Phage Endolysins as Potential Antimicrobials against Multidrug Resistant Vibrio alginolyticus and Vibrio parahaemolyticus: Current Status of Research and Challenges Ahead
Abstract
1. Introduction
2. Antibiotic Resistance in Vibrios alginolyticus and V. parahaemolyticus
Treatment Costs for Vibriosis
3. Bacteriophage Endolysins-‘the Holy Grail’ to Control Food Borne Pathogens
4. Gram-Negative Endolysins as Antimicrobials
Basic Structure and Function
5. Vibrio Phage Endolysins
5.1. Lysqdvp001 and Its Homologues
5.1.1. Structure, Function and Physiochemical Properties
5.1.2. LysVPMS1
5.1.3. LysVPp1
5.2. cwlQ- First Recombinant Endolysin with Holin Assistance
6. Challenges of Endolysin Engineering and Delivery
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Barbarossa, V.; Kucisec-Tepes, N.; Aldova, E.; Matek, D.; Stipoljev, F. Ilizarov technique in the treatment of chronic osteomyelitis caused by Vibrio alginolyticus. Croat. Med. J. 2002, 43, 346–349. [Google Scholar] [PubMed]
- Feingold, M.H.; Kumar, M.L. Otitis media associated with Vibrio alginolyticus in a child with pressure equalizing tubes. Pediatr. Infect. Dis. J. 2004, 23, 475–476. [Google Scholar] [CrossRef] [PubMed]
- Li, X.C.; Xiang, Z.Y.; Xu, X.M.; Yan, W.H.; Ma, J.M. Endophthalmitis caused by Vibrio alginolyticus. J. Clin. Microbiol. 2009, 47, 3379–3381. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Cheng, J.; Wu, Q.; Zhang, J.; Xie, T. Prevalence, characterization, and antibiotic susceptibility of Vibrio parahaemolyticus isolated from retail aquatic products in North China. BMC Microbiol. 2016, 16, 32. [Google Scholar] [CrossRef] [PubMed]
- Elmahdi, S.; DaSilva, L.V.; Parveen, S. Antibiotic resistance of Vibrio parahaemolyticus and Vibrio vulnificus in various countries: A review. Food Microbiol. 2016, 57, 128–134. [Google Scholar] [CrossRef]
- Romero, J.; Feijoó, C.G.; Navarrete, P. Antibiotics in aquaculture–use, abuse and alternatives. In Health and Environment in Aquaculture; InTechOpen: London, UK, 2012. [Google Scholar]
- Daniels, N.A.; Shafaie, A. A Review of Pathogenic Vibrio Infections for Clinicians. Infect. Med. 2000, 17, 665–685. [Google Scholar]
- CDC. Cholera and Other Vibrio Illness Surveillance (COVIS). 2016. Available online: https://www.cdc.gov/vibrio/surveillance.html (accessed on 28 January 2019).
- Kumar, S.; Singh, B. Ran Overview of Mechanisms and Emergence of Antimicrobials Drug Resistance. 2013. Available online: http://krishi.icar.gov.in/jspui/handle/123456789/11178 (accessed on 5 February 2019).
- Munita, J.M.; Arias, C.A. Mechanisms of antibiotic resistance. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef]
- Miller, S.I. Antibiotic resistance and regulation of the gram-negative bacterial outer membrane barrier by host innate immune molecules. mBio 2016, 7, e01541-16. [Google Scholar] [CrossRef]
- Partridge, S.R. Analysis of antibiotic resistance regions in Gram-negative bacteria. FEMS Microbiol. Rev. 2011, 35, 820–855. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Chen, X.; Yu, F.; Wu, M.; Wang, R.; Zheng, S.; Zhu, J. Serology, virulence, antimicrobial susceptibility and molecular characteristics of clinical Vibrio parahaemolyticus strains circulating in southeastern China from 2009 to 2013. Clin. Microbiol. Infect. 2016, 22, 258.e9–258.e16. [Google Scholar] [CrossRef]
- Hooper, D.C. Mechanisms of action of antimicrobials: Focus on fluoroquinolones. Clin. Infect. Dis. 2001, 32 (Suppl. 1), S9–S15. [Google Scholar] [CrossRef]
- Kitaoka, M.; Miyata, S.T.; Unterweger, D.; Pukatzki, S. Antibiotic resistance mechanisms of Vibrio cholerae. J. Med. Microbiol. 2011, 60, 397–407. [Google Scholar] [CrossRef] [PubMed]
- Wong, H.C.; Wang, T.Y.; Yang, C.W.; Tang, C.T.; Ying, C.; Wang, C.H.; Chang, W.H. Characterization of a lytic vibriophage VP06 of Vibrio parahaemolyticus. Res. Microbiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Morris, J.G., Jr.; Tenney, J. Antibiotic therapy for Vibrio vulnificus infection. JAMA 1985, 253, 1121–1122. [Google Scholar] [CrossRef] [PubMed]
- Kitiyodom, S.; Khemtong, S.; Wongtavatchai, J.; Chuanchuen, R. Characterization of antibiotic resistance in Vibrio spp. isolated from farmed marine shrimps (Penaeus monodon). FEMS Microbiol. Ecol. 2010, 72, 219–227. [Google Scholar] [CrossRef] [PubMed]
- Ye, L.; Li, R.; Lin, D.; Zhou, Y.; Fu, A.; Ding, Q.; Chen, S. Characterization of an IncA/C multidrug resistance plasmid in Vibrio alginolyticus. Antimicrob. Agents Chemother. 2016, 60, 3232–3235. [Google Scholar] [CrossRef] [PubMed]
- Ralston, E.P.; Kite-Powell, H.; Beet, A. An estimate of the cost of acute health effects from food-and water-borne marine pathogens and toxins in the USA. J. Water Health 2009, 9, 680–694. [Google Scholar] [CrossRef]
- O’Flaherty, S.; Ross, R.P.; Coffey, A. Bacteriophage and their lysins for elimination of infectious bacteria. FEMS Microbiol. Rev. 2009, 33, 801–819. [Google Scholar] [CrossRef]
- Young, R. Phage lysis: Do we have the whole story yet? Curr. Opin. Microbiol. 2013, 16, 790–797. [Google Scholar] [CrossRef]
- Berry, J.D.; Rajaure, M.; Pang, T.; Young, R. The spanin complex is essential for lambda lysis. J. Bacteriol. 2012, 194, 5667–5674. [Google Scholar] [CrossRef]
- Young, R.; Way, S.; Yin, J.; Syvanen, M. Transposition mutagenesis of bacteriophage lambda: A new gene affecting cell lysis. J. Mol. Biol. 1979, 132, 307–322. [Google Scholar] [CrossRef]
- Krupovič, M.; Cvirkaitė-Krupovič, V.; Bamford, D.H. Identification and functional analysis of the Rz/Rz1-like accessory lysis genes in the membrane-containing bacteriophage PRD1. Mol. Microbiol. 2008, 68, 492–503. [Google Scholar] [CrossRef] [PubMed]
- Berry, J.; Savva, C.; Holzenburg, A.; Young, R. The lambda spanin components Rz and Rz1 undergo tertiary and quaternary rearrangements upon complex formation. Protein Sci. 2010, 19, 1967–1977. [Google Scholar] [CrossRef] [PubMed]
- Nelson, D.; Loomis, L.; Fischetti, V.A. Prevention and elimination of upper respiratory colonization of mice by group A streptococci by using a bacteriophage lytic enzyme. Proc. Natl. Acad. Sci. USA 2001, 98, 4107–4112. [Google Scholar] [CrossRef] [PubMed]
- Donovan, D.M.; Dong, S.; Garrett, W.; Rousseau, G.M.; Moineau, S.; Pritchard, D.G. Peptidoglycan hydrolase fusions maintain their parental specificities. Appl. Environ. Microbol. 2006, 72, 2988–2996. [Google Scholar] [CrossRef] [PubMed]
- Beveridge, T.J. Structures of gram-negative cell walls and their derived membrane vesicles. J. Bacteriol. 1999, 181, 4725–4733. [Google Scholar]
- Larpin, Y.; Oechslin, F.; Moreillon, P.; Resch, G.; Entenza, J.M.; Mancini, S. In vitro characterization of PlyE146, a novel phage lysin that targets Gram-negative bacteria. PLoS ONE 2018, 13, e0192507. [Google Scholar] [CrossRef] [PubMed]
- Low, L.Y.; Yang, C.; Perego, M.; Osterman, A.; Liddington, R. Role of net charge on catalytic domain and influence of cell wall binding domain on bactericidal activity, specificity, and host range of phage lysins. J. Biol. Chem. 2011, 286, 34391–34403. [Google Scholar] [CrossRef]
- Chen, H.-M.; Li, Y.; Panda, T.; Buehler, F.U.; Ford, C.; Reilly, P.J. Effect of replacing helical glycine residues with alanines on reversible and irreversible stability and production of Aspergillus awamori glucoamylase. Protein Eng. 1996, 9, 499–505. [Google Scholar] [CrossRef]
- Briers, Y.; Lavigne, R.; Volckaert, G.; Hertveldt, K. A standardized approach for accurate quantification of murein hydrolase activity in high-throughput assays. J. Biochem. Biophys. Methods 2007, 70, 531–533. [Google Scholar] [CrossRef]
- Sanz-Gaitero, M.; Keary, R.; Garcia-Doval, C.; Coffey, A.; van Raaij, M.J. Crystallization of the CHAP domain of the endolysin from Staphylococcus aureus bacteriophage K. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2013, 69, 1393–1396. [Google Scholar] [CrossRef] [PubMed]
- Becker, S.C.; Swift, S.; Korobova, O.; Schischkova, N.; Kopylov, P.; Donovan, D.M.; Abaev, I. Lytic activity of the staphylolytic Twort phage endolysin CHAP domain is enhanced by the SH3b cell wall binding domain. FEMS Microbiol. Lett. 2015, 362, 1. [Google Scholar] [CrossRef] [PubMed]
- Walmagh, M.; Boczkowska, B.; Grymonprez, B.; Briers, Y.; Drulis-Kawa, Z.; Lavigne, R. Characterization of five novel endolysins from Gram-negative infecting bacteriophages. Appl. Microbiol. Biotechnol. 2013, 97, 4369–4375. [Google Scholar] [CrossRef]
- Oliveira, H.; Melo, L.D.; Santos, S.B.; Nóbrega, F.L.; Ferreira, E.C.; Cerca, N.; Azeredo, J.; Kluskens, L.D. Molecular aspects and comparative genomics of bacteriophage endolysins. J. Virol. 2013, 87, 4558–4570. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Yu, J.; Wei, H. Engineered bacteriophage lysins as novel anti-infectives. Front. Microbiol. 2014, 5, 542. [Google Scholar] [CrossRef]
- Idelevich, E.A.; von Eiff, C.; Friedrich, A.W.; Iannelli, D.; Xia, G.; Peters, G.; Peschel, A.; Wanninger, I.; Becker, K. In vitro activity against Staphylococcus aureus of a novel antimicrobial agent, PRF-119, a recombinant chimeric bacteriophage endolysin. Antimicrob. Agents Chemother. 2011, 55, 4416–4419. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, S.; Proença, D.; Cantante, C.; Silva, F.A.; Leandro, C.; Lourenço, S.; Milheiriço, C.; de Lencastre, H.; Cavaco-Silva, P.; Pimentel, M.; et al. Novel chimerical endolysins with broad antimicrobial activity against methicillin-resistant Staphylococcus aureus. Microb. Drug Resist. 2012, 18, 333–343. [Google Scholar] [CrossRef]
- Mao, J.; Schmelcher, M.; Harty, W.J.; Foster-Frey, J.; Donovan, D.M. Chimeric Ply187 endolysin kills Staphylococcus aureus more effectively than the parental enzyme. FEMS Microbiol. Lett. 2013, 1, 30–36. [Google Scholar] [CrossRef]
- Dong, Q.; Wang, J.; Yang, H.; Wei, C.; Yu, J.; Zhang, Y.; Huang, Y.; Zhang, X.E.; Wei, H. Construction of a chimeric lysin Ply187N-V12C with extended lytic activity against staphylococci and streptococci. Microb. Biotechnol. 2014. [Google Scholar] [CrossRef]
- Yang, H.; Zhang, Y.; Yu, J.; Huang, Y.; Zhang, X.E.; Wei, H. Novel chimeric lysin with high-level antimicrobial activity against methicillin-resistant Staphylococcus aureus in vitro and in vivo. Antimicrob. Agents Chemother. 2014, 58, 536–542. [Google Scholar] [CrossRef]
- Yang, H.; Zhang, H.; Wang, J.; Yu, J.; Wei, H. A novel chimeric lysin with robust antibacterial activity against planktonic and biofilm methicillin-resistant Staphylococcus aureus. Sci. Rep. 2017, 7, 40182. [Google Scholar] [CrossRef]
- Briers, Y.; Walmagh, M.; Grymonprez, B.; Biebl, M.; Pirnay, J.P.; Defraine, V.; Michiels, J.; Cenens, W.; Aertsen, A.; Miller, S.; et al. Art-175 is a highly efficient antibacterial against multidrug-resistant strains and persisters of Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2014, 58, 3774–3784. [Google Scholar] [CrossRef]
- Lukacik, P.; Barnard, T.J.; Keller, P.W.; Chaturvedi, K.S.; Seddiki, N.; Fairman, J.W.; Noinaj, N.; Kirby, T.L.; Henderson, J.P.; Steven, A.C.; et al. Structural engineering of a phage lysin that targets gram-negative pathogens. Proc. Natl. Acad. Sci. USA 2012, 109, 9857–9862. [Google Scholar] [CrossRef] [PubMed]
- Zampara, A.; Sørensen, M.C.; Grimon, D.; Antenucci, F.; Briers, Y.; Brøndsted, L. Innolysins: A novel approach to engineer endolysins to kill Gram-negative bacteria. BioRxiv 2018, 408948. [Google Scholar] [CrossRef]
- Crothers-Stomps, C.; Høj, L.; Bourne, D.G.; Hall, M.R.; Owens, L. Isolation of lytic bacteriophage againstVibrio harveyi. J. Appl. Microbiol. 2010, 108, 1744–1750. [Google Scholar] [CrossRef] [PubMed]
- Rong, R.; Lin, H.; Wang, J.; Khan, M.N.; Li, M. Reductions of Vibrio parahaemolyticus in oysters after bacteriophage application during depuration. Aquaculture 2014, 418, 171–176. [Google Scholar] [CrossRef]
- Zhang, J.; Cao, Z.; Xu, Y.; Li, X.; Li, H.; Wu, F.; Jin, L. Complete genomic sequence of the Vibrio alginolyticus lytic bacteriophage PVA1. Arch. Virol. 2014, 159, 3447–3451. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Li, X.; Zhang, J.; Wang, X.; Wang, L.; Cao, Z.; Xu, Y. Use of phages to control Vibrio splendidus infection in the juvenile sea cucumber Apostichopus japonicus. Fish Shellfish Immunol. 2016, 54, 302–311. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.G.; Giri, S.S.; Jun, J.W.; Yun, S.; Kim, H.J.; Kim, S.W.; Kang, J.W.; Han, S.J.; Jeong, D.; Park, S.C. Concentration-dependent reduction of planktonic-and biofilm-state Vibrio alginolyticus by the bacteriophage pVa-21. BioRxiv 2018, 322933. [Google Scholar] [CrossRef]
- Skliros, D.; Kalatzis, P.G.; Katharios, P.; Flemetakis, E. Comparative functional genomic analysis of two vibrio phages reveals complex metabolic interactions with the host cell. Front. Microbiol. 2016, 7, 1807. [Google Scholar] [CrossRef]
- Lal, T.M.; Sano, M.; Hatai, K.; Ransangan, J. Complete genome sequence of a giant Vibrio phage ValKK3infectingVibrioalginolyticus. Genomicsdata 2016, 8, 37. [Google Scholar] [CrossRef]
- Skliros, D.; Kalatzis, P.G.; Flemetakis, E. Biotechnology, Agricultural University of Athens, Iera Odos, Athens, Attika 11855, Greece Comparative genomics of small Myoviridae Vibriophages (GenBank accession number: MG640035.1). 2017. [Google Scholar]
- Kokkari, C.; Sarropoulou, E.; Bastias, R.; Mandalakis, M.; Katharios, P. Isolation and characterization of a novel bacteriophage infecting Vibrio alginolyticus. Arch. Microbiol. 2018, 200, 707–718. [Google Scholar] [CrossRef] [PubMed]
- Luo, P.; Yun, L.; Li, Y.; Tian, Y.; Liu, Q.; Huang, W.; Hu, C. Complete genomic sequence of the Vibrio alginolyticus bacteriophage Vp670 and characterization of the lysis-related genes, cwlQ and holA. BMC Genom. 2018, 19, 741. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Xing, S.; Fu, K.; Zhao, S.; Liu, J.; Tong, Y.; Zhou, L. Genomic and biological characterization of the Vibrio alginolyticus-infecting “Podoviridae” bacteriophage, vB_ValP_IME271. Virus Genes 2019, 1–9. [Google Scholar] [CrossRef]
- Wang, W.; Li, M.; Lin, H.; Wang, J.; Mao, X. The Vibrio parahaemolyticus-infecting bacteriophage qdvp001: Genome sequence and endolysin with a modular structure. Arch. Virol. 2016, 161, 2645–2652. [Google Scholar] [CrossRef] [PubMed]
- Wong, M.H.Y.; Liu, M.; Wan, H.Y.; Chen, S. Characterization of Extended Spectrum β-lactamase Producing Vibrio parahaemolyticus. Antimicrob. Agents Chemother. 2012, 56, 4026–4028. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Jin, Y.; Lin, H.; Wang, J.; Jiang, X. Complete Genome of a Novel Lytic Vibrio parahaemolyticus Phage VPp1 and Characterization of Its Endolysin for Antibacterial Activities. J. Food Prot. 2018, 81, 1117–1125. [Google Scholar] [CrossRef] [PubMed]
- Zermeño-Cervantes, L.A.; Makarov, R.; Lomelí-Ortega, C.O.; Martínez-Díaz, S.F.; Cardona-Félix, C.S. Recombinant Lys VPMS 1 as an endolysin with broad lytic activity against Vibrio parahaemolyticus strains associated to acute hepatopancreatic necrosis disease. Aquac. Res. 2018, 49, 1723–1726. [Google Scholar] [CrossRef]
- Lal, T.M.; Ransangan, J. Complete genome sequence of VpKK5, a novel Vibrio parahaemolyticus lytic siphophage. Genome Announc. 2015, 3. [Google Scholar] [CrossRef]
- Midorikawa, Y.; Sano, M. okyo University of Marine Science and Technology; Konan 4-5-7, Minato-ku, Tokyo, Japan. Complete Genome Sequence of Vibrio parahaemolyticus Bacteriophage pTD1 (GenBank accession number: AP017972.1). 2017. [Google Scholar]
- Fischetti, V.A. Bacteriophage lysins as effective antibacterials. Curr. Opin. Microbiol. 2008, 11, 393–400. [Google Scholar] [CrossRef]
- Jun, S.Y.; Jung, G.M.; Son, J.S.; Yoon, S.J.; Choi, Y.J.; Kang, S.H. Comparison of the antibacterial properties of phage endolysins SAL-1 and LysK. Antimicrob. Agents Chemother. 2011, 55, 1764–1767. [Google Scholar] [CrossRef]
- Jun, S.Y.; Jung, G.M.; Yoon, S.J.; Oh, M.D.; Choi, Y.J.; Lee, W.J.; Kong, J.C.; Seol, J.G.; Kang, S.H. Antibacterial properties of a pre-formulated recombinant phage endolysin, SAL-1. Int. J. Antimicrob. Agents 2013, 41, 156–161. [Google Scholar] [CrossRef] [PubMed]
- Jun, S.Y.; Jang, I.J.; Yoon, S.; Jang, K.; Yu, K.S.; Cho, J.Y.; Kang, S.H. Pharmacokinetics and tolerance of the phage endolysin-based candidate drug SAL200 after a single intravenous administration among healthy volunteers. Antimicrob. Agents Chemother. 2017, 61, e02629-16. [Google Scholar] [CrossRef]
- Totté, J.E.; van Doorn, M.B.; Pasmans, S.G. Successful treatment of chronic Staphylococcus aureus-related dermatoses with the topical endolysin Staphefekt SA. 100: A report of 3 cases. Case Rep. Dermatol. 2017, 9, 19–25. [Google Scholar] [CrossRef]
- Loeffler, J.M.; Djurkovic, S.; Fischetti, V.A. Phage lytic enzyme Cpl-1 as a novel antimicrobial for pneumococcal bacteremia. Infect. Immun. 2003, 71, 6199–6204. [Google Scholar] [CrossRef] [PubMed]
- Barrera-Rivas, C.I.; Valle-Hurtado, N.A.; González-Lugo, G.M.; Baizabal-Aguirre, V.M.; Bravo-Patiño, A.; Cajero-Juárez, M.; Valdez-Alarcón, J.J. Bacteriophage Therapy: An Alternative for the Treatment of Staphylococcus aureus Infections in Animals and Animal Models; Frontiers in Staphylococcus aureus; InTechOpen: London, UK, 2017. [Google Scholar] [CrossRef]
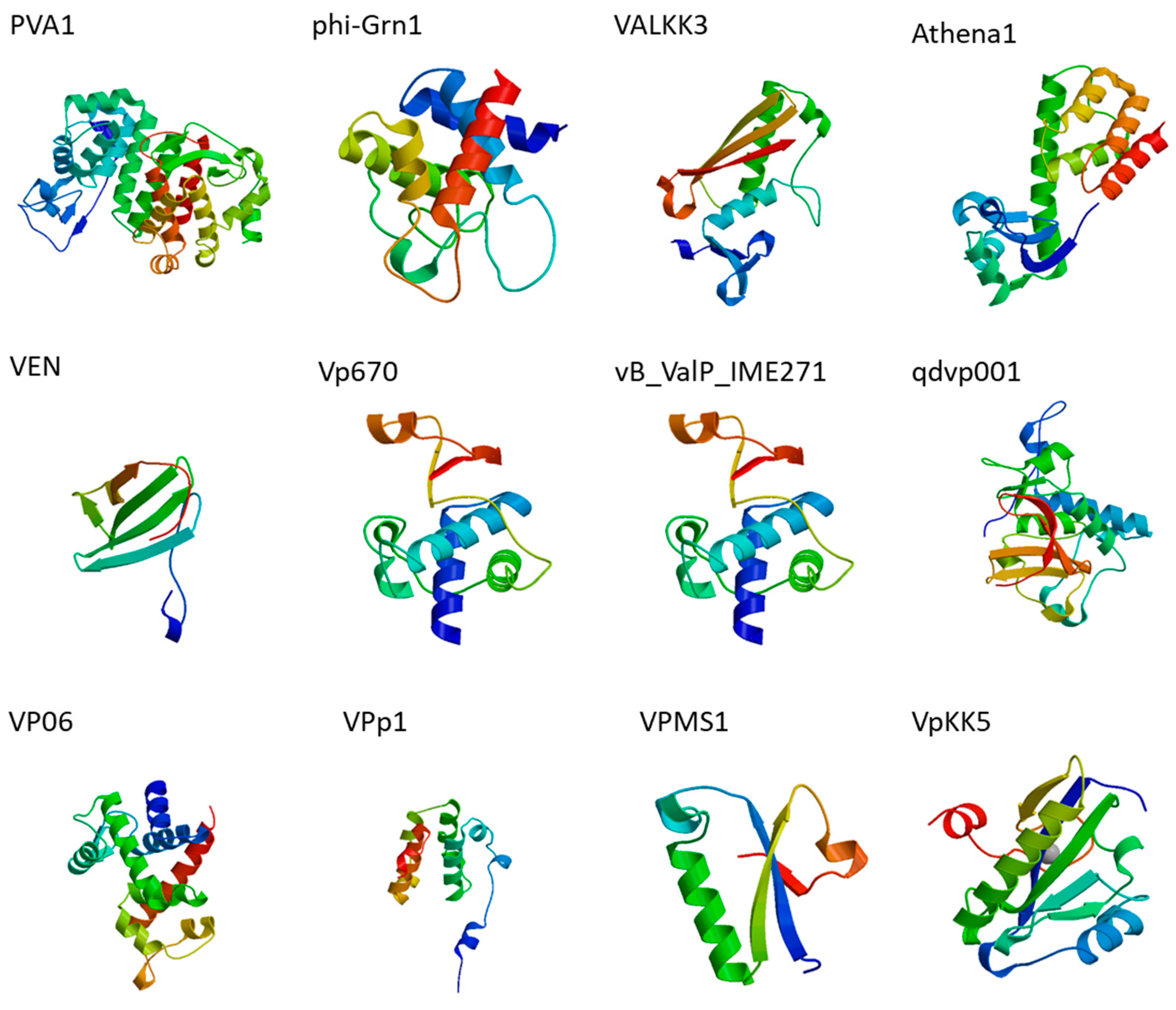
| Bacteria | Phage | Putative Endolysins/Predicted Orfs | Features | Reference |
|---|---|---|---|---|
| Vibrio alginolyticus | PVA1 | gp60 | Putative lysozyme family protein * | [50] |
| phi-Grn1 | phiGrn1_0012 | SLT domain protein/endolysin * | [53] | |
| ValKK3 | ORF304 | Tail lysozyme | [54] | |
| Athena1 | Cds006 | Protein with lysozyme activity * | [55] | |
| VEN | gp50 | Cell wall hydrolase-like protein * | [56] | |
| Vp670 | cwlQ | endolysin | [57] | |
| vB_ValP_IME271 | CDS64 | endolysin | [58] | |
| V. parahaemolyticus | qdvp001 | Lysqdvp001 | modular endolysin | [59] |
| VP06 | PP_00050 | membrane-bound lytic murein transglycosylase * | [60] | |
| VPp1 | LysVPp1 | endolysin | [61] | |
| VPMS1 | LysVPMS1 | endolysin | [62] | |
| VpKK5 | ORF62 | N-acetylmuramoyl-l-alanine amidase * | [63] | |
| pTD1 | BAW98403.1 | Endolysin * | [64] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Matamp, N.; Bhat, S.G. Phage Endolysins as Potential Antimicrobials against Multidrug Resistant Vibrio alginolyticus and Vibrio parahaemolyticus: Current Status of Research and Challenges Ahead. Microorganisms 2019, 7, 84. https://doi.org/10.3390/microorganisms7030084
Matamp N, Bhat SG. Phage Endolysins as Potential Antimicrobials against Multidrug Resistant Vibrio alginolyticus and Vibrio parahaemolyticus: Current Status of Research and Challenges Ahead. Microorganisms. 2019; 7(3):84. https://doi.org/10.3390/microorganisms7030084
Chicago/Turabian StyleMatamp, Nandita, and Sarita G. Bhat. 2019. "Phage Endolysins as Potential Antimicrobials against Multidrug Resistant Vibrio alginolyticus and Vibrio parahaemolyticus: Current Status of Research and Challenges Ahead" Microorganisms 7, no. 3: 84. https://doi.org/10.3390/microorganisms7030084
APA StyleMatamp, N., & Bhat, S. G. (2019). Phage Endolysins as Potential Antimicrobials against Multidrug Resistant Vibrio alginolyticus and Vibrio parahaemolyticus: Current Status of Research and Challenges Ahead. Microorganisms, 7(3), 84. https://doi.org/10.3390/microorganisms7030084




