Broad Environmental Tolerance for a Salicola Host-Phage Pair Isolated from the Cargill Solar Saltworks, Newark, CA, USA
Abstract
1. Introduction
2. Materials and Methods
2.1. Field Sampling
2.2. Host Cultivation and Isolation
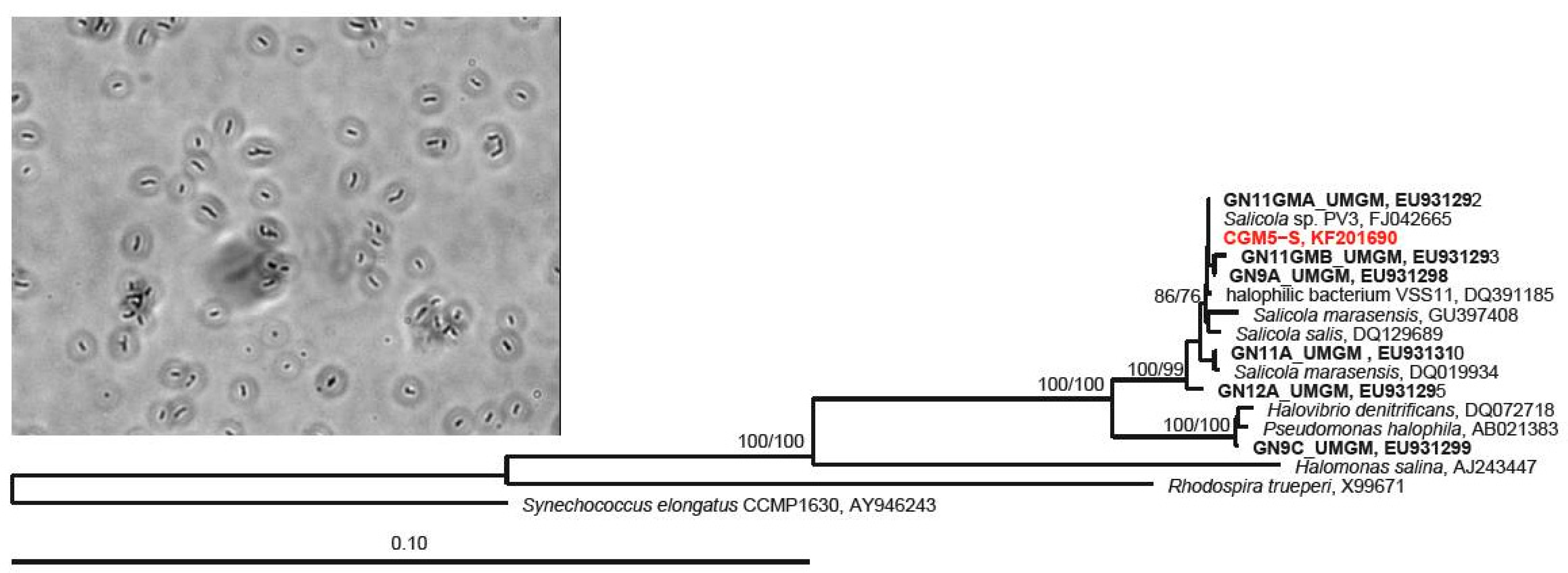
2.3. 16S rRNA Gene Host Identification and Phylogenetic Analysis
2.4. Physicochemical Tolerance Experiments
2.5. Microscopic and Biochemical Host Characterization
2.6. Phage Isolation and Enumeration
2.7. Generating Large-Scale Viral Stocks
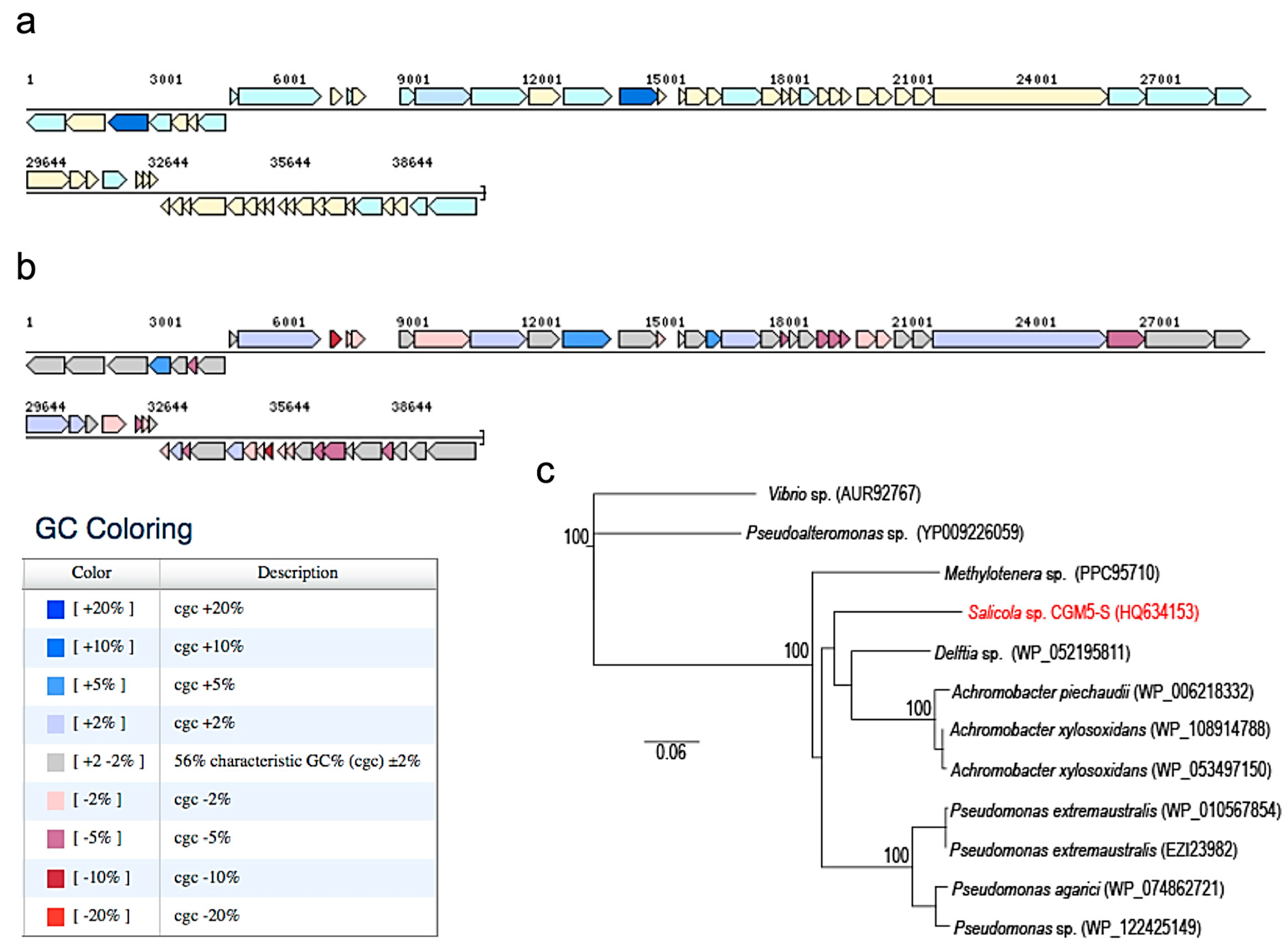
2.8. CGφ29 Genome
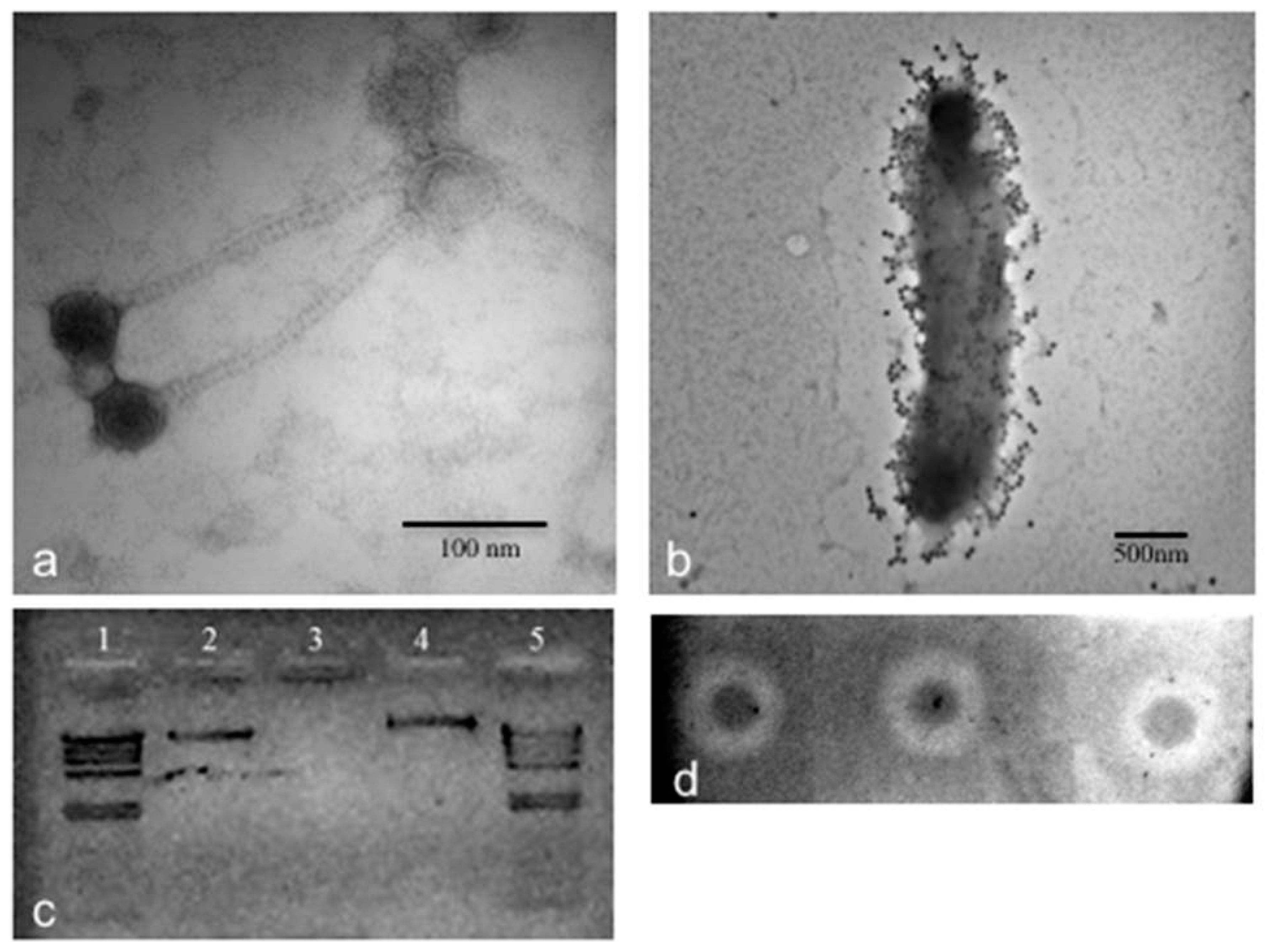
2.9. TEM of CGφ29 Phage
2.10. CGφ29 Host Range
2.11. Chloroform Sensitivity and Environmental Tolerance of CGφ29 Phage
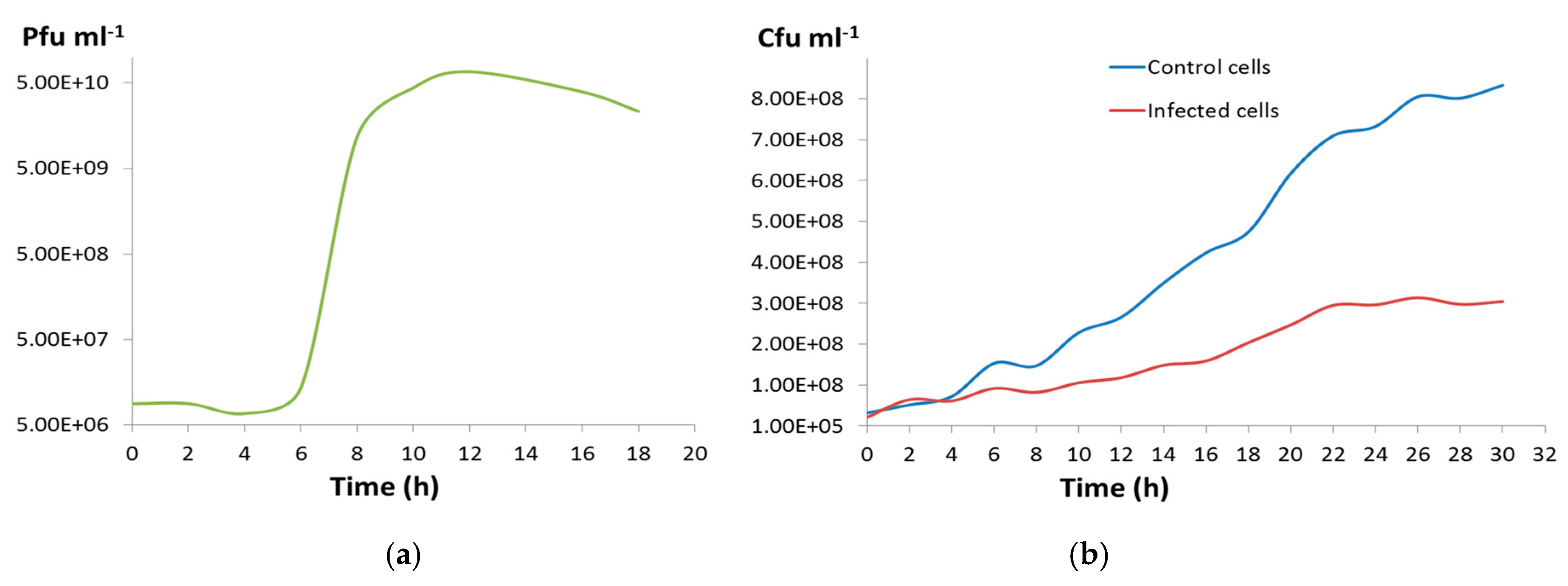
2.12. One-Step Growth Curve
2.13. Statistical Analysis
3. Results and Discussion
3.1. Host Characterization
3.2. Phage CGφ29 Characterization
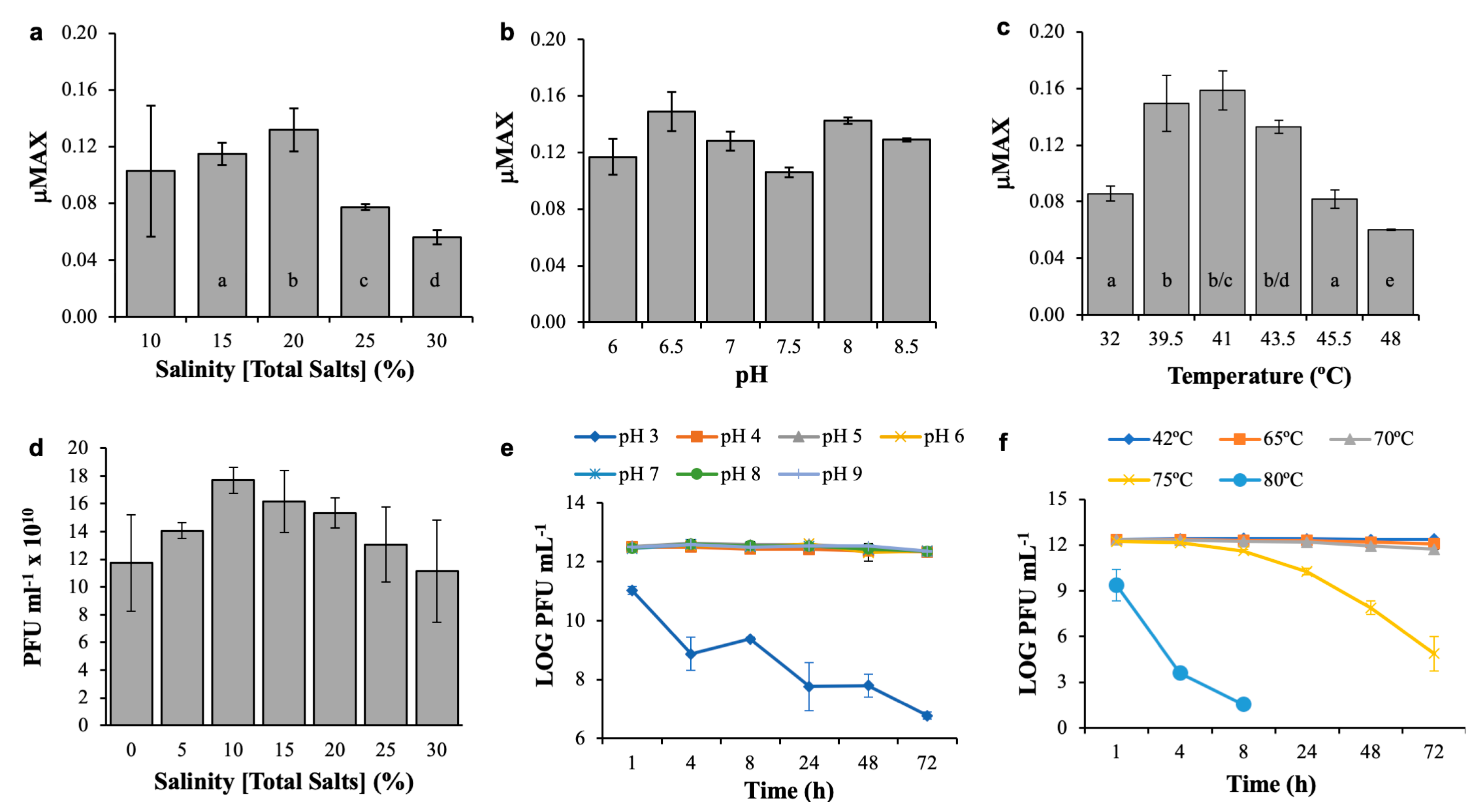
3.3. Host and Phage Are Both Broadly Tolerant of Environmental Conditions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Acheson, N. Fundamentals of Molecular Virology; John Wiley and Sons: Hoboken, NJ, USA, 2007. [Google Scholar]
- Flint, S.J.; Enquist, L.W.; Racaniello, V.R.; Skalka, A.M. Principles of Virology v.1 Molecular Biology; ASM Press: Washington, DC, USA, 2009; Volume 1, p. 569. [Google Scholar]
- Suttle, C.A. Viruses in the sea. Nature 2005, 437, 356–361. [Google Scholar] [CrossRef] [PubMed]
- Van Hannen, E.J.; Zwart, G.; van Agterveld, M.P.; Gons, H.J.; Ebert, J.; Laanbroek, H.J. Changes in bacterial and eukaryotic community structure after mass lysis of filamentous cyanobacteria associated with viruses. Appl. Environ. Microbiol. 1999, 65, 795–801. [Google Scholar] [PubMed]
- Middelboe, M. Bacterial growth rate and marine virus-host dynamics. Microb. Ecol. 2000, 40, 114–124. [Google Scholar] [CrossRef] [PubMed]
- Middelboe, M.; Hagstrom, A.; Blackburn, N.; Sinn, B.; Fischer, U.; Borch, N.H.; Pinhassi, J.; Simu, K.; Lorenz, M.G. Effects of bacteriophages on the population dynamics of four strains of pelagic marine bacteria. Microb. Ecol. 2001, 42, 395–406. [Google Scholar] [CrossRef] [PubMed]
- Weinbauer, M.G.; Rassoulzadegan, F. Are viruses driving microbial diversification and diversity? Environ. Microbiol. 2004, 6, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Wommack, K.E.; Colwell, R.R. Virioplankton: Viruses in aquatic ecosystems. Microbiol. Mol. Biol. Rev. 2000, 64, 69–114. [Google Scholar] [CrossRef] [PubMed]
- Mojica, K.D.; Brussaard, C.P. Factors affecting virus dynamics and microbial host–virus interactions in marine environments. FEMS Microbiol. Ecol. 2014, 89, 495–515. [Google Scholar] [CrossRef] [PubMed]
- Fuhrman, J.A.; Schwalbach, M. Viral influence on aquatic bacterial communities. Biol. Bull. 2003, 204, 192–195. [Google Scholar] [CrossRef]
- Calvo, C.; García de la Paz, A.; Bejar, V.; Quesada, E.; Ramos-Cormenzana, A. Isolation and characterization of phage F9-11 from a lysogenic Deleya halophila strain. Curr. Microbiol. 1988, 17, 49–53. [Google Scholar] [CrossRef]
- Kauri, T.; Ackermann, H.-W.; Goel, U.; Kushner, D. A bacteriophage of a moderately halophilic bacterium. Arch. Microbiol. 1991, 156, 435–438. [Google Scholar] [CrossRef]
- Uchida, K.; Kanbe, C. Occurrence of bacteriophages lytic for Pediococcus halophilus, a halophilic lactic-acid bacterium, in soy sauce fermentation. J. Gen. Appl. Microbiol. 1993, 39, 429–437. [Google Scholar] [CrossRef]
- Aalto, A.P.; Bitto, D.; Ravantti, J.J.; Bamford, D.H.; Huiskonen, J.T.; Oksanen, H.M. Snapshot of virus evolution in hypersaline environments from the characterization of a membrane-containing Salisaeta icosahedral phage 1. Proc. Natl. Acad. Sci. USA 2012, 109, 7079–7084. [Google Scholar] [CrossRef]
- Atanasova, N.S.; Roine, E.; Oren, A.; Bamford, D.H.; Oksanen, H.M. Global network of specific virus-host interactions in hypersaline environments. Environ. Microbiol. 2012, 14, 426–440. [Google Scholar] [CrossRef] [PubMed]
- Kristjansson, J.K.; Hreggvidsson, G.O. Ecology and habitats of extremophiles. World J. Microbiol. Biotechnol. 1995, 11, 17–25. [Google Scholar] [CrossRef]
- Grant, W.D.; Gemmell, R.T.; McGenity, T.J. Halobacteria: The evidence for longevity. Extremophiles 1998, 2, 279–287. [Google Scholar] [CrossRef] [PubMed]
- Javor, B.J. Hypersaline Environments; Springer: Berlin, Germany, 1989. [Google Scholar]
- Oren, A. Diversity of halophilic microorganisms: Environments, phylogeny, physiology, and applications. J. Ind. Microbiol. Biotechnol. 2002, 28, 56–63. [Google Scholar] [CrossRef]
- Ventosa, A.; Nieto, J.J. Biotechnological applications and potentialities of halophilic microorganisms. World J. Microbiol. Biotechnol. 1995, 11, 85–94. [Google Scholar] [CrossRef] [PubMed]
- Baati, H.; Guermazi, S.; Amdouni, R.; Gharsallah, N.; Sghir, A.; Ammar, E. Prokaryotic diversity of a Tunisian multipond solar saltern. Extremophiles 2008, 12, 505–518. [Google Scholar] [CrossRef] [PubMed]
- Benlloch, S.; Lopez-Lopez, A.; Casamayor, E.O.; Øvreås, L.; Goddard, V.; Daae, F.L.; Smerdon, G.; Massana, R.; Joint, I.; Thingstad, F.; et al. Prokaryotic genetic diversity throughout the salinity gradient of a coastal solar saltern. Environ. Microbiol. 2002, 4, 349–360. [Google Scholar] [CrossRef] [PubMed]
- Burns, D.G.; Camakaris, H.M.; Janssen, P.H.; Dyall-Smith, M.L. Combined use of cultivation-dependent and cultivation-independent methods indicates that members of most haloarchaeal groups in an Australian crystallizer pond are cultivable. Appl. Environ. Microbiol. 2004, 70, 5258–5265. [Google Scholar] [CrossRef]
- Javor, B.J. Industrial microbiology of solar salt production. J. Ind. Microbiol. Biotechnol. 2002, 28, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Maturrano, L.; Valens-Vadell, M.; Rossello-Mora, R.; Anton, J. Salicola marasensis gen. nov., sp nov., an extremely halophilic bacterium isolated from the Maras solar salterns in Peru. Int. J. Syst. Evol. Microbiol. 2006, 56, 1685–1691. [Google Scholar] [CrossRef] [PubMed]
- Oren, A. Halophilic Microorganisms and their Environments; Kluwer Academic: Boston, MA, USA, 2002; Volume 5, p. 575. [Google Scholar]
- Asker, D. Haloferax alexandrinus sp. nov., an extremely halophilic canthaxanthin-producing archaeon from a solar saltern in Alexandria (Egypt). Int. J. Syst. Evol. Microbiol. 2002, 52, 729–738. [Google Scholar] [CrossRef] [PubMed]
- Burns, D.G.; Camakaris, H.M.; Janssen, P.H.; Dyall-Smith, M.L. Cultivation of Walsby’s square haloarchaeon. FEMS Microbiol. Lett. 2004, 238, 469–473. [Google Scholar] [CrossRef] [PubMed]
- Papke, R.T.; Zhaxybayeva, O.; Feil, E.J.; Sommerfeld, K.; Muise, D.; Doolittle, W.F. Searching for species in haloarchaea. Proc. Natl. Acad. Sci. USA 2007, 104, 14092–14097. [Google Scholar] [CrossRef]
- Sabet, S.; Diallo, L.; Hays, L.; Jung, W.; Dillon, J. Characterization of halophiles isolated from solar salterns in Baja California, Mexico. Extremophiles 2009, 13, 643–656. [Google Scholar] [CrossRef]
- Antón, J.; Oren, A.; Benlloch, S.; Rodriguez-Valera, F.; Amann, R.; Rossello-Mora, R. Salinibacter ruber gen. nov., sp nov., a novel, extremely halophilic member of the Bacteria from saltern crystallizer ponds. Int. J. Syst. Evol. Microbiol. 2002, 52, 485–491. [Google Scholar] [CrossRef]
- De Lourdes Moreno, M.; Garcia, M.T.; Ventosa, A.; Mellado, E. Characterization of Salicola sp. IC10, a lipase- and protease-producing extreme halophile. FEMS Microbiol. Ecol. 2009, 68, 59–71. [Google Scholar] [CrossRef]
- Kindzierski, V.; Raschke, S.; Knabe, N.; Siedler, F.; Scheffer, B.; Pfluger-Grau, K.; Pfeiffer, F.; Oesterhelt, D.; Marin-Sanguino, A.; Kunte, H.J. Osmoregulation in the halophilic bacterium Halomonas elongata: A case study for integrative systems biology. PLoS ONE 2017, 12, e0168818. [Google Scholar] [CrossRef]
- Wais, A.C.; Daniels, L.L. Populations of bacteriophage infecting Halobacterium in a transient brine pool. FEMS Microbiol. Ecol. 1985, 31, 323–326. [Google Scholar] [CrossRef]
- Nuttall, S.D.; Dyall-Smith, M.L. Hf1 and Hf2-Novel Bacteriophages of Halophilic Archaea. Virology 1993, 197, 678–684. [Google Scholar] [CrossRef]
- Bath, C.; Dyall-Smith, M.L. His1, an archaeal virus of the Fuselloviridae family that infects Haloarcula hispanica. J. Virol. 1998, 72, 9392–9395. [Google Scholar]
- Guixa-Boixareu, N.; Calderon-Paz, J.I.; Heldal, M.; Bratbak, G.; Pedros-Alio, C. Viral lysis and bacterivory as prokaryotic loss factors along a salinity gradient. Aquat. Microb. Ecol. 1996, 11, 215–227. [Google Scholar] [CrossRef]
- Diez, B.; Anton, J.; Guixa-Boixereu, N.; Pedros-Alio, C.; Rodriguez-Valera, F. Pulsed-field gel electrophoresis analysis of virus assemblages present in a hypersaline environment. Int. Microbiol. 2000, 3, 159–164. [Google Scholar]
- Santos, F.; Meyerdierks, A.; Peña, A.; Rosselló-Mora, R.; Amann, R.; Antón, J. Metagenomic approach to the study of halophages: the environmental halophage 1. Environ. Microbiol. 2007, 9, 1711–1723. [Google Scholar] [CrossRef] [PubMed]
- Santos, F.; Moreno-Paz, M.; Meseguer, I.; Lopez, C.; Rosselló-Mora, R.; Parro, V.; Antón, J. Metatranscriptomic analysis of extremely halophilic viral communities. ISME J. 2011, 5, 1621–1633. [Google Scholar] [CrossRef]
- Santos, F.; Yarza, P.; Parro, V.; Briones, C.; Antón, J. The metavirome of a hypersaline environment. Environ. Microbiol. 2010, 12, 2965–2976. [Google Scholar] [CrossRef] [PubMed]
- Sime-Ngando, T.; Lucas, S.; Robin, A.; Tucker, K.P.; Colombet, J.; Bettarel, Y.; Desmond, E.; Gribaldo, S.; Forterre, P.; Breitbart, M.; et al. Diversity of virus-host systems in hypersaline Lake Retba, Senegal. Environ. Microbiol. 2011, 13, 1956–1972. [Google Scholar] [CrossRef]
- Boujelben, I.; Yarza, P.; Almansa, C.; Villamor, J.; Maalej, S.; Anton, J.; Santos, F. Virioplankton community structure in Tunisian solar salterns. Appl. Environ. Microbiol. 2012, 78, 7429–7437. [Google Scholar] [CrossRef]
- Bettarel, Y.; Bouvier, T.; Bouvier, C.; Carré, C.; Desnues, A.; Domaizon, I.; Jacquet, S.; Robin, A.; Sime-Ngando, T. Ecological traits of planktonic viruses and prokaryotes along a full-salinity gradient. FEMS Microbiol. Ecol. 2011, 76, 360–372. [Google Scholar] [CrossRef] [PubMed]
- Litchfield, C.D.; Irby, A.; Kis-Papo, T.; Oren, A. Comparisons of the polar lipid and pigment profiles of two solar salterns located in Newark, California, U.S.A., and Eilat, Israel. Extremophiles 2000, 4, 259–265. [Google Scholar] [CrossRef] [PubMed]
- Okeke, B.C.; Giblin, T.; Frankenberger, W.T. Reduction of perchlorate and nitrate by salt tolerant bacteria. Environ. Pollut. 2002, 118, 357–363. [Google Scholar] [CrossRef]
- Pesenti, P.T.; Sikaroodi, M.; Gillevet, P.M.; Sanchez-Porro, C.; Ventosa, A.; Litchfield, C.D. Halorubrum californiense sp nov., an extreme archaeal halophile isolated from a crystallizer pond at a solar salt plant in California, USA. Int. J. Syst. Evol. Microbiol. 2008, 58, 2710–2715. [Google Scholar] [CrossRef] [PubMed]
- Shirazian, P.; Asad, S.; Amoozegar, M.A. The potential of halophilic and halotolerant bacteria for the production of antineoplastic enzymes: L-asparaginase and L-glutaminase. EXCLI J. 2016, 15, 268–279. [Google Scholar] [CrossRef] [PubMed]
- De Lourdes Moreno, M.; Perez, D.; Garcia, M.T.; Mellado, E. Halophilic bacteria as a source of novel hydrolytic enzymes. Life 2013, 3, 38–51. [Google Scholar] [CrossRef]
- Kukkaro, P.; Bamford, D.H. Virus-host interactions in environments with a wide range of ionic strengths. Environ. Microbiol. Rep. 2009, 1, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Kivela, H.M.; Roine, E.; Kukkaro, P.; Laurinavicius, S.; Somerharju, P.; Bamford, D.H. Quantitative dissociation of archaeal virus SH1 reveals distinct capsid proteins and a lipid core. Virology 2006, 356, 4–11. [Google Scholar] [CrossRef]
- Pietila, M.K.; Laurinavicius, S.; Sund, J.; Roine, E.; Bamford, D.H. The single-stranded DNA genome of novel archaeal virus Halorubrum pleomorphic virus 1 is enclosed in the envelope decorated with glycoprotein spikes. J. Virol. 2010, 84, 788–798. [Google Scholar] [CrossRef]
- Pietila, M.K.; Roine, E.; Paulin, L.; Kalkkinen, N.; Bamford, D.H. An ssDNA virus infecting archaea: A new lineage of viruses with a membrane envelope. Mol. Microbiol. 2009, 72, 307–319. [Google Scholar] [CrossRef]
- Porter, K.; Tang, S.L.; Chen, C.P.; Chiang, P.W.; Hong, M.J.; Dyall-Smith, M. PH1: An archaeovirus of Haloarcula hispanica related to SH1 and HHIV-2. Archaea 2013, 2013, 456318. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, J.; Wang, Y.; Zhang, Z.; Oksanen, H.M.; Bamford, D.H.; Chen, X. Identification and characterization of SNJ2, the first temperate pleolipovirus integrating into the genome of the SNJ1-lysogenic archaeal strain. Mol. Microbiol. 2015, 98, 1002–1020. [Google Scholar] [CrossRef] [PubMed]
- Demina, T.A.; Atanasova, N.S.; Pietila, M.K.; Oksanen, H.M.; Bamford, D.H. Vesicle-like virion of Haloarcula hispanica pleomorphic virus 3 preserves high infectivity in saturated salt. Virology 2016, 499, 40–51. [Google Scholar] [CrossRef] [PubMed]
- Demina, T.A.; Pietila, M.K.; Svirskaite, J.; Ravantti, J.J.; Atanasova, N.S.; Bamford, D.H.; Oksanen, H.M. Archaeal Haloarcula californiae Icosahedral Virus 1 highlights conserved elements in icosahedral membrane-containing DNA viruses from extreme environments. MBio 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Pagaling, E.; Haigh, R.D.; Grant, W.D.; Cowan, D.A.; Jones, B.E.; Ma, Y.; Ventosa, A.; Heaphy, S. Sequence analysis of an archaeal virus isolated from a hypersaline lake in Inner Mongolia, China. BMC Genom. 2007, 8, 410. [Google Scholar] [CrossRef]
- Pietilä, M.K.; Atanasova, N.S.; Oksanen, H.M.; Bamford, D.H. Modified coat protein forms the flexible spindle-shaped virion of haloarchaeal virus His1. Environ. Microbiol. 2013, 15, 1674–1686. [Google Scholar] [CrossRef]
- Jiang, S.; Steward, G.; Jellison, R.; Chu, W.; Choi, S. Abundance, distribution, and diversity of viruses in alkaline, hypersaline Mono Lake, California. Microb. Ecol. 2004, 47, 9–17. [Google Scholar] [CrossRef]
- Shen, P.S.; Domek, M.J.; Sanz-Garcia, E.; Makaju, A.; Taylor, R.M.; Hoggan, R.; Culumber, M.D.; Oberg, C.J.; Breakwell, D.P.; Prince, J.T.; et al. Sequence and structural characterization of Great Salt Lake bacteriophage CW02, a member of the T7-like supergroup. J. Virol. 2012, 86, 7907–7917. [Google Scholar] [CrossRef]
- Dyall-Smith, M. The Halohandbook: Protocols for Haloarchaeal Genetics. Dyall-Smith, M., Ed.; v.7.1. ed. 2009, p. 79. Available online: http://www.haloarchaea.com/resources/halohandbook/ (accessed on 1 September 2009).
- Muyzer, G.; Teske, A.; Wirsen, C.O.; Jannasch, H.W. Phylogenetic-relationships of Thiomicrospira species and their identification in deep-sea hydrothermal vent samples by denaturing gradient gel-electrophoresis of 16S rDNA fragments. Arch. Microbiol. 1995, 164, 165–172. [Google Scholar] [CrossRef]
- DeLong, E.F. Archaea in coastal marine environments. Proc. Natl. Acad. Sci. USA 1992, 89, 5685–5689. [Google Scholar] [CrossRef]
- Ludwig, W.; Strunk, O.; Westram, R.; Richter, L.; Meier, H.; Kumar, Y.; Buchner, A.; Lai, T.; Steppi, S.; Jobb, G.; et al. ARB: A software environment for sequence data. Nucleic Acids Res. 2004, 32, 1363–1371. [Google Scholar] [CrossRef]
- Pruesse, E.; Quast, C.; Knittel, K.; Fuchs, B.M.; Ludwig, W.; Peplies, J.; Gloeckner, F.O. SILVA: A comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007, 35, 7188–7196. [Google Scholar] [CrossRef] [PubMed]
- Swofford, D.L. PAUP*: Phylogenetic Analysis Using Parsimony (and Other Methods), V. 4.0, Beta 8, 4.0; Sinauer Associates: Champaign, IL, USA, 1998. [Google Scholar]
- Dussault, H.P. An improved technique for staining red halophilic bacteria. J. Bacteriol. 1955, 70, 484–485. [Google Scholar] [PubMed]
- Adams, M. Bacteriophages; Interscience Publishers, Inc.: New York, NY, USA, 1959. [Google Scholar]
- Henn, M.R.; Sullivan, M.B.; Stange-Thomann, N.; Osburne, M.S.; Berlin, A.M.; Kelly, L.; Yandava, C.; Kodira, C.; Zeng, Q.; Weiand, M.; et al. Analysis of high-throughput sequencing and annotation strategies for phage genomes. PLoS One 2010, 5, e9083. [Google Scholar] [CrossRef] [PubMed]
- Chen, I.A.; Chu, K.; Palaniappan, K.; Pillay, M.; Ratner, A.; Huang, J.; Huntemann, M.; Varghese, N.; White, J.R.; Seshadri, R.; et al. IMG/M v.5.0: An integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 2019, 47, D666–D677. [Google Scholar] [CrossRef]
- Chow, M.S.; Rouf, M.A. Isolation and partial characterization of two Aeromonas hydrophila bacteriophages. Appl. Environ. Microbiol. 1983, 45, 1670–1676. [Google Scholar]
- Hedi, A.; Sadfi, N.; Fardeau, M.L.; Rebib, H.; Cayol, J.L.; Ollivier, B.; Boudabous, A. Studies on the biodiversity of halophilic microorganisms isolated from El-Djerid Salt Lake (Tunisia) under aerobic conditions. Int. J. Microbiol. 2009, 2009, 731786. [Google Scholar] [CrossRef] [PubMed]
- Kharroub, K.; Aguilera, M.; Quesada, T.; Morillo, J.A.; Ramos-Cormenzana, A.; Boulharouf, A.; Monteoliva-Sanchez, M. Salicola salis sp nov., an extremely halophilic bacterium isolated from Ezzemoul sabkha in Algeria. Int. J. Syst. Evol. Microbiol. 2006, 56, 2647–2652. [Google Scholar] [CrossRef]
- Vahed, S.Z.; Forouhandeh, H.; Hassanzadeh, S.; Klenk, H.P.; Hejazi, M.A.; Hejazi, M.S. Isolation and characterization of halophilic bacteria from Urmia Lake in Iran. Microbiology 2011, 80, 834–841. [Google Scholar] [CrossRef]
- Antón, J.; Llobet-Brossa, E.; Rodriguez-Valera, F.; Amann, R. Fluorescence in situ hybridization analysis of the prokaryotic community inhabiting crystallizer ponds. Environ. Microbiol. 1999, 1, 517–523. [Google Scholar] [CrossRef]
- Villamor, J.; Ramos-Barbero, M.D.; Gonzalez-Torres, P.; Gabaldon, T.; Rossello-Mora, R.; Meseguer, I.; Martinez-Garcia, M.; Santos, F.; Anton, J. Characterization of ecologically diverse viruses infecting co-occurring strains of cosmopolitan hyperhalophilic Bacteroidetes. ISME J. 2018, 12, 424–437. [Google Scholar] [CrossRef]
- Porter, K.; Kukkaro, P.; Bamford, J.K.H.; Bath, C.; Kivela, H.M.; Dyall-Smith, M.L.; Bamford, D.H. SHI: A novel, spherical halovirus isolated from an Australian hypersaline lake. Virology 2005, 335, 22–33. [Google Scholar] [CrossRef] [PubMed]
- Goel, U.; Kauri, T.; Ackermann, H.-W.; Kushner, D.J. A moderately halophilic Vibrio from a Spanish saltern and its lytic bacteriophage. Can. J. Microbiol. 1996, 42, 1015–1023. [Google Scholar] [CrossRef]
- Vogelsang-Wenke, H.; Oesterhelt, D. Isolation of a halobacterial phage with a fully cytosine-methylated genome. Mol. Gen. Genet. 1988, 211, 407–414. [Google Scholar] [CrossRef]
- Bath, C.; Cukalac, T.; Porter, K.; Dyall-Smith, M.L. His1 and His2 are distantly related, spindle-shaped haloviruses belonging to the novel virus group, Salterprovirus. Virology 2006, 350, 228–239. [Google Scholar] [CrossRef] [PubMed]
- Torsvik, T.; Dundas, I. Persisting phage infection in Halobacterium salinarium str. 1. J. Gen. Virol. 1980, 47, 29–36. [Google Scholar] [CrossRef]
- Pauling, C. Bacteriophages of Halobacterium halobium: isolated from fermented fish sauce and primary characterization. Can. J. Microbiol. 1982, 28, 916–921. [Google Scholar] [CrossRef] [PubMed]
- Mesbah, N.; Wiegel, J. Halophilic thermophiles: A novel group of extremophiles. In Microbial Diversity: Current Perspectives and Potential Applications; Satyanaryana, T., Johri, B.N., Eds.; I.K. International: New Delhi, India, 2005; pp. 91–118. [Google Scholar]
- Schneider, J.; Herrmann, A. Saltworks–natural laboratories for microbiological and geochemical investigations during the evaporation of seawater. In Proceedings of the Fifth International Symposium on Salt, Las Vegas, NV, USA, 13–18 October 1985; pp. 371–381. [Google Scholar]
- Schnabel, H.; Zillig, W.; Pfaffle, M.; Schnabel, R.; Michel, H.; Delius, H. Halobacterium halobium phage ΦH. EMBO J. 1982, 1, 87–92. [Google Scholar] [CrossRef]
- Cox, J.; Schubert, A.M.; Travisano, M.; Putonti, C. Adaptive evolution and inherent tolerance to extreme thermal environments. BMC Evol. Biol. 2010, 10, 75. [Google Scholar] [CrossRef]
- Breitbart, M.; Rohwer, F. Here a virus, there a virus, everywhere the same virus? Trends Microbiol. 2005, 13, 278–284. [Google Scholar] [CrossRef]
| Strains | CGM5-S (USA) | 9-A-U (Mexico) | S. marasensis and Related Isolates (Peru) | S. salis (Algeria) | 7SPE Isolates (Tunisia) | TBZ Isolates (Iran) | IC10 (Spain) |
|---|---|---|---|---|---|---|---|
| Salinity (%) | |||||||
| Optimum | 20 | 15 | 15 | 15–20 | NR | 2/3 grew at 20 | 20–25 |
| Range | 15–30 | 10–30 | 10–30 | 10–25 | 5–25 | NR | 15–30 |
| pH | |||||||
| Optimum | 6.5 | NR | 7 | 7–7.5 | NR | 2/3 grew at 9 | 8 |
| Range | 6–8.5 | NR | 6–8 | 6–9 | 6–7.5 | NR | 5–8.5 |
| Temperature (˚C) | |||||||
| Optimum | 41 | NR | 35 | 37 | 37 | NR | 40 |
| Range | 32–48 | NR | 20–37 | 30–45 | NR | NR | 28–40 |
| Virus | CGϕ29 | SCTP-1 | SCTP-2 | ϕD-86 | F9-11 |
|---|---|---|---|---|---|
| Source | Cargill Saltern (USA) | Saltern (Italy) | Saltern (Italy) | Soy sauce | Hypersaline soil (Spain) |
| Host | Salicola sp. CGM5-S | Salicola sp. PV3 | Salicola sp. PV3 | Pedicoccus halophilus | Deleya halophila |
| Salinity (%) | |||||
| Tolerance range | 0–30 | 0–26 | 0–26 | 0–15 | 0–30 |
| Maximum exposure time | 1 y | 15 min | 18 min | 1 h | 45 d |
| pH | |||||
| Tolerance range | 3-9 | NR | NR | 4.5-10.7 | NR |
| Maximum exposure time | 72 h | NR | NR | 1 h | NR |
| Temperature (˚C) | |||||
| Maximum temperature tolerated | 75 | NR | NR | 50 | NR |
| Maximum exposure time | 72 h | NR | NR | 1 h | NR |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodela, M.L.; Sabet, S.; Peterson, A.; Dillon, J.G. Broad Environmental Tolerance for a Salicola Host-Phage Pair Isolated from the Cargill Solar Saltworks, Newark, CA, USA. Microorganisms 2019, 7, 106. https://doi.org/10.3390/microorganisms7040106
Rodela ML, Sabet S, Peterson A, Dillon JG. Broad Environmental Tolerance for a Salicola Host-Phage Pair Isolated from the Cargill Solar Saltworks, Newark, CA, USA. Microorganisms. 2019; 7(4):106. https://doi.org/10.3390/microorganisms7040106
Chicago/Turabian StyleRodela, Meghan L., Shereen Sabet, Allison Peterson, and Jesse G. Dillon. 2019. "Broad Environmental Tolerance for a Salicola Host-Phage Pair Isolated from the Cargill Solar Saltworks, Newark, CA, USA" Microorganisms 7, no. 4: 106. https://doi.org/10.3390/microorganisms7040106
APA StyleRodela, M. L., Sabet, S., Peterson, A., & Dillon, J. G. (2019). Broad Environmental Tolerance for a Salicola Host-Phage Pair Isolated from the Cargill Solar Saltworks, Newark, CA, USA. Microorganisms, 7(4), 106. https://doi.org/10.3390/microorganisms7040106





