Feeding Essential Oils to Neonatal Holstein Dairy Calves Results in Increased Ruminal Prevotellaceae Abundance and Propionate Concentrations
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. SCFA Analysis
2.3. Microbial DNA Isolation and PCR Amplification
2.4. Computational Analysis of PCR Generated 16S rRNA Amplicon Sequences
2.5. Computational Analysis for Alpha and Beta Diversity
2.6. Statistical Analyses
2.7. Accession Numbers for Next Generation Sequencing Data
3. Results
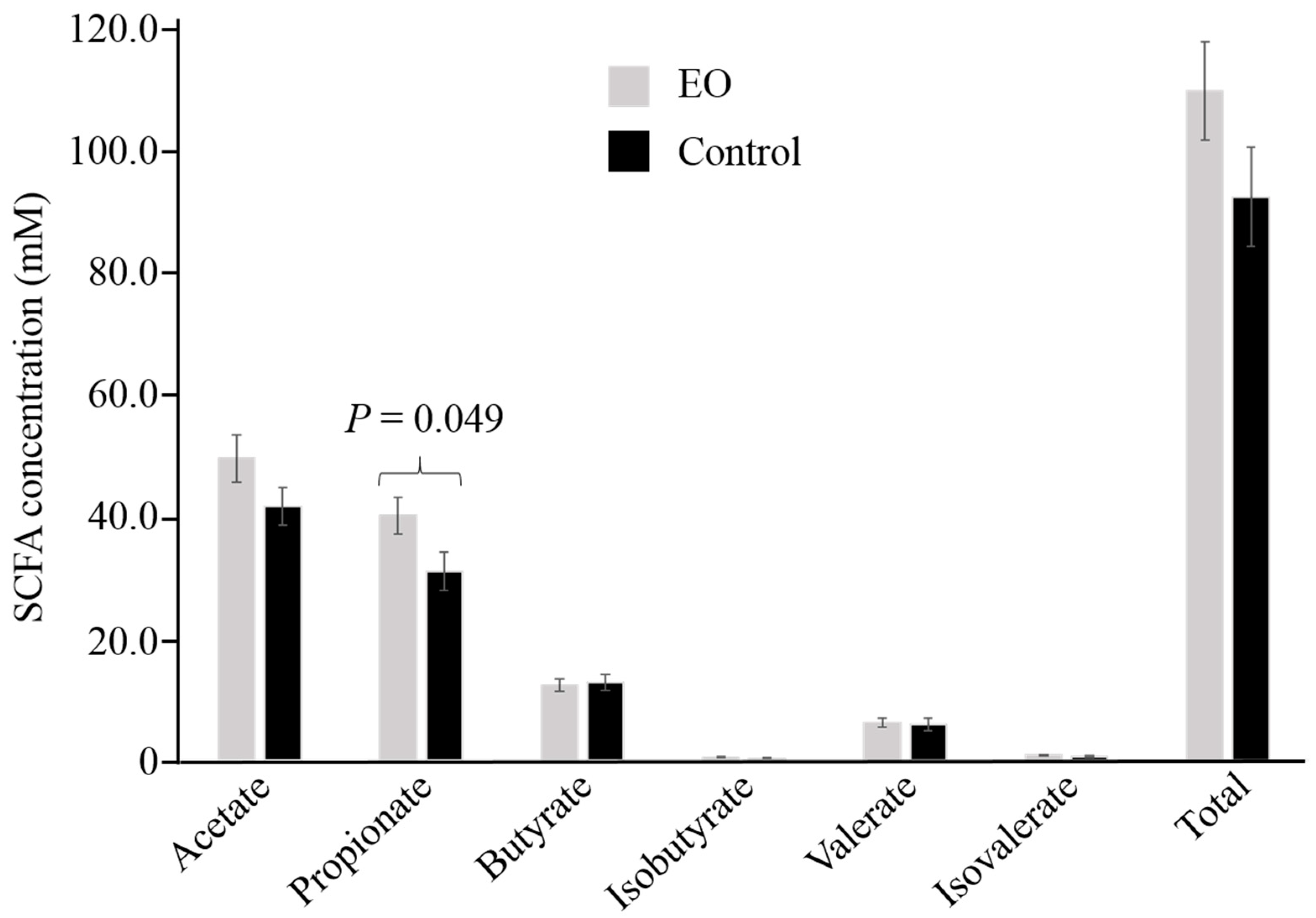
3.1. Comparative Analysis of Ruminal SCFA between EO Supplemented and Non-Supplemented Diets
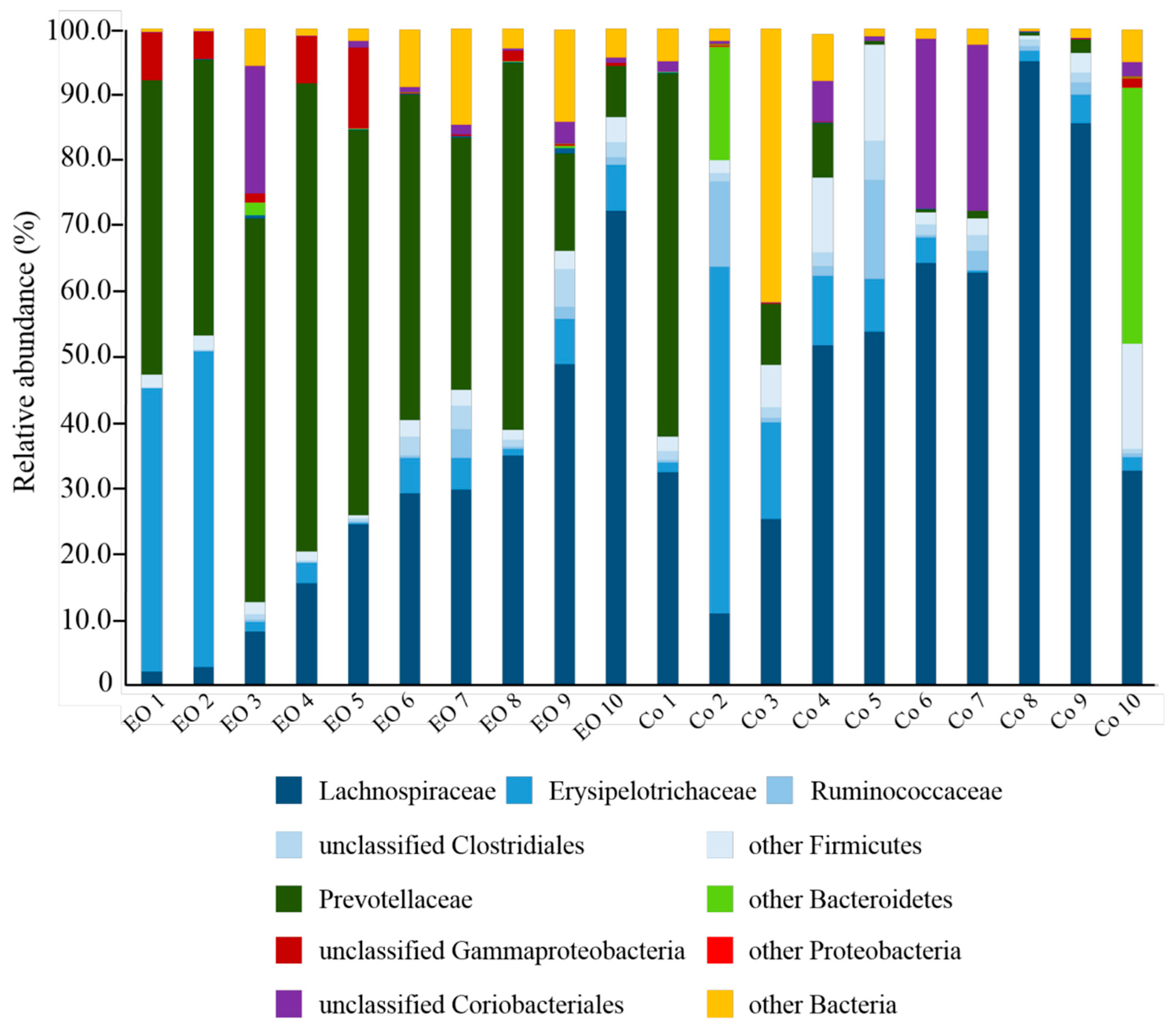
3.2. Effects of EO on the Taxonomic Composition of Ruminal Bacteria in Growing Calves
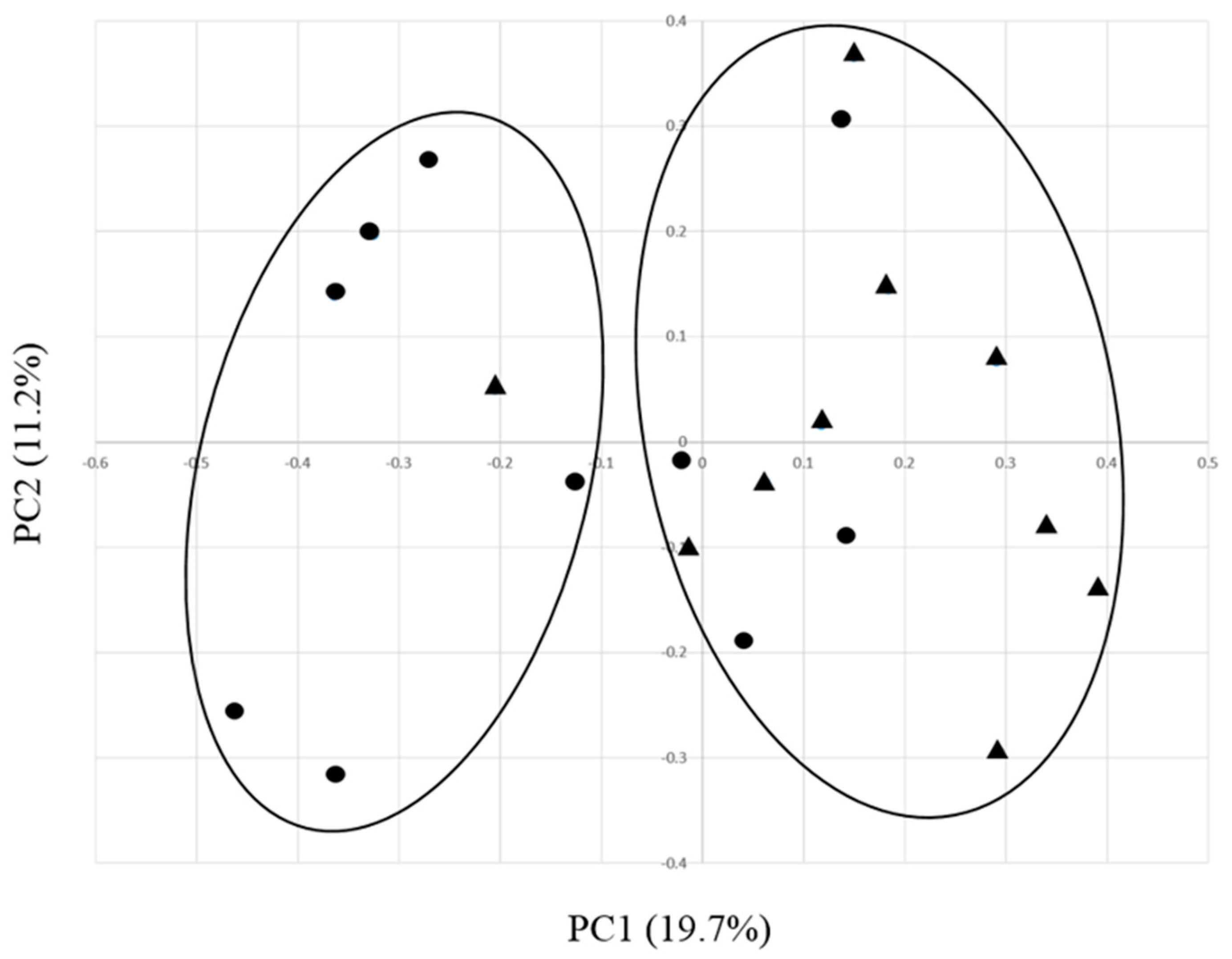
3.3. Effects of EO on the Ruminal Bacterial Community Structure in Growing Calves
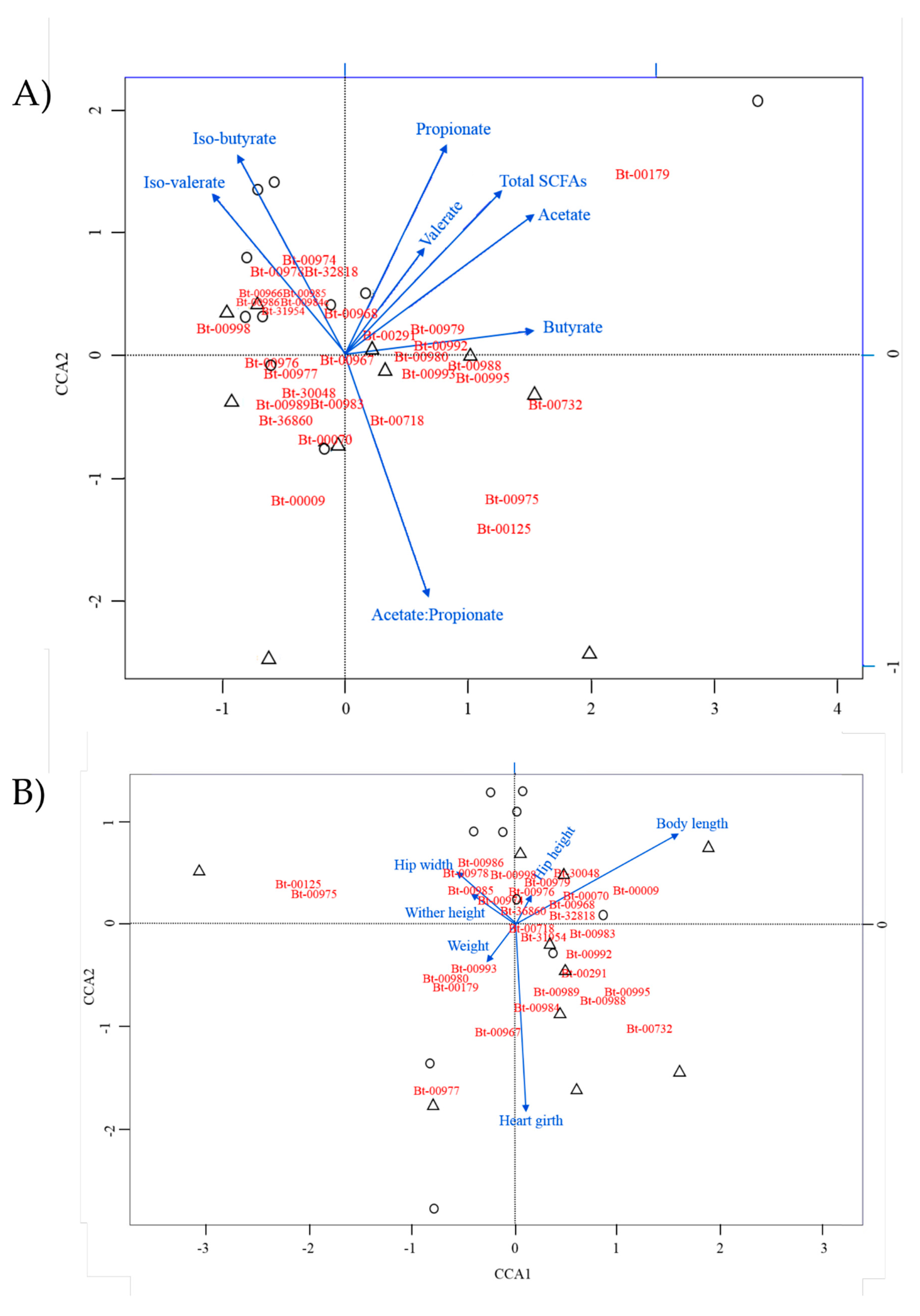
3.4. Identification of Potential Associations between Main OTUs and Ruminant Performance Parameters
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Gustafson, R.H.; Bowen, R.E. Antibiotic use in animal agriculture. J. Appl. Microbiol. 1997, 83, 531–541. [Google Scholar] [CrossRef] [PubMed]
- Allen, H.K.; Levine, U.Y.; Looft, T.; Bandrick, M.; Casey, T.A. Treatment, promotion, commotion: Antibiotic alternatives in food-producing animals. Trends Microbiol. 2013, 21, 114–119. [Google Scholar] [CrossRef] [PubMed]
- Cheng, G.; Hao, H.; Xie, S.; Wang, X.; Dai, M.; Huang, L.; Yuan, Z. Antibiotic alternatives: The substitution of antibiotics in animal husbandry? Front. Microbiol. 2014, 5, 217. [Google Scholar] [CrossRef] [PubMed]
- Patra, A.K.; Saxena, J. A new perspective on the use of plant secondary metabolites to inhibit methanogenesis in the rumen. Phytochem 2010, 71, 1198–1222. [Google Scholar] [CrossRef] [PubMed]
- Khiaosa-ard, R.; Zebeli, Q. Meta-analysis of the effects of essential oils and their bioactive compounds on rumen fermentation characteristics and feed efficiency in ruminants. J. Anim. Sci. 2013, 91, 1819–1830. [Google Scholar] [CrossRef]
- Meyer, N.F.; Erickson, G.E.; Klopfenstein, T.J.; Greenquist, M.A.; Luebbe, M.K.; Williams, P.; Engstrom, M.A. Effect of essential oils, tylosin, and monensin on finishing steer performance, carcass characteristics, liver abscesses, ruminal fermentation, and digestibility. J. Anim. Sci. 2009, 87, 2346–2354. [Google Scholar] [CrossRef]
- Tager, L.R.; Krause, K.M. Effects of essential oils on rumen fermentation, milk production, and feeding behavior in lactating dairy cows. J. Dairy Sci. 2011, 94, 2455–2464. [Google Scholar] [CrossRef] [PubMed]
- McIntosh, F.M.; Williams, P.; Losa, R.; Wallace, R.J.; Beeve, D.A.; Newbold, C.J. Effects of essential oils on ruminal microorganisms and their protein metabolism. Appl. Environ. Microbiol. 2003, 69, 5011–5014. [Google Scholar] [CrossRef] [PubMed]
- Calsamiglia, S.; Busquet, M.; Cardozo, P.W.; Castillejos, L.; Ferret, A. Invited review: Essential oils as modifiers of rumen microbial fermentation. J. Dairy Sci. 2007, 90, 2580–2595. [Google Scholar] [CrossRef] [PubMed]
- Hill, T.M.; Aldrich, J.M.; Schotterbeck, R.L.; Bateman, H.G. Apex plant botanicals for neonatal calf milk replacers and starters. Prof. Anim. Sci. 2007, 23, 521–526. [Google Scholar] [CrossRef]
- Jami, E.; Israel, A.; Kotser, A.; Mizrahi, I. Exploring the bovine rumen bacterial community from birth to adulthood. ISME J. 2013, 7, 1069–1079. [Google Scholar] [CrossRef] [PubMed]
- Malmuthuge, N.; Guan, L.L. Understanding host-microbial interactions in rumen: Searching the best opportunity for microbiota manipulation. J. Anim. Sci. Biotechnol. 2017, 8, 8. [Google Scholar] [CrossRef] [PubMed]
- Froehlich, K.A.; Abdelsalam, K.W.; Chase, C.; Koppien-Fox, J.; Casper, D.P. Evaluation of essential oils and prebiotics for newborn dairy calves. J. Anim. Sci. 2017, 95, 3772–3782. [Google Scholar] [CrossRef] [PubMed]
- Dion, C.; Chappuis, E.; Ripoll, C. Does larch arabinogalactan enhance immune function? A review of mechanistic and clinical trials. Nut. Metab. 2016, 13, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.; Morrison, M. Improved extraction of PCR-quality community DNA from digesta and fecal samples. Biotechniques 2004, 36, 808–812. [Google Scholar] [CrossRef] [PubMed]
- Edwards, U.; Rogall, T.; Blöcker, H.; Emde, M.; Böttger, E.C. Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res. 1989, 17, 7843–7853. [Google Scholar] [CrossRef] [PubMed]
- Lane, D.J.; Pace, B.; Olsen, G.J.; Stahl, D.A.; Sogin, M.L.; Pace, N.R. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc. Natl. Acad. Sci. USA 1985, 82, 6955–6959. [Google Scholar] [CrossRef] [PubMed]
- Opdahl, L.; Gonda, M.; St-Pierre, B. Identification of Uncultured Bacterial Species from Firmicutes, Bacteroidetes and CANDIDATUS Saccharibacteria as Candidate Cellulose Utilizers from the Rumen of Beef Cows. Microorganisms 2018, 6, 17. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef] [PubMed]
- Parte, A.C. LPSN—list of prokaryotic names with standing in nomenclature. Nucleic Acids Res. 2014, 42, D613–D616. [Google Scholar] [CrossRef] [PubMed]
- Yanez-Ruiz, D.R.; Abecia, L.; Newbold, C.J. Manipulating rumen microbiome and fermentation through interventions during early life: A review. Front Microbiol. 2015, 6, 1133. [Google Scholar] [CrossRef] [PubMed]
- Meale, S.J.; Chaucheyras-Durand, F.; Berends, H.; Guan, L.L.; Steele, M.A. From pre- to postweaning: Transformation of the young calf’s gastrointestinal tract. J. Dairy Sci. 2017, 100, 5984–5995. [Google Scholar] [CrossRef] [PubMed]
- Steele, M.A.; Penner, G.B.; Chaucheyras-Durand, F.; Guan, L.L. Development and physiology of the rumen and the lower gut: Targets for improving gut health. J. Dairy Sci. 2016, 99, 4955–4966. [Google Scholar] [CrossRef] [PubMed]
- Cosentino, S.; Tuberoso, C.I.G.; Pisano, B.; Satta, M.; Mascia, V.; Arzedi, E.; Palmas, F. In-vitro antimicrobial activity and chemical composition of Sardinian Thymus essential oils. Lett. Appl. Microbiol. 1999, 29, 130–135. [Google Scholar] [CrossRef]
- Benchaar, C.; Calsamiglia, S.; Chaves, A.V.; Fraser, G.R.; Colombatto, D.; McAllister, T.A.; Beauchemin, K.A. A review of plant-derived essential oils in ruminant nutrition and production. Anim. Feed Sci. Technol. 2008, 145, 209–228. [Google Scholar] [CrossRef]
- Patra, A.K.; Yu, Z. Effects of essential oils on methane production and fermentation by, and abundance and diversity of, rumen microbial populations. Appl. Environ. Microbiol. 2012, 78, 4271–4280. [Google Scholar] [CrossRef] [PubMed]
- Bergman, E.N. Energy contribution of volatile fatty acids from the gastrointestinal tract in various species. Physiol. Rev. 1990, 70, 567–590. [Google Scholar] [CrossRef]
- McLeod, K.R.; Baldwin, R.L. Effects of diet forage:concentrate ratio and metabolizable energy intake on isolated rumen epithelial cell metabolism in vitro. J. Anim. Sci. 2000, 78, 771–783. [Google Scholar] [CrossRef]
- Tamate, H.; McGilliard, A.D.; Jacobson, N.L.; Getty, R. Effect of various dietaries on the anatomical development of the stomach in the calf. J. Dairy Sci. 1962, 45, 408–420. [Google Scholar] [CrossRef]
- Gonzalez-Garcia, R.A.; McCubbin, T.; Navone, L.; Stowers, C.; Nielsen, L.; Marcellin, E. Microbial propionic acid production. Fermentation 2017, 3, 21. [Google Scholar] [CrossRef]
- Reichardt, N.; Duncan, S.H.; Young, P.; Belenguer, A.; McWilliam Leitch, C.; Scott, K.P.; Flint, H.J.; Louis, P. Phylogenetic distribution of three pathways for propionate production within the human gut microbiota. ISME J. 2014, 8, 1323–1335. [Google Scholar] [CrossRef] [PubMed]
- Russell, J.B.; Baldwin, R.L. Comparison of maintenance energy expenditures and growth yields among several rumen bacteria grown on continuous culture. Appl. Environ. Microbiol. 1979, 37, 537–543. [Google Scholar] [PubMed]
- Strobel, H.J. Vitamin B12-dependent propionate production by the ruminal bacterium Prevotella ruminicola 23. Appl. Environ. Microbiol. 1992, 58, 2331–2333. [Google Scholar] [PubMed]
- Russell, J.B.; Dombrowski, D.B. Effect of pH on the efficiency of growth by pure cultures of rumen bacteria in continuous culture. Appl. Environ. Microbiol. 1980, 39, 604–610. [Google Scholar] [PubMed]
- Dehority, B.A. Characterization of several bovine rumen bacteria isolated with a xylan medium. J. Bacteriol. 1966, 91, 1724–1729. [Google Scholar]
- Wallnofer, P.; Baldwin, R.L. Pathway of propionate formation in Bacteroides ruminicola. J. Bacteriol. 1967, 93, 504–505. [Google Scholar] [PubMed]
- Dias, J.; Marcondes, M.I.; Noronha, M.F.; Resende, R.T.; Machado, F.S.; Mantovani, H.C.; Dill-McFarland, K.A.; Suen, G. Effect of pre-weaning diet on the ruminal archaeal, bacterial, and fungal communities of dairy calves. Front. Microbiol. 2017, 8, 1553. [Google Scholar] [CrossRef] [PubMed]
- Malmuthuge, N.; Griebel, P.J.; Guan, L.L. Taxonomic identification of commensal bacteria associated with the mucosa and digesta throughout the gastrointestinal tracts of preweaned calves. Appl. Environ. Microbiol. 2014, 80, 2021–2028. [Google Scholar] [CrossRef]
- Li, R.W.; Connor, E.E.; Li, C.; Baldwin Vi, R.L.; Sparks, M.E. Characterization of the rumen microbiota of pre-ruminant calves using metagenomic tools. Environ. Microbiol. 2012, 14, 129–139. [Google Scholar] [CrossRef] [PubMed]
- Patra, A.K.; Yu, Z. Essential oils affect populations of some rumen bacteria in vitro as revealed by microarray (RumenBactArray) analysis. Front. Microbiol. 2015, 6, 297. [Google Scholar] [CrossRef] [PubMed]
- Benchaar, C.; Petit, H.V.; Berthiaume, R.; Ouellet, D.R.; Chiquette, J.; Chouinard, P.Y. Effects of essential oils on digestion, ruminal fermentation, rumen microbial populations, milk production, and milk composition in dairy cows fed alfalfa silage or corn silage. J. Dairy Sci. 2007, 90, 886–897. [Google Scholar] [CrossRef]
- Patra, A.K. Effects of essential oils on rumen fermentation, microbial ecology and ruminant production. Asian J. Anim. Vet. Adv. 2011, 2011, 1. [Google Scholar] [CrossRef]
- Dorman, H.J.D.; Deans, S.G. Antimicrobial agents from plants: Antibacterial activity of plant volatile oils. J. Appl. Microbiol. 2000, 88, 308–316. [Google Scholar] [CrossRef] [PubMed]
- Burt, S. Essential oils: Their antibacterial properties and potential applications in foods—A review. Int J. Food Microbiol. 2004, 94, 223–253. [Google Scholar] [CrossRef] [PubMed]
- Lambert, R.J.W.; Skandamis, P.N.; Coote, P.J.; Nychas, G.-J.E. A study of the minimum inhibitory concentration and mode of action of oregano essential oil, thymol and carvacrol. J. Appl. Microbiol. 2001, 91, 453–462. [Google Scholar] [CrossRef] [PubMed]
- Ultee, A.; Bennik, M.H.J.; Moezelaar, R. The phenolic hydroxyl group of carvacrol is essential for action against the food-borne pathogen Bacillus cereus. Appl. Environ. Microbiol. 2002, 68, 1561–1568. [Google Scholar] [CrossRef]
- Braun, H.-S.; Schrapers, K.T.; Mahlkow-Nerge, K.; Stumpff, F.; Rosendahl, J. Dietary supplementation of essential oils in dairy cows: Evidence for stimulatory effects on nutrient absorption. Animal 2019, 13, 518–523. [Google Scholar] [CrossRef]
- Paster, B.J.; Canale-Parola, E. Physiological diversity of rumen spirochetes. Appl. Environ. Microbiol. 1982, 43, 686–693. [Google Scholar] [PubMed]
- Vince, A.J.; McNeil, N.I.; Wager, J.D.; Wrong, O.M. The effect of lactulose, pectin, arabinogalactan and cellulose on the production of organic acids and metabolism of ammonia by intestinal bacteria in a faecal incubation system. Br. J. Nutr. 1990, 60, 17–26. [Google Scholar] [CrossRef]
| Taxonomic Affiliation | EO | Control | P-Value |
|---|---|---|---|
| Firmicutes | 43.68 ± 6.92 | 73.22 ± 6.79 | 0.0069 * |
| Lachnospiraceae | 26.87 ± 6.92 | 51.53 ± 8.44 | 0.0364 * |
| Erysipelotrichaceae | 12.11 ± 5.65 | 9.99 ± 4.97 | 0.7812 |
| Ruminococcaceae | 0.91 ± 0.42 | 3.71 ± 1.74 | 0.1349 |
| unclassified Clostridiales | 1.72 ± 0.59 | 1.94 ± 0.48 | 0.7802 |
| Other Firmicutes | 2.07 ± 0.28 | 6.04 ± 1.82 | 0.0453 * |
| Bacteroidetes | 44.63 ± 6.28 | 13.45 ± 6.02 | 0.0021 * |
| Prevotellaceae | 44.20 ± 6.27 | 9.70 ± 5.94 | 0.0009 * |
| unclassified Bacteroidales | 0.18 ± 0.07 | 0.06 ± 0.02 | 0.1359 |
| Other Bacteroidetes | 0.26 ± 0.20 | 5.64 ± 4.08 | 0.2048 |
| Proteobacteria | 3.51 ± 1.32 | 0.25 ± 0.17 | 0.0246 * |
| unclassified Gammaproteobacteria | 3.49 ± 1.32 | 0.17 ± 0.13 | 0.0222 * |
| Other Proteobacteria | 0.02 ± 0.01 | 0.08 ± 0.05 | 0.2111 |
| Actinobacteria | 2.77 ± 1.88 | 6.37 ± 3.28 | 0.3531 |
| Coriobacteriales | 2.75 ± 1.88 | 6.27 ± 3.28 | 0.3638 |
| Other Bacteria | 5.41 ± 1.70 | 6.72 ± 3.94 | 0.7643 |
| Index | EO | Control | P-Value |
|---|---|---|---|
| Chao1 | 484 ± 48 | 543 ± 80 | 0.5375 |
| OTUs * | 206 ± 19 | 219 ± 25 | 0.6760 |
| Shannon | 3.18 ± 0.25 | 3.27 ± 0.24 | 0.8052 |
| Simpson | 0.16 ± 0.04 | 0.14 ± 0.03 | 0.7518 |
| Coverage (%) | 91.5 ± 0.8 | 90.7 ± 1.2 | 0.5723 |
| OTUs | EO | Control | P-Value | Closest Valid Taxon (id%) |
|---|---|---|---|---|
| Bacteriodetes | ||||
| SD_Bt-00966 a | 19.51 ± 5.32 | 2.70 ± 1.80 | 0.008 | P. ruminicola (97%) |
| SD_Bt-00976 a | 4.74 ± 1.32 | 8.01 ± 5.02 | 0.536 | P. ruminicola (90%) |
| SD_Bt-00979 a | 2.35 ± 2.10 | 0.02 ± 0.01 | 0.281 | P. salivae (89%) |
| SD_Bt-00985 a | 1.92 ± 1.02 | 0.18 ± 0.08 | 0.105 | P. salivae (89%) |
| SD_Bt-00986 a | 0.91 ± 0.34 | 0.22 ± 0.13 | 0.080 | P. ruminicola (95%) |
| SD_Bt-32818 a | 1.11 ± 0.79 | 0.08 ± 0.05 | 0.212 | P. multisaccharivorax (93%) |
| Total | 30.55 | 11.22 | ||
| Proteobacteria | ||||
| SD_Bt-00978 b | 3.44 ± 1.30 | 0.17 ± 0.13 | 0.022 | Haemophilus influenzae (84%) |
| Actinobacteria | ||||
| SD_Bt-00967 | 1.96 ± 1.37 | 4.98 ± 2.69 | 0.331 | Olsenella umbonata (99%) |
| OTUs | EO | Control | P-Value | Closest Valid Taxon (id%) |
|---|---|---|---|---|
| SD_Bt-00009 a | 4.15 ± 1.47 | 7.57 ± 5.85 | 0.577 | Butyrivibrio hungatei (91%) |
| SD_Bt-00070 a | 1.74 ± 0.54 | 1.71 ± 0.97 | 0.982 | Clostridium aminophilum (91%) |
| SD_Bt-00179 a | 3.54 ± 3.26 | 0.26 ± 0.17 | 0.329 | Lachnospira pectinoschiza (89%) |
| SD_Bt-00291 a | 0.96 ± 0.55 | 2.60 ± 1.34 | 0.271 | Coprococcus catus (90%) |
| SD_Bt-00718 a | 0.92 ± 0.34 | 1.41 ± 0.45 | 0.400 | Eisenbergiella tayi (92%) |
| SD_Bt-00968 a | 0.84 ± 0.38 | 2.17 ± 1.46 | 0.389 | Butyrivibrio fibrisolvens (90%) |
| SD_Bt-00977 a | 4.51 ± 3.06 | 0.64 ± 0.43 | 0.227 | Butyrivibrio fibrisolvens (91%) |
| SD_Bt-00980 a | 0.52 ± 0.23 | 6.56 ± 4.38 | 0.185 | Butyrivibrio fibrisolvens (89%) |
| SD_Bt-00983 a | 0.05 ± 0.03 | 3.76 ± 1.95 | 0.073 | Butyrivibrio fibrisolvens (91%) |
| SD_Bt-00988 a | 0.48 ± 0.19 | 1.64 ± 0.85 | 0.198 | Lachnospira multipara (91%) |
| SD_Bt-00993 a | 0.27 ± 0.15 | 1.00 ± 0.63 | 0.269 | Clostridium bolteae (87%) |
| SD_Bt-00998 a | 0.15 ± 0.08 | 1.50 ± 1.39 | 0.347 | Clostridium lavalense (90%) |
| SD_Bt-30048 a | 1.08 ± 0.46 | 1.40 ± 0.71 | 0.718 | Butyrivibrio fibrisolvens (91%) |
| SD_Bt-31954 a | 0.50 ± 0.24 | 1.87 ± 0.94 | 0.176 | Butyrivibrio fibrisolvens (90%) |
| SD_Bt-00974 b | 8.85 ± 5.45 | 1.14 ± 0.55 | 0.176 | Kandleria vitulina (89%) |
| SD_Bt-00975 b | 0.48 ± 0.35 | 5.16 ± 5.00 | 0.363 | Catenibacterium mitsuokai (88%) |
| SD_Bt-00989 b | 0.53 ± 0.29 | 1.71 ± 0.89 | 0.225 | Eubacterium cylindroides (92%) |
| SD_Bt-00992 b | 0.62 ± 0.29 | 1.01 ± 0.50 | 0.509 | Solobacterium moorei (91%) |
| SD_Bt-00125 c | 0.03 ± 0.01 | 1.25 ± 1.21 | 0.324 | Ruminococcus albus (90%) |
| SD_Bt-00995 c | 0.63 ± 0.33 | 1.80 ± 1.29 | 0.390 | Ruminococcus albus (86%) |
| SD_Bt-00732 d | 0.08 ± 0.05 | 1.11 ± 1.05 | 0.338 | Mogibacterium pumilum (92%) |
| SD_Bt-00984 d | 1.25 ± 0.59 | 1.55 ± 0.98 | 0.797 | Syntrophococcus sucromutans (91%) |
| SD_Bt-36860 e | 0.12 ± 0.06 | 1.72 ± 1.00 | 0.129 | Dialister succinatiphilus (99%) |
| Total | 32.94 | 50.63 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Poudel, P.; Froehlich, K.; Casper, D.P.; St-Pierre, B. Feeding Essential Oils to Neonatal Holstein Dairy Calves Results in Increased Ruminal Prevotellaceae Abundance and Propionate Concentrations. Microorganisms 2019, 7, 120. https://doi.org/10.3390/microorganisms7050120
Poudel P, Froehlich K, Casper DP, St-Pierre B. Feeding Essential Oils to Neonatal Holstein Dairy Calves Results in Increased Ruminal Prevotellaceae Abundance and Propionate Concentrations. Microorganisms. 2019; 7(5):120. https://doi.org/10.3390/microorganisms7050120
Chicago/Turabian StylePoudel, Prakash, Kelly Froehlich, David Paul Casper, and Benoit St-Pierre. 2019. "Feeding Essential Oils to Neonatal Holstein Dairy Calves Results in Increased Ruminal Prevotellaceae Abundance and Propionate Concentrations" Microorganisms 7, no. 5: 120. https://doi.org/10.3390/microorganisms7050120
APA StylePoudel, P., Froehlich, K., Casper, D. P., & St-Pierre, B. (2019). Feeding Essential Oils to Neonatal Holstein Dairy Calves Results in Increased Ruminal Prevotellaceae Abundance and Propionate Concentrations. Microorganisms, 7(5), 120. https://doi.org/10.3390/microorganisms7050120






