Tissue- and Population-Level Microbiome Analysis of the Wasp Spider Argiope bruennichi Identified a Novel Dominant Bacterial Symbiont
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Sample Preparation
2.3. DNA Extraction and Illumina Amplicon Sequencing
2.4. Sequence Processing
2.5. Data Analysis and Visualization
3. Results
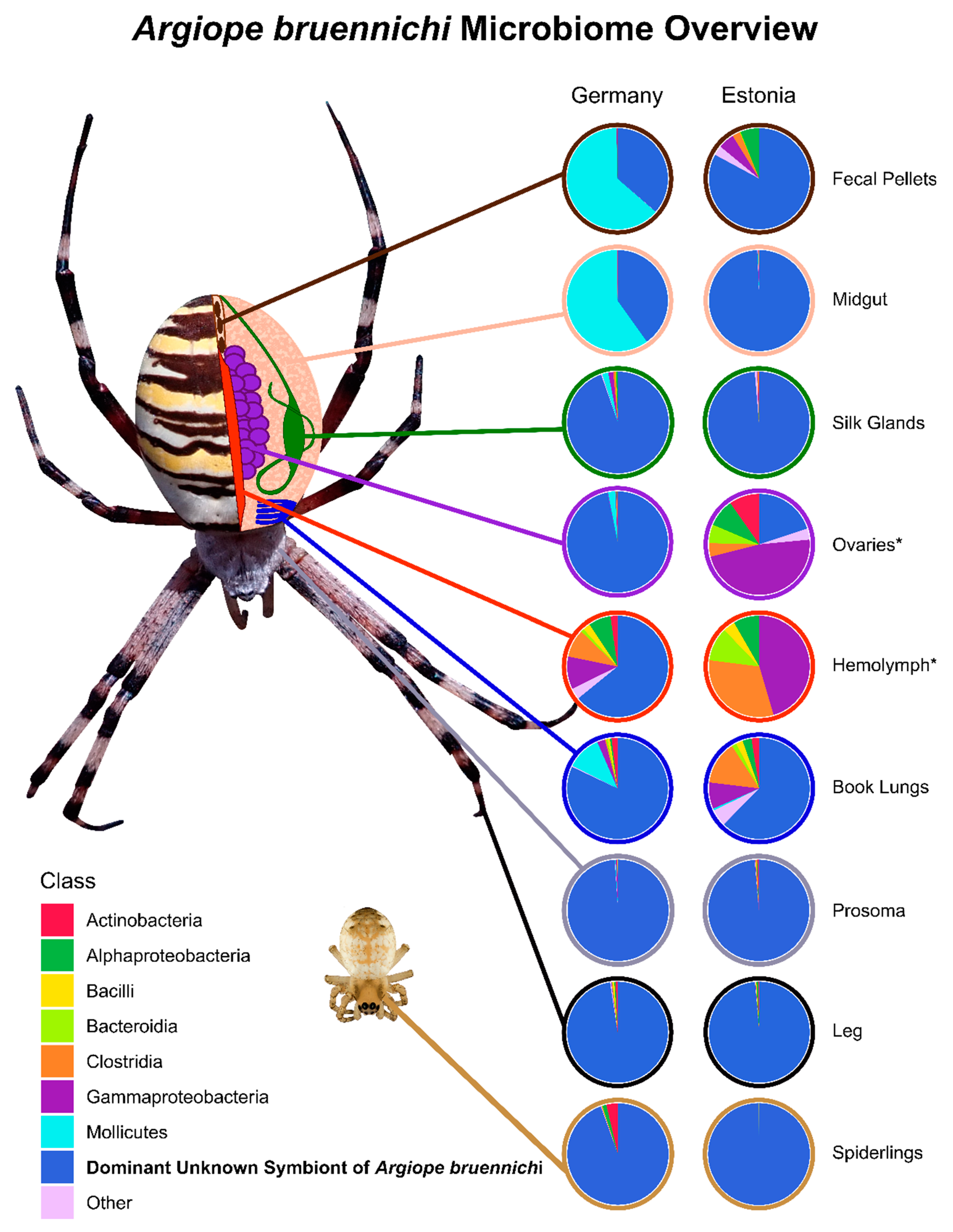
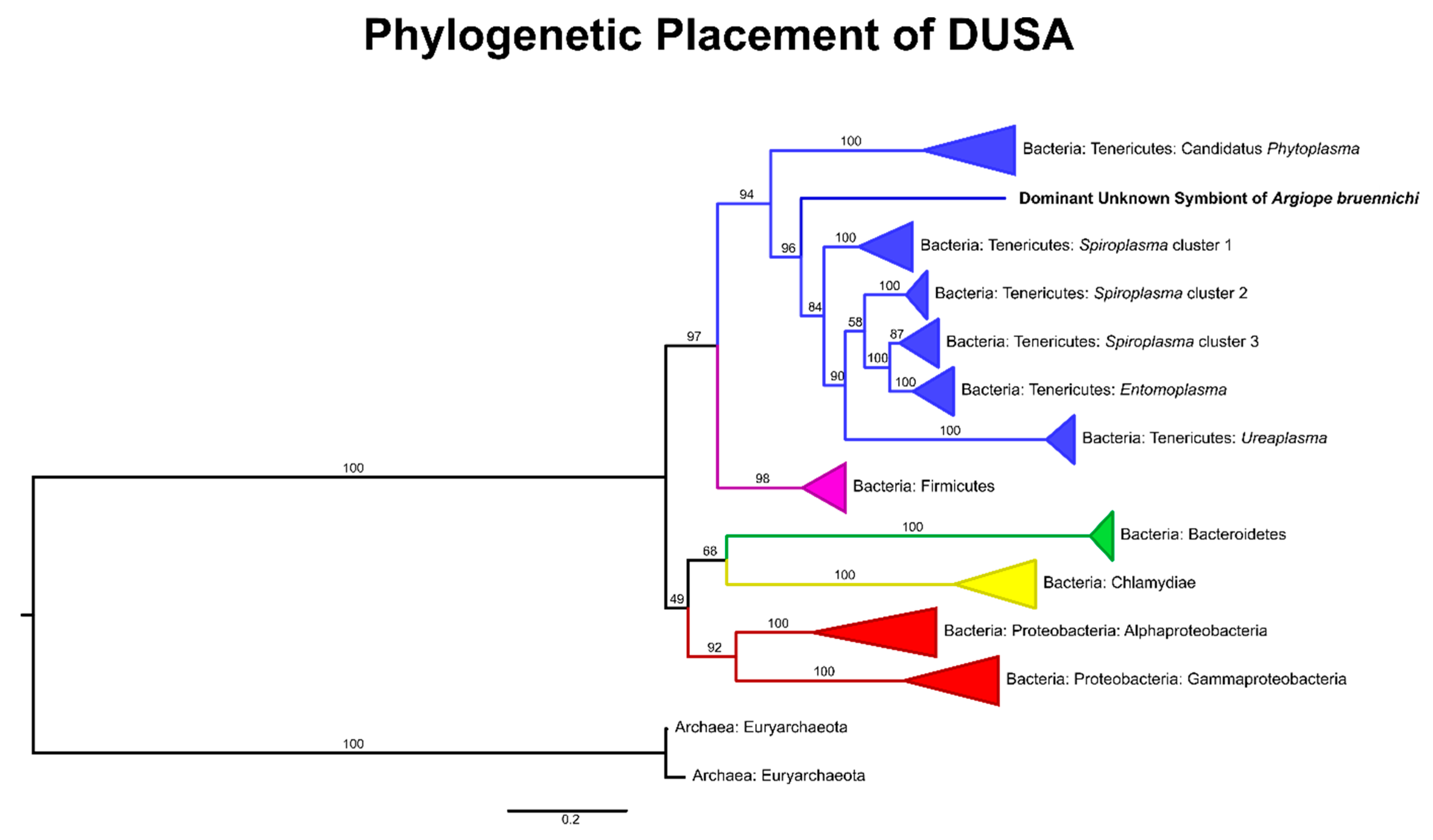
3.1. A Bacterial Symbiont in Argiope bruennichi
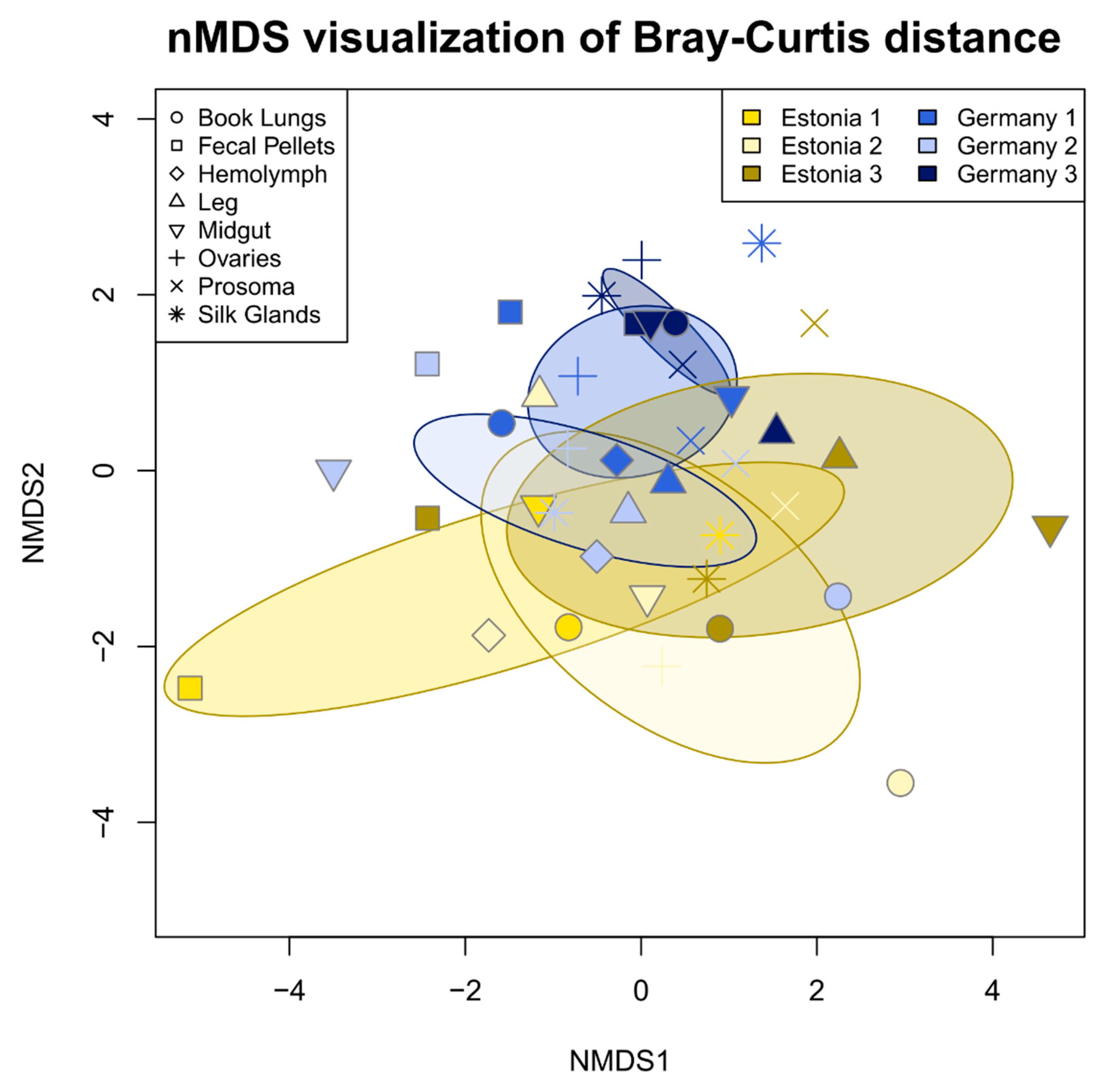
3.2. Tissue Localization and Population Differentiation
3.3. Vertical Transmission
4. Discussion
4.1. An Unknown Symbiont Dominates the Argiope bruennichi Microbiome
4.2. The Argiope bruennichi Microbiome Varies between Individuals and Populations, but Not between Tissues
4.3. Evidence of Vertical Transmission of DUSA
4.4. Implications for Future Studies of Argiope bruennichi and Beyond
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- McFall-Ngai, M.; Hadfield, M.G.; Bosch, T.C.G.; Carey, H.V.; Domazet-Lošo, T.; Douglas, A.E.; Dubilier, N.; Eberl, G.; Fukami, T.; Gilbert, S.F.; et al. Animals in a bacterial world, a new imperative for the life sciences. Proc. Natl. Acad. Sci. USA 2013, 110, 3229–3236. [Google Scholar] [CrossRef]
- Margulis, L. Symbiogenesis and symbionticism. In Symbiosis as a Source of Evolutionary Innovation: Speciation and Morphogenesis; Margulis, L., Fester, R., Eds.; MIT Press: Cambridge, MA, USA, 1991; pp. 1–14. ISBN 9780262132695. [Google Scholar]
- Bordenstein, S.R.; Theis, K.R. Host biology in light of the microbiome: Ten principles of holobionts and hologenomes. PLoS Biol. 2015, 13, e1002226. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, E.; Zilber-Rosenberg, I. Microbes drive evolution of animals and plants: The hologenome concept. MBio 2016, 7, e01395-15. [Google Scholar] [CrossRef]
- Zilber-Rosenberg, I.; Rosenberg, E. Role of microorganisms in the evolution of animals and plants: The hologenome theory of evolution. FEMS Microbiol. Rev. 2008, 32, 723–735. [Google Scholar] [CrossRef]
- Baumann, P.; Baumann, L.; Lai, C.-Y.; Rouhbakhsh, D.; Moran, N.A.; Clark, M.A. Genetics, physiology, and evolutionary relationships of the genus Buchnera: intracellular symbionts of aphids. Annu. Rev. Microbiol. 1995, 49, 55–94. [Google Scholar] [CrossRef] [PubMed]
- Douglas, A.E. Reproductive failure and the free amino acid pools in pea aphids (Acyrthosiphon pisum) lacking symbiotic bacteria. J. Insect Physiol. 1996, 42, 247–255. [Google Scholar] [CrossRef]
- Wilkinson, T.L.; Koga, R.; Fukatsu, T. Role of host nutrition in symbiont regulation: Impact of dietary nitrogen on proliferation of obligate and facultative bacterial endosymbionts of the pea aphid Acyrthosiphon pisum. Appl. Environ. Microbiol. 2007, 73, 1362–1366. [Google Scholar] [CrossRef]
- Brinza, L.; Viñuelas, J.; Cottret, L.; Calevro, F.; Rahbé, Y.; Febvay, G.; Duport, G.; Colella, S.; Rabatel, A.; Gautier, C.; et al. Systemic analysis of the symbiotic function of Buchnera aphidicola, the primary endosymbiont of the pea aphid Acyrthosiphon pisum. C. R. Biol. 2009, 332, 1034–1049. [Google Scholar] [CrossRef]
- Burstein, D.; Sun, C.L.; Brown, C.T.; Sharon, I.; Anantharaman, K.; Probst, A.J.; Thomas, B.C.; Banfield, J.F. Major bacterial lineages are essentially devoid of CRISPR-Cas viral defence systems. Nat. Commun. 2016, 7, 10613. [Google Scholar] [CrossRef] [PubMed]
- Engel, P.; Moran, N.A. The gut microbiota of insects—Diversity in structure and function. FEMS Microbiol. Rev. 2013, 37, 699–735. [Google Scholar] [CrossRef]
- Hammer, T.J.; Janzen, D.H.; Hallwachs, W.; Jaffe, S.P.; Fierer, N. Caterpillars lack a resident gut microbiome. Proc. Natl. Acad. Sci. USA 2017, 114, 9641–9646. [Google Scholar] [CrossRef] [PubMed]
- Marchesi, J.R.; Ravel, J. The vocabulary of microbiome research: A proposal. Microbiome 2015, 3, 31. [Google Scholar] [CrossRef] [PubMed]
- Cho, I.; Blaser, M.J. The human microbiome: at the interface of health and disease. Nat. Rev. Genet. 2012, 13, 260–270. [Google Scholar] [CrossRef] [PubMed]
- Nishijima, S.; Suda, W.; Oshima, K.; Kim, S.-W.; Hirose, Y.; Morita, H.; Hattori, M. The gut microbiome of healthy Japanese and its microbial and functional uniqueness. DNA Res. 2016, 23, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Ursell, L.K.; Clemente, J.C.; Rideout, J.R.; Gevers, D.; Caporaso, J.G.; Knight, R. The interpersonal and intrapersonal diversity of human-associated microbiota in key body sites. J. Allergy Clin. Immunol. 2012, 129, 1204–1208. [Google Scholar] [CrossRef]
- Shoemark, D.K.; Allen, S.J. The microbiome and disease: Reviewing the links between the oral microbiome, aging, and Alzheimer’s disease. J. Alzheimer’s Dis. 2015, 43, 725–738. [Google Scholar] [CrossRef]
- Shropshire, J.D.; Bordenstein, S.R. Speciation by symbiosis: The microbiome and behavior. MBio 2016, 7, e01785. [Google Scholar] [CrossRef]
- Kozek, W.J.; Rao, R.U. The discovery of Wolbachia in arthropods and nematodes—A historical perspective. Issues Infect. Dis. 2007, 5, 1–14. [Google Scholar]
- Werren, J.H.; Windsor, D.; Guo, L. Distribution of Wolbachia among neotropical arthropods. Proc. R. Soc. B Biol. Sci. 1995, 262, 197–204. [Google Scholar]
- Bourtzis, K.; O’Neill, S.L. Wolbachia infections and arthropod reproduction. Bioscience 1998, 48, 287–293. [Google Scholar] [CrossRef]
- Werren, J.H.; Baldo, L.; Clark, M.E. Wolbachia: master manipulators of invertebrate biology. Nat. Rev. Microbiol. 2008, 6, 741–751. [Google Scholar] [CrossRef] [PubMed]
- Hurst, G.D.; Jiggins, F.M. Male-killing bacteria in insects: mechanisms, incidence, and implications. Emerg. Infect. Dis. 2000, 6, 329–336. [Google Scholar] [CrossRef] [PubMed]
- Engelstädter, J.; Hurst, G.D.D. The impact of male-killing bacteria on host evolutionary processes. Genetics 2007, 175, 245–254. [Google Scholar] [CrossRef] [PubMed]
- Stouthamer, R.; Breeuwer, J.A.J.; Hurst, G.D.D. Wolbachia pipientis: microbial manipulator of arthropod reproduction. Annu. Rev. Microbiol. 1999, 53, 71–102. [Google Scholar] [CrossRef]
- Montllor, C.B.; Maxmen, A.; Purcell, A.H. Facultative bacterial endosymbionts benefit pea aphids Acyrthosiphon pisum under heat stress. Ecol. Entomol. 2002, 27, 189–195. [Google Scholar] [CrossRef]
- Fischer, C.; Trautman, E.P.; Crawford, J.M.; Stabb, E.V.; Handelsman, J.; Broderick, N.A. Metabolite exchange between microbiome members produces compounds that influence Drosophila behavior. eLife 2017, 6, e18855. [Google Scholar] [CrossRef]
- Roughgarden, J.; Gilbert, S.F.; Rosenberg, E.; Zilber-Rosenberg, I.; Lloyd, E.A. Holobionts as Units of Selection and a Model of Their Population Dynamics and Evolution. Biol. Theory 2018, 13, 44–65. [Google Scholar] [CrossRef]
- Veneti, Z.; Clark, M.E.; Karr, T.L.; Savakis, C.; Bourtzis, K. Heads or tails: Host-parasite interactions in the Drosophila-Wolbachia system. Appl. Environ. Microbiol. 2004, 70, 5366–5372. [Google Scholar] [CrossRef]
- Moran, N.A.; Baumann, P. Bacterial endosymbionts in animals. Curr. Opin. Microbiol. 2000, 3, 270–275. [Google Scholar] [CrossRef]
- Dobson, S.L.; Bourtzis, K.; Braig, H.R.; Jones, B.F.; Zhou, W.; Rousset, F.; O’Neill, S.L. Wolbachia infections are distributed throughout insect somatic and germ line tissues. Insect Biochem. Mol. Biol. 1999, 29, 153–160. [Google Scholar] [CrossRef]
- McGraw, E.A.; O’Neill, S.L. Wolbachia pipientis: intracellular infection and pathogenesis in Drosophila. Curr. Opin. Microbiol. 2004, 7, 67–70. [Google Scholar] [CrossRef] [PubMed]
- Buchner, P. Studien an intracellularen Symbionten V. die symbiontischen Einrichtungen der Zikaden. Z. Morphol. Ökologie Tiere 1925, 4, 88–245. [Google Scholar] [CrossRef]
- Douglas, A.E. Nutritional interactions in insect-microbial symbioses: aphids and their symbiotic bacteria Buchnera. Annu. Rev. Entomol. 1998, 43, 17–37. [Google Scholar] [CrossRef] [PubMed]
- Shigenobu, S.; Watanabe, H.; Hattori, M.; Sakaki, Y.; Ishikawa, H. Genome sequence of the endocellular bacterial symbiont of aphids Buchnera sp. APS. Nature 2000, 407, 81–86. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Daugherty, S.C.; Van Aken, S.E.; Pai, G.H.; Watkins, K.L.; Khouri, H.; Tallon, L.J.; Zaborsky, J.M.; Dunbar, H.E.; Tran, P.L.; et al. Metabolic complementarity and genomics of the dual bacterial symbiosis of sharpshooters. PLoS Biol. 2006, 4, 1079–1092. [Google Scholar] [CrossRef] [PubMed]
- Hosokawa, T.; Koga, R.; Kikuchi, Y.; Meng, X.-Y.; Fukatsu, T. Wolbachia as a bacteriocyte-associated nutritional mutualist. Proc. Natl. Acad. Sci. USA 2010, 107, 769–774. [Google Scholar] [CrossRef] [PubMed]
- Nikoh, N.; Hosokawa, T.; Moriyama, M.; Oshima, K.; Hattori, M.; Fukatsu, T. Evolutionary origin of insect-Wolbachia nutritional mutualism. Proc. Natl. Acad. Sci. USA 2014, 111, 10257–10262. [Google Scholar] [CrossRef]
- Van Leuven, J.T. The Evolution of Nutritional Co-Endosymbionts in Cicadas. Ph.D. Thesis, University of Montana, Missoula, MT, USA, 2015. [Google Scholar]
- Rio, R.V.M.; Attardo, G.M.; Weiss, B.L. Grandeur alliances: symbiont metabolic integration and obligate arthropod hematophagy. Trends Parasitol. 2016, 32, 739–749. [Google Scholar] [CrossRef]
- Zhang, L.; Yun, Y.; Hu, G.; Peng, Y. Insights into the bacterial symbiont diversity in spiders. Ecol. Evol. 2018, 8, 4899–4906. [Google Scholar] [CrossRef]
- Duron, O.; Bouchon, D.; Boutin, S.; Bellamy, L.; Zhou, L.; Engelstädter, J.; Hurst, G.D. The diversity of reproductive parasites among arthropods: Wolbachia do not walk alone. BMC Biol. 2008, 6, 27. [Google Scholar] [CrossRef]
- Duron, O.; Hurst, G.D.D.; Hornett, E.A.; Josling, J.A.; Engelstädter, J. High incidence of the maternally inherited bacterium Cardinium in spiders. Mol. Ecol. 2008, 17, 1427–1437. [Google Scholar] [CrossRef] [PubMed]
- Vanthournout, B.; Hendrickx, F. Endosymbiont dominated bacterial communities in a dwarf spider. PLoS ONE 2015, 10, e0117297. [Google Scholar] [CrossRef] [PubMed]
- Vanthournout, B.; Swaegers, J.; Hendrickx, F. Spiders do not escape reproductive manipulations by Wolbachia. BMC Evol. Biol. 2011, 11, 15. [Google Scholar] [CrossRef] [PubMed]
- Goodacre, S.L.; Martin, O.Y.; Bonte, D.; Hutchings, L.; Woolley, C.; Ibrahim, K.; George Thomas, C.; Hewitt, G.M. Microbial modification of host long-distance dispersal capacity. BMC Biol. 2009, 7, 32. [Google Scholar] [CrossRef] [PubMed]
- Krehenwinkel, H.; Graze, M.; Rödder, D.; Tanaka, K.; Baba, Y.G.; Muster, C.; Uhl, G. A phylogeographical survey of a highly dispersive spider reveals eastern Asia as a major glacial refugium for Palaearctic fauna. J. Biogeogr. 2016, 43, 1583–1594. [Google Scholar] [CrossRef]
- Schneider, J.; Uhl, G.; Herberstein, M.E. Cryptic Female Choice within the Genus Argiope: A Comparative Approach. In Cryptic Female Choice in Arthropods: Patterns, Mechanisms and Prospects; Peretti, A., Aisenberg, A., Eds.; Springer International Publishing: Cham, Switzerland, 2015; pp. 55–77. ISBN 9783319178943. [Google Scholar]
- Fromhage, L.; Uhl, G.; Schneider, J.M. Fitness consequences of sexual cannibalism in female Argiope bruennichi. Behav. Ecol. Sociobiol. 2003, 55, 60–64. [Google Scholar] [CrossRef]
- Schneider, J.M.; Fromhage, L.; Uhl, G. Extremely short copulations do not affect hatching success in Argiope bruennichi (Araneae, Araneidae). J. Arachnol. 2005, 33, 663–669. [Google Scholar] [CrossRef]
- Welke, K.W.; Schneider, J.M. Males of the orb-web spider Argiope bruennichi sacrifice themselves to unrelated females. Biol. Lett. 2010, 6, 585–588. [Google Scholar] [CrossRef]
- Guttmann, R. Zur Arealentwicklung und Ökologie der Wespenspinne (Argiope bruennichi) in der Bundesrepublik Deutschland und den angrenzenden Ländern (Araneae). Bonner Zool. Beiträge 1979, 30, 454–486. [Google Scholar]
- Kumschick, S.; Fronzek, S.; Entling, M.H.; Nentwig, W. Rapid spread of the wasp spider Argiope bruennichi across Europe: A consequence of climate change? Clim. Chang. 2011, 109, 319–329. [Google Scholar] [CrossRef][Green Version]
- Krehenwinkel, H.; Tautz, D. Northern range expansion of European populations of the wasp spider Argiope bruennichi is associated with global warming-correlated genetic admixture and population-specific temperature adaptations. Mol. Ecol. 2013, 22, 2232–2248. [Google Scholar] [CrossRef] [PubMed]
- Krehenwinkel, H.; Rödder, D.; Tautz, D. Eco-genomic analysis of the poleward range expansion of the wasp spider Argiope bruennichi shows rapid adaptation and genomic admixture. Glob. Chang. Biol. 2015, 21, 4320–4332. [Google Scholar] [CrossRef] [PubMed]
- Yun, Y.; Peng, Y.; Liu, F.; Lei, C. Wolbachia screening in spiders and assessment of horizontal transmission between predator and prey. Neotrop. Entomol. 2011, 40, 164–169. [Google Scholar] [PubMed]
- Becker, H. Verhaltensbiologie der Wespenspinne. Biol. Unserer Zeit 1981, 11, 86–90. [Google Scholar] [CrossRef]
- Wilson, R.S. The control of dragline spinning in the garden spider. J. Cell Sci. 1962, 104, 557–571. [Google Scholar]
- Vollrath, F.; Knight, D.P.; Hu, X.W. Silk production in a spider involves acid bath treatment. Proc. R. Soc. Lond. Ser. B Biol. Sci. 1998, 265, 817–820. [Google Scholar] [CrossRef]
- Vollrath, F.; Porter, D. Spider silk as archetypal protein elastomer. Soft Matter 2006, 2, 377. [Google Scholar] [CrossRef]
- Ko, F.K.; Wan, L.Y. Engineering properties of spider silk. In Handbook of Properties of Textile and Technical Fibres; Woodhead Publishing: Sawston, UK, 2018; pp. 185–220. ISBN 9780081012727. [Google Scholar]
- Urich, T.; Lanzén, A.; Qi, J.; Huson, D.H.; Schleper, C.; Schuster, S.C. Simultaneous assessment of soil microbial community structure and function through analysis of the meta-transcriptome. PLoS ONE 2008, 3, e2527. [Google Scholar] [CrossRef]
- Walters, W.; Hyde, E.R.; Berg-Lyons, D.; Ackermann, G.; Humphrey, G.; Parada, A.; Gilbert, J.A.; Jansson, J.K.; Caporaso, J.G.; Fuhrman, J.A.; et al. Improved Bacterial 16S rRNA Gene (V4 and V4-5) and Fungal Internal Transcribed Spacer Marker Gene Primers for Microbial Community Surveys. mSystems 2016, 1, e00009-15. [Google Scholar] [CrossRef]
- Sahlin, K.; Tomaszkiewicz, M.; Makova, K.D.; Medvedev, P. Deciphering highly similar multigene family transcripts from Iso-Seq data with IsoCon. Nat. Commun. 2018, 9, 4601. [Google Scholar] [CrossRef]
- Callahan, B.J.; Mcmurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019; Available online: https://www.R-project.org/. (accessed on 26 November 2019).
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef] [PubMed]
- Pruesse, E.; Quast, C.; Knittel, K.; Fuchs, B.M.; Ludwig, W.; Rg Peplies, J.; GlöCkner, F.O. SILVA: A comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007, 35, 7188–7196. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016; ISBN 978-3-319-24277-4. [Google Scholar]
- Oksanen, J.; Blanchet, G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. 2018. Available online: https://cran.r-project.org/web/packages/vegan/index.html (accessed on 26 November 2019).
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A. FigTree: Tree Figure Drawing Tool 2018. Available online: http://tree.bio.ed.ac.uk/software/figtree/ (accessed on 26 November 2019).
- Brown, D.R. Tenericutes. In Bergey’s Manual of Systematics of Archaea and Bacteria; John Wiley & Sons, Ltd.: Chichester, UK, 2018; pp. 1–3. [Google Scholar]
- Whitman, W.B. Bergey’s Manual Trust. In Bergey’s Manual of Systematics of Archaea and Bacteria; John Wiley and Sons Inc.: Hoboken, NJ, USA, 2015; ISBN 9781118960608. [Google Scholar]
- Tully, J.G.; Rose, D.L.; Carle, P.; Bove, J.M.; Hackett, K.J.; Whitcomb, R.F. Acholeplasma entomophilum sp. nov. from gut contents of a wide range of host insects. Int. J. Syst. Bacteriol. 1988, 38, 164–167. [Google Scholar] [CrossRef]
- Sapountzis, P.; Zhukova, M.; Shik, J.Z.; Schiott, M.; Boomsma, J.J. Reconstructing the functions of endosymbiotic Mollicutes in fungus-growing ants. eLife 2018, 7, e39209. [Google Scholar] [CrossRef]
- Meeus, I.; Vercruysse, V.; Smagghe, G. Molecular detection of Spiroplasma apis and Spiroplasma melliferum in bees. J. Invertebr. Pathol. 2012, 109, 172–174. [Google Scholar] [CrossRef]
- Xie, J.; Tiner, B.; Vilchez, I.; Mateos, M. Effect of the Drosophila endosymbiont Spiroplasma on parasitoid wasp development and on the reproductive fitness of wasp-attacked fly survivors. Evol. Ecol. 2011, 25, 1065–1079. [Google Scholar] [CrossRef]
- Moran, N.A.; von Dohlen, C.D.; Baumann, P. Faster evolutionary rates in endosymbiotic bacteria than in cospeciating insect hosts. J. Mol. Evol. 1995, 41, 727–731. [Google Scholar] [CrossRef]
- Moran, N.A.; McCutcheon, J.P.; Nakabachi, A. Genomics and evolution of heritable bacterial symbionts. Annu. Rev. Genet. 2008, 42, 165–190. [Google Scholar] [CrossRef] [PubMed]
- Delaney, N.F.; Balenger, S.; Bonneaud, C.; Marx, C.J.; Hill, G.E.; Ferguson-Noel, N.; Tsai, P.; Rodrigo, A.; Edwards, S.V. Ultrafast evolution and loss of CRISPRs following a host shift in a novel wildlife pathogen, Mycoplasma gallisepticum. PLoS Genet. 2012, 8, e1002511. [Google Scholar] [CrossRef]
- Bennett, G.M.; McCutcheon, J.P.; McDonald, B.R.; Moran, N.A. Lineage-specific patterns of genome deterioration in obligate symbionts of sharpshooter leafhoppers. Genome Biol. Evol. 2016, 8, 296–301. [Google Scholar] [CrossRef] [PubMed]
- Wernegreen, J.J. Genome evolution in bacterial endosymbionts of insects. Nat. Rev. Genet. 2002, 3, 850–861. [Google Scholar] [CrossRef]
- Mouches, C.; Bové, J.M.; Albisetti, J. Pathogenicity of Spiroplasma apis and other spiroplasmas for honey-bees in Southwestern France. Ann. l’Institut Pasteur Microbiol. 1984, 135, 151–155. [Google Scholar] [CrossRef]
- Seemüller, E.; Garnier, M.; Schneider, B. Mycoplasmas of Plants and Insects. In Molecular Biology and Pathogenicity of Mycoplasmas; Springer US: Boston, MA, USA, 2002; pp. 91–115. [Google Scholar]
- Krehenwinkel, H.; Kennedy, S.; Pekár, S.; Gillespie, R.G. A cost-efficient and simple protocol to enrich prey DNA from extractions of predatory arthropods for large-scale gut content analysis by Illumina sequencing. Methods Ecol. Evol. 2017, 8, 126–134. [Google Scholar] [CrossRef]
- Wood, S.F. Importance of feeding and defecation times of insect vectors in transmission of Chagas’ disease. J. Econ. Entomol. 1951, 44, 52–54. [Google Scholar] [CrossRef]
- Werren, J.H.; Zhang, W.; Guo, L.R. Evolution and phylogeny of Wolbachia: Reproductive parasites of arthropods. Proc. R. Soc. Lond. Ser. B Biol. Sci. 1995, 261, 55–63. [Google Scholar]
- Williamson, D.L.; Sakaguchi, B.; Hackett, K.J.; Whitcomb, R.F.; Tully, J.G.; Carle, P.; Bové, J.M.; Adams, J.R.; Konai, M.; Henegar, R.B. Spiroplasma poulsonii sp. nov., a new species associated with male- lethality in Drosophila willistoni, a neotropical species of fruit fly. Int. J. Syst. Bacteriol. 1999, 49, 611–618. [Google Scholar] [CrossRef]
- Sakaguchi, B.; Poulson, D.F. Distribution of “sex-ratio” agent in tissues of Drosophila willistoni. Genetics 1961, 46, 1665–1676. [Google Scholar] [PubMed]
- Wright, S.; Goodacre, S.L. Evidence for antimicrobial activity associated with common house spider silk. BMC Res. Notes 2012, 5, 326. [Google Scholar] [CrossRef] [PubMed]
- White, J.A.; Styer, A.; Rosenwald, L.C.; Curry, M.M.; Welch, K.D.; Athey, K.J.; Chapman, E.G. Endosymbiotic bacteria are prevalent and diverse in agricultural spiders. Microb. Ecol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, G.; Yun, Y.; Peng, Y. Bacterial community of a spider, Marpiss magister (Salticidae). 3Biotech 2017, 7, 371. [Google Scholar] [CrossRef] [PubMed]
| Query Sequence | GenBank NR Best Match: Taxonomy (Accession number): Sequence Identity % | GenBank Bacteria & Archaea Best Match: Taxonomy (Accession Number): Sequence Identity % | Silva SSU 138 NR: Phylum; Class; Order; Family: Sequence Identity % |
|---|---|---|---|
| ASV V4 region (248bp) | Uncultured prokaryote clone Otu01661 (MG853790.1): 84.3% | Holdemania filiformis strain J1-31B-1 (NR_029335.1): 79.92% | Firmicutes; Erysipelotrichia; Erysipelotrichales; Erysipelotrichaceae: 78.7% |
| Near full-length 16S gene (1492bp) | Mycoplasma sp. (e.g., Biomphalaria glabrata) (CP013128.1): 82.3% | Spiroplasma eriocheiris CCTCC M 207170 strain CRAB (NR_125517.1): 80.79% | Tenericutes; Mollicutes; Entomoplasmatales; Spiroplasmataceae: 79.2% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sheffer, M.M.; Uhl, G.; Prost, S.; Lueders, T.; Urich, T.; Bengtsson, M.M. Tissue- and Population-Level Microbiome Analysis of the Wasp Spider Argiope bruennichi Identified a Novel Dominant Bacterial Symbiont. Microorganisms 2020, 8, 8. https://doi.org/10.3390/microorganisms8010008
Sheffer MM, Uhl G, Prost S, Lueders T, Urich T, Bengtsson MM. Tissue- and Population-Level Microbiome Analysis of the Wasp Spider Argiope bruennichi Identified a Novel Dominant Bacterial Symbiont. Microorganisms. 2020; 8(1):8. https://doi.org/10.3390/microorganisms8010008
Chicago/Turabian StyleSheffer, Monica M., Gabriele Uhl, Stefan Prost, Tillmann Lueders, Tim Urich, and Mia M. Bengtsson. 2020. "Tissue- and Population-Level Microbiome Analysis of the Wasp Spider Argiope bruennichi Identified a Novel Dominant Bacterial Symbiont" Microorganisms 8, no. 1: 8. https://doi.org/10.3390/microorganisms8010008
APA StyleSheffer, M. M., Uhl, G., Prost, S., Lueders, T., Urich, T., & Bengtsson, M. M. (2020). Tissue- and Population-Level Microbiome Analysis of the Wasp Spider Argiope bruennichi Identified a Novel Dominant Bacterial Symbiont. Microorganisms, 8(1), 8. https://doi.org/10.3390/microorganisms8010008





