Polyphasic Characterization of Yeasts and Lactic Acid Bacteria Metabolic Contribution in Semi-Solid Fermentation of Chinese Baijiu (Traditional Fermented Alcoholic Drink): Towards the Design of a Tailored Starter Culture
Abstract
1. Introduction
2. Materials and Methods
2.1. Screening of Yeast and LAB Isolates for Fermentation
2.1.1. Strain Cultivation
2.1.2. Inoculum Preparation
2.1.3. Assays of α-Amylase and Glucoamylase Activity
2.1.4. Glucose Consumption Assay
2.1.5. Ethanol Tolerance Assay
2.2. Fermentation Trials
2.2.1. Inoculum Preparation
2.2.2. Preparation of Semi-Solid Fermentation Medium
2.2.3. Lab-Scale Fermentation Trials
2.3. Detection of Acetic Acid and Lactic Acid Content
2.4. GC–MS Analysis of Volatile Compounds
2.5. Statistical Analysis
3. Results
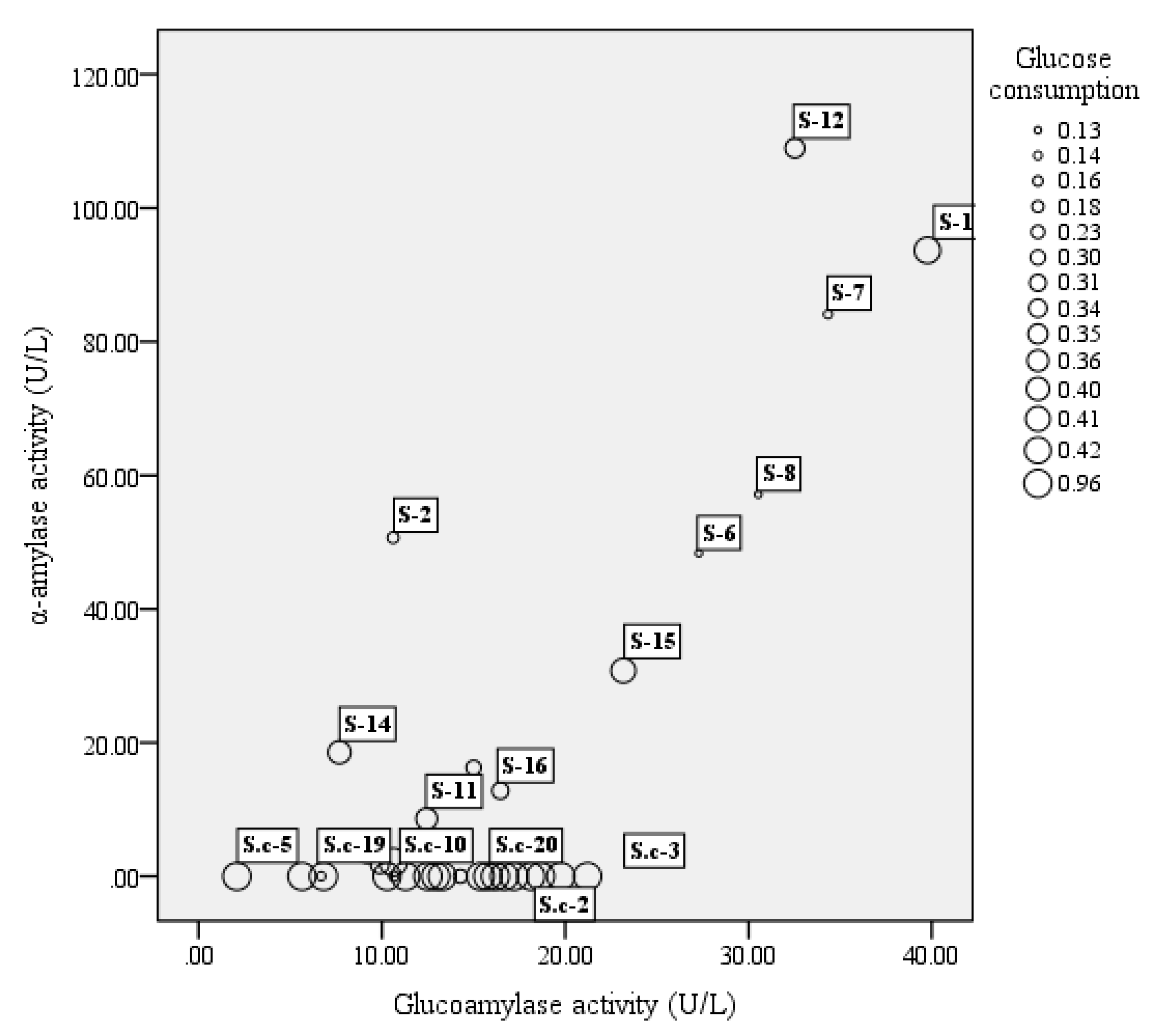
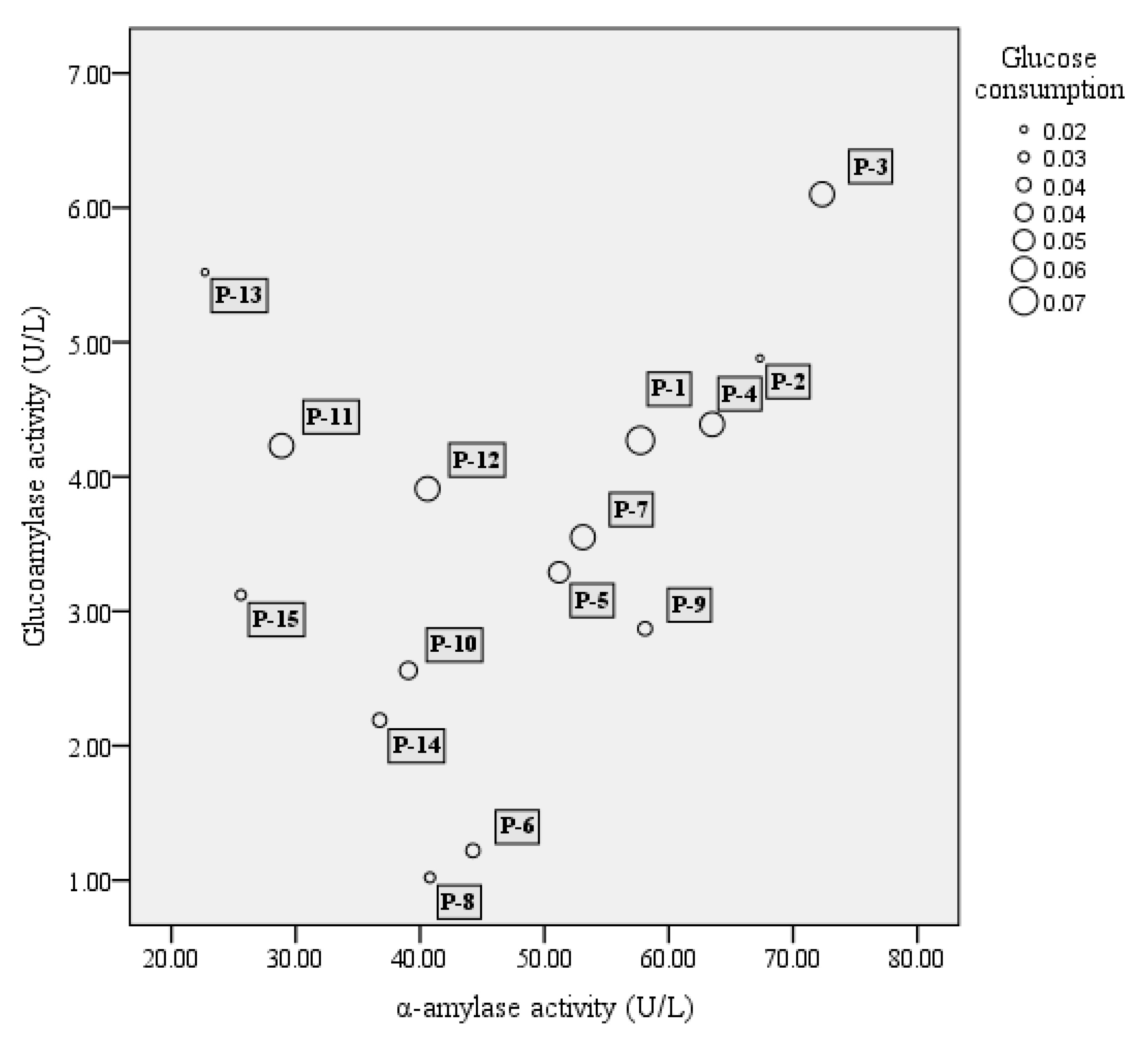
3.1. Strain Screening for Activity of α-Amylase, Glucoamylase, and Glucose Utilization
3.2. Strain Screening for Ethanol Tolerance
3.3. Design of Fermentation Trials with Different Strain Mixtures
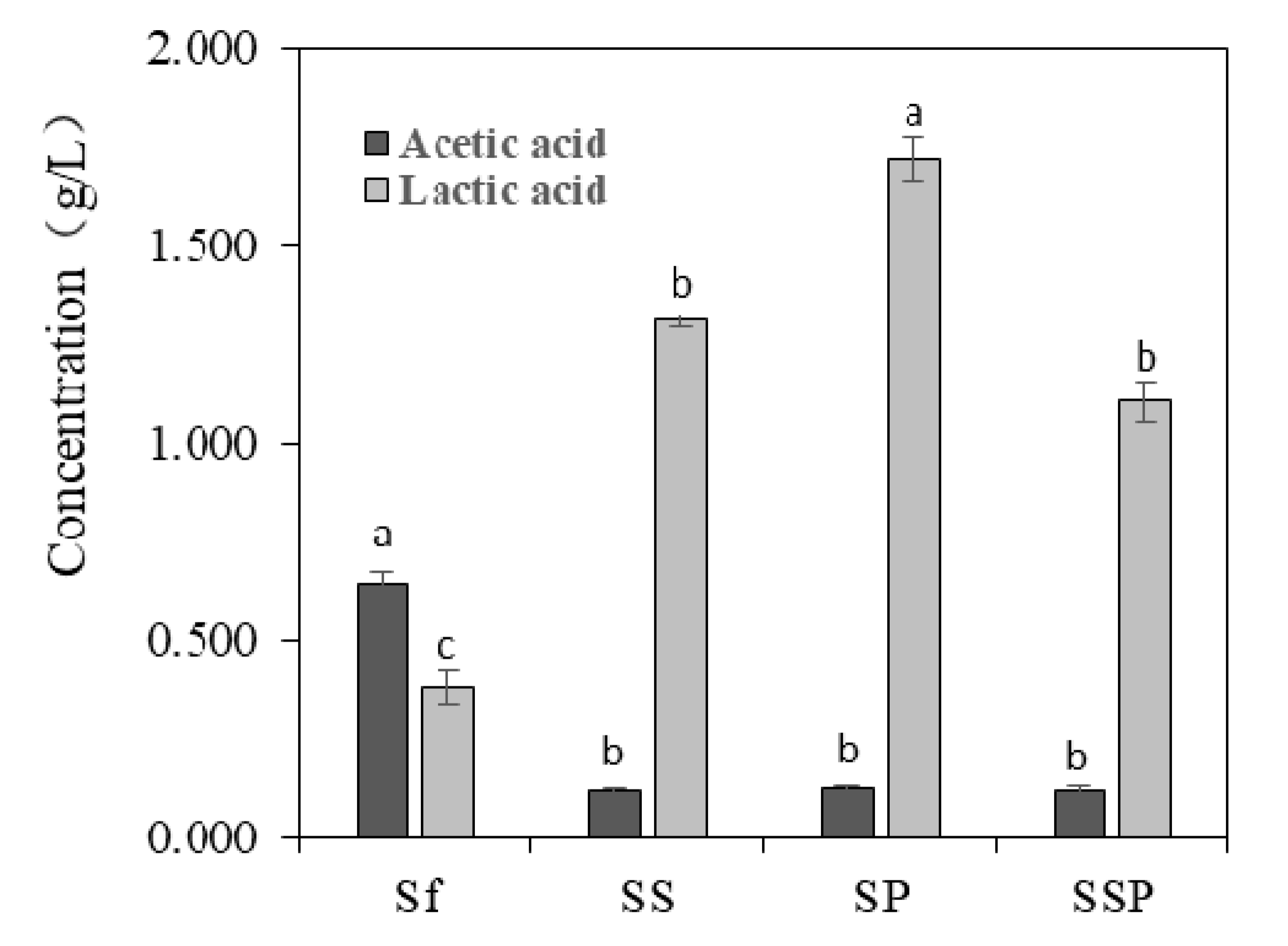
3.4. Content of Acetic Acid and Lactic Acid
3.5. The Metabolic Profiles in Different Fermentation Trials
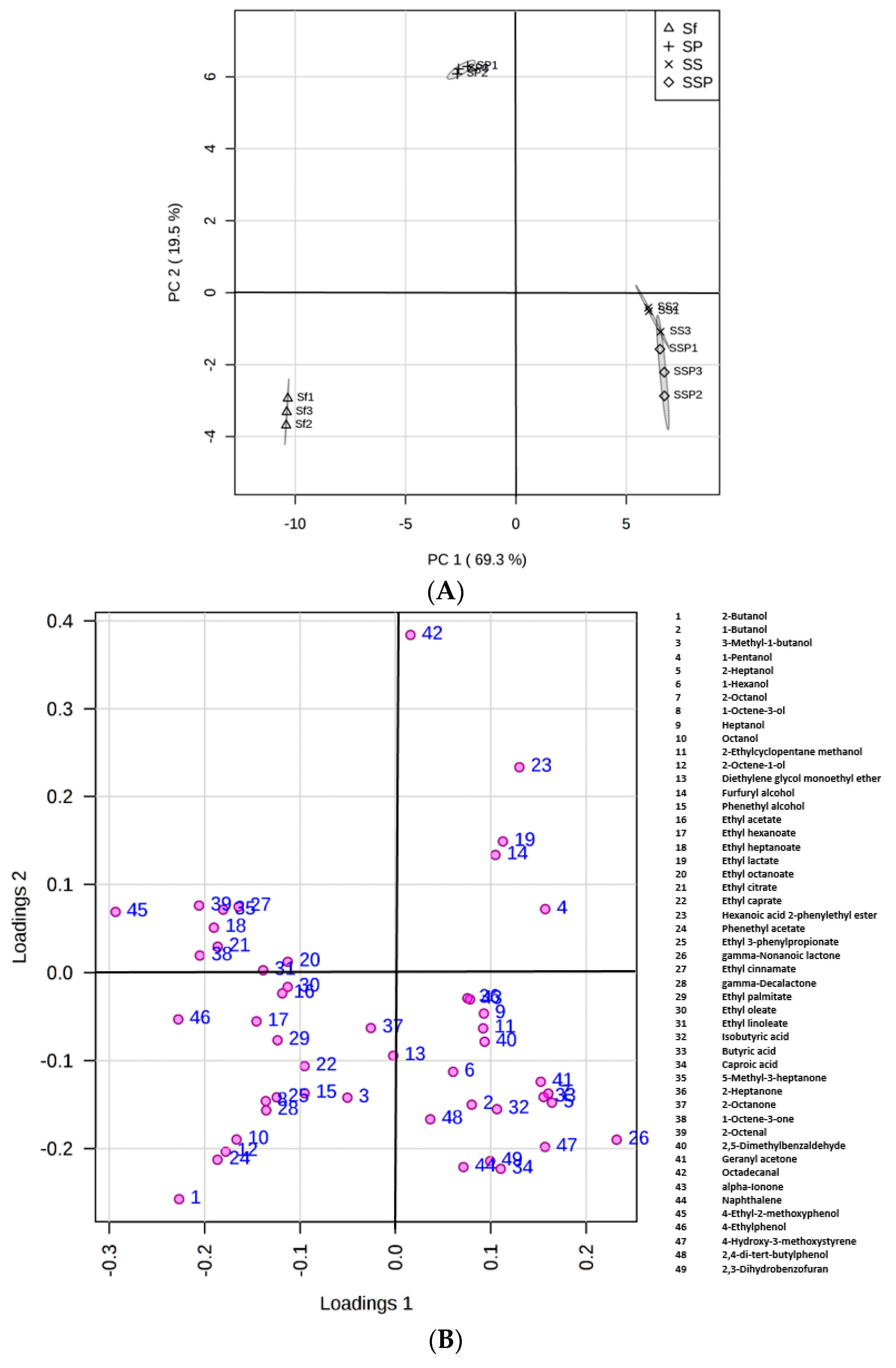
3.6. Statistical Analysis of Flavor Compounds
4. Discussion
4.1. Contributions of Tested Strains in Baijiu Fermentation
4.1.1. S. fibuligera
4.1.2. S. cerevisiae
4.1.3. P. acidilactici
4.2. Interactions of Selected Strains in Baijiu Fermentation
5. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Du, H.; Fan, W.; Xu, Y. Characterization of geosmin as source of earthy odor in different aroma type Chinese liquors. J. Agric. Food Chem. 2011, 59, 8331–8337. [Google Scholar] [CrossRef]
- Liu, H.; Sun, B. Effect of Fermentation Processing on the Flavor of Baijiu. J. Agric. Food Chem. 2018, 66, 5425–5432. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Li, Y.J.; Huang, J.; Hu, Y.; Zhang, Z.Y. Classification, production process and nutritional value of Chinese liquor. Ind. Rev. 2007, 1, 21–24. [Google Scholar]
- Zheng, X.W.; Han, B.Z. Baijiu, Chinese liquor: History, classification and manufacture. J. Ethn. Foods 2016, 3, 19–25. [Google Scholar] [CrossRef]
- Li, P.; Li, S.; Cheng, L.; Luo, L. Analyzing the relation between the microbial diversity of Daqu and the turbidity spoilage of traditional Chinese vinegar. Appl. Microbiol. Biotechnol. 2014, 98, 6073–6084. [Google Scholar] [CrossRef]
- Zha, M.S.; Sun, B.G.; Wu, Y.P.; Yin, S.; Wang, C.T. Improving flavor metabolism of Saccharomyces cerevisiae by mixed culture with Wickerhamomyces anomalus for Chinese Baijiu making. J. Biosci. Bioeng. 2018, 126, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Tabassum, R.; Khaliq, S.; Rajoka, M.I.; Agblevor, F. Solid State Fermentation of a Raw Starch Digesting Alkaline Alpha-Amylase from Bacillus licheniformis RT7PE1 and Its Characteristics. Biotechnol. Res. Int. 2014, 2014, 495384. [Google Scholar] [CrossRef]
- Wang, Y.; Yuan, B.; Ji, Y.; Li, H. Hydrolysis of hemicellulose to produce fermentable monosaccharides by plasma acid. Carbohydr. Polym. 2013, 97, 518–522. [Google Scholar] [CrossRef]
- Kim, H.S.; Kim, N.R.; Choi, W. Total fatty acid content of the plasma membrane of Saccharomyces cerevisiae; is more responsible for ethanol tolerance than the degree of unsaturation. Biotechnol. Lett. 2011, 33, 509–515. [Google Scholar] [CrossRef]
- Wu, Q.; Ling, J.; Xu, Y. Starter culture selection for making Chinese sesame-flavored liquor based on microbial metabolic activity in mixed-culture fermentation. Appl. Environ. Microbiol. 2014, 80, 4450–4459. [Google Scholar] [CrossRef]
- Fan, W.L.; Shen, H.; Xu, Y. Quantification of volatile compounds in Chinese soy sauce aroma type liquor by stir bar sorptive extraction and gas chromatography–mass spectrometry. J. Sci. Food Agric. 2011, 91, 1187–1198. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.X.; Wu, Z.Y.; Zhang, Q.S.; Wang, R.; Li, H. Combination of newly developed high quality Fuqu with traditional Daqu for Luzhou-flavor liquor brewing. World J. Microbiol. Biotechnol. 2009, 25, 1721–1726. [Google Scholar] [CrossRef]
- Kaneuchi, C.; Seki, M.; Komagata, K. Production of succinic acid from citric acid and related acids by Lactobacillus strains. Appl. Environ. Microbiol. 1988, 54, 3053–3056. [Google Scholar]
- Rojas, V.; Gil, J.V.; Piñaga, F.; Manzanares, P. Studies on acetate ester production by non-Saccharomyces wine yeasts. Int. J. Food Microbiol. 2011, 70, 283–289. [Google Scholar] [CrossRef]
- Fan, W.; Qian, M.C. Characterization of aroma compounds of Chinese ‘‘Wuliangye” and ‘‘Jiannanchun” liquors by aroma extraction dilution analysis. J. Agric. Food Chem. 2006, 54, 2695–2704. [Google Scholar] [CrossRef] [PubMed]
- Niu, Y.W.; Yao, Z.M.; Xiao, Q.; Xiao, Z.B.; Ma, N.; Zhu, J.C. Characterization of the key aroma compounds in different light aroma type Chinese liquors by GC-olfactometry, GC-FPD, quantitative measurements, and aroma recombination. Food Chem. 2017, 233, 204–215. [Google Scholar] [CrossRef] [PubMed]
- Stephan, A.; Steinhart, H. Quantification and sensory studies of character impact odorants of different soybean lecithins. J. Agric. Food Chem. 1999, 47, 4357–4364. [Google Scholar] [CrossRef]
- Zhao, D.; Li, Y.H.; Xiang, S.Q. Determination of aromatic compounds in distiller’s grains and in liquor by GC-MS. Liq.-Mak. Sci. Technol. 2006, 10, 92–95. [Google Scholar]
- Nie, Z.; Zheng, Y.; Wang, M.; Han, Y.; Wang, Y.; Luo, J.; Niu, D. Exploring microbial succession and diversity during solid-state fermentation of Tianjin duliu mature vinegar. Bioresour. Technol. 2013, 148, 325–333. [Google Scholar] [CrossRef]
- Saelim, K.; Dissara, Y.; H-Kittikun, A. Saccharification of cassava starch by Saccharomycopsis fibuligera YCY1 isolated from Loog-Pang (rice cake starter). Songklanakarin J. Sci. Technol. 2008, 30, 65–71. [Google Scholar]
- Chen, L.; Chi, Z.M.; Chi, Z.; Li, M. Amylase production by Saccharomycopsis fibuligera A11 in solid-state fermentation for hydrolysis of cassava starch. Appl. Biochem. Biotechnol. 2010, 162, 252–263. [Google Scholar] [CrossRef]
- Chi, Z.M.; Chi, Z.; Liu, G.L.; Wang, F.; Ju, L.; Zhang, T. Saccharomycopsis fibuligera and its applications in biotechnology. Biotechnol. Adv. 2009, 27, 423–431. [Google Scholar] [CrossRef]
- Wang, H.Y.; Gao, Y.B.; Fan, Q.W.; Xu, Y. Characterization and comparison of microbial community of different typical Chinese liquor Daqus by PCR-DGGE. Lett. Appl. Microbiol. 2011, 53, 134–140. [Google Scholar] [CrossRef]
- Liu, C.; Xie, G.F.; Sun, J.Q.; Zou, H.J.; Wang, R.Q.; Qian, B.; Zang, W.; Zhou, J.D.; Shou, H.R.; Bai, F.Y. Research progress on the microorganisms of Chinese ricewine. Sci. Technol. Food Ind. 2016, 8, 374–380. [Google Scholar]
- Yan, S.M.; Yan, H.W.; Chen, L. Diversity of culturable fungi in different sweet alcoholic drink’s starters. China Brew. 2015, 120, 48–54. [Google Scholar]
- Ismaya, W.T.; Hasan, K.; Kardi, I.; Zainuri, A.; Rahmawaty, R.I.; Permanahadi, S.; Viera, B.V.; Harinanto, G.; Gaffar, S.; Natalia, D.; Subroto, T.; Soemitro, S. Chemical modification of Saccharomycopsis fibuligera R64 α-amylase to improve its stability against thermal, chelator, and proteolytic inactivation. Appl. Biochem. Biotechnol. 2013, 170, 44–57. [Google Scholar] [CrossRef]
- Brimer, L.; Nout, M.J.R.; Tuncel, G. b-glycosidase (amygdalase and linamarase) from Endomyces fibuliger (LU677): formation and crude enzyme properties. Appl. Microbiol. Biotechnol. 1998, 49, 182–188. [Google Scholar] [CrossRef]
- Knox, A.M.; du Preez, J.C.; Kilian, S.G. Starch fermentation characteristics of Saccharomyces cerevisiae strains transformed with amylase genes from Lipomyces kononenkoae and Saccharomycopsis fibuligera. Enzyme Microb. Technol. 2004, 34, 453–460. [Google Scholar] [CrossRef]
- Li, X.R.; Ma, E.B.; Yan, L.Z.; Han, M.H.; Du, X.W.; Zhang, S.W.; Quan, Z.X. Bacterial and fungal diversity in the traditional Chinese liquor fermentation process. Int. J. Food Microbiol. 2011, 146, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.D.; Chen, M.Z.; Ban, S.D. Separating and identifying and function-exploring of the yeast strains in Moutai-flavor Daqu. Food Sci. 2017, 38, 51–57. [Google Scholar]
- Nout, M.J.R. Rich nutrition from the poorest e cereal fermentations in Africa and Asia. Food Microbiol. 2009, 26, 685–692. [Google Scholar] [CrossRef]
- Urso, R.; Rantsiou, K.; Dolci, P.; Rolle, L.; Comi, G.; Cocolin, L. Yeast biodiversity and dynamics during sweet wine production as determined by molecular methods. FEMS Yeast Res. 2008, 8, 1053–1062. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.W.; Yan, Z.; Han, B.Z. Complex microbiota of a Chinese “Fen” liquor fermentation starter (Fen-Daqu), revealed by culture-dependent and culture-independent methods. Food Microbiol. 2012, 31, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.G.; Hu, Y.G.; Ma, Y.H.; Du, X.W.; Ma, E.B.; Quan, Z.X.; Zhang, S.W. Variation of aromatic components in solid phase fermented grains during fermentation of Fen liquor. Food Sci. 2010, 31, 367–371. [Google Scholar]
- Dung, N.T.P.; Rombouts, F.M.; Nout, M.J.R. Characteristics of some traditional Vietnamese starch-based rice wine fermentation starters (men). LWT-Food Sci. Technol. 2007, 40, 130–135. [Google Scholar] [CrossRef]
- Jeyaram, K.; Singh, W.M.; Capece, A.; Romano, P. Molecular identification of yeast species associated with ‘Hamei’ e a traditional starter used for rice wine production in Manipur, India. Int. J. Food Microbiol. 2008, 124, 115–125. [Google Scholar] [CrossRef] [PubMed]
- Nyanga, L.K.; Nout, M.J.R.; Gadaga, T.H.; Theelen, B.; Boekhout, T.; Zwietering, M.H. Yeasts and lactic acid bacteria microbiota from masau (Ziziphus maur-itiana) fruits and their fermented fruit pulp in Zimbabwe. Int. J. Food Microbiol. 2007, 120, 159–166. [Google Scholar] [CrossRef]
- Sujaya, I.N.; Antara, N.S.; Sone, T.; Tamura, Y.; Aryanta, W.R.; Yokota, A.; Asano, K.; Tomita, F. Identification and characterization of yeasts in brem, a traditional Balinese rice wine. World J. Microbiol. Biotechnol. 2004, 20, 143–150. [Google Scholar] [CrossRef]
- Sun, Y.T. The fermentation characteristics of yeast species isolated from Fen-Daqu during alcoholic fermentation. Master’s Thesis, China Agricultral University, Beijing, China, May 2014. [Google Scholar]
- Nadaroglu, H.; Adiguzel, G.; Adiguzel, A.; Sönmez, Z. A thermostable-endo-β-(1,4)-mannanase from Pediococcus acidilactici (M17): Purification, characterization and its application in fruit juice clarification. Eur. Food Res. Technol. 2017, 243, 193–201. [Google Scholar] [CrossRef]
- Adiguzel, G.; Faiz, O.; Sisecioglu, M.; Sari, B.; Baltaci, O.; Akbulut, S.; Genc, B.; Adiguzel, A. A novel endo-β-1, 4-xylanase from Pediococcus acidilactici GC25; purification, characterization and application in clarification of fruit juices. Int. J. Biol. Macromol. 2019, 129, 571–578. [Google Scholar] [CrossRef]
- Wang, W.; Zhao, H.; Chen, F.G. Isolation & preliminary identification of lactic acid Bacteria in Luzhou-flavor pit mud. Liqour-Mak. Sci. Technol. 2006, 4, 29–31. [Google Scholar]
- Xing, M.Y. Community struction and function of lactic acid bacteria during fermentation process for sesame-flavor liquor making. Master’s Thesis, Jiangnan University, Wuxi, China, June 2017. [Google Scholar]
- Du, H.; Xing, M.Y.; Xu, Y. Isolation of lactic acid bacteria and their characteristic of carbon utilization during the liquor-making process of Chinese roasted sesame-like flavor liquor. Food Ferment. Ind. 2018, 44, 13–18. [Google Scholar]
- Huffer, S.; Clark, M.E.; Ning, J.C.; Blanch, H.W.; Clark, D.S. Role of alcohols in growth, lipid composition, and membrane fluidity of yeasts, bacteria, and archaea. Appl. Environ. Microbiol. 2011, 77, 6400–6408. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Qureshi, N. How microbes tolerate ethanol and butanol. New Biotechnol. 2009, 26, 117–121. [Google Scholar] [CrossRef]
- Wang, G.; Ning, J.; Zhao, J.C.; Hang, F.; Tian, F.W.; Zhao, J.X.; Chen, Y.Q.; Zhang, H.; Chen, W. Partial characterization of an anti-listeria substance produced by Pediococcus acidilactici P9. Int. Dairy J. 2014, 34, 275–279. [Google Scholar] [CrossRef]
- Tristezza, M.; Feo, L.D.; Tufariello, M.; Grieco, F.; Capozzi, V.; Spano, G.; Mita, G.; Grieco, F. Simultaneous inoculation of yeasts and lactic acid bacteria: Effects on fermentation dynamics and chemical composition of Negroamaro wine. LWT-Food Sci. Technol. 2016, 66, 406–412. [Google Scholar] [CrossRef]
| Microbial Isolates | Growth on Medium 1 Containing Ethanol | |||
|---|---|---|---|---|
| 0 | 4% | 8% | 10% | |
| S.c-2, 3, 4, 5, 8, 9, 10, 11, 12, 14, 15, 16, 17, 18, 19, 20 | + | + | + | + |
| S.c-1, 7 | + | + | + | W |
| S.c-6, 13 | + | + | + | – |
| S-3, 4, 6, 10, 12 | + | + | + | – |
| S-14 | + | + | W | – |
| S-1, 2, 5, 7, 8, 9, 11, 13, 15, 16 | + | + | – | – |
| P-4 | + | + | + | W |
| P-15 | + | + | W | – |
| P-1, 2, 3, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14 | + | + | – | – |
| Compounds | Concentration (mg/L) 1 | |||
|---|---|---|---|---|
| Sf | SP | SS | SSP | |
| Alcohols | ||||
| 2-Butanol | 6.60 ± 0.80 a | – b | – b | – b |
| 1-Butanol | 0.33 ± 0.06 b | 0.17 ± 0.04 c | 1.04 ± 0.11 a | 0.45 ± 0.05 b |
| 3-Methyl-1-butanol | 31.56 ± 1.15 a | 13.82 ± 2.94 c | 14.41 ± 1.18 c | 22.69 ± 3.84 b |
| 1-Pentanol | – c | 0.12 ± 0.02 b | 0.15 ± 0.05 b | 0.36 ± 0.06 a |
| 2-Heptanol | – c | – c | 0.13 ± 0.02 b | 0.37 ± 0.09 a |
| 1-Hexanol | 0.46 ± 0.04 bc | 0.36 ± 0.05 c | 0.55 ± 0.05 b | 0.87 ± 0.17 a |
| 2-Octanol | – c | – c | 0.14 ± 0.01 b | 0.24 ± 0.04 a |
| 1-Octene-3-ol | 1.41 ± 0.33 a | 0.20 ± 0.06 b | 0.16 ± 0.02 b | 0.21 ± 0.07 b |
| Heptanol | 0.20 ± 0.02 c | 0.25 ± 0.04 bc | 0.38 ± 0.10 b | 0.54 ± 0.11 a |
| Octanol | 0.46 ± 0.04 a | – b | – b | – b |
| 2-Ethylcyclopentane methanol | 0.19 ± 0.02 c | 0.22 ± 0.04 bc | 0.31 ± 0.04 b | 0.61 ± 0.08 a |
| 2-Octene-1-ol | 0.74 ± 0.21 a | – b | – b | – b |
| Diethylene glycol monoethyl ether | 2.10 ± 0.35 b | 1.47 ± 0.27 bc | 1.14 ± 0.08 c | 3.16 ± 0.92 a |
| Furfuryl alcohol | 0.11 ± 0.04 c | 0.69 ± 0.17 b | 0.28 ± 0.04 c | 0.98 ± 0.11 a |
| Phenethyl alcohol | 27.75 ± 0.99 a | 8.08 ± 1.80 b | 7.98 ± 0.11 b | 9.86 ± 0.80 b |
| Esters | ||||
| Ethyl acetate | 1.32 ± 0.08 a | 0.67 ± 0.04 b | 0.33 ± 0.05 c | 0.42 ± 0.07 c |
| Ethyl hexanoate | 1.07 ± 0.07 a | 0.31 ± 0.04 b | 0.11 ± 0.04 c | 0.19 ± 0.05 c |
| Ethyl heptanoate | 0.42 ± 0.07 a | 0.16 ± 0.02 b | – c | – c |
| Ethyl lactate | – d | 0.19 ± 0.02 b | 0.07 ± 0.04 c | 0.30 ± 0.05 a |
| Ethyl octanoate | 0.39 ± 0.03 a | 0.26 ± 0.05 b | 0.09 ± 0.02 d | 0.16 ± 0.02 c |
| Ethyl citrate | 0.37 ± 0.02 a | 0.12 ± 0.02 b | – c | – c |
| Ethyl caprate | 0.06 ± 0.02 a | – b | –b | – b |
| Hexanoic acid 2-phenylethyl ester | – c | 0.50 ± 0.05 a | 0.19 ± 0.02b | 0.30 ± 0.05 b |
| Phenethyl acetate | 1.02 ± 0.17 a | – b | – b | – b |
| Ethyl 3-phenylpropionate | 0.12 ± 0.02 a | – b | – b | – b |
| γ-Nonanoic lactone | – c | – c | 1.62 ± 0.00b | 2.27 ± 0.05 a |
| Ethyl cinnamate | 0.40 ± 0.06 a | 0.21 ± 0.01b | 0.07 ± 0.01c | – d |
| γ-Decalactone | 0.18 ± 0.07 a | – b | – b | – b |
| Ethyl palmitate | 2.55 ± 0.30 a | 0.84 ± 0.07 b | 0.51 ± 0.02 c | 0.65 ± 0.11 bc |
| Ethyl oleate | 0.88 ± 0.15 a | 0.49 ± 0.12 b | 0.12 ± 0.02 c | 0.39 ± 0.09 b |
| Ethyl linoleate | 0.83 ± 0.16 a | 0.39 ± 0.04 b | 0.16 ± 0.02 c | 0.16 ± 0.02 c |
| Acids | ||||
| Isobutyric acid | 0.14 ± 0.04 b | 0.09 ± 0.02 b | 0.45 ± 0.06 a | 0.39 ± 0.09 a |
| Butyric acid | – c | – c | 0.11 ± 0.04 b | 0.29 ± 0.08 a |
| Caproic acid | 0.22 ± 0.09 b | 0.07 ± 0.03 b | 0.65 ± 0.13 a | 0.82 ± 0.11 a |
| Aldehydes and Ketones | ||||
| 2-Octenal | 0.70 ± 0.13 a | 0.28 ± 0.07 b | – c | – c |
| 2,5-Dimethylbenzaldehyde | 0.67 ± 0.07 c | 0.69 ± 0.13 c | 1.30 ± 0.10 b | 1.66 ± 0.32 a |
| Octadecanal | – c | 1.11 ± 0.30 a | 0.17 ± 0.04 b | – c |
| 5-Methyl-3-heptanone | 0.32 ± 0.09 a | 0.16 ± 0.02 b | – c | – c |
| 2-Heptanone | –b | –b | 0.12 ± 0.02 a | –b |
| 2-Octanone | 0.24 ± 0.04 ab | 0.19 ± 0.05 b | 0.13 ± 0.02 b | 0.28 ± 0.04 a |
| 1-Octene-3-one | 0.71 ± 0.15 a | 0.16 ± 0.02 b | – c | – c |
| Geranyl acetone | –b | –b | 0.15 ± 0.05 a | 0.15 ± 0.04 a |
| alpha-Ionone | –b | –b | 0.14 ± 0.04 a | – b |
| Aromatics | ||||
| Naphthalene | 0.19 ± 0.05 b | 0.07 ± 0.03 c | 0.23 ± 0.02 b | 0.50 ± 0.11 a |
| 4-Ethyl-2-methoxyphenol | 27.75 ± 2.56 a | 2.40 ± 0.31 b | – c | – c |
| 4-Ethylphenol | 9.83 ± 1.15 a | 0.71 ± 0.14 b | 0.10 ± 0.04 b | 0.11 ± 0.06 b |
| 4-Hydroxy-3-methoxystyrene | 1.55 ± 0.40 b | 0.63 ± 0.07 b | 7.55 ± 1.27 a | 13.00 ± 2.86 a |
| 2,4-di-tert-butylphenol | 4.78 ± 1.29 b | 2.46 ± 0.59 c | 4.32 ± 0.78 b | 7.29 ± 1.48 a |
| 2,3-Dihydrobenzofuran | 1.99 ± 0.09 c | 0.78 ± 0.04 c | 4.09 ± 0.48 b | 7.13 ± 1.81 a |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, R.; Sui, L.; Zhang, J.; Hu, J.; Liu, P. Polyphasic Characterization of Yeasts and Lactic Acid Bacteria Metabolic Contribution in Semi-Solid Fermentation of Chinese Baijiu (Traditional Fermented Alcoholic Drink): Towards the Design of a Tailored Starter Culture. Microorganisms 2019, 7, 147. https://doi.org/10.3390/microorganisms7050147
Ma R, Sui L, Zhang J, Hu J, Liu P. Polyphasic Characterization of Yeasts and Lactic Acid Bacteria Metabolic Contribution in Semi-Solid Fermentation of Chinese Baijiu (Traditional Fermented Alcoholic Drink): Towards the Design of a Tailored Starter Culture. Microorganisms. 2019; 7(5):147. https://doi.org/10.3390/microorganisms7050147
Chicago/Turabian StyleMa, Rufei, Lu Sui, Jingsheng Zhang, Jinrong Hu, and Ping Liu. 2019. "Polyphasic Characterization of Yeasts and Lactic Acid Bacteria Metabolic Contribution in Semi-Solid Fermentation of Chinese Baijiu (Traditional Fermented Alcoholic Drink): Towards the Design of a Tailored Starter Culture" Microorganisms 7, no. 5: 147. https://doi.org/10.3390/microorganisms7050147
APA StyleMa, R., Sui, L., Zhang, J., Hu, J., & Liu, P. (2019). Polyphasic Characterization of Yeasts and Lactic Acid Bacteria Metabolic Contribution in Semi-Solid Fermentation of Chinese Baijiu (Traditional Fermented Alcoholic Drink): Towards the Design of a Tailored Starter Culture. Microorganisms, 7(5), 147. https://doi.org/10.3390/microorganisms7050147





