Oxidative Stress Response of Aspergillus oryzae Induced by Hydrogen Peroxide and Menadione Sodium Bisulfite
Abstract
1. Introduction
2. Materials and Methods
2.1. Effect of Oxidative Stress on Aspergillus oryzae Growth
2.2. Determination of Reduced Glutathione Levels
2.3. Measurement of Intracellular Fatty Acid Content
2.4. RNA Extraction and Illumina Sequencing
2.5. Differentially Expressed Genes Analysis
2.6. Calculations and Statistical Analyses
3. Results
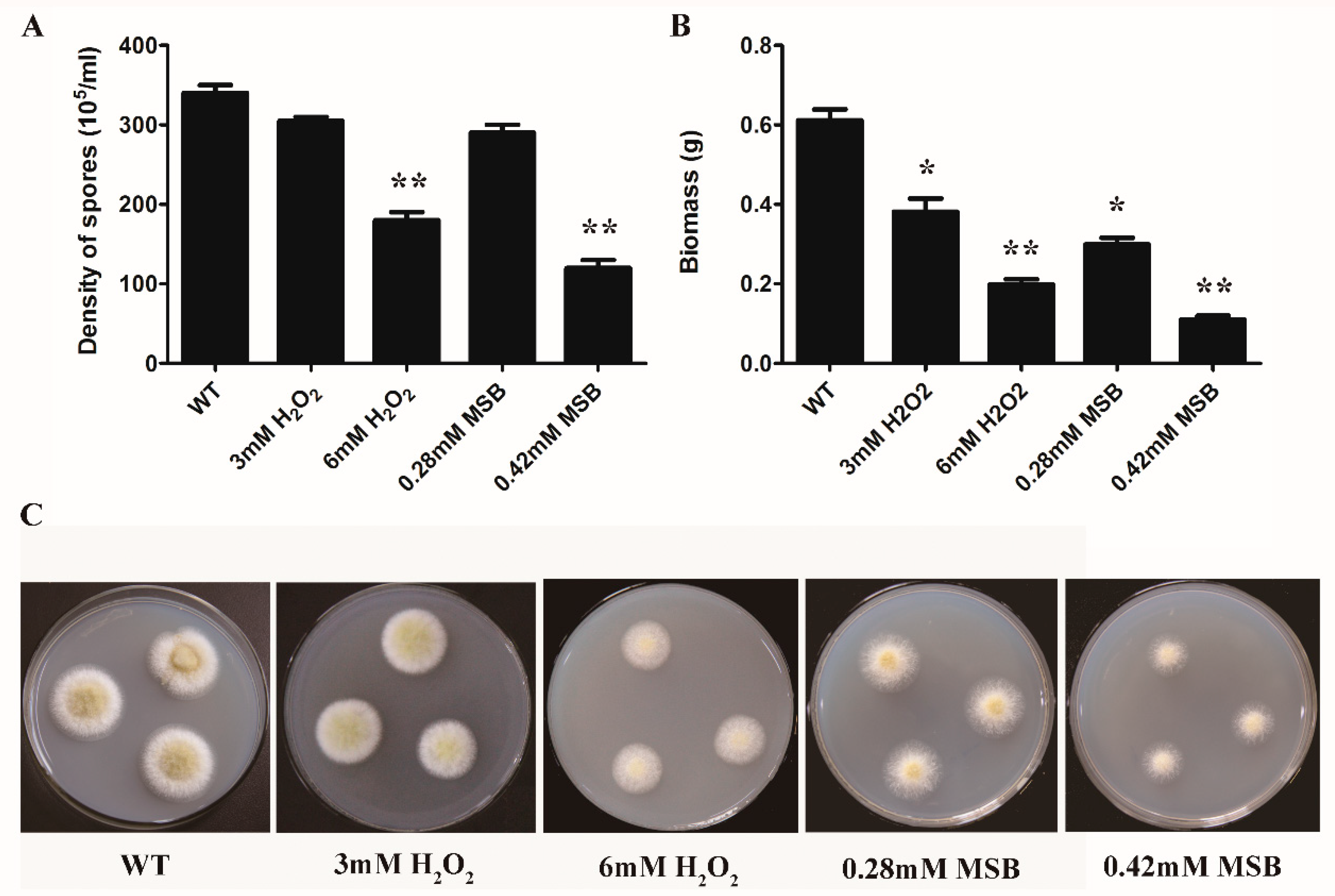
3.1. Effect of Oxidant Species and Concentration on A. oryzae Cell Growth
3.2. Global Analysis of A. oryzae Transcriptome under Oxidative Stress
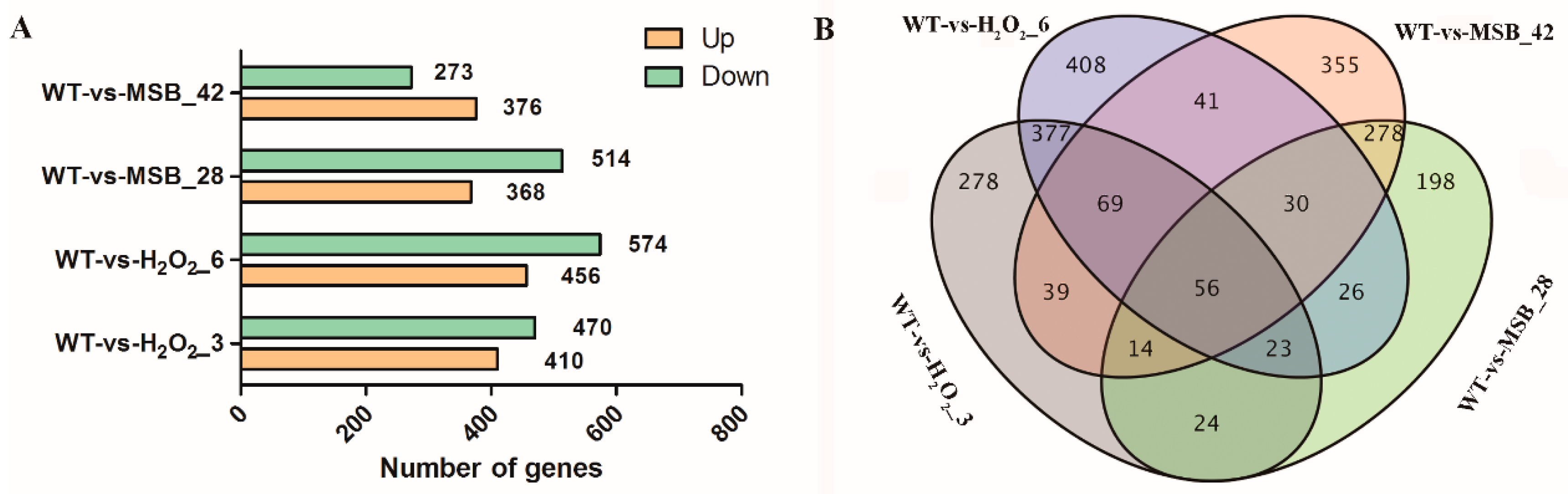
3.3. Differentially Expressed genes Analysis of A. oryzae Transcriptome under Oxidative Stress
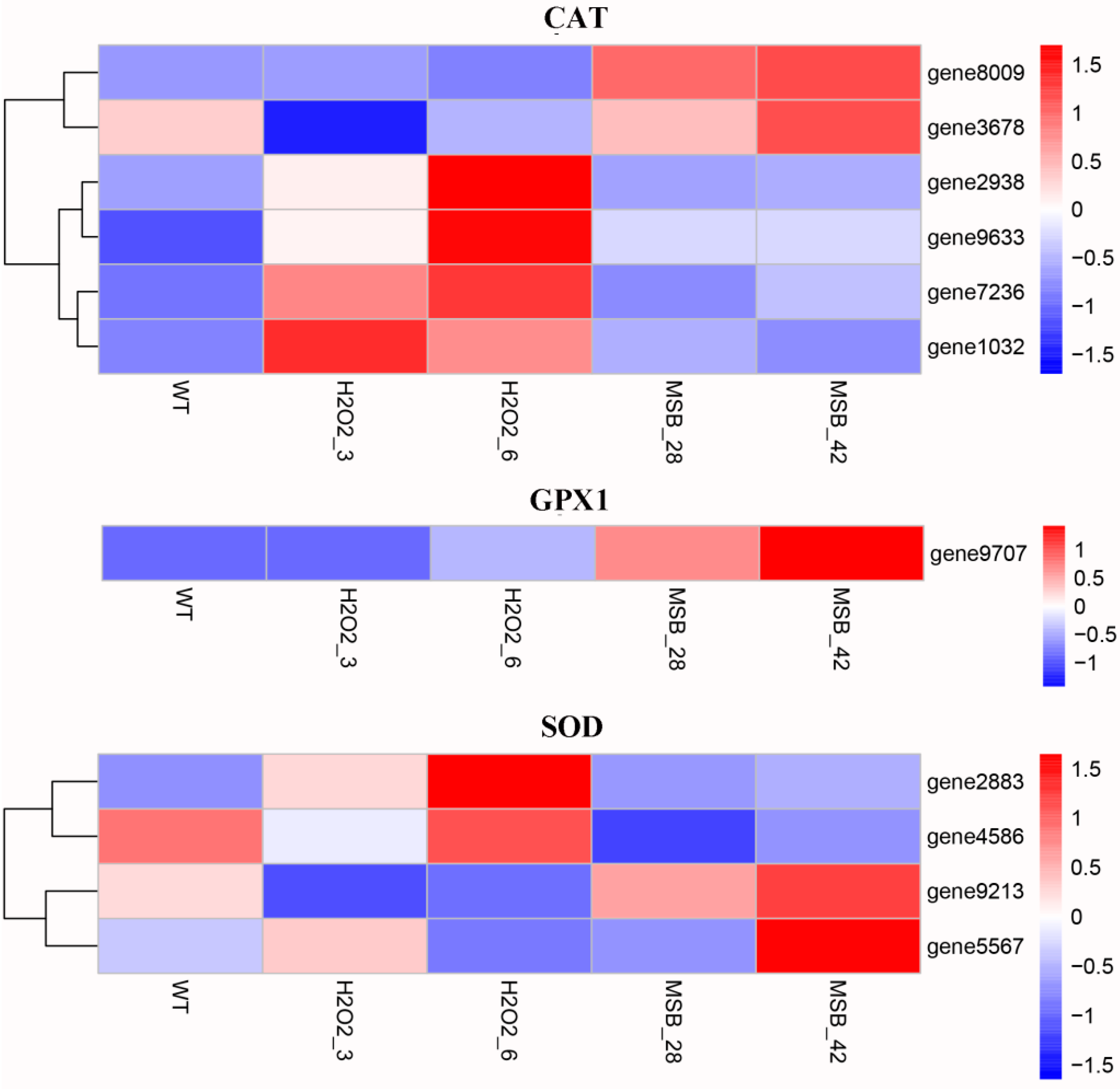
3.4. Expression Analysis of Genes Involved in Oxidoreduction under Oxidative Stress
3.5. Expression Analysis of Transcription Factors Possibly Involved in Oxidative Stress Response
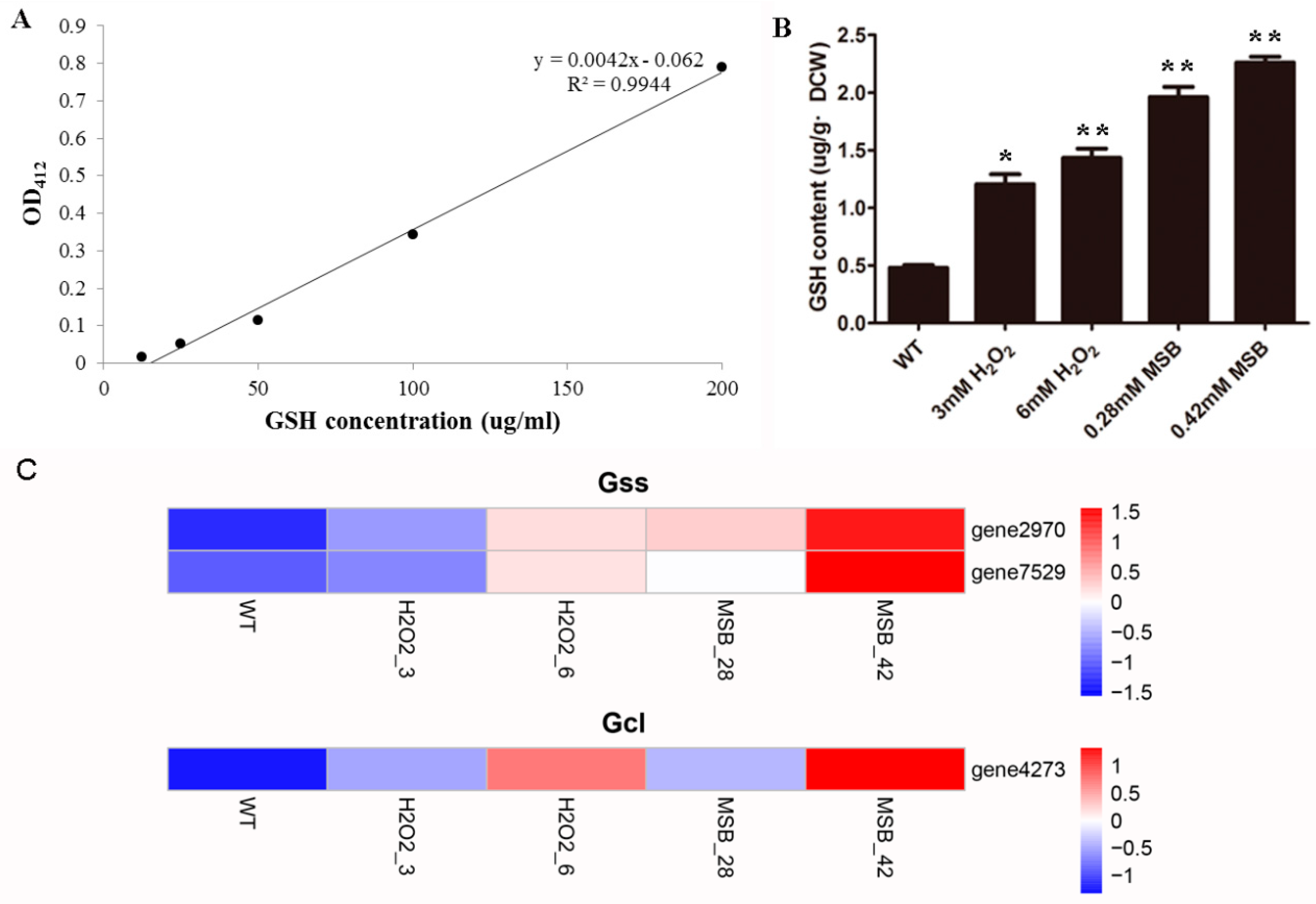
3.6. Effects of Oxidant on A. oryzae GSH
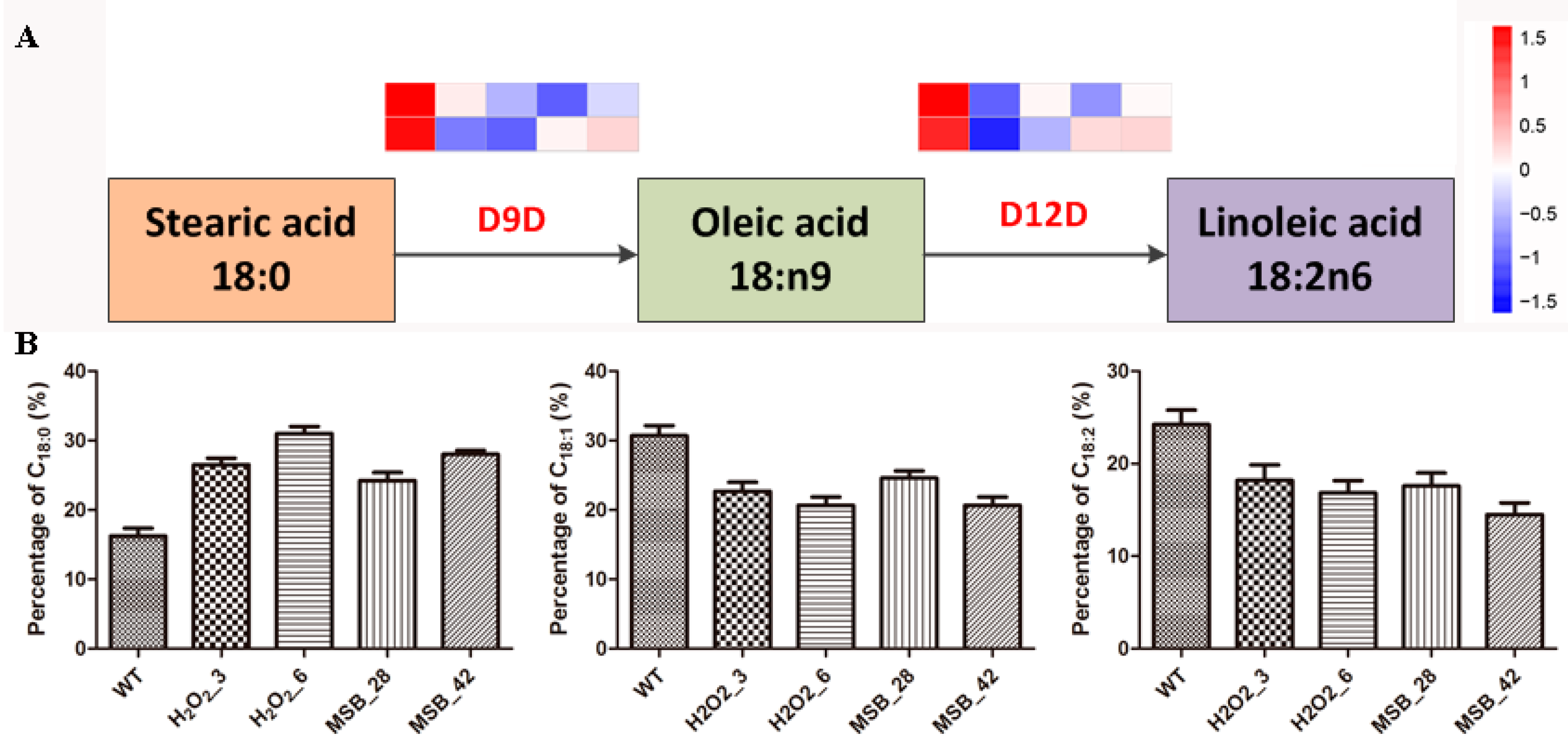
3.7. Change in Linoleic acid Biosynthesis under Oxidative Stress
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- He, B.; Li, H.; Hu, Z.; Zhang, Y.; Sun, M.; Qiu, S.; Zeng, B. Difference in microbial community and taste compounds between Mucor-type and Aspergillus-type Douchi during koji-making. Food Res. Int. (Ott. Ont.) 2019, 121, 136–143. [Google Scholar] [CrossRef] [PubMed]
- Machida, M.; Asai, K.; Sano, M.; Tanaka, T.; Kumagai, T.; Terai, G.; Kusumoto, K.-I.; Arima, T.; Akita, O.; Kashiwagi, Y.; et al. Genome sequencing and analysis of Aspergillus oryzae. Nature 2005, 438, 1157. [Google Scholar] [CrossRef] [PubMed]
- He, B.; Tu, Y.Y.; Jiang, C.M.; Zhang, Z.; Li, Y.K.; Zeng, B. Functional Genomics of Aspergillus oryzae: Strategies and Progress. Microorganisms 2019, 7, 103. [Google Scholar] [CrossRef] [PubMed]
- Zadrąg-Tęcza, R.; Maślanka, R.; Bednarska, S.; Kwolek-Mirek, M. Response Mechanisms to Oxidative Stress in Yeast and Filamentous Fungi. In Stress Response Mechanisms in Fungi: Theoretical and Practical Aspects; Skoneczny, M., Ed.; Springer International Publishing: Cham, Switzerland, 2018. [Google Scholar]
- Kreiner, M.; Harveya, L.M.; Mcneila, B. Oxidative stress response of a recombinant Aspergillus niger to exogenous menadione and H2O2 addition. Enzym. Microb. Technol. 2002, 30, 346–353. [Google Scholar] [CrossRef]
- Zhao, H.; Chen, J.; Liu, J.; Han, B. Transcriptome analysis reveals the oxidative stress response in Saccharomyces cerevisiae. Rsc Adv. 2015, 5, 22923–22934. [Google Scholar] [CrossRef]
- Lópezmirabal, H.R.; Thorsen, M.; Kiellandbrandt, M.C.; Toledano, M.B.; Winther, J.R. Cytoplasmic glutathione redox status determines survival upon exposure to the thiol-oxidant 4,4′-dipyridyl disulfide. Fems Yeast Res. 2010, 7, 391–403. [Google Scholar] [CrossRef] [PubMed]
- Suo, Y.; Liu, Y.; Zhou, X.; Huang, Y.; Shi, C.; Matthews, K.; Shi, X. Impact of sod on the expression of stress-related genes in Listeria monocytogenes 4b G with/without paraquat treatment. J. Food Sci. 2015, 79, M1745–M1749. [Google Scholar] [CrossRef] [PubMed]
- Barker, M.G.; Brimage, L.J.E.; Smart, K.A. Effect of Cu,Zn superoxide dismutase disruption mutation on replicative senescence in Saccharomyces cerevisiae. Fems Microbiol. Lett. 2010, 177, 199–204. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Nishimoto, T.; Watanabe, T.; Furuta, M.; Kataoka, M.; Kishida, M. Roles of Catalase and Trehalose in the Protection from Hydrogen Peroxide Toxicity in Saccharomyces cerevisiae. Biocontrol Sci. 2016, 21, 179–182. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Grant, C.M.; Perrone, G.; Dawes, I.W. Glutathione and Catalase Provide Overlapping Defenses for Protection against Hydrogen Peroxide in the Yeast Saccharomyces cerevisiae. Biochem. Biophys. Res. Commun. 1998, 253, 893–898. [Google Scholar] [CrossRef] [PubMed]
- Dalto, D.B.; Matte, J.J. Pyridoxine (Vitamin B6) and the Glutathione Peroxidase System; a Link between One-Carbon Metabolism and Antioxidation. Nutrients 2017, 9, 189. [Google Scholar] [CrossRef] [PubMed]
- Mulford, K.E.; Fassler, J.S. Association of the Skn7 and Yap1 Transcription Factors in the Saccharomyces cerevisiae Oxidative Stress Response. Eukaryot. Cell 2011, 10, 761–769. [Google Scholar] [CrossRef] [PubMed]
- Carmel-Harel, O.; Stearman, R.; Gasch, A.P.; Botstein, D.; Brown, P.O.; Storz, G. Role of thioredoxin reductase in the Yap1p-dependent response to oxidative stress in Saccharomyces cerevisiae. Mol. Microbiol. 2010, 39, 595–605. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.H.; Tsai, H.C.; Yu, P.L.; Chung, K.R. A Major Facilitator Superfamily Transporter-Mediated Resistance to Oxidative Stress and Fungicides Requires Yap1, Skn7, and MAP Kinases in the Citrus Fungal Pathogen Alternaria alternata. PLoS ONE 2017, 12, e0169103. [Google Scholar] [CrossRef] [PubMed]
- Yi, D.G.; Kim, M.J.; Ji, E.C.; Lee, J.; Jung, J.; Huh, W.K.; Chung, W.H. Yap1 and Skn7 genetically interact with Rad51 in response to oxidative stress and DNA double-strand break in Saccharomyces cerevisiae. Free Radic. Biol. Med. 2016, 101, 424–433. [Google Scholar] [CrossRef]
- Ilyas, S.; Rehman, A. Oxidative stress, glutathione level and antioxidant response to heavy metals in multi-resistant pathogen, Candida tropicalis. Environ. Monit. Assess. 2015, 187, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Grant, C.M.; Maciver, F.H.; Dawes, I.W. Glutathione is an essential metabolite required for resistance to oxidative stress in the yeast Saccharomyces cerevisiae. Curr. Genet. 1996, 29, 511–515. [Google Scholar] [CrossRef]
- Kikugawa, K.; Yasuhara, Y.; Ando, K.; Koyama, K.; Hiramoto, K.; Suzuki, M. Protective Effect of Supplementation of Fish Oil with High n−3 Polyunsaturated Fatty Acids against Oxidative Stress-Induced DNA Damage of Rat Liver in Vivo. J. Agric. Food Chem. 2003, 51, 6073–6079. [Google Scholar] [CrossRef]
- Lagarde, M.; Guichardant, M.; Bernoudhubac, N.; Calzada, C.; Véricel, E. Oxygenation of polyunsaturated fatty acids and oxidative stress within blood platelets. Biochim. Et Biophys. Acta (Bba) - Mol. Cell Biol. Lipids 2018, 1863, 651–656. [Google Scholar] [CrossRef]
- Belozerskaya, T.A.; Gessler, N.N. Oxidative stress and differentiation in Neurospora crassa. Microbiology 2006, 75, 427–431. [Google Scholar] [CrossRef]
- Li, Q.; Mcneil, B.; Harvey, L.M. Adaptive response to oxidative stress in the filamentous fungus Aspergillus niger B1-D. Free Radic. Biol. Med. 2008, 44, 394–402. [Google Scholar] [CrossRef]
- Emri, T.; Szarvas, V.; Orosz, E.; Antal, K.; Park, H.S.; Han, K.H.; Yu, J.H.; Pócsi, I. Core oxidative stress response in Aspergillus nidulans. BMC Genom. 2015, 16, 478. [Google Scholar] [CrossRef]
- Emilie, F.; Cécile-Marie, A.D.; Laurence, D.; David, H.; Eduardo, D.C. Transcriptomic insights into the oxidative response of stress-exposed Aspergillus fumigatus. Curr. Pharm. Des. 2013, 19, 3713–3737. [Google Scholar]
- He, B.; Long, M.; Hu, Z.; Li, H.; Ai, M.; Long, C.; Zeng, B. Deep sequencing analysis of transcriptomes in Aspergillus oryzae in response to salinity stress. Appl. Microbiol. Biotechnol. 2018, 102, 897–906. [Google Scholar] [CrossRef]
- Vries, R.P.D.; Riley, R.; Wiebenga, A.; Aguilar-Osorio, G.; Amillis, S.; Uchima, C.A.; Anderluh, G.; Asadollahi, M.; Askin, M.; Barry, K. Comparative genomics reveals high biological diversity and specific adaptations in the industrially and medically important fungal genus Aspergillus. Genome Biol. 2017, 18, 28. [Google Scholar] [CrossRef]
- Rahman, I.; Kode, A.; Biswas, S. Assay for quantitative determination of glutathione and glutathione disulfide levels using enzymatic recycling method. Nat. Protoc. 2007, 1, 3159–3165. [Google Scholar] [CrossRef]
- Wu, X.; Tong, Y.; Shankar, K.; Baumgardner, J.N.; Kang, J.; Badeaux, J.; Badger, T.M.; Ronis, M.J. Lipid fatty acid profile analyses in liver and serum in rats with nonalcoholic steatohepatitis using improved gas chromatography-mass spectrometry methodology. J. Agric. Food Chem. 2011, 59, 747–754. [Google Scholar] [CrossRef]
- Kim, D.; Salzberg, S.L. TopHat-Fusion: An algorithm for discovery of novel fusion transcripts. Genome Biol. 2011, 12, R72. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Zuo, Z.; Yang, L.; Chen, S.; Ye, C.; Han, Y.; Wang, S.; Ma, Y. Effects of nitrogen nutrients on the volatile organic compound emissions from Microcystis aeruginosa. Ecotoxicol. Environ. Saf. 2018, 161, 214–220. [Google Scholar] [CrossRef]
- Bai, Z.; Harvey, L.; Mcneil, B. Physiological responses of chemostat cultures of Aspergillus niger (B1-D) to simulated and actual oxidative stress. Biotechnol. Bioeng. 2003, 82, 691–701. [Google Scholar] [CrossRef]
- He, B.; Tu, Y.; Hu, Z.; Ma, L.; Dai, J.; Cheng, X.; Li, H.; Liu, L.; Zeng, B. Genome-wide identification and expression profile analysis of the HOG gene family in Aspergillus oryzae. World J. Microb. Biot. 2018, 34, 35. [Google Scholar] [CrossRef]
- Bin, W.; Guangwu, G.; Chao, W.; Ying, L.; Xiaoning, W.; Mouming, Z.; Yong, G.; Minghui, H.; Yong, Z.; Li, P. Survey of the transcriptome of Aspergillus oryzae via massively parallel mRNA sequencing. Nucleic Acids Res. 2010, 38, 5075–5087. [Google Scholar]
- He, B.; Hu, Z.; Long, M.; Li, H.; Ai, M.; Han, J.; Zeng, B. Transcriptome analysis of different growth stages of Aspergillus oryzae reveals dynamic changes of distinct classes of genes during growth. BMC Microbiol. 2018, 18, 12. [Google Scholar] [CrossRef]
- Rocha, M.C.; Godoy, K.F.D.; Bannitzfernandes, R.; Fabri, J.H.T.M.; Barbosa, M.M.F.; Castro, P.A.D.; Almeida, F.; Goldman, G.H.; Cunha, A.F.D.; Netto, L.E.S. Analyses of the three 1-Cys Peroxiredoxins from Aspergillus fumigatus reveal that cytosolic Prx1 is central to H2O2 metabolism and virulence. Sci. Rep. 2018, 8, 12314. [Google Scholar] [CrossRef]
- Wang, M.; Sun, X.; Yu, D.; Xu, J.; Chung, K.; Li, H. Genomic and transcriptomic analyses of the tangerine pathotype of Alternaria alternata in response to oxidative stress. Sci. Rep. 2016, 6, 32437. [Google Scholar] [CrossRef]
- Angelova, M.B.; Pashova, S.B.; Spasova, B.K.; Vassilev, S.V.; Slokoska, L.S. Oxidative stress response of filamentous fungi induced by hydrogen peroxide and paraquat. Mycol. Res. 2005, 109, 150–158. [Google Scholar] [CrossRef]
- Mitchell, D.A.; Marshall, T.K.; Deschenes, R.J. Vectors for the inducible overexpression of glutathione S-transferase fusion proteins in yeast. Yeast 2010, 9, 715–722. [Google Scholar] [CrossRef]
- Lee, J.C.; Straffon, M.J.; Jang, T.Y.; Higgins, V.J.; Grant, C.M.; Dawes, I.W. The essential and ancillary role of glutathione in Saccharomyces cerevisiae analysed using a grande gsh1 disruptant strain. Fems Yeast Res. 2001, 1, 57–65. [Google Scholar]
- Andrišić, L.; Collinson, E.J.; Tehlivets, O.; Perak, E.; Žarković, T.; Dawes, I.W.; Žarković, N.; Gasparović, A.Č. Transcriptional and antioxidative responses to endogenous polyunsaturated fatty acid accumulation in yeast. Mol. Cell. Biochem. 2015, 399, 27–37. [Google Scholar] [CrossRef]
- Cipak, A.; Jaganjac, M.; Tehlivets, O.; Kohlwein, S.D.; Zarkovic, N. Adaptation to oxidative stress induced by polyunsaturated fatty acids in yeast. Bba - Mol. Cell Biol. Lipids 2008, 1781, 283–287. [Google Scholar] [CrossRef] [PubMed]
- Ianutsevich, E.A.; Danilova, O.A.; Groza, N.V.; Tereshina, V.M. Membrane lipids and cytosol carbohydrates in Aspergillus niger under osmotic, oxidative, and cold impact. Microbiology 2016, 85, 302–310. [Google Scholar] [CrossRef]

| Samples | WT | 3 mM H2O2 | 6 mM H2O2 | 0.28 mM MSB | 0.42 mM MSB |
|---|---|---|---|---|---|
| Clean Reads | 40,788,832 | 41,412,228 | 41,102,850 | 43,613,782 | 39,908,124 |
| GC Content | 52.28% | 52.47% | 52.26% | 51.64% | 51.88% |
| % ≥ Q30 | 94.27% | 94.33% | 94.40% | 94.36% | 94.35% |
| Mapped reads | 36,552,991 (89.62%) | 34,900,808 (84.28%) | 34,852,216 (84.79%) | 38,343,753 (87.92%) | 35,212,300 (88.23%) |
| Unique mapped | 36,452,891 (89.37%) | 34,787,458 (84.00%) | 34,722,583 (84.48%) | 38,241,527 (87.68%) | 35,122,814 (88.01%) |
| Multiple mapped | 100,100 (0.25%) | 113,350 (0.27%) | 129,633 (0.32%) | 102,226 (0.23%) | 89,486 (0.22%) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shao, H.; Tu, Y.; Wang, Y.; Jiang, C.; Ma, L.; Hu, Z.; Wang, J.; Zeng, B.; He, B. Oxidative Stress Response of Aspergillus oryzae Induced by Hydrogen Peroxide and Menadione Sodium Bisulfite. Microorganisms 2019, 7, 225. https://doi.org/10.3390/microorganisms7080225
Shao H, Tu Y, Wang Y, Jiang C, Ma L, Hu Z, Wang J, Zeng B, He B. Oxidative Stress Response of Aspergillus oryzae Induced by Hydrogen Peroxide and Menadione Sodium Bisulfite. Microorganisms. 2019; 7(8):225. https://doi.org/10.3390/microorganisms7080225
Chicago/Turabian StyleShao, Huanhuan, Yayi Tu, Yijing Wang, Chunmiao Jiang, Long Ma, Zhihong Hu, Jiangfan Wang, Bin Zeng, and Bin He. 2019. "Oxidative Stress Response of Aspergillus oryzae Induced by Hydrogen Peroxide and Menadione Sodium Bisulfite" Microorganisms 7, no. 8: 225. https://doi.org/10.3390/microorganisms7080225
APA StyleShao, H., Tu, Y., Wang, Y., Jiang, C., Ma, L., Hu, Z., Wang, J., Zeng, B., & He, B. (2019). Oxidative Stress Response of Aspergillus oryzae Induced by Hydrogen Peroxide and Menadione Sodium Bisulfite. Microorganisms, 7(8), 225. https://doi.org/10.3390/microorganisms7080225





