Community- and Hospital-Acquired Klebsiella pneumoniae Urinary Tract Infections in Portugal: Virulence and Antibiotic Resistance
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Isolates
2.2. Antimicrobial Susceptibility Testing and Phenotypic Detection of Extended-Spectrum β-lactamase (ESBL) Production
2.3. Detection of Resistance and Virulence Genes
2.4. Multilocus Sequence Typing
2.5. PCR-based Replicon Typing
2.6. Statistical Analysis
2.7. Ethical Approval
3. Results
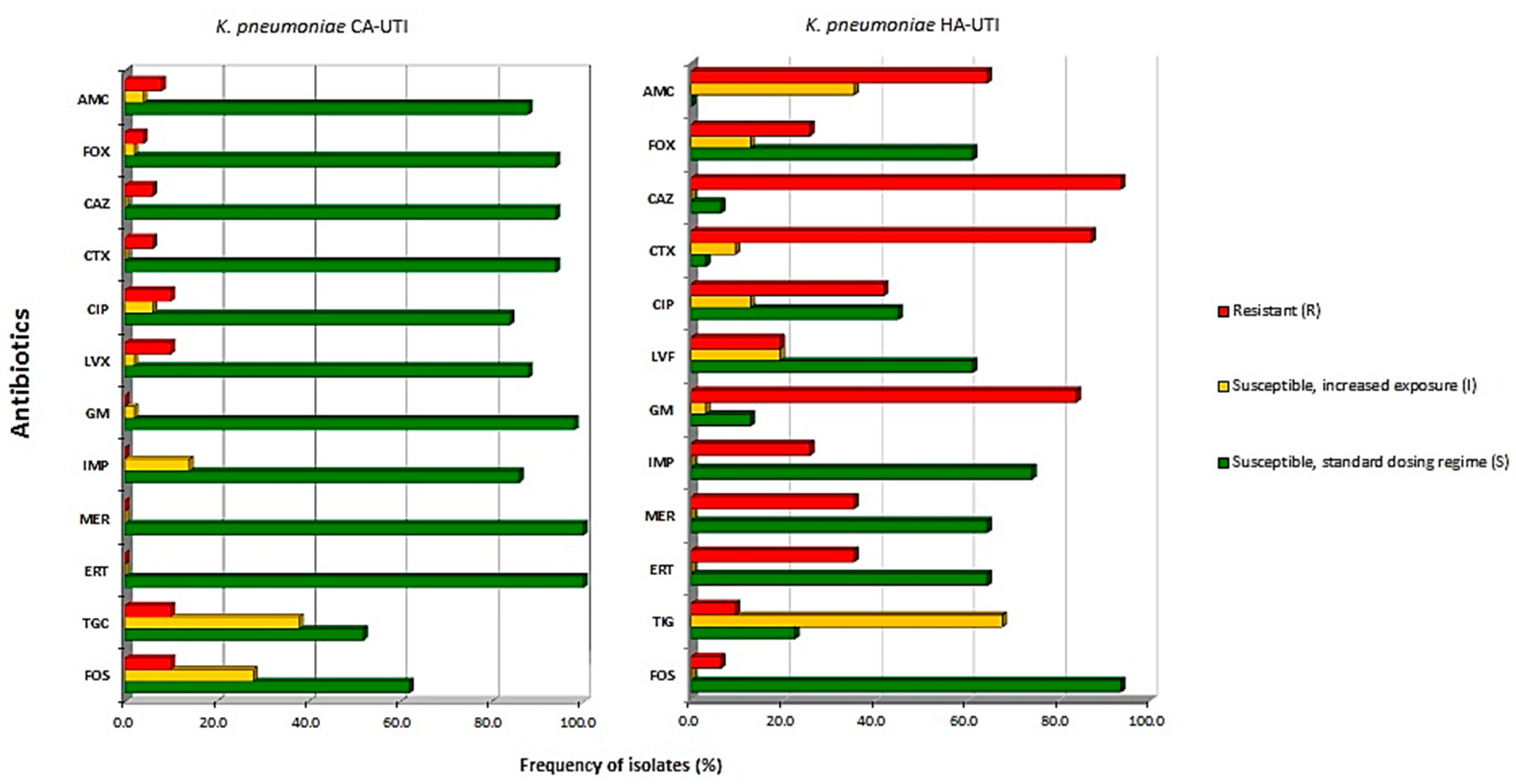
3.1. Antimicrobial Susceptibility
3.2. Identification of the β-Lactamases
3.3. Virulence Genes
3.4. MLST Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Podschun, R.; Ullmann, U. Klebsiella spp. as nosocomial pathogens: Epidemiology, taxonomy, typing methods, and pathogenicity factors. Clin. Microbiol. Rev. 1998, 11, 589–603. [Google Scholar] [CrossRef]
- Keynan, Y.; Rubinstein, E. The changing face of Klebsiella pneumoniae infections in the community. Int. J. Antimicrob. Agents 2007, 30, 385–389. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.H.; Wang, M.C.; Tseng, C.C.; Ko, W.C.; Wu, A.B.; Zheng, P.X.; Wu, J.J. Clinical and microbiological characteristics of Klebsiella pneumoniae isolates causing community-acquired urinary tract infections. Infection 2010, 38, 459–464. [Google Scholar] [CrossRef]
- WHO. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics. Available online: https://www.who.int/medicines/publications/global-priority-list-antibiotic-resistant-bacteria/en/ (accessed on 27 February 2017).
- Navon-Venezia, S.; Kondratyeva, K.; Carattoli, A. Klebsiella pneumoniae: A major worldwide source and shuttle for antibiotic resistance. FEMS Microbiol. Rev. 2017, 41, 252–275. [Google Scholar] [CrossRef]
- Butler, C.C. Antibiotics: Responding to a Global Challenge. Antibiotics 2012, 1, 14–16. [Google Scholar] [CrossRef] [PubMed]
- Kock, R.; Siemer, P.; Esser, J.; Kampmeier, S.; Berends, M.S.; Glasner, C.; Arends, J.P.; Becker, K.; Friedrich, A.W. Defining Multidrug Resistance of Gram-Negative Bacteria in the Dutch-German Border Region-Impact of National Guidelines. Microorganisms 2018, 6, 11. [Google Scholar] [CrossRef]
- Russo, R.; Kolesnikova, I.; Kim, T.; Gupta, S.; Pericleous, A.; Kadouri, D.E.; Connell, N.D. Susceptibility of Virulent Yersinia pestis Bacteria to Predator Bacteria in the Lungs of Mice. Microorganisms 2018, 7, 2. [Google Scholar] [CrossRef] [PubMed]
- Hennequin, C.; Robin, F. Correlation between antimicrobial resistance and virulence in Klebsiella pneumoniae. Eur. J. Clin. Microbiol. Infect. Dis. 2016, 35, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Russo, T.A.; Shon, A.S.; Beanan, J.M.; Olson, R.; MacDonald, U.; Pomakov, A.O.; Visitacion, M.P. Hypervirulent K. pneumoniae secretes more and more active iron-acquisition molecules than “classical“ K. pneumoniae thereby enhancing its virulence. PLoS ONE 2011, 6, e26734. [Google Scholar] [CrossRef]
- Khaertynov, K.S.; Anokhin, V.A.; Rizvanov, A.A.; Davidyuk, Y.N.; Semyenova, D.R.; Lubin, S.A.; Skvortsova, N.N. Virulence Factors and Antibiotic Resistance of Klebsiella pneumoniae Strains Isolated From Neonates With Sepsis. Front. Med. 2018, 5, 225. [Google Scholar] [CrossRef]
- Ferreira, R.L.; da Silva, B.C.M.; Rezende, G.S.; Nakamura-Silva, R.; Pitondo-Silva, A.; Campanini, E.B.; Brito, M.C.A.; da Silva, E.M.L.; Freire, C.C.M.; da Cunha, A.F.; et al. High Prevalence of Multidrug-Resistant Klebsiella pneumoniae Harboring Several Virulence and beta-Lactamase Encoding Genes in a Brazilian Intensive Care Unit. Front. Microbiol. 2018, 9, 3198. [Google Scholar] [CrossRef]
- Li, B.; Zhao, Y.; Liu, C.; Chen, Z.; Zhou, D. Molecular pathogenesis of Klebsiella pneumoniae. Future Microbiol. 2014, 9, 1071–1081. [Google Scholar] [CrossRef]
- Caneiras, C.; Lito, L.; Mayoralas-Alises, S.; Diaz-Lobato, S.; Melo-Cristino, J.; Duarte, A. Virulence and resistance determinants of Klebsiella pneumoniae isolated from a Portuguese tertiary university hospital centre over a 31-year period. Enferm. Infec. Microbiol. Clin. 2018. [Google Scholar] [CrossRef]
- Caneiras, C.; Calisto, F.; Jorge da Silva, G.; Lito, L.; Melo-Cristino, J.; Duarte, A. First Description of Colistin and Tigecycline-Resistant Acinetobacter baumannii Producing KPC-3 Carbapenemase in Portugal. Antibiotics 2018, 7, 96. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial disk susceptibility tests; approved standard. In Document M2-A11, 11th ed.; CLSI: Wayne, PA, USA, 2014. [Google Scholar]
- EUCAST Proposes to Retain Susceptibility Categories “S, I, and R” but to Change the Definitions to “Susceptible, Standard Dosing Regimen”, “Susceptible, Increased Exposure”, and “Resistant”. European Committee on Antimicrobial Susceptibility Testing (EUCAST), 2018. Available online: http://www.eucast.org/ (accessed on 28 February 2019).
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.C.; Koh, T.H.; Lee, N.; Fung, C.P.; Chang, F.Y.; Tsai, Y.K.; Ip, M.; Siu, L.K. Genotypes and virulence in serotype K2 Klebsiella pneumoniae from liver abscess and non-infectious carriers in Hong Kong, Singapore and Taiwan. Gut Pathog. 2014, 6, 21. [Google Scholar] [CrossRef]
- Compain, F.; Babosan, A.; Brisse, S.; Genel, N.; Audo, J.; Ailloud, F.; Kassis-Chikhani, N.; Arlet, G.; Decre, D. Multiplex PCR for detection of seven virulence factors and K1/K2 capsular serotypes of Klebsiella pneumoniae. J. Clin. Microbiol. 2014, 52, 4377–4380. [Google Scholar] [CrossRef] [PubMed]
- Siu, L.K.; Yeh, K.M.; Lin, J.C.; Fung, C.P.; Chang, F.Y. Klebsiella pneumoniae liver abscess: A new invasive syndrome. Lancet Infect. Dis. 2012, 12, 881–887. [Google Scholar] [CrossRef]
- Wang, X.; Xie, Y.; Li, G.; Liu, J.; Li, X.; Tian, L.; Sun, J.; Ou, H.Y.; Qu, H. Whole-Genome-Sequencing characterization of bloodstream infection-causing hypervirulent Klebsiella pneumoniae of capsular serotype K2 and ST374. Virulence 2018, 9, 510–521. [Google Scholar] [CrossRef] [PubMed]
- Wyres, K.L.; Wick, R.R.; Judd, L.M.; Froumine, R.; Tokolyi, A.; Gorrie, C.L.; Lam, M.M.C.; Duchene, S.; Jenney, A.; Holt, K.E. Distinct evolutionary dynamics of horizontal gene transfer in drug resistant and virulent clones of Klebsiella pneumoniae. PLoS Genet. 2019, 15, e1008114. [Google Scholar] [CrossRef] [PubMed]
- Struve, C.; Roe, C.C.; Stegger, M.; Stahlhut, S.G.; Hansen, D.S.; Engelthaler, D.M.; Andersen, P.S.; Driebe, E.M.; Keim, P.; Krogfelt, K.A. Mapping the Evolution of Hypervirulent Klebsiella pneumoniae. MBio 2015, 6, e00630. [Google Scholar] [CrossRef]
- Wasfi, R.; Elkhatib, W.F.; Ashour, H.M. Molecular typing and virulence analysis of multidrug resistant Klebsiella pneumoniae clinical isolates recovered from Egyptian hospitals. Sci. Rep. 2016, 6, 38929. [Google Scholar] [CrossRef]
- Diancourt, L.; Passet, V.; Verhoef, J.; Grimont, P.A.; Brisse, S. Multilocus sequence typing of Klebsiella pneumoniae nosocomial isolates. J. Clin. Microbiol. 2005, 43, 4178–4182. [Google Scholar] [CrossRef]
- Carattoli, A.; Bertini, A.; Villa, L.; Falbo, V.; Hopkins, K.L.; Threlfall, E.J. Identification of plasmids by PCR-based replicon typing. J. Microbiol. Methods 2005, 63, 219–228. [Google Scholar] [CrossRef]
- Bialek-Davenet, S.; Criscuolo, A.; Ailloud, F.; Passet, V.; Jones, L.; Delannoy-Vieillard, A.S.; Garin, B.; Le Hello, S.; Arlet, G.; Nicolas-Chanoine, M.H.; et al. Genomic definition of hypervirulent and multidrug-resistant Klebsiella pneumoniae clonal groups. Emerg. Infect. Dis. 2014, 20, 1812–1820. [Google Scholar] [CrossRef] [PubMed]
- Ku, Y.H.; Chuang, Y.C.; Chen, C.C.; Lee, M.F.; Yang, Y.C.; Tang, H.J.; Yu, W.L. Klebsiella pneumoniae Isolates from Meningitis: Epidemiology, Virulence and Antibiotic Resistance. Sci. Rep. 2017, 7, 6634. [Google Scholar] [CrossRef]
- Melo, R.D.C.A.; de Barros, E.M.; Loureiro, N.G.; de Melo, H.R.; Maciel, M.A.; Souza Lopes, A.C. Presence of fimH, mrkD, and irp2 virulence genes in KPC-2-producing Klebsiella pneumoniae isolates in Recife-PE, Brazil. Curr. Microbiol. 2014, 69, 824–831. [Google Scholar] [CrossRef]
- Antimicrobial Consumption Database (ESAC-Net). Available online: https://ecdc.europa.eu/en/antimicrobial-consumption/database/country-overview (accessed on 7 September 2018).
- Data from the ECDC Surveillance Atlas-Antimicrobial Resistance. Available online: https://ecdc.europa.eu/en/antimicrobial-resistance/surveillance-and-disease-data/data-ecdc (accessed on 7 September 2018).
- Surveillance of Antimicrobial Resistance in Europe 2017. Available online: https://ecdc.europa.eu/en/publications-data/surveillance-antimicrobial-resistance-europe-2017 (accessed on 2 February 2019).
- Alfouzan, W.; Dhar, R.; Nicolau, D.P. In Vitro Activity of Newer and Conventional Antimicrobial Agents, Including Fosfomycin and Colistin, against Selected Gram-Negative Bacilli in Kuwait. Pathogens 2018, 7, 75. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, S.; Sengupta, M.; Sarker, T.K. Fosfomycin susceptibility among multidrug-resistant, extended-spectrum beta-lactamase-producing, carbapenem-resistant uropathogens. Indian J. Urol. 2017, 33, 149–154. [Google Scholar] [CrossRef]
- Endimiani, A.; Patel, G.; Hujer, K.M.; Swaminathan, M.; Perez, F.; Rice, L.B.; Jacobs, M.R.; Bonomo, R.A. In vitro activity of fosfomycin against blaKPC-containing Klebsiella pneumoniae isolates, including those nonsusceptible to tigecycline and/or colistin. Antimicrob. Agents Chemother. 2010, 54, 526–529. [Google Scholar] [CrossRef]
- Bandeira, M.; Carvalho, P.A.; Duarte, A.; Jordao, L. Exploring Dangerous Connections between Klebsiella pneumoniae Biofilms and Healthcare-Associated Infections. Pathogens 2014, 3, 720–731. [Google Scholar] [CrossRef]
- Johnson, J.G.; Murphy, C.N.; Sippy, J.; Johnson, T.J.; Clegg, S. Type 3 fimbriae and biofilm formation are regulated by the transcriptional regulators MrkHI in Klebsiella pneumoniae. J. Bacteriol. 2011, 193, 3453–3460. [Google Scholar] [CrossRef] [PubMed]
- Murphy, C.N.; Clegg, S. Klebsiella pneumoniae and type 3 fimbriae: Nosocomial infection, regulation and biofilm formation. Future Microbiol. 2012, 7, 991–1002. [Google Scholar] [CrossRef] [PubMed]
- Stahlhut, S.G.; Tchesnokova, V.; Struve, C.; Weissman, S.J.; Chattopadhyay, S.; Yakovenko, O.; Aprikian, P.; Sokurenko, E.V.; Krogfelt, K.A. Comparative structure-function analysis of mannose-specific FimH adhesins from Klebsiella pneumoniae and Escherichia coli. J. Bacteriol 2009, 191, 6592–6601. [Google Scholar] [CrossRef] [PubMed]
- Stahlhut, S.G.; Chattopadhyay, S.; Struve, C.; Weissman, S.J.; Aprikian, P.; Libby, S.J.; Fang, F.C.; Krogfelt, K.A.; Sokurenko, E.V. Population variability of the FimH type 1 fimbrial adhesin in Klebsiella pneumoniae. J. Bacteriol. 2009, 191, 1941–1950. [Google Scholar] [CrossRef] [PubMed]
- Autenrieth, I.; Hantke, K.; Heesemann, J. Immunosuppression of the host and delivery of iron to the pathogen: A possible dual role of siderophores in the pathogenesis of microbial infections? Med. Microbiol. Immunol. 1991, 180, 135–141. [Google Scholar] [CrossRef]
- Mei, Y.F.; Liu, P.P.; Wan, L.G.; Liu, Y.; Wang, L.H.; Wei, D.D.; Deng, Q.; Cao, X.W. Virulence and Genomic Feature of a Virulent Klebsiella pneumoniae Sequence Type 14 Strain of Serotype K2 Harboring blaNDM-5 in China. Front. Microbiol. 2017, 8, 335. [Google Scholar] [CrossRef]
- Pomakova, D.K.; Hsiao, C.B.; Beanan, J.M.; Olson, R.; MacDonald, U.; Keynan, Y.; Russo, T.A. Clinical and phenotypic differences between classic and hypervirulent Klebsiella pneumoniae: An emerging and under-recognized pathogenic variant. Eur. J. Clin. Microbiol. Infect. Dis. 2012, 31, 981–989. [Google Scholar] [CrossRef]
- Lin, Y.C.; Lu, M.C.; Tang, H.L.; Liu, H.C.; Chen, C.H.; Liu, K.S.; Lin, C.; Chiou, C.S.; Chiang, M.K.; Chen, C.M.; et al. Assessment of hypermucoviscosity as a virulence factor for experimental Klebsiella pneumoniae infections: Comparative virulence analysis with hypermucoviscosity-negative strain. BMC Microbiol. 2011, 11, 50. [Google Scholar] [CrossRef] [PubMed]
- Turton, J.F.; Perry, C.; Elgohari, S.; Hampton, C.V. PCR characterization and typing of Klebsiella pneumoniae using capsular type-specific, variable number tandem repeat and virulence gene targets. J. Med. Microbiol. 2010, 59, 541–547. [Google Scholar] [CrossRef] [PubMed]
- Brisse, S.; Passet, V.; Haugaard, A.B.; Babosan, A.; Kassis-Chikhani, N.; Struve, C.; Decre, D. wzi Gene sequencing, a rapid method for determination of capsular type for Klebsiella strains. J. Clin. Microbiol. 2013, 51, 4073–4078. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.J.; Lin, T.L.; Lin, Y.T.; Su, P.A.; Chen, C.T.; Hsieh, P.F.; Hsu, C.R.; Chen, C.C.; Hsieh, Y.C.; Wang, J.T. Identification of capsular types in carbapenem-resistant Klebsiella pneumoniae strains by WZC sequencing and implications for capsule depolymerase treatment. Antimicrob Agents Chemother. 2015, 59, 1038–1047. [Google Scholar] [CrossRef] [PubMed]
- Lawlor, M.S.; O’Connor, C.; Miller, V.L. Yersiniabactin is a virulence factor for Klebsiella pneumoniae during pulmonary infection. Infect. Immun. 2007, 75, 1463–1472. [Google Scholar] [CrossRef]
- Koczura, R.; Kaznowski, A. Occurrence of the Yersinia high-pathogenicity island and iron uptake systems in clinical isolates of Klebsiella pneumoniae. Microb. Pathog. 2003, 35, 197–202. [Google Scholar] [CrossRef]
- Bowers, J.R.; Kitchel, B.; Driebe, E.M.; MacCannell, D.R.; Roe, C.; Lemmer, D.; de Man, T.; Rasheed, J.K.; Engelthaler, D.M.; Keim, P.; et al. Genomic Analysis of the Emergence and Rapid Global Dissemination of the Clonal Group 258 Klebsiella pneumoniae Pandemic. PLoS ONE 2015, 10, e0133727. [Google Scholar] [CrossRef] [PubMed]
- Matheeussen, V.; Xavier, B.B.; Mermans, I.; De Weerdt, A.; Lammens, C.; Goossens, H.; Jansens, H.; Malhotra-Kumar, S. Emergence of colistin resistance during treatment of recurrent pneumonia caused by carbapenemase producing Klebsiella pneumoniae. Diagn. Microbiol. Infect. Dis. 2019. [Google Scholar] [CrossRef]
- Lam, M.M.C.; Wyres, K.L.; Wick, R.R.; Judd, L.M.; Fostervold, A.; Holt, K.E.; Lohr, I.H. Convergence of virulence and MDR in a single plasmid vector in MDR Klebsiella pneumoniae ST15. J. Antimicrob. Chemother. 2019. [Google Scholar] [CrossRef] [PubMed]
- De Man, T.J.B.; Lutgring, J.D.; Lonsway, D.R.; Anderson, K.F.; Kiehlbauch, J.A.; Chen, L.; Walters, M.S.; Sjolund-Karlsson, M.; Rasheed, J.K.; Kallen, A.; et al. Genomic Analysis of a Pan-Resistant Isolate of Klebsiella pneumoniae, United States 2016. MBio 2018, 9. [Google Scholar] [CrossRef]
| Isolate ID | Date of Isolation | β-Lactamase Produced | Antibiotics Resistance Profile | Virulence Profile | PBRT | MLST | |
|---|---|---|---|---|---|---|---|
| CA-UTI isolates | 201-010 | 2010 | TEM-156 | CIP, LVF | khe, mrkDV1 | FIA | NP |
| 201-075 | 2010 | SHV-11 | CIP, FOS | mrkDV2-4 | FIA, X, Y | NP | |
| 201-076 | 2010 | TEM-24 | LVF, FOS | No VG | X, A/C | NP | |
| 201-094 | 2010 | TEM-156 | AMC, FOS | mrkDV1 | W | NP | |
| 208-309 | 2010 | SHV-33 | NR | fimH, mrkDV2-4, iucC | FIA, Y | NP | |
| 212-193 | 2010 | CTX-M-15 | AMC, CAZ, CTX, CIP, LVF | khe, iucC | FIA, HI2 | ST15 | |
| HA-UTI isolates | 625 | 1980 | TEM-10 | GM | K2, fimH, khe, mrkDV1 | NT | NP |
| 683 | 1999 | TEM-10 | AMC, CTX, CAZ, GM | K2, fimH, khe, mrkDV1 | NT | NP | |
| 684 | 1999 | TEM-10 | AMC, CAZ, GM | K2, fimH, khe, mrkDV1 | NT | NP | |
| 712 | 1999 | TEM-24 | AMC, CTX, CAZ | fimH, khe, mrkDV1 | NT | NP | |
| 721 | 1999 | TEM-10 | AMC, CTX, CAZ, GM | fimH, khe, mrkDV1 | NT | NP | |
| 732 | 2000 | TEM-10 | AMC, CTX, CAZ, GM | fimH, khe, mrkDV1 | NT | NP | |
| 749 | 2001 | TEM-24 | AMC, FOX, CTX, CAZ, CIP | fimH, khe | NT | NP | |
| 770 | 2001 | TEM-24 | AMC, FOX, CTX, CAZ, CIP | fimH, khe | NT | NP | |
| 775 | 2001 | TEM-10, CTX-M-15 | CAZ, GM | fimH, khe, mrkDV1 | HI1 | ST15 | |
| 847 | 2003 | CTX-M-15 | AMC, CTX, CAZ, GM, CIP | fimH, khe, mrkDV1 | HI1 | ST15 | |
| 931 | 2004 | CTX-M-15 | CTX, CAZ, GM, CIP | fimH, khe, mrkDV1 | HI1 | ST15 | |
| 1119 | 2005 | CTX-M-15 | CTX, CAZ, GM, CIP | fimH, khe, mrkDV1 | HI1 | ST15 | |
| 1263 | 2007 | CTX-M-15 | GM, CIP | fimH, khe, mrkDV1 | HI1 | ST15 | |
| 2323 | 2008 | CTX-M-15 | AMC, CTX, CAZ, GM | fimH, khe, mrkDV1 | HI1 | ST15 | |
| 2325 | 2008 | CTX-M-15 | AMC, CTX, CAZ, GM | fimH, mrkDV1 | HI1 | ST15 | |
| 2386 | 2008 | CTX-M-15 | CTX, CAZ, GM | fimH, khe, mrkDV1 | HI1 | ST15 | |
| 2394 | 2008 | CTX-M-15 | CTX, CAZ, GM | fimH, khe, mrkDV1 | HI1 | ST15 | |
| 2398 | 2008 | CTX-M-15 | CTX, CAZ, GM, CIP | fimH, mrkDV1 | HI1 | ST15 | |
| 2400 | 2008 | CTX-M-15 | CTX, CAZ, GM | fimH, khe, mrkDV1 | HI1 | ST15 | |
| 2414 | 2008 | CTX-M-15 | CTX, CAZ, GM, CIP | fimH, khe, mrkDV1 | HI1 | ST15 | |
| 2909 | 2009 | KPC-3, SHV-11 | AMC, CTX, CAZ, IMP, GM, MEM, ERT | K2, fimH, khe, mrkDV1 | F | ST14 | |
| 2954 | 2010 | KPC-3 | AMC, FOX CTX, CAZ, IMP, GM, LVF, TIG, MEM, ERT | K2, fimH, khe, mrkDV1 | A/C, F | ST14 | |
| 3108 | 2010 | KPC-3 | AMC, CTX, CAZ, TIG, MEM, ERT | khe, mrkDV1, iucC | FIA, F | ST14 | |
| 29078 | 2010 | KPC-3, CTX-M-15 | AMC, FOX, CTX, CAZ, IMP, GM, MEM, ERT, FOS | fimH, khe, mrkDV1,iucC | FIA, F | ST14 | |
| 43201 | 2010 | KPC-3, CTX-M-15 | AMC, FOX, CTX, CAZ, IMP, GM, CIP, LVF, MEM, ERT | K2, fimH, khe, mrkDV1,iucC | A/C | ST14 | |
| 61095 | 2011 | KPC-3, CTX-M-15 | AMC, CTX, CAZ, IMP, GM, CIP, LVF, MEM, ERT, TIG | K2, fimH, mrkDV1 | A/C, F | ST14 | |
| 72562 | 2011 | KPC-3 | AMC, FOX, CTX, CAZ, MEM, ERT | fimH, mrkDV1,iucC | A/C | ST14 | |
| 87582 | 2012 | KPC-3 | AMC, FOX, CTX, CAZ, IMP, GM, CIP, LVF, MEM, ERT, FOS | fimH, khe, mrkDV1,iucC | F | ST14 | |
| 91488 | 2012 | KPC-3, CTX-M-15 | CTX, CAZ, GM, CIP, LVF, MEM, ERT | K2, fimH, khe, mrkDV1 | A/C, F | ST14 | |
| 22073 | 2013 | KPC-3 | AMC, FOX, CTX, CAZ, IMP, GM, CIP, LVF, MEM, ERT | K2, fimH, khe, mrkDV1 | A/C | ST14 | |
| 31149 | 2013 | KPC-3 | AMC, CTX, CAZ, IMP, GM, MEM, ERT | K2, fimH, mrkDV1,iucC | A/C, F | ST14 |
| Virulence Factor | Target Gene | Nr. of CA-UTI Isolates (%) | Nr. of HA-UTI Isolates (%) | Total (%) | |
|---|---|---|---|---|---|
| (n = 50) | (n = 31) | (n = 81) | |||
| Fimbrial adhesins | Type 1 | fimH | 20 (40.0) | 31 (100.0) * | 51 (63.0) |
| Type 3, variant 1 | mrkDV1 | 10 (20.0) | 30 (96.8) * | 40 (49.3) | |
| Type 3, variant 2-4 | mrkDV2-4 | 31 (62.0) | 0 * | 31 (38.2) | |
| Toxin | Haemolysin | khe | 23 (46.0) | 26 (83.9) * | 49 (60.5) |
| Capsular type | K2 serotype | K2A | 3 (6.0) | 10 (32.2) * | 13 (16.0) |
| Siderophore | Aerobactin | iucC | 10 (20.0) | 5 (16.1) | 15 (18.5) |
| Protectins or invasins | Mucoviscosity phenotype | magA | 0 | 0 | 0 |
| Regulator of mucoid phenotype | rmpA | 0 | 0 | 0 | |
| Nr. of Virulence Genes | Virulence Profile (VP) | Virulence Genes | Nr. Isolates (%) | ||
|---|---|---|---|---|---|
| CA-UTI n = 50 (100) | HA-UTI n = 31 (100) | Total n = 81 (100) | |||
| 0 VF | 1 | No VF | 8 (16.0) | 0 | 8 (9.9) |
| 1 VF | 2 | khe | 5 (10.0) | 0 | 5 (6.2) |
| 3 | fimH | 1 (2.0) | 0 | 1 (1.2) | |
| 4 | mrkDV1 | 2 (4.0) | 0 | 2 (2.5) | |
| 5 | mrkDV2-4 | 3 (6.0) | 0 | 3 (3.7) | |
| 2 VF | 6 | fimH, khe | 2 (4.0) | 2 (6.3) | 4 (4.9) |
| 7 | fimH, mrkDV1 | 0 | 3 (9.4) | 3 (3.7) | |
| 8 | khe, iucC | 1 (2.0) | 0 | 1 (1.2) | |
| 9 | iucC, mrkDV2-4 | 2 (4.0) | 0 | 2 (2.5) | |
| 10 | khe, mrkDV1 | 2 (4.0) | 0 | 2 (2.5) | |
| 11 | khe, mrkDV2-4 | 5 (10.0) | 0 | 5 (6.2) | |
| 12 | fimH, mrkDV2-4 | 5 (10.0) | 0 | 5 (6.2) | |
| 13 | fimH, khe, mrkDV1 | 2 (4.0) | 13 (40.6) | 15 (18.5) | |
| 14 | fimH, khe, iucC | 1 (2.0) | 0 | 1 (1.2) | |
| 15 | fimH, khe, mrkDV2-4 | 2 (4.0) | 0 | 2 (2.5) | |
| 3 VF | 16 | fimH, iucC, mrkDV1 | 2 (4.0) | 1 (3.1) | 3 (3.7) |
| 17 | fimH, iucC, mrkDV2-4 | 4 (8.0) | 0 | 4 (4.9) | |
| 18 | khe, mrkDV1, iucC | 0 | 1 (3.1) | 1 (1.2) | |
| 19 | K2, khe, mrkDV1 | 2 (4.0) | 0 | 2 (2.5) | |
| 20 | K2, fimH, mrkDV1 | 0 | 1 (3.1) | 1 (1.2) | |
| 21 | K2, fimH, khe, mrkDV2-4 | 1 (2.0) | 0 | 1 (1.2) | |
| 4 VF | 22 | K2, fimH, khe, mrkDV1 | 0 | 7 (21.9) | 7 (8.6) |
| 23 | fimH, khe, mrkDV1, iucC | 0 | 2 (6.3) | 2 (2.5) | |
| 5 VF | 24 | K2, fimH, khe, mrkDV1, iucC | 0 | 1 (3.1) | 1 (1.2) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Caneiras, C.; Lito, L.; Melo-Cristino, J.; Duarte, A. Community- and Hospital-Acquired Klebsiella pneumoniae Urinary Tract Infections in Portugal: Virulence and Antibiotic Resistance. Microorganisms 2019, 7, 138. https://doi.org/10.3390/microorganisms7050138
Caneiras C, Lito L, Melo-Cristino J, Duarte A. Community- and Hospital-Acquired Klebsiella pneumoniae Urinary Tract Infections in Portugal: Virulence and Antibiotic Resistance. Microorganisms. 2019; 7(5):138. https://doi.org/10.3390/microorganisms7050138
Chicago/Turabian StyleCaneiras, Cátia, Luis Lito, José Melo-Cristino, and Aida Duarte. 2019. "Community- and Hospital-Acquired Klebsiella pneumoniae Urinary Tract Infections in Portugal: Virulence and Antibiotic Resistance" Microorganisms 7, no. 5: 138. https://doi.org/10.3390/microorganisms7050138
APA StyleCaneiras, C., Lito, L., Melo-Cristino, J., & Duarte, A. (2019). Community- and Hospital-Acquired Klebsiella pneumoniae Urinary Tract Infections in Portugal: Virulence and Antibiotic Resistance. Microorganisms, 7(5), 138. https://doi.org/10.3390/microorganisms7050138





