, S.; Hofmann, J.; Hamprecht, K.; Obermeier, P.; Rath, B.; Hausmann, F.; Dobner, T.; et al. Molecular evolution of human adenovirus (HAdV) species C. Sci. Rep. 2019, 9, 1039. [Google Scholar] [CrossRef] [PubMed]
- Lenaerts, L.; De Clercq, E.; Naesens, L. Clinical features and treatment of adenovirus infections. Rev. Med. Virol. 2008, 18, 357–374. [Google Scholar] [CrossRef] [PubMed]
- Lichtenstein, D.L.; Wold, W.S.M. Experimental infections of humans with wild-type adenoviruses and with replication-competent adenovirus vectors: Replication, safety, and transmission. Cancer Gene Ther. 2004, 11, 819–829. [Google Scholar] [CrossRef]
- Lion, T. Adenovirus infections in immunocompetent and immunocompromised patients. Clin. Microbiol. Rev. 2014, 27, 441–462. [Google Scholar] [CrossRef]
- Saha, B.; Parks, R.J. Recent advances in novel antiviral therapies against human adenovirus. Microorganisms 2020, 8, 1284. [Google Scholar] [CrossRef]
- Chamberlain, J.M.; Sortino, K.; Sethna, P.; Bae, A.; Lanier, R.; Bambara, R.A.; Dewhurst, S. Cidofovir diphosphate inhibits adenovirus 5 DNA polymerase via both nonobligate chain termination and direct inhibition, and polymerase mutations confer cidofovir resistance on intact virus. Antimicrob. Agents Chemother. 2018, 63, e01925-18. [Google Scholar] [CrossRef]
- Takamatsu, A.; Tagashira, Y.; Hasegawa, S.; Honda, H. Disseminated adenovirus infection in a patient with a hematologic malignancy: A case report and literature review. Future Sci. OA 2019, 5, FSO412. [Google Scholar] [CrossRef]
- Izzedine, H.; Launay-Vacher, V.; Deray, G. Antiviral drug-induced nephrotoxicity. Am. J. Kidney Dis. 2005, 45, 804–817. [Google Scholar] [CrossRef]
- Nagafuji, K.; Aoki, K.; Henzan, H.; Kato, K.; Miyamoto, T.; Eto, T.; Nagatoshi, Y.; Ohba, T.; Obama, K.; Gondo, H.; et al. Cidofovir for treating adenoviral hemorrhagic cystitis in hematopoietic stem cell transplant recipients. Bone Marrow Transplant. 2004, 34, 909–914. [Google Scholar] [CrossRef]
- Sudhindra, P.; Knoll, B.; Nog, R.; Singh, N.; Dhand, A. brincidofovir (CMX001) for the treatment of severe adenoviral pneumonia in kidney transplant recipient. Cureus 2019, 11, e5296. [Google Scholar] [CrossRef] [PubMed]
- Lopez, S.M.C.; Michaels, M.G.; Green, M. Adenovirus infection in pediatric transplant recipients: Are effective antiviral agents coming our way? Curr. Opin. Organ. Transplant. 2018, 23, 395–399. [Google Scholar] [CrossRef] [PubMed]
- Florescu, D.F.; Schaenman, J.M.; AST Infectious Diseases Community of Practice. Adenovirus in solid organ transplant recipients: Guidelines from the American Society of Transplantation Infectious Diseases Community of Practice. Clin. Transplant. 2019, 33, e13527. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Cespedes, J.; Moyer, C.L.; Whitby, L.R.; Boger, D.L.; Nemerow, G.R. Inhibition of adenovirus replication by a trisubstituted piperazin-2-one derivative. Antivir. Res. 2014, 108, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Hartline, C.B.; Keith, K.A.; Eagar, J.; Harden, E.A.; Bowlin, T.L.; Prichard, M.N. A standardized approach to the evaluation of antivirals against DNA viruses: Orthopox-, adeno-, and herpesviruses. Antivir. Res. 2018, 159, 104–112. [Google Scholar] [CrossRef]
- Saha, B.; Varette, O.; Stanford, W.L.; Diallo, J.-S.; Parks, R.J. Development of a novel screening platform for the identification of small molecule inhibitors of human adenovirus. Virology 2019, 538, 24–34. [Google Scholar] [CrossRef]
- Saha, B.; Parks, R.J. Identification of human adenovirus replication inhibitors from a library of small molecules targeting cellular epigenetic regulators. Virology 2020. [Google Scholar] [CrossRef]
- Hussein, I.T.M.; Brooks, J.; Bowlin, T.L. The discovery and development of filociclovir for the prevention and treatment of human cytomegalovirus-related disease. Antivir. Res. 2020, 176, 104710. [Google Scholar] [CrossRef]
- Saha, B.; Parks, R.J. Histone deacetylase inhibitor suberoylanilide hydroxamic acid suppresses human adenovirus gene expression and replication. J. Virol. 2019, 93, e00088-19. [Google Scholar] [CrossRef]
- Lee, W.-H.; Loo, C.-Y.; Bebawy, M.; Luk, F.; Mason, R.S.; Rohanizadeh, R. Curcumin and its derivatives: Their application in neuropharmacology and neuroscience in the 21st century. Curr. Neuropharmacol. 2013, 11, 338–378. [Google Scholar] [CrossRef]
- Kotha, R.R.; Luthria, D.L. Curcumin: Biological, pharmaceutical, nutraceutical, and analytical aspects. Molecules 2019, 24, 2930. [Google Scholar] [CrossRef] [PubMed]
- Liczbiński, P.; Michałowicz, J.; Bukowska, B. Molecular mechanism of curcumin action in signaling pathways: Review of the latest research. Phytother. Res. 2020. [Google Scholar] [CrossRef] [PubMed]
- Ismail, N.I.; Othman, I.; Abas, F.H.; Lajis, N.; Naidu, R. Mechanism of apoptosis induced by curcumin in colorectal cancer. Int. J. Mol. Sci. 2019, 20, 2454. [Google Scholar] [CrossRef]
- Kim, S.-R.; Park, H.-J.; Bae, Y.-H.; Ahn, S.-C.; Wee, H.-J.; Yun, I.; Jang, H.-O.; Bae, M.-K.; Bae, S.-K. Curcumin down-regulates visfatin expression and inhibits breast cancer cell invasion. Endocrinology 2012, 153, 554–563. [Google Scholar] [CrossRef] [PubMed]
- Clark, C.A.; McEachern, M.D.; Shah, S.H.; Rong, Y.; Rong, X.; Smelley, C.L.; Caldito, G.C.; Abreo, F.W.; Nathan, C.O. Curcumin inhibits carcinogen and nicotine-induced mammalian target of rapamycin pathway activation in head and neck squamous cell carcinoma. Cancer Prev. Res. 2010, 3, 1586–1595. [Google Scholar] [CrossRef]
- Wilken, R.; Veena, M.S.; Wang, M.B.; Srivatsan, E.S. Curcumin: A review of anti-cancer properties and therapeutic activity in head and neck squamous cell carcinoma. Mol. Cancer 2011, 10, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Sidhu, G.S.; Singh, A.K.; Thaloor, D.; Banaudha, K.K.; Patnaik, G.K.; Srimal, R.C.; Maheshwari, R.K. Enhancement of wound healing by curcumin in animals. Wound Repair Regen. 1998, 6, 167–177. [Google Scholar] [CrossRef]
- Tyagi, P.; Singh, M.; Kumari, H.; Kumari, A.; Mukhopadhyay, K. Bactericidal activity of curcumin I is associated with damaging of bacterial membrane. PLoS ONE 2015, 10, e0121313. [Google Scholar] [CrossRef]
- Praditya, D.; Kirchhoff, L.; Brüning, J.; Rachmawati, H.; Steinmann, J.; Steinmann, E. Anti-infective properties of the golden spice curcumin. Front. Microbiol. 2019, 10, 1–16. [Google Scholar] [CrossRef]
- Richart, S.M.; Li, Y.L.; Mizushina, Y.; Chang, Y.Y.; Chung, T.Y.; Chen, G.H.; Tzen, J.T.; Shia, K.S.; Hsu, W.L. Synergic effect of curcumin and its structural analogue (Monoacetylcurcumin) on anti-influenza virus infection. J. Food Drug Anal. 2018, 26, 1015–1023. [Google Scholar] [CrossRef]
- Mazumder, A.; Raghavan, K.; Weinstein, J.; Kohn, K.W.; Pommier, Y. Inhibition of human immunodeficiency virus type-1 integrase by curcumin. Biochem. Pharmacol. 1995, 49, 1165–1170. [Google Scholar] [CrossRef]
- Ali, A.; Banerjea, A.C. Curcumin inhibits HIV-1 by promoting Tat protein degradation. Sci. Rep. 2016, 6, 27539. [Google Scholar] [CrossRef] [PubMed]
- Dai, J.; Gu, L.; Su, Y.; Wang, Q.; Zhao, Y.; Chen, X.; Deng, H.; Li, W.; Wang, G.; Li, K. Inhibition of curcumin on influenza A virus infection and influenzal pneumonia via oxidative stress, TLR2/4, p38/JNK MAPK and NF-κB pathways. Int. Immunopharmacol. 2018, 54, 177–187. [Google Scholar] [CrossRef] [PubMed]
- Balasubramanian, A.; Pilankatta, R.; Teramoto, T.; Sajith, A.M.; Nwulia, E.; Kulkarni, A.; Padmanabhan, R. Inhibition of dengue virus by curcuminoids. Antivir. Res. 2019, 162, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Crisostomo, L.; Soriano, A.M.; Frost, J.R.; Olanubi, O.; Mendez, M.; Pelka, P. The influence of E1A C-terminus on adenovirus replicative cycle. Viruses 2017, 9, 387. [Google Scholar] [CrossRef]
- Yang, M.; Lee, G.; Si, J.; Lee, S.-J.; You, H.J.; Ko, G. Curcumin shows antiviral properties against norovirus. Molecules 2016, 21, 1401. [Google Scholar] [CrossRef]
- Mounce, B.C.; Cesaro, T.; Carrau, L.; Vallet, T.; Vignuzzi, M. Curcumin inhibits Zika and chikungunya virus infection by inhibiting cell binding. Antivir. Res. 2017, 142, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Wen-Tsai, J.; Hung, J.L. PI3K-Akt signaling and viral infection. Recent Pat. Biotechnol. 2008, 2, 218–226. [Google Scholar] [CrossRef]
- Li, E.; Stupack, D.; Klemke, R.; Cheresh, D.A.; Nemerow, G.R. Adenovirus endocytosis via alpha(v) integrins requires phosphoinositide-3-OH kinase. J. Virol. 1998, 72, 2055–2061. [Google Scholar] [CrossRef]
- Saha, B.; Parks, R.J. Human adenovirus type 5 vectors deleted of early region 1 (E1) undergo limited expression of early replicative E2 proteins and DNA replication in non-permissive cells. PLoS ONE 2017, 12, e0181012. [Google Scholar] [CrossRef]
- Binder, A.M.; Biggs, H.M.; Haynes, A.K.; Chommanard, C.; Lu, X.; Erdman, D.D.; Watson, J.T.; Gerber, S.I. Human adenovirus surveillance-United States, 2003–2016. Mmwr. Morb. Mortal. Wkly. Rep. 2017, 66, 1039–1042. [Google Scholar] [CrossRef] [PubMed]
- Adamson, C.S.; Nevels, M.M. Bright and early: Inhibiting human cytomegalovirus by targeting major immediate-early gene expression or protein function. Viruses 2020, 12, 110. [Google Scholar] [CrossRef] [PubMed]
- Ramachandran, C.; Rodriguez, S.; Ramachandran, R.; Raveendran Nair, P.K.; Fonseca, H.; Khatib, Z.; Escalon, E.; Melnick, S.J. Expression profiles of apoptotic genes induced by curcumin in human breast cancer and mammary epithelial cell lines. Anticancer Res. 2005, 25, 3293–3302. [Google Scholar] [PubMed]
- Ullman, A.J.; Hearing, P. Cellular proteins PML and Daxx mediate an innate antiviral defense antagonized by the adenovirus E4 ORF3 protein. J. Virol. 2008, 82, 7325–7335. [Google Scholar] [CrossRef] [PubMed]
- Schreiner, S.; Wimmer, P.; Sirma, H.; Everett, R.D.; Blanchette, P.; Groitl, P.; Dobner, T. Proteasome-dependent degradation of Daxx by the viral E1B-55K protein in human adenovirus-infected cells. J. Virol. 2010, 84, 7029–7038. [Google Scholar] [CrossRef]
- Somsakeesit, L.-o.; Senawong, T.; Kumboonma, P.; Saenglee, S.; Samankul, A.; Senawong, G.; Yenjai, C.; Phaosiri, C. Influence of side-chain changes on histone deacetylase inhibitory and cytotoxicity activities of curcuminoid derivatives. Bioorganic Med. Chem. Lett. 2020, 30, 127171. [Google Scholar] [CrossRef]
- Balasubramanyam, K.; Varier, R.A.; Altaf, M.; Swaminathan, V.; Siddappa, N.B.; Ranga, U.; Kundu, T.K. Curcumin, a novel p300/CREB-binding protein-specific inhibitor of acetyltransferase, represses the acetylation of histone/nonhistone proteins and histone acetyltransferase-dependent chromatin transcription. J. Biol. Chem. 2004, 279, 51163–51171. [Google Scholar] [CrossRef]
- Ferrari, R.; Pellegrini, M.; Horwitz, G.A.; Xie, W.; Berk, A.J.; Kurdistani, S.K. Epigenetic reprogramming by adenovirus e1a. Science 2008, 321, 1086–1088. [Google Scholar] [CrossRef]
- Radhakrishna Pillai, G.; Srivastava, A.S.; Hassanein, T.I.; Chauhan, D.P.; Carrier, E. Induction of apoptosis in human lung cancer cells by curcumin. Cancer Lett. 2004, 208, 163–170. [Google Scholar] [CrossRef]
- Lestari, M.L.A.D.; Indrayanto, G. Chapter Three-Curcumin. In Profiles of Drug Substances, Excipients and Related Methodology; Brittain, H.G., Ed.; Academic Press: Cambridge, MA, USA, 2014; Volume 39, pp. 113–204. [Google Scholar]
- Kumari, N.; Kulkarni, A.A.; Lin, X.; McLean, C.; Ammosova, T.; Ivanov, A.; Hipolito, M.; Nekhai, S.; Nwulia, E. Inhibition of HIV-1 by curcumin A, a novel curcumin analog. Drug Des. Dev. 2015, 9, 5051–5060. [Google Scholar] [CrossRef]
- Alves, T.F.; Chaud, M.V.; Grotto, D.; Jozala, A.F.; Pandit, R.; Rai, M.; dos Santos, C.A. Association of silver nanoparticles and curcumin solid dispersion: Antimicrobial and antioxidant properties. AAPS Pharmscitech. 2018, 19, 225–231. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.X.; Li, C.M.; Huang, C.Z. Curcumin modified silver nanoparticles for highly efficient inhibition of respiratory syncytial virus infection. Nanoscale 2016, 8, 3040–3048. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-J.; Chang, L.; Chu, H.-W.; Lin, H.-J.; Chang, P.-C.; Wang, R.Y.L.; Unnikrishnan, B.; Mao, J.-Y.; Chen, S.-Y.; Huang, C.-C. High amplification of the antiviral activity of curcumin through transformation into carbon quantum dots. Small 2019, 15, 1902641. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jennings, M.R.; Parks, R.J. Antiviral Effects of Curcumin on Adenovirus Replication. Microorganisms 2020, 8, 1524. https://doi.org/10.3390/microorganisms8101524
Jennings MR, Parks RJ. Antiviral Effects of Curcumin on Adenovirus Replication. Microorganisms. 2020; 8(10):1524. https://doi.org/10.3390/microorganisms8101524
Chicago/Turabian StyleJennings, Morgan R., and Robin J. Parks. 2020. "Antiviral Effects of Curcumin on Adenovirus Replication" Microorganisms 8, no. 10: 1524. https://doi.org/10.3390/microorganisms8101524
APA StyleJennings, M. R., & Parks, R. J. (2020). Antiviral Effects of Curcumin on Adenovirus Replication. Microorganisms, 8(10), 1524. https://doi.org/10.3390/microorganisms8101524



