Abstract
Specific microbial profiles and changes in intestinal microbiota have been widely demonstrated to be associated with the pathogenesis of a number of extra-intestinal (obesity and metabolic syndrome) and intestinal (inflammatory bowel disease) diseases as well as other metabolic disorders, such as non-alcoholic fatty liver disease and type 2 diabetes. Thus, maintaining a healthy gut ecosystem could aid in avoiding the early onset and development of these diseases. Furthermore, it is mandatory to evaluate the alterations in the microbiota associated with pathophysiological conditions and how to counteract them to restore intestinal homeostasis. This review highlights and critically discusses recent literature focused on identifying changes in and developing gut microbiota-targeted interventions (probiotics, prebiotics, diet, and fecal microbiota transplantation, among others) for the above-mentioned pathologies. We also discuss future directions and promising approaches to counteract unhealthy alterations in the gut microbiota. Altogether, we conclude that research in this field is currently in its infancy, which may be due to the large number of factors that can elicit such alterations, the variety of related pathologies, and the heterogeneity of the population involved. Further research on the effects of probiotics, prebiotics, or fecal transplantations on the composition of the human gut microbiome is necessary.
1. Introduction
Health is defined as “the state of the organism when it functions optimally without evidence of disease,” and the words “microbes” or “microorganism” are surprisingly missing in this definition. Currently, sequencing technologies (e.g., next-generation sequencing technologies) have stimulated new research that relates the microbial communities that reside in our gut with a number of physiological conditions. In 2014, the term microbiota was defined as the “full collection of microbes (bacteria, fungi, and viruses, among others) that naturally exist within a particular biological niche,” an estimated 500–1000 species [1,2,3] that may have a tremendous impact on human health.
The gut microbiota is regulated by an enormous number of factors, such as microbiological factors, host characteristics, diet patterns, and environmental variables [4]. Some protective, structural, and metabolic functions have been reported for gut microbiota [5], and these functions are associated with the regulation of homeostasis and host health. Host defenses against pathogens are in part mediated through the activity of the gut microbiota, requiring an intimate interpretation of the current microenvironment to discriminate between commensal and transient bacteria [6,7].
The intestinal epithelium is constantly exposed to high levels of food and bacterial antigens. Under normal physiological conditions, the intestinal epithelial monolayer facilitates a controlled and selective flux of components between the lumen and the underlying mucosa [8]. The intestine and the gut-associated lymphoid tissue are essential components of the immune defense system, protecting the host from foreign antigens and pathogens while tolerating commensal bacteria and dietary antigens. Antigen-presenting cell populations in the gut dictate in part the homeostasis between tolerance and immunity in the intestine, and the dysregulation of this balance can contribute to the pathogenesis of numerous inflammatory conditions [9]. All of the aforementioned cells are involved in the host response, and their function principally involves the maintenance of homeostasis.
Endogenous and exogenous factors influence the gut microbiota [10,11] including the mode of delivery of a neonate [12], host genetic features [13], host immune response [14], diet [15] (including dietary supplements, breast-feeding, and formula-feeding), xenobiotics (including antibiotics) and other drugs [16,17], infections [18], diurnal rhythm [19], and environmental microbial exposures [20,21]. Despite evidence linking dysbiosis of the gut microbiota with disease manifestations at sites distant from the gut, most studies have not explored mechanisms outside the affected site, nor have they considered the effect of the microbiota and its varied products on the multitude of molecular pathways potentially involved [21]. Overall, our current understanding of the precise relationships between the human gut microbiome and disease remains limited. Case–control studies often report disease-associated microbial dysbiosis, defined as the alteration in terms of the diversity, quantity, and stability of resident commensal communities relative to the community found in healthy individuals [3,22].
Probiotics are defined as “live microorganisms that confer a health benefit to the host when administered in adequate amounts, although dead bacteria and bacterial molecular components may also exhibit probiotic properties” [23], whereas a prebiotic is a non-viable food component that confers a health benefit to the host and is associated with the modulation of the intestinal microbiota [3]. The combination of prebiotics and probiotics is referred to as synbiotics [3,24,25].
A healthy host–microorganism balance contributes to optimally perform metabolic and immune functions as well as prevent disease development. Indeed, alterations in the delicate host–microbe relationship is nowadays recognized as associated with a wide variety of diseases such as cancer, neurological, ophthalmological, premature newborn, extra-intestinal and intestinal diseases, and metabolic disorders, such as non-alcoholic fatty liver disease and type 2 diabetes (T2D) [26]. However, the specific changes in the intestinal microbiome that are associated with these pathologies remain unelucidated.
To shed some light on this matter, a comprehensive search of the relevant literature reported during the last five years was performed in multiple electronic databases, including MEDLINE (PubMed), EMBASE, and the Cochrane Library. MEDLINE was searched through PubMed for scientific articles in English using the terms “microbiota” combined with “obesity,” “inflammatory bowel disease,” “non-alcoholic fatty liver disease,” “insulin resistance syndrome,” and “diabetes mellitus type 2”. The gut microbiota changes associated with the following conditions are reviewed: (1) obesity—antibiotics and obesity, gut microbiota and children’s obesity, bariatric surgery and the gut microbiota, potential gut microbiota biomarkers in obesity, and clinical trials; (2) inflammatory bowel disease—ulcerative colitis, and Crohn’s disease; (3) non-alcoholic fatty liver disease; (4) insulin resistance syndrome; and (5) diabetes mellitus type 2.
2. Gut Microbiota Changes in Particular Pathologies
2.1. Obesity
The prevalence of obesity is increasing worldwide [27,28] and currently represents a major health problem in both adults and children. The etiology of obesity has been associated with diverse factors, such as dietary, environmental, educational, and genetic factors. However, these factors do not fully explain the global incremental rise in obesity, and microbiota traits have recently been shown to play a causative role in obesity [29,30], demonstrating the potential of the microbiota as a therapeutic target in obesity. Here we present evidence-based studies linking host gut bacteria to obesity.
2.1.1. Antibiotics and Obesity
Antibiotic exposure has an overwhelming impact on the gut microbiota [31]. Epidemiological studies have shown that the administration of antibiotics during childhood is associated with a major risk of obesity [32,33]. Well-known studies have demonstrated the correlation between obesity and the intestinal microbiota balance [34,35]. These investigations showed an obesity-associated decrease in the phylum Bacteroidetes and an increase in the phylum Firmicutes, alterations in the gut microbiota that have been confirmed in animal models that exhibit fat accumulation due to an alteration of host metabolism [36,37]. Early life exposure to antibiotics has also been suggested to lead to obesity [38]. Interestingly, a decrease in the number of antiobesogenic bacteria (bifidobacteria and Bacteroides) after repeated exposure to antibiotics in neonates has been described [39]. Moreover, several studies have reported a correlation between childhood body mass index (BMI) and the risk of developing obesity later in life with antibiotics consumption [32,33,40,41,42,43]. Clearly, as revealed from extensive cohort, multicenter studies, gut microbiome alterations induced by antibiotics may promote obesity in adulthood, emphasizing the need for additional research on the relationships among antibiotic exposure, the intestinal microbiota balance, and the susceptibility to develop obesity.
2.1.2. Gut Microbiota and Childhood Obesity
The aforementioned alterations in gut bacteria have been very recently confirmed in children. An elevated abundance of Firmicutes and a decreased abundance of Bacteroidetes have been shown to correlate with overweight and obesity [44]. Interestingly, Bacteroidetes have been shown to be a better predictor of BMI than Firmicutes, possibly due to a higher variability of Firmicutes. This phenomenon may explain the results of a microbiome profiling study of eight obese children in whom the phylum Bacteroidetes exhibited the aforementioned tendency while Firmicutes showed little variation compared to the controls [45]. The levels of short chain fatty acids (SCFAs), the end products of fermentation of dietary fibers by the anaerobic intestinal microbiota, have been shown to be higher in obese children [44,46,47], suggesting that gut dysbiosis and augmented intestinal fermentation must be considered to be among the factors involved in the etiology of infant obesity. Interestingly, the use of prebiotics has been shown in a recent study to alter the intestinal microbiota and reduce body fat in obese or overweight children [48]. Personalized nutrition [49,50], the use of prebiotics, probiotics, postbiotics, and synbiotics [51], fecal microbiota transplantation (FMT) [52,53], dietary education [54], and physical activity [55] constitute the primary approaches aimed at using the gut microbiota as a therapeutic target for obesity in children. In a prospective study of 70 children within a four-year window, pre-obese dysbiotic gut and dietary habits were correlated with excessive weight gaining [56].
2.1.3. Bariatric Surgery and the Gut Microbiota
The gut microbiota has been shown to mediate the metabolic benefits and weight loss observed after bariatric surgery. Bariatric surgery alters the expression of several genes involved in metabolic pathways, which induces changes in the gut microbiota composition [57,58,59,60]. Given the relationship between the gut microbiota and the metabolic health of an individual, there is a growing interest in understanding the modifications that occur in intestinal microbiota after bariatric surgery and how these changes lead to weight loss and a better metabolic profile. Microbiome analyses following Roux-en-Y gastric bypass (RYGB) or vertical banded gastroplasty have shown similar long-term effects on gut microbiota and fat loss [61]. In mice, RYGB has been shown to induce a rapid and maintained increase in the presence of Gammaproteobacteria (Escherichia) and Verrucomicrobia (Akkermansia) [62]. A study of microbiome modifications in 13 obese patients after bariatric surgery reported an increase in gut microbial diversity and identified alterations in the relative abundances of 31 species [63]. These changes in microbiota occurred in parallel with weight loss and functional and metabolic improvements: (i) an increased potential to assimilate multiple energy sources using transporters and phosphotransferase system, (ii) a better use of aerobic respiration, and (iii) the use of amino acids and fatty acids as energy sources.
Very recently, an investigation of a cohort of lean and obese individuals identified a distinct causal role of the gut microbiota in the development of obesity [64]. Liu et al. associated changes in microbiota composition of some obesity-related microbial species with circulating metabolites. In particular, the abundance of a glutamate-fermenting commensal species, Bacteroides thetaiotaomicron, was notably decreased in obese individuals and was inversely correlated with serum glutamate concentration. Furthermore, these authors reported that, after bariatric surgery, several microbial enzymatic functions become more similar to those of lean controls, including pathways involved in carbohydrate fermentation, citrate cycle, glycosaminoglycan degradation, and lipopolysaccharide (LPS) synthesis, as well as the production of aromatic amino acid and branched chain amino acids (BCAA), suggesting that weight-loss intervention by bariatric surgery reversed, at least partially, the reported obesity-associated microbial and metabolic alterations. Very recently, decreased gut microbial gene richness (MGR) has been associated with severe obesity [22] and, although bariatric surgery increased MGR one year after surgery, most RYGB patients remained with low MGR one year post-surgery, which emphasizes the need of additional strategies to improve the dysbiosis in severe obesity. Table 1 summarizes the changes in gut microbiota associated with obesity and microbiota-targeted interventions for this disease.

Table 1.
Microbiota changes associated with obesity.
2.1.4. Potential Gut Microbiota Biomarkers of Obesity
The use of metagenomics analyses to link obesity and the gut microbiota allows for the specific bacterial strains to be identified as potential biomarkers of obesity and of the development or progression obesity. Dominant bacterial phyla that are consistently identified in the guts of normal individuals include Firmicutes, Bacteroidetes, and Actinobacteria, with Verrucomicrobia and Proteobacteria being present at lower abundances [65,66]. It has been recently described that microbiota markers in adolescent and adult obese patients show different age-dependent traits [67]. In particular, Faecalibacterium prausnitzii and Actinomyces have been assigned to the microbiota of obese adolescents, whereas Bacteroides caccae, Barnesiellaceae, Parabacteroides, Rikenellaceae, and Oscillospira have been assigned to the microbiota of adolescents with normal weights. F. prausnitzii participates in the fermentation of non-absorbed carbohydrates, and its abundance in the guts of obese adolescents may contribute to increased energy recovery, leading to a higher dietary energy intake that may in turn contribute to the lower success of weight loss diets reported for individuals with a higher abundance of F. prausnitzii [68]. Other reports associate the butyrate production capacity of F. prausnitzii with a healthy state [69,70,71], an apparent contradiction that emphasizes the need of future studies using broader cohorts to identify distinct microbial biomarkers that can unequivocally associate obesity with a given microbiota profile.
2.1.5. Clinical Trials
Of the 123 active clinical trials annotated to date in ClinicalTrials.gov (not finished or completed) that consider the gut microbiota as a therapeutic target in obesity, 55 (44.7%) employ dietary supplements or modifications in the diets of individuals in their interventions, 21 (17.07%) propose the use of probiotics, prebiotics, or synbiotics, 15 (12.2%) analyze the microbiota after bariatric surgery, 9 (7.3%) perform fecal transplantation interventions, 12 (9.7%) institute changes in patient lifestyle (ranging from exercise to mindfulness), 4 (3.2%) use approved drugs to adjust the gut microbiota, and 7 (5.7%) are strictly observational analyses. The important role of the gut microbiota in the alteration of the physiological systems involved in obesity is reflected by the wide variety of therapies based on counteracting their pathological modifications.
2.2. Inflammatory Bowel Disease
Crohn’s disease (CD) and ulcerative colitis (UC) are the primary pathological conditions associated with IBD and are described as a chronic and non-specific inflammation of the gastrointestinal tract [72]. There is some information regarding the correlation between changes in the gut microbiota and the development of IBD [73,74]. Regarding microbiota profiles, some studies have reported lower relative abundances of Alistipes finegoldi and Alistipes putredinis, as well as Firmicutes, Tenericutes, and Bacteroidetes in patients with IBD [75].
2.2.1. Ulcerative Colitis
The gut microbiota associated UC is characterized by a high ratio of B. fragilis/F. prausnitzii and a low abundance of butyrate-producing bacteria (BPB) [76]. Some studies have described a decrease in bifidobacteria [73], Akkermansia municiphila [77], and Clostridium clusters IV, XIVa, and XVIII [78] in patients with UC compared with the control groups. Two recent studies have reported an increased diversity in Clostridium cluster XIVa in UC patients [77] as well as increases in the abundances of Bacteroidetes, Bacilli, Proteobacteria, and Clostridium clusters IX and XI [78]. Recently, Bacteroidetes has been reported to be absent in patients with active UC [79]. These conflicting results regarding the Bacteroidetes phylum might be explained by the different status of the UC disease.
FMT is currently being considered as a potential future therapeutic approach to restore normal intestinal microbiota. FMT-treated patients exhibit different outcomes with respect to their gut microbiota composition depending on whether they respond to the transplant. Thus, non-responders show significantly higher levels of Bacteroidetes versus responders [80,81]. In addition, patients in remission have augmented levels of Barnesiella spp., Parabacteroides spp., Clostridium cluster IV, and Ruminococcus species [82,83].
Interesting changes in intestinal microbiota have been described after pharmacological treatments. Patients treated with mesalazine exhibited decreases in the abundances in Bifidobacterium and Lactobacillus and increases in the abundances of Klebsiella, Proteus, Citrobacter, and hemolytic Escherichia coli [84]. After treatment with andecaliximab, the presence of Clostridia and Akkermansia was observed in patients with active UC [85]. Patients treated with vedolizumab and those with non-remission showed increased levels of Streptococcus salivarius [86], and corticosteroid treatment was observed to be related to an increase in the abundances of bifidobacteria and Clostridium and a decrease in that of Faecalibacterium [87].
Regarding the use of probiotics, Matsuoka et al. have described that the administration of Bifidobacterium breve Yakult to UC patients barely had an effect, with the only change being in the relative abundance of Clostridium leptum, which significantly increased [88]. However, Ananthakrishnan et al. have suggested that early clinical remission of UC patients could be predicted by microbial composition only at baseline with a weaker influence at the level of the species or genus [87].
As for the impact of diet on the gut microbiota of UC patients, a low fat diet has been reported to increase the presence of Bacteroidetes after four weeks of intervention [89].
2.2.2. Crohn’s Disease
Recent investigations have described lower levels of Clostridium coccoides, Clostridium leptum and Faecalibacterium prausnitzii and a higher abundance of Escherichia coli in patients with CD [90].
A significant increase in Escherichia coli was reported in patients with CD subjected to FMT [91]. Another study investigating FMT in CD patients described decreases in the abundances of Bacteroides, Roseburia, Phascolarctobacterium, and Eubacterium along with increases in those of Bilophila, Streptococcus, Clostridium, and Paraprevotella [92].
Infliximab treatment has been reported to increase the abundance of Clostridiales in responders versus relapsing patients [93]. Similarly, ustekinumab has been shown to increase Faecalibacterium in responders compared with relapsing patients [94].
Diet is an important variable in the treatment of patients with CD. The administration of fermentable oligosaccharides, disaccharides, monosaccharides and polyols was observed to increase the levels of Clostridium cluster XIVa and Akkermansia municiphila, whereas that of Faecalibacterium prausnitzii was unaffected [95]. With the objective of evaluating differences in the microbial profiles associated with exclusive enteral nutrition and corticosteroid administration, the authors of one study observed a decrease in Prevotella, bifidobacteria, and Enterobacteriaceae in both assayed treatments [87]. Table 2 summarizes changes in gut microbiota associated with intestinal diseases and microbiota-targeted interventions for these diseases.

Table 2.
Microbiota changes associated with inflammatory bowel disease.
2.3. Non-Alcoholic Fatty Liver Disease
Non-alcoholic fatty liver disease (NAFLD) is characterized by an incremental increase in fat accumulation in the form of micro- and macrovacuoles of lipids in hepatocytes [96]. This disease is closely related to obesity and metabolic syndrome and exhibits features associated with abdominal obesity, insulin resistance, glucose intolerance, and T2D [97,98].
In addition to general overnutrition, specific diet factors, such as an elevated intake of sugar and/or fat, and genetic factors are crucial for its pathogenesis and progression [99]. Thus, an elevated body weight appears to not be the only risk factor associated with NAFLD [100], although alterations in gut microbiota have been reported as promoting the development of NAFLD by mediating inflammation, insulin resistance, bile acids, and choline metabolism [101]. Nevertheless, neither the pathophysiology of NAFLD nor the gut microbiota alterations in patients suffering for NAFLD have been completely characterized, and there are currently no effective drug therapies for NAFLD [102], with weight loss via diet and lifestyle modifications being the most successful care recommendations [103].
Host responses to changes in microbiota metabolism lead to an overwhelming presence of gut-derived microbial products and the activation of innate immunity and inflammation, resulting in the development of NAFLD. The presence of the dysbiotic microbiota and an altered intestinal barrier, possibly associated with disruption of tight-junctions (“leaky gut”), promotes the translocation of several bacterial products into the portal circulation. The interaction of bacterial products with toll-like receptors (TLRs) on the hepatic cell surface promotes the progression from simple steatosis to inflammation and fibrosis of the liver. For instance, the presence of H. pylori has been shown to induce gastric atrophy, with consequent acid losses predisposing small intestinal bacterial overgrowth, leaky gut, and portal endotoxin translocation [104].
Thus, interventions targeting the gut–liver axis through modulation of the gut microbiota by diet and/or pharmacological strategies appear to be a safe and sustainable tool to manage NAFLD. Manipulation of the gut microbiota has been primarily achieved through the use of probiotics, prebiotics, or symbiotic supplementation as well as through dietary intervention. For instance, very recently, Wang et al. reported that the alterations in the levels of bile acids (BAs) by gut bacteria through an isocaloric Mediterranean dietary intervention in 48 NAFLD patients did not alter fecal BA levels. Fecal BA levels and the microbiome do not appear to be responsible for the improvement in hepatic steatosis observed during dietary intervention in these patients [105]. In this respect and contrary to expectations, a study performed with 100 morbidly obese NAFLD patients undergoing laparoscopic sleeve gastrectomy surgery plus probiotic supplementation (Bio-25, Supherb, Israel) for six months showed higher abundances of Actinobacteria and Collinsella in the experimental group compared with the placebo. Although the alpha-diversity values never returned to the baseline levels after the intervention, an incremental improvement was observed at Month 6 compared with Month 0, and reductions were also observed at Month 12 vs. Month 6. However, the probiotic treatment did not have any added benefits beyond the influence of the surgery itself with respect to hepatic improvement, inflammatory or clinical outcomes [106]. Conversely, a prospective study that included 42 patients with NAFLD observed that those subjects receiving prebiotics supplements and lifestyle modification (i.e., diet and exercise) showed significant improvement in parameters related to liver function compared with those that only modified their lifestyle [102]. Similarly, prebiotic supplementation with inulin following metronidazole therapy plus a very low calorie diet was shown to ameliorate decreased liver function by reducing the levels of alanine aminotransferase in NAFLD subjects [107].
The potential benefits of probiotics administration have also been evaluated by Kobyliak et al., who reported that the co-administration of a probiotic with omega-3 in T2D patients with NAFLD improved serum lipid levels and metabolic profiles and reduced liver fat levels and the chronic systemic inflammatory state [108]. Along this line, the same group showed similar results after the administration of a concentrated biomass of 14 probiotic bacterial genera (including Bifidobacterium, Lactobacillus, Lactococcus, and Propionibacterium) [108]. However, in the abovementioned studies, details regarding the changes in gut microbiota associated with the probiotics/prebiotics supplementation were not described. In contrast, Ahn et al. [109] performed a study in which 68 NAFLD obese patients were randomly divided into prebiotic (receiving a mixture of probiotics Lactobacillus acidophilus, Lactobacillus rhamnosus, Lactobacillus paracasei, Pediococcus pentosaceus, Bifidobacterium lactis, and Bifidobacterium breve) or placebo groups. In the probiotics group, but not in the control group, a significant increase in all strains except Lactobacillus paracasei was observed as well as higher ratio Bacteroidetes/Firmicutes. Those patients that presented fatty liver improvement also showed increased abundances of Ruminococcaceae-2, Lachnospiraceae-2, Coprococcus, Lachnospiraceae-1, Ruminococcus, and Dorea. Additionally, patients with successful weight loss presented increases in the abundances of Ruminococcaceae-2, Ruminococcaceae-1, Clostridiales-2, Lachnospiraceae-2, and Coprococcus. All of these changes were associated with fatty liver improvements.
The activation of inflammation due to both dysbiosis and alterations in intestinal permeability via TLRs signaling in hepatocytes induces progression from simple steatosis to non-alcoholic steato-hepatitis (NASH). This progression in dysbiotic NASH patients may be explained by the cytotoxicity associated with the increase in primary fecal BA, the primary/secondary fecal BA ratio, and plasma and hepatic BA concentrations [104]. Specifically, for NASH patients, therapies based on prebiotic supplementation consisting of the ingestion of oligofructose for 24 weeks improved liver histology independently of body weight. In addition, oligofructose increased the abundance of Bifidobacterium, the incremental increase in which inversely associated with obesity and plasma LPS [110]. Moreover, a 12-week treatment with a probiotic cocktail significantly altered the microbial structure in the feces of the experimental group, increasing towards the normal range without significantly altering the abundances of pathogenic enterobacteria and causing a significant decrease in liver inflammation without other adverse events [111].
Similarly, Alisi et al. reported changes in gut microbiota of obese children associated with the improvement of steatohepatitis and BMI after supplementation with VSL#3, a mixture of eight probiotic strains (Streptococcus thermophilus, bifidobacteria (B. breve, B. infantis, and B. longum), Lactobacillus acidophilus, L. plantarum, L. paracasei, and L. delbrueckii subsp. bulgaricus) for four months [112]. In contrast, synbiotic supplementation only had positive effects on anthropometric parameters associated with NASH (BMI, waist circumference, or uric acid levels) but not on gut permeability [113]. Most authors have concluded that modulation of gut microbiota brings added benefits by itself or in combination with other interventions (i.e., diet, exercise, or bariatric surgery) to NAFLD patients [106,108,114]. However, the heterogeneity of strategies (the use of pre- or probiotics, the length of treatment, the grade of steatosis, the presence of inflammation, etc.) makes it mandatory to understand and completely define what the precise changes causing the imbalance are and/or the changes in gut microbiota that lead from hepatic health to fatty liver disease. Along this line, evidence that the gut microbiota is altered in pediatric NAFLD and obese patients was investigated using targeted metagenomics and metabolomics (Table 3). The authors concluded that the combination of a low abundance of Oscillospira with high levels of 2-butanone may be a specific intestinal metagenomic and metabolomic profile for liver steatosis in children. Moreover, pediatric NASH patients presented high relative abundances of Lachnospiraceae, Ruminococcus, and Dorea. In addition, Blautia was extremely high in NASH but not in NAFLD or obese children, while Oscillospira was significantly less abundant in the NAFLD, NASH, and obese patients compared with controls, suggesting that changes in the gut microbiota are associated with the severity of fatty liver disease [45]. In this regard, differences in the microbiota of adult patients with biopsy-proven NAFLD, HCC, and fibrosis were also assessed. In this study, the composition of gut bacteria was determined by sequencing of 16S rRNA gene, the results of which showed decreased abundances of Bacteroidetes and Faecalibacterium prausnitzii in NAFLD patients compared with the healthy controls.

Table 3.
Microbiota changes associated with non-alcoholic fatty liver disease.
Furthermore, endotoxin levels were observed to be higher in NAFLD patients with severe fibrosis than in those with mild fibrosis, revealing that the mechanism of fibrotic progression via the endotoxin in NAFLD may be strongly associated with gut permeability. The latest results suggest that the distinct composition of the gut microbiota among NAFLD and HCC patients may offer a target for intervention or a marker for disease [115]. In addition, a recent study by Lelouvier et al. attempted to associate microbiota dysbiosis from blood and feces with LF and BMI > 40 kg/m2 patients in two different geographic populations. Only in one of the studied populations, specific bacterial taxonomic profiles and their functions were associated with LF. This finding suggests that there is a potential interaction between bacterial communities and an environmental factor specific to the geographical area in obese patients [116].
In summary, available studies use a large variety of interventions (probiotic, prebiotics, symbiotic supplementation, diet, or other approaches) and different patient baselines and treatment times (from weeks to months), making it difficult to draw clear conclusions and highlighting the need for further research in this field. From our point of view, the analysis of the gut microbiota under unhealthy conditions remains to be defined, as well as how it is affected by diet, BMI, or age as well as others factors, such as the grade of steatosis and geographical environment.
2.4. Insulin Resistance Syndrome
Insulin resistance syndrome (IRS) or metabolic syndrome is characterized by hyperinsulinemia and an increased prevalence of obesity, hypertension, dyslipidemia, and T2D [117]. NAFLD is the hepatic component of IRS [97].
IRS is currently a global burden, with 20–25% of the adult population worldwide suffering from this disease [118]. In addition to genetic, lifestyle, and nutritional factors [119,120,121], alterations in gut microbiota (dysbiosis) are also associated with IRS status [118,122,123,124]. The primary contributors to the development of IRS are as follows: LPS from the gut microbiota, which can induce a chronic subclinical inflammatory process through activation of TLR4; a reduction in circulating SCFA; alterations in the secretion of glucagon-like peptide-1 (GLP-1) mediated by bile acids; and an increased circulation of branched-chain amino acids (BCAA) [125].
The results of a metagenomic analysis performed with 58 IRS patients from Kazakhstan revealed differences in gut microbiota composition of these patients compared with the healthy controls (e.g., significant reductions in the Firmicutes/Bacteroidetes ratio, bifidobacteria, and Subdoligranulum and increased Prevotella). The results of this study disagree with previously published studies that were primarily performed with European subjects in whom an increased Firmicutes/Bacteroidetes ratio was observed, a discrepancy that is probably due to the dietary, lifestyle, and geographical features of the Kazakh population [126].
With the purpose of reversing the different IRS-associated changes that occur in the gut microbiota and reduce the impact of this disease, different approaches are being investigated to increase the therapeutic strategies available to combat this syndrome (Table 4). Among these strategies are dietary changes, nutritional supplementation, and the administration of prebiotics, probiotics, FMT [81,127,128,129,130,131,132,133], and antibiotics [134]. Haro et al. evaluated the consumption of both a Mediterranean and a low-fat diet in 138 IRS patients for two years. Firstly, they observed higher abundances of the genera Bacteroides, Eubacterium, and Lactobacillus and reduced abundances of B. fragilis group members, P. distasonis, B. thetaiotaomicron, F. prausnitzii, F. nucleatum, B. longum, B. adolescentis, the R. flavefaciens subgroup, and E. rectale in IRS patients at time 0. After the intervention, only the Mediterranean diet had induced significant changes (increased abundances of P. distasonis, B. thetaiotaomicron, F. prausnitzii, B. adolescentis, and B. longum) in IRS patients, suggesting that, although IRS persisted after the intervention, a Mediterranean diet may contribute to partially restoring the beneficial components of the gut microbiota [135]. These authors also performed a similar study (an evaluation of the effect of Mediterranean and low-fat diet for two years) in 33 obese patients with severe IRS versus 32 obese patients that did not fulfill IRS criteria and 41 normal-weight subjects. In this study, they observed a significant decrease in the Firmicutes/Bacteroidetes ratio after the consumption of the low-fat diet, but only a decreasing trend for the Mediterranean diet. Additionally, the abundances of the genera Bacteroides, Prevotella, and Faecalibacterium increased in IRS patients for both diets, Mediterranean and low-fat, and reduced abundances of the genera Streptococcus and Clostridium were also observed in the low-fat diet group. In contrast, the group that received the Mediterranean diet did not exhibit changes in the abundance of these genera but did exhibit increases in those of the genera Roseburia and Ruminococcus and in the bacterial species P. distasonis and F. prausnitzii. Regarding the effect of these diets on metabolic parameters, the triglyceride levels decreased in the IRS group and remained unchanged in the other groups. These results suggest that the chronic intake of Mediterranean or low-fat diets partially ameliorates the gut microbiome dysbiosis in severe IRS patients but that this improvement depends on the degree of the metabolic dysfunction [136].

Table 4.
Microbiota changes associated with insulin resistance syndrome.
Salonen et al. reported on changes in the gut microbiota of 14 IRS patients fed four different diets in a sequential fashion for 10 weeks as follows: (1) a standard diet for weight maintenance (M) for one week; (2) a diet with a high content of type 3 resistant starch (RS) for three weeks; (3) a diet with a high content of non-starch polysaccharides (NSP) for three weeks; and, (4) a weight loss diet with high protein and medium carbohydrate levels (WL) for another two weeks. The main results after the different diet interventions were as follows: the RS diet increased Ruminococcus-related bacterial phylotypes and the microbiota diversity, the NSP diet primarily led to increases in the abundance of Lachnospiraceae, and the WL diet reduced the bifidobacteria. Notably, the dietary responsiveness of the microbiota varied substantially among patients and was inversely associated with their diversity. This result suggests that individuals can be stratified into responders and non-responders based on the features of their intestinal microbiota. A positive correlation between fecal bifidobacteria and plasma insulin was also detected in IRS patients [137].
Another strategy to modulate the gut microbiota composition through nutritional supplementation in IRS was described in a study by Moreno-Indias et al. performed with 10 IRS patients. The intake of two red wines of equal composition except for the ethanol content (regular and ethanol-free red wines) for 30 days significantly increased the number of fecal bifidobacteria, Lactobacillus and butyrate-producing bacteria (Faecalibacterium prausnitzii and Roseburia) at the expense of less desirable groups of bacteria, such as LPS producers (Escherichia coli and Enterobacter cloacae). These changes were attributed to the polyphenols present in both wines, and were associated with a reduction in IRS risk markers, triglycerides, HDL-cholesterol, glucose, and glutamic pyruvic transaminase [138]. A study by Ni et al. observed higher proportions of Lactobacillus and Bifidobacterium and reduced abundances of Anaerostipes, Coprococcus, and Ruminococcus in 12 elderly IRS patients compared with healthy individuals of the same age. A four-week treatment with Yangyin Tiluo Decoction (a Chinese Herbal Formula) decreased the abundance of the order Bacteroidales. Lipoprotein(a) levels were also reduced in plasma in correlation with the abundance of Acinetobacter species [139].
Whole grain has been described to increase the abundances of Faecalibacterium prausnitzii, Prevotella copri, and Clostridiales and decrease those of Bacteroides thetaiotaomicron compared with a refined grain diet. Whole grain ingestion reduced body weight and systemic low-grade inflammation but did not alter insulin sensitivity [140].
With respect to interventions with prebiotics and probiotics, the intake of barley β-glucans for four weeks by 27 IRS patients induced an increase in the abundance of Agathobacter rectalis and decreased that of Coriobacteriales and Clostridiales and was associated with a reduction in total plasma cholesterol [141]. In a 12-week study, Stadlbauer et al. evaluated the effect of Lactobacillus casei Shirota in 28 IRS patients (13 patients took β-glucans and 15 did not). Despite inducing an increase in the abundances of Parabacteroides and Lactobacillus casei Shirota, the supplementation did not restore gut microbiota composition, the gut barrier or the Bacteroidetes/Firmicutes ratio [142].
Smits et al. tested an FMT-based strategy in 20 patients with IRS [134], where patients received feces from vegan donors and another 10 patients received their own feces. Feces from vegans induced subtle changes, such as increased abundances of several microbial groups belonging to the Lachnospiraceae family, including bacteria related to Bryantella formatexigens and Megamonas hypermegale as well as L. bovis two weeks after transplantation. However, FMT did not induce any improvement in parameters related to vascular inflammation. Furthermore, no changes occurred in the group of patients that was subjected to autologous transplantation [134].
With regard to the use of antibiotics, the effect of oral vancomycin treatment (500 mg for seven days) on the gut microbiota in 10 IRS patients was described by Vrieze et al. [143]. The primary findings of this study were a reduction in fecal microbial diversity, a decrease in Gram-positive bacteria (especially Firmicutes) with a concomitant increment in Gram-negative bacteria (especially Proteobacteria). Vancomycin also decreased the levels of fecal secondary bile acids with a simultaneous postprandial increase in primary circulating bile acids, which was associated with an altered abundance of Firmicutes. With respect to metabolic effects, the administration of vancomycin reduced peripheral insulin sensitivity [143].
Overall, findings supporting the beneficial effects of various interventions (diet, prebiotics, probiotics, FMT, etc.) on gut microbiota as a strategy to treat IRS appear to be scarce and modest. In addition, the persistence of these changes remains to be evaluated. In our view, both the number of studies and enrolled patients are small, and the inter-individual variation between patients should also be considered. Consequently, further research in this area of study is required to address the aforementioned weaknesses.
2.5. Diabetes Mellitus Type II
Over 60 million people are diagnosed with T2D in Europe [144]. According to WHO, diabetes will be the seventh leading cause of death in 2030 [145]. Factors such as a healthy diet, regular physical activity, and body weight maintenance at normal levels prevent T2D development, delay its progression, and help mitigate its pathological consequences. Prediabetes and high blood glucose levels maintained over time without symptomatology are associated with overweight and obesity. Because prediabetes leads to the development of diabetes in 70% of cases, a prevention strategy is highly recommended in this state. T2D could be linked to the gut microbiota composition, and it is directly responsible for the induction of a low-grade inflammatory state. The composition of the gut microbiota also plays a significant role in the development of pre-diabetes conditions such as resistance to insulin. The main alterations in gut microbiota associated with T2D have a significantly lower prevalence of Firmicutes and enrichment of Bacteroidetes and Proteobacteria [140].
Growing evidence indicates that functional foods, such as prebiotics, probiotics, and postbiotics, may be used in the prevention and treatment of diabetes [146,147,148]. Table 5 summarizes microbiota-targeted interventions for T2D.

Table 5.
Microbiota changes associated with diabetes mellitus type II.
Totum63 is an association of plant principles for the treatment of T2D and prediabetes. In db/db mice, a well-established obesity-induced animal model of TD2 in which mice carry a mutation in the leptin receptor gene, Totum63 delayed the defect in insulin secretion, suggesting a protective role on pancreatic beta-cells. Furthermore, in a phase I/II clinical trial Totum63 administration also induced improvements in glucose levels and insulin response to a standardized breakfast in healthy participants. Moreover, Totum63 treatment has been described to prevent the dysbiosis induced by a high fat diet [149].
Metformin is the most frequent treatment used for T2D, and this drug has been reported to significantly decrease the abundance of opportunistic bacteria [150] and significantly increase that of beneficial bacteria, particularly Blautia spp. Moreover, a Chinese herbal formula has been reported to increase Blautia and Faecalibacterium abundances in an open label study in T2D patients. Both treatments were linked with diabetes improvements with respect to glucose levels, a homeostasis model assessment of insulin resistance (HOMA-IR), and lipid homeostasis [151].
Canfora et al. studied the effects of GOS in 44 obese/overweight pediatric individuals [152]. The supplementation of diets with GOS increased the abundance of Bifidobacterium species in feces, while the microbial richness and diversity in fecal samples was unaffected. No differences in fecal or fasting plasma SCFA concentrations or in the systemic concentrations of gut-derived hormones, incretins, LPS-binding protein, or other markers of inflammation were observed. In addition, no significant alterations in peripheral and adipose tissue insulin sensitivity, body composition, or energy and substrate metabolism were observed [152].
The microbiome may also be involved in determining how glycemic control impacts weight change for different diets. The administration of SCFAs has been reported to result in a wide range of health benefits, including improvements in blood lipid profiles, glucose homeostasis, and body composition as well as in reduced body weight [148]. High butyrate levels were observed to be associated with beneficial effects on insulin sensitivity and glucose levels [153,154]. A randomized, double-blind study was carried out with 81 T2D patients divided into two three-month treatment groups one group took a placebo, while the other group followed a reduced-energy diet comprising high-fiber, polyphenol-rich, and vegetable-protein functional foods. Intervention with the last diet significantly modified fecal microbiota compared with the placebo, increasing the abundances of F. prausnitzii and A. muciniphila and decreasing that of P. copri.
SCFAs are produced by various human gut microbes. SCFAs act as an energy source to the colonic epithelium and are also sensed by host signaling pathways that modulate appetite and inflammation. A deficiency in gut SCFAs is associated with type 2 diabetes. Zhao et al. observed that a high-fiber diet promoted the growth of SCFA-producing organisms in diabetic humans [147,155]. Furthermore, the high-fiber diet induced changes in the entire gut microbe community and correlated with elevated levels of glucagon-like peptide-1, a decrease in acetylated hemoglobin levels, and improved blood–glucose regulation [147,155].
In a clinical trial with T2D patients, the relative abundance of A. muciniphila, a mucin-associated bacterium with anti-inflammatory properties, increased after inulin and sodium butyrate prebiotic supplementation [156,157,158].
A type of black tea from China contains some polysaccharides and polyphenols that improve glucose and lipid metabolism in prediabetic subjects by modifying the gut microbiota (decreasing Faecalibacterium and Oscillospira). In particular, GLP-1 and GLP-2 serum levels increased after 12 weeks in the group of patients that drank the tea compared with those who received a placebo [159].
Another dietary resistant starch-based prebiotic has been shown to positively affect metabolic health markers. In humans, resistant starch fermentation may enhance the abundances of specific gut microbiota taxa, such as Bifidobacterium, Ruminococcus bromii, and Eubacterium rectale, with associated improvements in insulin response, body fat reserve, and cholesterol levels [160]. Transglucosidase (TGD) produces oligosaccharides from starch in the human intestinal tract. Shimozato et al. observed that TGD administration to patients with T2D for 12 weeks significantly affected the fecal microbiota content, increasing Clostridium cluster IV, Clostridium subcluster XIVa, Clostridium cluster XVIII, and fecal pH and decreasing the order Lactobacillales in patients with bowel movement disorder compared with the controls. The TGD treatment significantly improved bowel movements compared with the placebo treatment, and this effect was not observed in patients without bowel movement disorder [161].
Several lines of clinical evidence suggest that the use of probiotics as alternative therapeutics against T2D increases the quality of life [146,162,163,164]. Some clinical trials were reviewed to confirm the beneficial effects and deepen understanding of the underlying mechanisms [165], resulting in a combination of life-style intervention and probiotic supplementation in prediabetic adolescents being proposed [163]. The efficacy of probiotics appears to depend on the bacterial strain and the study outcome. Lactobacillus casei strain Shirota-fermented milk was investigated in T2D pathologies, the results of which confirmed that this probiotic reduced bacterial translocation in Japanese individuals with T2D [146]. Furthermore, the administration of Lactobacillus reuteri to T2D patients has been shown to improve insulin sensitivity [162] (Table 5). As for postbiotics, defined as secreted factors, cellular components and metabolites of bacteria that, upon delivery to a host, may exert biological effects on it. Microbial components from live or dead bacteria can also be considered a source of postbiotics. Examples of postbiotics such as microbial-derived short-chain fatty acids or flavonoids may directly influence host feeding behavior, energy metabolism, insulin secretion, and insulin sensitivity. These postbiotic properties should be considered in obesity-related metabolic disease and represent a promising alternative to current treatments [166].
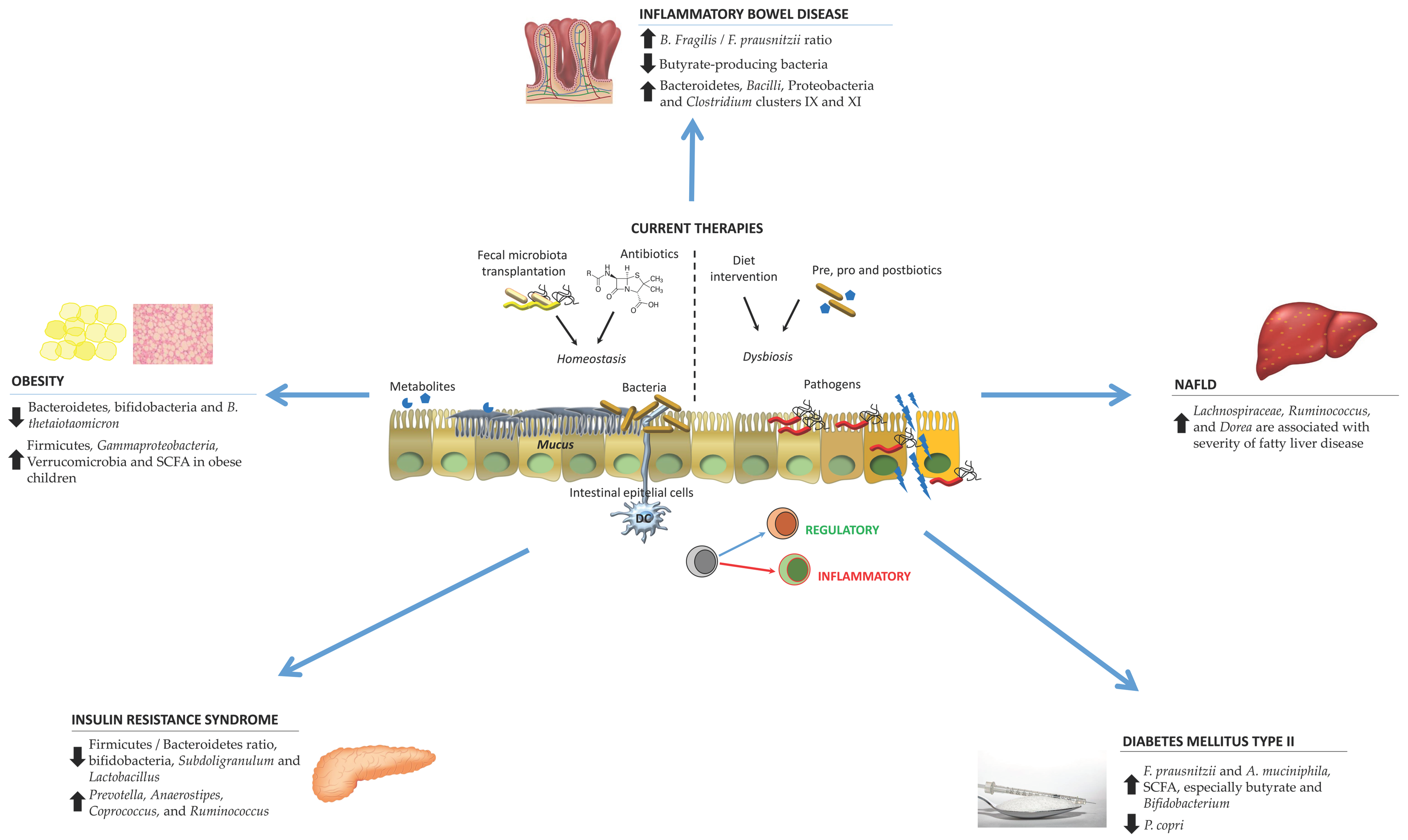
Figure 1 summarizes the microbiota alterations in particular pathologies as well as current therapies.
Figure 1.
Schematic representation of the microbiota alterations in various disorders and the current therapies to counteract their effects. Abbreviations. DC: dendritic cells; NAFLD: non-alcoholic fatty liver disease; SCFA: short-chain fatty acids.
3. Future Directions
Maintaining a healthy gut ecosystem is essential, and alterations in the diversity, quantity, and stability of a healthy microbiota has been associated with obesity, IBD, and IRS as well as other metabolic disorders such as NAFLD and T2D. Since the prevalence of these diseases continues to rise worldwide, the modulation of the gut microbiota through various interventions (e.g., prebiotics, probiotics, and FMT) is a research area of increasing interest given their potential as tools to prevent or treat these diseases. However, conflicting results and limited efficacy have been reported to date. The different durations and doses of the interventions, as well as the technology used for sequencing the microbiome makes comparisons among studies difficult. 16S approaches for microbiome analyses are well suited for analysis of a large number of samples, i.e., multiple patients, longitudinal studies, and so on. However, this approach offers limited taxonomical and functional resolution. Moreover, it should be pointed out that using primers for different regions of the 16S rRNA gene may lead to discordant results due not only to the distinct binding affinities for the corresponding flanking conserved regions, but also due to the resolution of each variable region across taxa [167]. In contrast, shotgun metagenomics is usually more expensive but offers increased resolution, enabling a more specific taxonomic and functional classification of sequences, as well as the discovery of new bacterial genes and genomes [168].
FMT, which has yielded promising results for treating certain disorders, such as Clostridium difficile infections, is a clear example of this issue and should be addressed in future studies. Stool donor preparation and the transplantation method (endoscopy, nasointestinal tubes, or capsules) must be standardized, and the long-term potential adverse effects should be studied in greater detail.
Among the next-generation beneficial microbes that have been identified, Akkermansia muciniphila and Faecalibacterium prausnitzii are promising candidates. A. muciniphila has been reported to be inversely associated with obesity, diabetes, cardiometabolic diseases, and low-grade inflammation. Besides the numerous correlations observed, a large body of evidence has demonstrated the causal beneficial impact of this bacterium in a variety of preclinical models [169]. Many studies have shown that F. prausnitzii abundance is reduced in different intestinal disorders. It has been proposed that F. prausnitzii monitoring may therefore serve as a biomarker to assist in gut diseases diagnostics [170]. Testing these strategies in many more clinical trials and with great numbers of human subjects is clearly needed.
Author Contributions
All authors participated in the bibliographic search, discussion, and writing of the manuscript.
Funding
Ongoing research is funded by grant PI-0538-2017, Junta de Andalucía, Spain (to L.F.).
Acknowledgments
Julio Plaza-Diaz is part of the Unit of Excellence on Exercise and Health (UCEES), Units of Excellence, Excellence actions, Plan Propio de Investigación 2016, University of Granada.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Gilbert, J.A.; Krajmalnik-Brown, R.; Porazinska, D.L.; Weiss, S.J.; Knight, R. Toward effective probiotics for autism and other neurodevelopmental disorders. Cell 2013, 155, 1446–1448. [Google Scholar] [CrossRef] [PubMed]
- Plaza-Diaz, J.; Ruiz-Ojeda, F.J.; Vilchez-Padial, L.M.; Gil, A. Evidence of the Anti-Inflammatory Effects of Probiotics and Synbiotics in Intestinal Chronic Diseases. Nutrients 2017, 9, 555. [Google Scholar] [CrossRef] [PubMed]
- Plaza-Diaz, J.; Ruiz-Ojeda, F.J.; Gil-Campos, M.; Gil, A. Immune-Mediated Mechanisms of Action of Probiotics and Synbiotics in Treating Pediatric Intestinal Diseases. Nutrients 2018, 10, 42. [Google Scholar] [CrossRef] [PubMed]
- Plaza-Diaz, J.; Ruiz-Ojeda, F.J.; Gil-Campos, M.; Gil, A. Mechanisms of action of probiotics. Adv. Nutr. 2019, 10, S49–S66. [Google Scholar] [CrossRef] [PubMed]
- Bermudez-Brito, M.; Plaza-Diaz, J.; Munoz-Quezada, S.; Gomez-Llorente, C.; Gil, A. Probiotic mechanisms of action. Ann. Nutr. Metab. 2012, 61, 160–174. [Google Scholar] [CrossRef] [PubMed]
- Belkaid, Y.; Hand, T.W. Role of the microbiota in immunity and inflammation. Cell 2014, 157, 121–141. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Ashida, H.; Ogawa, M.; Yoshikawa, Y.; Mimuro, H.; Sasakawa, C. Bacterial interactions with the host epithelium. Cell Host Microbe 2010, 8, 20–35. [Google Scholar] [CrossRef] [PubMed]
- Ghadimi, D.; Vrese, M.; Heller, K.J.; Schrezenmeir, J. Effect of natural commensal-origin DNA on toll-like receptor 9 (TLR9) signaling cascade, chemokine IL-8 expression, and barrier integritiy of polarized intestinal epithelial cells. Inflamm. Bowel Dis. 2010, 16, 410–427. [Google Scholar] [CrossRef] [PubMed]
- Artis, D. Epithelial-cell recognition of commensal bacteria and maintenance of immune homeostasis in the gut. Nat. Rev. Immunol. 2008, 8, 411–420. [Google Scholar] [CrossRef] [PubMed]
- Zhernakova, A.; Kurilshikov, A.; Bonder, M.J.; Tigchelaar, E.F.; Schirmer, M.; Vatanen, T.; Mujagic, Z.; Vila, A.V.; Falony, G.; Vieira-Silva, S.; et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 2016, 352, 565–569. [Google Scholar] [CrossRef] [PubMed]
- Falony, G.; Joossens, M.; Vieira-Silva, S.; Wang, J.; Darzi, Y.; Faust, K.; Kurilshikov, A.; Bonder, M.J.; Valles-Colomer, M.; Vandeputte, D.; et al. Population-level analysis of gut microbiome variation. Science 2016, 352, 560–564. [Google Scholar] [CrossRef] [PubMed]
- Backhed, F.; Roswall, J.; Peng, Y.; Feng, Q.; Jia, H.; Kovatcheva-Datchary, P.; Li, Y.; Xia, Y.; Xie, H.; Zhong, H.; et al. Dynamics and Stabilization of the Human Gut Microbiome during the First Year of Life. Cell Host Microbe 2015, 17, 690–703. [Google Scholar] [CrossRef] [PubMed]
- Goodrich, J.K.; Waters, J.L.; Poole, A.C.; Sutter, J.L.; Koren, O.; Blekhman, R.; Beaumont, M.; Van Treuren, W.; Knight, R.; Bell, J.T.; et al. Human genetics shape the gut microbiome. Cell 2014, 159, 789–799. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Charbonnier, L.M.; Noval Rivas, M.; Georgiev, P.; Li, N.; Gerber, G.; Bry, L.; Chatila, T.A. MyD88 Adaptor-Dependent Microbial Sensing by Regulatory T Cells Promotes Mucosal Tolerance and Enforces Commensalism. Immunity 2015, 43, 289–303. [Google Scholar] [CrossRef] [PubMed]
- David, L.A.; Maurice, C.F.; Carmody, R.N.; Gootenberg, D.B.; Button, J.E.; Wolfe, B.E.; Ling, A.V.; Devlin, A.S.; Varma, Y.; Fischbach, M.A.; et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 2014, 505, 559–563. [Google Scholar] [CrossRef] [PubMed]
- Cho, I.; Yamanishi, S.; Cox, L.; Methe, B.A.; Zavadil, J.; Li, K.; Gao, Z.; Mahana, D.; Raju, K.; Teitler, I.; et al. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature 2012, 488, 621–626. [Google Scholar] [CrossRef] [PubMed]
- Maurice, C.F.; Haiser, H.J.; Turnbaugh, P.J. Xenobiotics shape the physiology and gene expression of the active human gut microbiome. Cell 2013, 152, 39–50. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, A.; Ahmed, A.M.; Subramanian, S.; Griffin, N.W.; Drewry, L.L.; Petri, W.A., Jr.; Haque, R.; Ahmed, T.; Gordon, J.I. Members of the human gut microbiota involved in recovery from Vibrio cholerae infection. Nature 2014, 515, 423–426. [Google Scholar] [CrossRef] [PubMed]
- Thaiss, C.A.; Zeevi, D.; Levy, M.; Zilberman-Schapira, G.; Suez, J.; Tengeler, A.C.; Abramson, L.; Katz, M.N.; Korem, T.; Zmora, N.; et al. Transkingdom control of microbiota diurnal oscillations promotes metabolic homeostasis. Cell 2014, 159, 514–529. [Google Scholar] [CrossRef] [PubMed]
- Fujimura, K.E.; Demoor, T.; Rauch, M.; Faruqi, A.A.; Jang, S.; Johnson, C.C.; Boushey, H.A.; Zoratti, E.; Ownby, D.; Lukacs, N.W.; et al. House dust exposure mediates gut microbiome Lactobacillus enrichment and airway immune defense against allergens and virus infection. Proc. Natl. Acad. Sci. USA 2014, 111, 805–810. [Google Scholar] [CrossRef] [PubMed]
- Lynch, S.V.; Pedersen, O. The Human Intestinal Microbiome in Health and Disease. N. Engl. J. Med. 2016, 375, 2369–2379. [Google Scholar] [CrossRef] [PubMed]
- Aron-Wisnewsky, J.; Prifti, E.; Belda, E.; Ichou, F.; Kayser, B.D.; Dao, M.C.; Verger, E.O.; Hedjazi, L.; Bouillot, J.L.; Chevallier, J.M.; et al. Major microbiota dysbiosis in severe obesity: Fate after bariatric surgery. Gut 2019, 68, 70–82. [Google Scholar] [CrossRef] [PubMed]
- Plaza-Diaz, J.; Robles-Sanchez, C.; Abadia-Molina, F.; Moron-Calvente, V.; Saez-Lara, M.J.; Ruiz-Bravo, A.; Jimenez-Valera, M.; Gil, A.; Gomez-Llorente, C.; Fontana, L. Adamdec1, Ednrb and Ptgs1/Cox1, inflammation genes upregulated in the intestinal mucosa of obese rats, are downregulated by three probiotic strains. Sci. Rep. 2017, 7, 1939. [Google Scholar] [CrossRef] [PubMed]
- Pineiro, M.; Asp, N.G.; Reid, G.; Macfarlane, S.; Morelli, L.; Brunser, O.; Tuohy, K. FAO Technical meeting on prebiotics. J. Clin. Gastroenterol. 2008, 42 Pt 2 (Suppl. 3), S156–S159. [Google Scholar] [CrossRef]
- Saez-Lara, M.J.; Robles-Sanchez, C.; Ruiz-Ojeda, F.J.; Plaza-Diaz, J.; Gil, A. Effects of Probiotics and Synbiotics on Obesity, Insulin Resistance Syndrome, Type 2 Diabetes and Non-Alcoholic Fatty Liver Disease: A Review of Human Clinical Trials. Int. J. Mol. Sci. 2016, 17, 928. [Google Scholar] [CrossRef] [PubMed]
- Rinninella, E.; Raoul, P.; Cintoni, M.; Franceschi, F.; Miggiano, G.A.D.; Gasbarrini, A.; Mele, M.C. What is the Healthy Gut Microbiota Composition? A Changing Ecosystem across Age, Environment, Diet, and Diseases. Microorganisms 2019, 7, 14. [Google Scholar] [CrossRef] [PubMed]
- Bischoff, S.C.; Boirie, Y.; Cederholm, T.; Chourdakis, M.; Cuerda, C.; Delzenne, N.M.; Deutz, N.E.; Fouque, D.; Genton, L.; Gil, C.; et al. Towards a multidisciplinary approach to understand and manage obesity and related diseases. Clin. Nutr. 2017, 36, 917–938. [Google Scholar] [CrossRef] [PubMed]
- Engin, A. The Definition and Prevalence of Obesity and Metabolic Syndrome. Adv. Exp. Med. Biol. 2017, 960, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Gerard, P. Gut microbiota and obesity. Cell Mol. Life Sci. 2016, 73, 147–162. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.J.; Gerasimidis, K.; Edwards, C.A.; Shaikh, M.G. Role of Gut Microbiota in the Aetiology of Obesity: Proposed Mechanisms and Review of the Literature. J. Obes. 2016, 2016, 7353642. [Google Scholar] [CrossRef] [PubMed]
- Turta, O.; Rautava, S. Antibiotics, obesity and the link to microbes—What are we doing to our children? BMC Med. 2016, 14, 57. [Google Scholar] [CrossRef] [PubMed]
- Trasande, L.; Blustein, J.; Liu, M.; Corwin, E.; Cox, L.M.; Blaser, M.J. Infant antibiotic exposures and early-life body mass. Int. J. Obes. 2013, 37, 16–23. [Google Scholar] [CrossRef] [PubMed]
- Ajslev, T.A.; Andersen, C.S.; Gamborg, M.; Sorensen, T.I.; Jess, T. Childhood overweight after establishment of the gut microbiota: The role of delivery mode, pre-pregnancy weight and early administration of antibiotics. Int. J. Obes. 2011, 35, 522–529. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Hamady, M.; Yatsunenko, T.; Cantarel, B.L.; Duncan, A.; Ley, R.E.; Sogin, M.L.; Jones, W.J.; Roe, B.A.; Affourtit, J.P.; et al. A core gut microbiome in obese and lean twins. Nature 2009, 457, 480–484. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Microbial ecology: Human gut microbes associated with obesity. Nature 2006, 444, 1022–1023. [Google Scholar] [CrossRef] [PubMed]
- Ridaura, V.K.; Faith, J.J.; Rey, F.E.; Cheng, J.; Duncan, A.E.; Kau, A.L.; Griffin, N.W.; Lombard, V.; Henrissat, B.; Bain, J.R.; et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 2013, 341, 1241214. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Backhed, F.; Fulton, L.; Gordon, J.I. Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 2008, 3, 213–223. [Google Scholar] [CrossRef] [PubMed]
- Yallapragada, S.G.; Nash, C.B.; Robinson, D.T. Early-Life Exposure to Antibiotics, Alterations in the Intestinal Microbiome, and Risk of Metabolic Disease in Children and Adults. Pediatr. Ann. 2015, 44, e265–e269. [Google Scholar] [CrossRef] [PubMed]
- Reinhardt, C.; Reigstad, C.S.; Bäckhed, F. Intestinal microbiota during infancy and its implications for obesity. J. Pediatr. Gastroenterol. Nutr. 2009, 48, 249–256. [Google Scholar] [CrossRef] [PubMed]
- Ignacio, A.; Fernandes, M.; Rodrigues, V.; Groppo, F.; Cardoso, A.; Avila-Campos, M.; Nakano, V. Correlation between body mass index and faecal microbiota from children. Clin. Microbiol. Infect. 2016, 22, 258. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, B.S.; Pollak, J.; Bailey-Davis, L.; Hirsch, A.G.; Cosgrove, S.E.; Nau, C.; Kress, A.M.; Glass, T.A.; Bandeen-Roche, K. Antibiotic use and childhood body mass index trajectory. Int. J. Obes. 2016, 40, 615–621. [Google Scholar] [CrossRef] [PubMed]
- Bailey, L.C.; Forrest, C.B.; Zhang, P.; Richards, T.M.; Livshits, A.; DeRusso, P.A. Association of antibiotics in infancy with early childhood obesity. JAMA Pediatr. 2014, 168, 1063–1069. [Google Scholar] [CrossRef] [PubMed]
- Murphy, R.; Stewart, A.W.; Braithwaite, I.; Beasley, R.; Hancox, R.J.; Mitchell, E.A.; Group, I.P.T.S. Antibiotic treatment during infancy and increased body mass index in boys: An international cross-sectional study. Int. J. Obes. 2014, 38, 1115–1119. [Google Scholar] [CrossRef] [PubMed]
- Riva, A.; Borgo, F.; Lassandro, C.; Verduci, E.; Morace, G.; Borghi, E.; Berry, D. Pediatric obesity is associated with an altered gut microbiota and discordant shifts in Firmicutes populations. Environ. Microbiol. 2017, 19, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Del Chierico, F.; Nobili, V.; Vernocchi, P.; Russo, A.; Stefanis, C.; Gnani, D.; Furlanello, C.; Zandona, A.; Paci, P.; Capuani, G.; et al. Gut microbiota profiling of pediatric nonalcoholic fatty liver disease and obese patients unveiled by an integrated meta-omics-based approach. Hepatology 2017, 65, 451–464. [Google Scholar] [CrossRef] [PubMed]
- Murugesan, S.; Nirmalkar, K.; Hoyo-Vadillo, C.; Garcia-Espitia, M.; Ramirez-Sanchez, D.; Garcia-Mena, J. Gut microbiome production of short-chain fatty acids and obesity in children. Eur. J. Clin. Microbiol. Infect. Dis. 2018, 37, 621–625. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Fan, C.; Li, P.; Lu, Y.; Chang, X.; Qi, K. Short Chain Fatty Acids Prevent High-fat-diet-induced Obesity in Mice by Regulating G Protein-coupled Receptors and Gut Microbiota. Sci. Rep. 2016, 6, 37589. [Google Scholar] [CrossRef] [PubMed]
- Nicolucci, A.C.; Hume, M.P.; Martinez, I.; Mayengbam, S.; Walter, J.; Reimer, R.A. Prebiotics Reduce Body Fat and Alter Intestinal Microbiota in Children Who Are Overweight or With Obesity. Gastroenterology 2017, 153, 711–722. [Google Scholar] [CrossRef] [PubMed]
- Sonnenburg, J.L.; Backhed, F. Diet-microbiota interactions as moderators of human metabolism. Nature 2016, 535, 56–64. [Google Scholar] [CrossRef] [PubMed]
- Christensen, L.; Roager, H.M.; Astrup, A.; Hjorth, M.F. Microbial enterotypes in personalized nutrition and obesity management. Am. J. Clin. Nutr. 2018, 108, 645–651. [Google Scholar] [CrossRef] [PubMed]
- Barengolts, E. Gut Microbiota, Prebiotics, Probiotics, and Synbiotics in Management of Obesity and Prediabetes: Review of Randomized Controlled Trials. Endocr. Pract. 2016, 22, 1224–1234. [Google Scholar] [CrossRef] [PubMed]
- Carlucci, C.; Petrof, E.O.; Allen-Vercoe, E. Fecal Microbiota-based Therapeutics for Recurrent Clostridium difficile Infection, Ulcerative Colitis and Obesity. EBioMedicine 2016, 13, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Marotz, C.A.; Zarrinpar, A. Treating Obesity and Metabolic Syndrome with Fecal Microbiota Transplantation. Yale J. Biol. Med. 2016, 89, 383–388. [Google Scholar] [PubMed]
- Zhang, C.; Yin, A.; Li, H.; Wang, R.; Wu, G.; Shen, J.; Zhang, M.; Wang, L.; Hou, Y.; Ouyang, H.; et al. Dietary Modulation of Gut Microbiota Contributes to Alleviation of Both Genetic and Simple Obesity in Children. EBioMedicine 2015, 2, 968–984. [Google Scholar] [CrossRef] [PubMed]
- Bai, J.; Hu, Y.; Bruner, D.W. Composition of gut microbiota and its association with body mass index and lifestyle factors in a cohort of 7-18 years old children from the American Gut Project. Pediatr. Obes. 2018. [Google Scholar] [CrossRef] [PubMed]
- Rampelli, S.; Guenther, K.; Turroni, S.; Wolters, M.; Veidebaum, T.; Kourides, Y.; Molnar, D.; Lissner, L.; Benitez-Paez, A.; Sanz, Y.; et al. Pre-obese children’s dysbiotic gut microbiome and unhealthy diets may predict the development of obesity. Commun. Biol. 2018, 1, 222. [Google Scholar] [CrossRef] [PubMed]
- Nicoletti, C.F.; Cortes-Oliveira, C.; Pinhel, M.A.S.; Nonino, C.B. Bariatric Surgery and Precision Nutrition. Nutrients 2017, 9, 974. [Google Scholar] [CrossRef] [PubMed]
- Magouliotis, D.E.; Tasiopoulou, V.S.; Sioka, E.; Chatedaki, C.; Zacharoulis, D. Impact of Bariatric Surgery on Metabolic and Gut Microbiota Profile: A Systematic Review and Meta-analysis. Obes. Surg. 2017, 27, 1345–1357. [Google Scholar] [CrossRef] [PubMed]
- Anhe, F.F.; Varin, T.V.; Schertzer, J.D.; Marette, A. The Gut Microbiota as a Mediator of Metabolic Benefits after Bariatric Surgery. Can. J. Diabetes 2017, 41, 439–447. [Google Scholar] [CrossRef] [PubMed]
- Zheng, S.; Shi, J.; Wu, X.; Peng, Z.; Xin, C.; Zhang, L.; Liu, Y.; Gao, M.; Xu, S.; Han, H.; et al. Presence of Torque teno sus virus 1 and 2 in porcine circovirus 3-positive pigs. Transbound. Emerg. Dis. 2018, 65, 327–330. [Google Scholar] [CrossRef] [PubMed]
- Tremaroli, V.; Karlsson, F.; Werling, M.; Stahlman, M.; Kovatcheva-Datchary, P.; Olbers, T.; Fandriks, L.; le Roux, C.W.; Nielsen, J.; Backhed, F. Roux-en-Y Gastric Bypass and Vertical Banded Gastroplasty Induce Long-Term Changes on the Human Gut Microbiome Contributing to Fat Mass Regulation. Cell Metab. 2015, 22, 228–238. [Google Scholar] [CrossRef] [PubMed]
- Liou, A.P.; Paziuk, M.; Luevano, J.M., Jr.; Machineni, S.; Turnbaugh, P.J.; Kaplan, L.M. Conserved shifts in the gut microbiota due to gastric bypass reduce host weight and adiposity. Sci. Transl. Med. 2013, 5, 178ra141. [Google Scholar] [CrossRef] [PubMed]
- Palleja, A.; Kashani, A.; Allin, K.H.; Nielsen, T.; Zhang, C.; Li, Y.; Brach, T.; Liang, S.; Feng, Q.; Jorgensen, N.B.; et al. Roux-en-Y gastric bypass surgery of morbidly obese patients induces swift and persistent changes of the individual gut microbiota. Genome Med. 2016, 8, 67. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Hong, J.; Xu, X.; Feng, Q.; Zhang, D.; Gu, Y.; Shi, J.; Zhao, S.; Liu, W.; Wang, X.; et al. Gut microbiome and serum metabolome alterations in obesity and after weight-loss intervention. Nat. Med. 2017, 23, 859–868. [Google Scholar] [CrossRef] [PubMed]
- Hollister, E.B.; Riehle, K.; Luna, R.A.; Weidler, E.M.; Rubio-Gonzales, M.; Mistretta, T.-A.; Raza, S.; Doddapaneni, H.V.; Metcalf, G.A.; Muzny, D.M. Structure and function of the healthy pre-adolescent pediatric gut microbiome. Microbiome 2015, 3, 36. [Google Scholar] [CrossRef] [PubMed]
- Huttenhower, C.; Gevers, D.; Knight, R.; Abubucker, S.; Badger, J.H.; Chinwalla, A.T.; Creasy, H.H.; Earl, A.M.; FitzGerald, M.G.; Fulton, R.S.; et al. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef]
- Del Chierico, F.; Abbatini, F.; Russo, A.; Quagliariello, A.; Reddel, S.; Capoccia, D.; Caccamo, R.; Ginanni Corradini, S.; Nobili, V.; De Peppo, F.; et al. Gut Microbiota Markers in Obese Adolescent and Adult Patients: Age-Dependent Differential Patterns. Front. Microbiol. 2018, 9, 1210. [Google Scholar] [CrossRef] [PubMed]
- Le Chatelier, E.; Nielsen, T.; Qin, J.; Prifti, E.; Hildebrand, F.; Falony, G.; Almeida, M.; Arumugam, M.; Batto, J.M.; Kennedy, S.; et al. Richness of human gut microbiome correlates with metabolic markers. Nature 2013, 500, 541–546. [Google Scholar] [CrossRef] [PubMed]
- Erickson, A.R.; Cantarel, B.L.; Lamendella, R.; Darzi, Y.; Mongodin, E.F.; Pan, C.; Shah, M.; Halfvarson, J.; Tysk, C.; Henrissat, B.; et al. Integrated metagenomics/metaproteomics reveals human host-microbiota signatures of Crohn’s disease. PLoS ONE 2012, 7, e49138. [Google Scholar] [CrossRef] [PubMed]
- Joossens, M.; Huys, G.; Cnockaert, M.; De Preter, V.; Verbeke, K.; Rutgeerts, P.; Vandamme, P.; Vermeire, S. Dysbiosis of the faecal microbiota in patients with Crohn’s disease and their unaffected relatives. Gut 2011, 60, 631–637. [Google Scholar] [CrossRef] [PubMed]
- Canani, R.B.; Costanzo, M.D.; Leone, L.; Pedata, M.; Meli, R.; Calignano, A. Potential beneficial effects of butyrate in intestinal and extraintestinal diseases. World J. Gastroenterol. 2011, 17, 1519–1528. [Google Scholar] [CrossRef] [PubMed]
- Baumgart, D.C.; Carding, S.R. Inflammatory bowel disease: Cause and immunobiology. Lancet 2007, 369, 1627–1640. [Google Scholar] [CrossRef]
- De Caro, G.; Gaiani, F.; Duranti, S.; Fugazza, A.; Madia, C.; Milani, C.; Mancabelli, L.; Turroni, F.; de’ Angelis, G.L.; Carra, M.C.; et al. Inflammatory Bowel Disease. Am. J. Gastroenterol. 2016, 111, S260–S336. [Google Scholar] [CrossRef]
- Suskind, D.L.; Brittnacher, M.J.; Wahbeh, G.; Shaffer, M.L.; Hayden, H.S.; Qin, X.; Singh, N.; Damman, C.J.; Hager, K.R.; Nielson, H.; et al. Fecal microbial transplant effect on clinical outcomes and fecal microbiome in active Crohn’s disease. Inflamm. Bowel Dis. 2015, 21, 556–563. [Google Scholar] [CrossRef] [PubMed]
- Walters, W.A.; Xu, Z.; Knight, R. Meta-analyses of human gut microbes associated with obesity and IBD. FEBS Lett. 2014, 588, 4223–4233. [Google Scholar] [CrossRef] [PubMed]
- Sitkin, S.; Pokrotnieks, J. Clinical Potential of Anti-inflammatory Effects of Faecalibacterium prausnitzii and Butyrate in Inflammatory Bowel Disease. Inflamm. Bowel Dis. 2018, in press. [Google Scholar] [CrossRef] [PubMed]
- James, S.L.; Christophersen, C.T.; Bird, A.R.; Conlon, M.A.; Rosella, O.; Gibson, P.R.; Muir, J.G. Abnormal fibre usage in UC in remission. Gut 2015, 64, 562–570. [Google Scholar] [CrossRef] [PubMed]
- Rossen, N.G.; Fuentes, S.; van der Spek, M.J.; Tijssen, J.G.; Hartman, J.H.; Duflou, A.; Lowenberg, M.; van den Brink, G.R.; Mathus-Vliegen, E.M.; de Vos, W.M.; et al. Findings From a Randomized Controlled Trial of Fecal Transplantation for Patients With Ulcerative Colitis. Gastroenterology 2015, 149, 110–118. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.; Clavel, T.; Smirnov, K.; Schmidt, A.; Lagkouvardos, I.; Walker, A.; Lucio, M.; Michalke, B.; Schmitt-Kopplin, P.; Fedorak, R.; et al. Oral versus intravenous iron replacement therapy distinctly alters the gut microbiota and metabolome in patients with IBD. Gut 2017, 66, 863–871. [Google Scholar] [CrossRef] [PubMed]
- Fuentes, S.; Rossen, N.G.; van der Spek, M.J.; Hartman, J.H.; Huuskonen, L.; Korpela, K.; Salojarvi, J.; Aalvink, S.; de Vos, W.M.; D’Haens, G.R.; et al. Microbial shifts and signatures of long-term remission in ulcerative colitis after faecal microbiota transplantation. ISME J. 2017, 11, 1877–1889. [Google Scholar] [CrossRef] [PubMed]
- Ishikawa, D.; Sasaki, T.; Osada, T.; Kuwahara-Arai, K.; Haga, K.; Shibuya, T.; Hiramatsu, K.; Watanabe, S. Changes in Intestinal Microbiota Following Combination Therapy with Fecal Microbial Transplantation and Antibiotics for Ulcerative Colitis. Inflamm. Bowel Dis. 2017, 23, 116–125. [Google Scholar] [CrossRef] [PubMed]
- Paramsothy, S.; Kamm, M.A.; Kaakoush, N.O.; Walsh, A.J.; van den Bogaerde, J.; Samuel, D.; Leong, R.W.; Connor, S.; Ng, W.; Paramsothy, R. Multidonor intensive faecal microbiota transplantation for active ulcerative colitis: A randomised placebo-controlled trial. Lancet 2017, 389, 1218–1228. [Google Scholar] [CrossRef]
- Lo, B.; Prosberg, M.V.; Gluud, L.L.; Chan, W.; Leong, R.W.; van der List, E.; van der Have, M.; Sarter, H.; Gower-Rousseau, C.; Peyrin-Biroulet, L.; et al. Systematic review and meta-analysis: Assessment of factors affecting disability in inflammatory bowel disease and the reliability of the inflammatory bowel disease disability index. Aliment. Pharmacol. Ther. 2018, 47, 6–15. [Google Scholar] [CrossRef] [PubMed]
- Dobrolyubova, E.; Ruchkina, I.; Parfenov, A.; Knyazev, O. Ulcerative colitis (UC) with IBS-like disorders: Particular qualities of clinical manifestations and medical therapy. United Eur. Gastroenterol. J. 2017, 5, A293. [Google Scholar] [CrossRef]
- LaMere, B.; Wendt, E.R.; Kanwar, B.; Lynch, S.V. Investigating the Microbiome in a Phase 1b Study of Andecaliximab in Ulcerative Colitis. In Proceedings of the XXV UEG Week 2017, Barcelona, Spain, 28 October–1 November 2017; p. 103. [Google Scholar]
- Ananthakrishnan, A.N.; Luo, C.; Yajnik, V.; Khalili, H.; Garber, J.J.; Stevens, B.W.; Cleland, T.; Xavier, R.J. Gut Microbiome Function Predicts Response to Anti-integrin Biologic Therapy in Inflammatory Bowel Diseases. Cell Host Microbe 2017, 21, 603–610. [Google Scholar] [CrossRef] [PubMed]
- World Congress of Pediatric Gastroenterology, Hepatology and Nutrition. J. Pediatr. Gastroenterol. Nutr. 2016, 63 (Suppl. 2), S14–S15. [CrossRef]
- Matsuoka, K.; Uemura, Y.; Kanai, T.; Kunisaki, R.; Suzuki, Y.; Yokoyama, K.; Yoshimura, N.; Hibi, T. Efficacy of Bifidobacterium breve Fermented Milk in Maintaining Remission of Ulcerative Colitis. Dig. Dis. Sci. 2018, 63, 1910–1919. [Google Scholar] [CrossRef] [PubMed]
- Phillips, M.C.; Quintero, M.A.; Martinez, A.; Kerman, D.; Deshpande, A.R.; Damas, O.; Pignac-Kobinger, J.; Santaolalla, R.; Knight, K.; Rodriguez, V. P069 Low fat diet improves quality of life and changes the microbiome in a catered, cross-over design intervention of uc patients with quiescent disease: results of a pilot study. Gastroenterology 2018, 154, S36. [Google Scholar] [CrossRef]
- Rajca, S.; Grondin, V.; Louis, E.; Vernier-Massouille, G.; Grimaud, J.C.; Bouhnik, Y.; Laharie, D.; Dupas, J.L.; Pillant, H.; Picon, L.; et al. Alterations in the intestinal microbiome (dysbiosis) as a predictor of relapse after infliximab withdrawal in Crohn’s disease. Inflamm. Bowel Dis. 2014, 20, 978–986. [Google Scholar] [CrossRef] [PubMed]
- Suskind, D.L. Reply to can fecal microbial transplant effectively treat Crohn’s disease? Inflamm. Bowel Dis. 2015, 21, E8. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.W.X.; Bu, C. Fecal Microbiota Transplant For Crohn’s Disease: A Prospective, Randomized Study In Chinese Population. In Proceedings of the XXV UEG Week Barcelona, Barcelona, Spain, 28 October–1 November 2017; p. 112. [Google Scholar]
- Zhou, Y.; Xu, Z.Z.; He, Y.; Yang, Y.; Liu, L.; Lin, Q.; Nie, Y.; Li, M.; Zhi, F.; Liu, S.; et al. Gut Microbiota Offers Universal Biomarkers across Ethnicity in Inflammatory Bowel Disease Diagnosis and Infliximab Response Prediction. mSystems 2018, 3. [Google Scholar] [CrossRef] [PubMed]
- Doherty, M.K.; Koumpouras, C.; Telesco, S.; Monast, C.S.; Brodmerkel, C.; Schloss, P.D. The Fecal Microbiome as a Tool for Monitoring and Predicting Response Outcomes in Ustekinumab-Treated, Anti-TNFî ‘Refractory Crohn’s Disease Patients: Results from the Certifi Study. Gastroenterology 2017, 152, S191. [Google Scholar] [CrossRef]
- Halmos, E.; Christophersen, C.; Bird, A.; Shepherd, S.; Muir, J.; Gibson, P. prebiotic effect of Fodmaps in patients with Crohn’s disease: A randomised controlled trial. J. Gastroenterol. Hepatol. 2015, 30, 157–158. [Google Scholar]
- Brunt, E.M.; Wong, V.W.; Nobili, V.; Day, C.P.; Sookoian, S.; Maher, J.J.; Bugianesi, E.; Sirlin, C.B.; Neuschwander-Tetri, B.A.; Rinella, M.E. Nonalcoholic fatty liver disease. Nat. Rev. Dis. Primers 2015, 1, 15080. [Google Scholar] [CrossRef] [PubMed]
- Anstee, Q.M.; Day, C.P. The genetics of NAFLD. Nat. Rev. Gastroenterol. Hepatol. 2013, 10, 645–655. [Google Scholar] [CrossRef] [PubMed]
- Fotbolcu, H.; Zorlu, E. Nonalcoholic fatty liver disease as a multi-systemic disease. World J. Gastroenterol. 2016, 22, 4079–4090. [Google Scholar] [CrossRef] [PubMed]
- Wiest, R.; Albillos, A.; Trauner, M.; Bajaj, J.S.; Jalan, R. Targeting the gut-liver axis in liver disease. J. Hepatol. 2017, 67, 1084–1103. [Google Scholar] [CrossRef] [PubMed]
- Engstler, A.J.; Aumiller, T.; Degen, C.; Durr, M.; Weiss, E.; Maier, I.B.; Schattenberg, J.M.; Jin, C.J.; Sellmann, C.; Bergheim, I. Insulin resistance alters hepatic ethanol metabolism: Studies in mice and children with non-alcoholic fatty liver disease. Gut 2016, 65, 1564–1571. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Zhou, Q.; Li, H. Gut Microbiota and Nonalcoholic Fatty Liver Disease: Insights on Mechanisms and Therapy. Nutrients 2017, 9, 1124. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.Y.; Li, L.; Yu, C.H.; Shen, Z.; Chen, L.H.; Li, Y.M. Effects of prebiotics on non-alcoholic fatty liver disease. In Proceedings of the XXXVIIth National Congress of Gastroenterology, Hepatology and Digestive Endoscopy, Bucharest, Romania, 22–24 June 2017. [Google Scholar]
- Lambert, J.E.; Parnell, J.A.; Eksteen, B.; Raman, M.; Bomhof, M.R.; Rioux, K.P.; Madsen, K.L.; Reimer, R.A. Gut microbiota manipulation with prebiotics in patients with non-alcoholic fatty liver disease: A randomized controlled trial protocol. BMC Gastroenterol. 2015, 15, 169. [Google Scholar] [CrossRef] [PubMed]
- Poeta, M.; Pierri, L.; Vajro, P. Gut–liver axis derangement in non-alcoholic fatty liver disease. Children 2017, 4, 66. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Properzi, C.; Liddle, C.; Melton, P.; Ariff, A.; O’Sullivan, T.; Sherriff, J.; Coulter, S.; Christophersen, C.; Morrison, M. Bile Acids, Hepatic Steatosis and Gut Microbiome in Patients Undergoing Dietary Intervention for Non-Alcoholic Fatty Liver Disease. In Hepatology; Wiley: Hoboken, NJ, USA, 2018; p. 972A. [Google Scholar]
- Sherf-Dagan, S.; Zelber-Sagi, S.; Zilberman-Schapira, G.; Webb, M.; Buch, A.; Keidar, A.; Raziel, A.; Sakran, N.; Goitein, D.; Goldenberg, N.; et al. Probiotics administration following sleeve gastrectomy surgery: A randomized double-blind trial. Int. J. Obes. 2018, 42, 147–155. [Google Scholar] [CrossRef] [PubMed]
- Orr, D.W.; Murphy, R. prebiotic supplementation with inulin following metronidazole therapy achieves greater Alt reduction in Non-alcoholic Fatty Liver Disease (NAFLD): A randomised double-blind placebo controlled trial: 2176. Hepatology 2015, 62, 1268A–1269A. [Google Scholar]
- Kobyliak, N.; Bosak, N.; Falalyeyeva, T.; Beregova, T.; Bodnar, P. Effect of a probiotic on fatty liver index and liver stiffness in NAFLD patients: Randomized clinical trial. J. Hepatol. 2017, 66, S426–S427. [Google Scholar] [CrossRef]
- Ahn, S.; Jun, D.; Kim, E.; Oh, H.; Jeong, J.; Sohn, J.; Jang, E. Change of microbiota in patients with improved fatty liver and obesity. J. Hepatol. 2018, 68, S838–S839. [Google Scholar] [CrossRef]
- Bomhof, M.R.; Parnell, J.A.; Ramay, H.R.; Crotty, P.; Rioux, K.P.; Probert, C.S.; Jayakumar, S.; Raman, M.; Reimer, R.A. Histological improvement of non-alcoholic steatohepatitis with a prebiotic: A pilot clinical trial. Eur. J. Nutr. 2018, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Manzhalii, E.; Virchenko, O.; Falalyeyeva, T.; Beregova, T.; Stremmel, W. Treatment efficacy of a probiotic preparation for non-alcoholic steatohepatitis: A pilot trial. J. Dig. Dis. 2017, 18, 698–703. [Google Scholar] [CrossRef] [PubMed]
- Alisi, A.; Bedogni, G.; Baviera, G.; Giorgio, V.; Porro, E.; Paris, C.; Giammaria, P.; Reali, L.; Anania, F.; Nobili, V. Randomised clinical trial: The beneficial effects of VSL#3 in obese children with non-alcoholic steatohepatitis. Aliment. Pharmacol. Ther. 2014, 39, 1276–1285. [Google Scholar] [CrossRef] [PubMed]
- Ferolla, S.M.; Couto, C.A.; Costa-Silva, L.; Armiliato, G.N.; Pereira, C.A.; Martins, F.S.; Ferrari Mde, L.; Vilela, E.G.; Torres, H.O.; Cunha, A.S.; et al. Beneficial Effect of Synbiotic Supplementation on Hepatic Steatosis and Anthropometric Parameters, But Not on Gut Permeability in a Population with Nonalcoholic Steatohepatitis. Nutrients 2016, 8, 397. [Google Scholar] [CrossRef] [PubMed]
- Asghari-Jafarabadi Rad PhD, M. The Effect of Probiotic and Conventional Yogurt Consumptions on Anthropometric Parameters in Individuals with Non Alcoholic Fatty Liver Disease. J. Babol Univ. Med. Sci. 2014, 16, 55–62. [Google Scholar]
- Kessoku, T.; Imajo, K.; Honda, Y.; Kato, T.; Ogawa, Y.; Tomeno, W.; Higurashi, T.; Yoneda, M.; Shimakawa, M.; Tanaka, Y. Characteristics of Fecal Microbiota in Japanese Patients with Nonalcoholic Fatty Liver Disease: A Connection among Gut-Permeability, Endotoxin and NAFLD. Gastroenterology 2017, 152, S1200. [Google Scholar] [CrossRef]
- Lelouvier, B.; Servant, F.; Paisse, S.; Brunet, A.C.; Benyahya, S.; Serino, M.; Valle, C.; Ortiz, M.R.; Puig, J.; Courtney, M.; et al. Changes in blood microbiota profiles associated with liver fibrosis in obese patients: A pilot analysis. Hepatology 2016, 64, 2015–2027. [Google Scholar] [CrossRef] [PubMed]
- Rao, G. Insulin resistance syndrome. Am. Fam. Physician 2001, 63, 1159–1163, 1165–1166. [Google Scholar] [PubMed]
- Mazidi, M.; Rezaie, P.; Kengne, A.P.; Mobarhan, M.G.; Ferns, G.A. Gut microbiome and metabolic syndrome. Diabetes Metab. Syndr. 2016, 10, S150–S157. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.E.; Walker, M. Genetics of Insulin Resistance and the Metabolic Syndrome. Curr. Cardiol. Rep. 2016, 18, 75. [Google Scholar] [CrossRef] [PubMed]
- Czech, M.P. Insulin action and resistance in obesity and type 2 diabetes. Nat. Med. 2017, 23, 804–814. [Google Scholar] [CrossRef] [PubMed]
- Mutie, P.M.; Giordano, G.N.; Franks, P.W. Lifestyle precision medicine: The next generation in type 2 diabetes prevention? BMC Med. 2017, 15, 171. [Google Scholar] [CrossRef] [PubMed]
- Federico, A.; Dallio, M.; Sarno, R.I.; Giorgio, V.; Miele, L. Gut microbiota, obesity and metabolic disorders. Minerva Gastroenterol. Dietol. 2017, 63, 337–344. [Google Scholar] [CrossRef] [PubMed]
- Festi, D.; Schiumerini, R.; Eusebi, L.H.; Marasco, G.; Taddia, M.; Colecchia, A. Gut microbiota and metabolic syndrome. World J. Gastroenterol. 2014, 20, 16079–16094. [Google Scholar] [CrossRef] [PubMed]
- Caricilli, A.M.; Saad, M.J. Gut microbiota composition and its effects on obesity and insulin resistance. Curr. Opin. Clin. Nutr. Metab. Care 2014, 17, 312–318. [Google Scholar] [CrossRef] [PubMed]
- Saad, M.J.; Santos, A.; Prada, P.O. Linking Gut Microbiota and Inflammation to Obesity and Insulin Resistance. Physiology 2016, 31, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Kushugulova, A.; Forslund, S.K.; Costea, P.I.; Kozhakhmetov, S.; Khassenbekova, Z.; Urazova, M.; Nurgozhin, T.; Zhumadilov, Z.; Benberin, V.; Driessen, M.; et al. Metagenomic analysis of gut microbial communities from a Central Asian population. BMJ Open 2018, 8, e021682. [Google Scholar] [CrossRef] [PubMed]
- Gareau, M.G.; Sherman, P.M.; Walker, W.A. Probiotics and the gut microbiota in intestinal health and disease. Nat. Rev. Gastroenterol. Hepatol. 2010, 7, 503–514. [Google Scholar] [CrossRef] [PubMed]
- Mikkelsen, K.H.; Frost, M.; Bahl, M.I.; Licht, T.R.; Jensen, U.S.; Rosenberg, J.; Pedersen, O.; Hansen, T.; Rehfeld, J.F.; Holst, J.J.; et al. Effect of Antibiotics on Gut Microbiota, Gut Hormones and Glucose Metabolism. PLoS ONE 2015, 10, e0142352. [Google Scholar] [CrossRef] [PubMed]
- Macfarlane, S.; Cleary, S.; Bahrami, B.; Reynolds, N.; Macfarlane, G.T. Synbiotic consumption changes the metabolism and composition of the gut microbiota in older people and modifies inflammatory processes: A randomised, double-blind, placebo-controlled crossover study. Aliment. Pharmacol. Ther. 2013, 38, 804–816. [Google Scholar] [CrossRef] [PubMed]
- Bruzzese, E.; Callegari, M.L.; Raia, V.; Viscovo, S.; Scotto, R.; Ferrari, S.; Morelli, L.; Buccigrossi, V.; Lo Vecchio, A.; Ruberto, E.; et al. Disrupted intestinal microbiota and intestinal inflammation in children with cystic fibrosis and its restoration with Lactobacillus GG: A randomised clinical trial. PLoS ONE 2014, 9, e87796. [Google Scholar] [CrossRef] [PubMed]
- Kang, D.W.; Adams, J.B.; Gregory, A.C.; Borody, T.; Chittick, L.; Fasano, A.; Khoruts, A.; Geis, E.; Maldonado, J.; McDonough-Means, S.; et al. Microbiota Transfer Therapy alters gut ecosystem and improves gastrointestinal and autism symptoms: An open-label study. Microbiome 2017, 5, 10. [Google Scholar] [CrossRef] [PubMed]
- Plaza-Diaz, J.; Fernandez-Caballero, J.A.; Chueca, N.; Garcia, F.; Gomez-Llorente, C.; Saez-Lara, M.J.; Fontana, L.; Gil, A. Pyrosequencing analysis reveals changes in intestinal microbiota of healthy adults who received a daily dose of immunomodulatory probiotic strains. Nutrients 2015, 7, 3999–4015. [Google Scholar] [CrossRef] [PubMed]
- Vandeputte, D.; Falony, G.; Vieira-Silva, S.; Wang, J.; Sailer, M.; Theis, S.; Verbeke, K.; Raes, J. Prebiotic inulin-type fructans induce specific changes in the human gut microbiota. Gut 2017, 66, 1968–1974. [Google Scholar] [CrossRef] [PubMed]
- Smits, L.P.; Kootte, R.S.; Levin, E.; Prodan, A.; Fuentes, S.; Zoetendal, E.G.; Wang, Z.; Levison, B.S.; Cleophas, M.C.P.; Kemper, E.M.; et al. Effect of Vegan Fecal Microbiota Transplantation on Carnitine- and Choline-Derived Trimethylamine-N-Oxide Production and Vascular Inflammation in Patients With Metabolic Syndrome. J. Am. Heart Assoc. 2018, 7, e008342. [Google Scholar] [CrossRef] [PubMed]
- Haro, C.; Garcia-Carpintero, S.; Alcala-Diaz, J.F.; Gomez-Delgado, F.; Delgado-Lista, J.; Perez-Martinez, P.; Rangel Zuniga, O.A.; Quintana-Navarro, G.M.; Landa, B.B.; Clemente, J.C.; et al. The gut microbial community in metabolic syndrome patients is modified by diet. J. Nutr. Biochem. 2016, 27, 27–31. [Google Scholar] [CrossRef] [PubMed]
- Haro, C.; Garcia-Carpintero, S.; Rangel-Zuniga, O.A.; Alcala-Diaz, J.F.; Landa, B.B.; Clemente, J.C.; Perez-Martinez, P.; Lopez-Miranda, J.; Perez-Jimenez, F.; Camargo, A. Consumption of Two Healthy Dietary Patterns Restored Microbiota Dysbiosis in Obese Patients with Metabolic Dysfunction. Mol. Nutr. Food Res. 2017, 61, 1700300. [Google Scholar] [CrossRef] [PubMed]
- Salonen, A.; Lahti, L.; Salojarvi, J.; Holtrop, G.; Korpela, K.; Duncan, S.H.; Date, P.; Farquharson, F.; Johnstone, A.M.; Lobley, G.E.; et al. Impact of diet and individual variation on intestinal microbiota composition and fermentation products in obese men. ISME J. 2014, 8, 2218–2230. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Indias, I.; Sanchez-Alcoholado, L.; Perez-Martinez, P.; Andres-Lacueva, C.; Cardona, F.; Tinahones, F.; Queipo-Ortuno, M.I. Red wine polyphenols modulate fecal microbiota and reduce markers of the metabolic syndrome in obese patients. Food Funct. 2016, 7, 1775–1787. [Google Scholar] [CrossRef] [PubMed]
- Ni, Y.; Mu, C.; He, X.; Zheng, K.; Guo, H.; Zhu, W. Characteristics of gut microbiota and its response to a Chinese Herbal Formula in elder patients with metabolic syndrome. Drug Discov. Ther. 2018, 12, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Roager, H.M.; Vogt, J.K.; Kristensen, M.; Hansen, L.B.S.; Ibrugger, S.; Maerkedahl, R.B.; Bahl, M.I.; Lind, M.V.; Nielsen, R.L.; Frokiaer, H.; et al. Whole grain-rich diet reduces body weight and systemic low-grade inflammation without inducing major changes of the gut microbiome: A randomised cross-over trial. Gut 2019, 68, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Velikonja, A.; Lipoglavsek, L.; Zorec, M.; Orel, R.; Avgustin, G. Alterations in gut microbiota composition and metabolic parameters after dietary intervention with barley beta glucans in patients with high risk for metabolic syndrome development. Anaerobe 2018, 55, 67–77. [Google Scholar] [CrossRef] [PubMed]
- Stadlbauer, V.; Leber, B.; Lemesch, S.; Trajanoski, S.; Bashir, M.; Horvath, A.; Tawdrous, M.; Stojakovic, T.; Fauler, G.; Fickert, P.; et al. Lactobacillus casei Shirota Supplementation Does Not Restore Gut Microbiota Composition and Gut Barrier in Metabolic Syndrome: A Randomized Pilot Study. PLoS ONE 2015, 10, e0141399. [Google Scholar] [CrossRef] [PubMed]
- Vrieze, A.; Out, C.; Fuentes, S.; Jonker, L.; Reuling, I.; Kootte, R.S.; van Nood, E.; Holleman, F.; Knaapen, M.; Romijn, J.A.; et al. Impact of oral vancomycin on gut microbiota, bile acid metabolism, and insulin sensitivity. J. Hepatol. 2014, 60, 824–831. [Google Scholar] [CrossRef] [PubMed]
- Burch, E.; Williams, L.T.; Makepeace, H.; Alston-Knox, C.; Ball, L. How Does Diet Change with A Diagnosis of Diabetes? Protocol of the 3D Longitudinal Study. Nutrients 2019, 11, 158. [Google Scholar] [CrossRef] [PubMed]
- Alwan, A. Global Status Report on Noncommunicable Diseases 2010; World Health Organization: Geneva, Switzerland, 2011. [Google Scholar]
- Sato, J.; Kanazawa, A.; Azuma, K.; Ikeda, F.; Goto, H.; Komiya, K.; Kanno, R.; Tamura, Y.; Asahara, T.; Takahashi, T.; et al. Probiotic reduces bacterial translocation in type 2 diabetes mellitus: A randomised controlled study. Sci. Rep. 2017, 7, 12115. [Google Scholar] [CrossRef] [PubMed]
- Medina-Vera, I.; Sanchez-Tapia, M.; Noriega-Lopez, L.; Granados-Portillo, O.; Guevara-Cruz, M.; Flores-Lopez, A.; Avila-Nava, A.; Fernandez, M.L.; Tovar, A.R.; Torres, N. A dietary intervention with functional foods reduces metabolic endotoxaemia and attenuates biochemical abnormalities by modifying faecal microbiota in people with type 2 diabetes. Diabetes Metab. 2018. [Google Scholar] [CrossRef] [PubMed]
- Hjorth, M.F.; Zohar, Y.; Hill, J.O.; Astrup, A. Personalized Dietary Management of Overweight and Obesity Based on Measures of Insulin and Glucose. Annu. Rev. Nutr. 2018, 38, 245–272. [Google Scholar] [CrossRef] [PubMed]
- Chavanelle, V.; Otero, Y.F.; Sirvent, P.; Cani, P.D.; Peltier, S. Pleiotropic Effects of Totum-63—Simultaneous Targeting of Multiple Diabetes Mediators. Am. Diabetes Assoc. 2018. [Google Scholar] [CrossRef]
- Elbere, I.; Kalnina, I.; Silamikelis, I.; Konrade, I.; Zaharenko, L.; Sekace, K.; Radovica-Spalvina, I.; Fridmanis, D.; Gudra, D.; Pirags, V.; et al. Association of metformin administration with gut microbiome dysbiosis in healthy volunteers. PLoS ONE 2018, 13, e0204317. [Google Scholar] [CrossRef] [PubMed]
- Tong, X.; Xu, J.; Lian, F.; Yu, X.; Zhao, Y.; Xu, L.; Zhang, M.; Zhao, X.; Shen, J.; Wu, S.; et al. Structural Alteration of Gut Microbiota during the Amelioration of Human Type 2 Diabetes with Hyperlipidemia by Metformin and a Traditional Chinese Herbal Formula: A Multicenter, Randomized, Open Label Clinical Trial. MBio 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Canfora, E.E.; van der Beek, C.M.; Hermes, G.D.A.; Goossens, G.H.; Jocken, J.W.E.; Holst, J.J.; van Eijk, H.M.; Venema, K.; Smidt, H.; Zoetendal, E.G.; et al. Supplementation of Diet With Galacto-oligosaccharides Increases Bifidobacteria, but Not Insulin Sensitivity, in Obese Prediabetic Individuals. Gastroenterology 2017, 153, 87–97. [Google Scholar] [CrossRef] [PubMed]
- Scorletti, E.; Afolabi, P.R.; Miles, E.A.; Smith, D.E.; Almehmadi, A.; Alshathry, A.; Moyses, H.E.; Clough, G.F.; Wright, M.; Patel, J.; et al. Design and rationale of the INSYTE study: A randomised, placebo controlled study to test the efficacy of a synbiotic on liver fat, disease biomarkers and intestinal microbiota in non-alcoholic fatty liver disease. Contemp. Clin. Trials 2018, 71, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Cornejo-Pareja, I.; Munoz-Garach, A.; Clemente-Postigo, M.; Tinahones, F.J. Importance of gut microbiota in obesity. Eur. J. Clin. Nutr. 2018. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Zhang, F.; Ding, X.; Wu, G.; Lam, Y.Y.; Wang, X.; Fu, H.; Xue, X.; Lu, C.; Ma, J.; et al. Gut bacteria selectively promoted by dietary fibers alleviate type 2 diabetes. Science 2018, 359, 1151–1156. [Google Scholar] [CrossRef] [PubMed]
- Roshanravan, N.; Mahdavi, R.; Alizadeh, E.; Ghavami, A.; Rahbar Saadat, Y.; Mesri Alamdari, N.; Alipour, S.; Dastouri, M.R.; Ostadrahimi, A. The effects of sodium butyrate and inulin supplementation on angiotensin signaling pathway via promotion of Akkermansia muciniphila abundance in type 2 diabetes; A randomized, double-blind, placebo-controlled trial. J. Cardiovasc. Thorac. Res. 2017, 9, 183–190. [Google Scholar] [CrossRef] [PubMed]
- Roshanravan, N.; Mahdavi, R.; Jafarabadi, M.A.; Alizadeh, E.; Ghavami, A.; Saadat, Y.R.; Alamdari, N.M.; Dastouri, M.R.; Alipour, S.; Ostadrahimi, A. The effects of sodium butyrate and high-performance inulin supplementation on the promotion of gut bacterium Akkermansia muciniphila growth and alterations in miR-375 and KLF5 expression in type 2 diabetic patients: A randomized, double-blind, placebo-controlled trial. Eur. J. Integr. Med. 2018, 18, 1–7. [Google Scholar]
- Mitchell, C.M.; Davy, B.M.; Halliday, T.M.; Hulver, M.W.; Neilson, A.P.; Ponder, M.A.; Davy, K.P. The effect of prebiotic supplementation with inulin on cardiometabolic health: Rationale, design, and methods of a controlled feeding efficacy trial in adults at risk of type 2 diabetes. Contemp. Clin. Trials 2015, 45, 328–337. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Yang, L.; Ma, M.; Liu, Y. Improving the metabolism of glucose and lipids in patients with prediabetes by affecting the gut microbiota. In Diabetes-Metabolism Research and Reviews; Wiley: Hoboken, NJ, USA, 2017. [Google Scholar]
- Martinez, I.; Kim, J.; Duffy, P.R.; Schlegel, V.L.; Walter, J. Resistant starches types 2 and 4 have differential effects on the composition of the fecal microbiota in human subjects. PLoS ONE 2010, 5, e15046. [Google Scholar] [CrossRef] [PubMed]
- Shimozato, A.; Sasaki, M.; Ogasawara, N.; Funaki, Y.; Ebi, M.; Goto, C.; Koikeda, S.; Joh, T.; Kasugai, K. Transglucosidase improves the bowel movements in type 2 diabetes mellitus patients: A preliminary randomized double-blind, placebo-controlled study. United Eur. Gastroenterol. J. 2017, 5, 898–907. [Google Scholar] [CrossRef] [PubMed]
- Mobini, R.; Tremaroli, V.; Stahlman, M.; Karlsson, F.; Levin, M.; Ljungberg, M.; Sohlin, M.; Berteus Forslund, H.; Perkins, R.; Backhed, F.; et al. Metabolic effects of Lactobacillus reuteri DSM 17938 in people with type 2 diabetes: A randomized controlled trial. Diabetes Obes. Metab. 2017, 19, 579–589. [Google Scholar] [CrossRef] [PubMed]
- Stefanaki, C.; Bacopoulou, F.; Michos, A. The impact of probiotics’ administration on glycemic control, body composition, gut microbiome, mitochondria, and other hormonal signals in adolescents with prediabetes—A randomized, controlled trial study protocol. Contemp. Clin. Trials Commun. 2018, 11, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Razmpoosh, E.; Javadi, A.; Ejtahed, H.S.; Mirmiran, P.; Javadi, M.; Yousefinejad, A. The effect of probiotic supplementation on glycemic control and lipid profile in patients with type 2 diabetes: A randomized placebo controlled trial. Diabetes Metab. Syndr. Clin. Res. Rev. 2019, 13, 175–182. [Google Scholar] [CrossRef] [PubMed]
- Samah, S.; Ramasamy, K.; Lim, S.M.; Neoh, C.F. Probiotics for the management of type 2 diabetes mellitus: A systematic review and meta-analysis. Diabetes Res. Clin. Pract. 2016, 118, 172–182. [Google Scholar] [CrossRef] [PubMed]
- Poznanski, S.M.; Barra, N.G.; Ashkar, A.A.; Schertzer, J.D. Immunometabolism of T cells and NK cells: Metabolic control of effector and regulatory function. Inflamm. Res. 2018, 67, 813–828. [Google Scholar] [CrossRef] [PubMed]
- Soergel, D.A.; Dey, N.; Knight, R.; Brenner, S.E. Selection of primers for optimal taxonomic classification of environmental 16S rRNA gene sequences. ISME J. 2012, 6, 1440–1444. [Google Scholar] [CrossRef] [PubMed]
- Franzosa, E.A.; Hsu, T.; Sirota-Madi, A.; Shafquat, A.; Abu-Ali, G.; Morgan, X.C.; Huttenhower, C. Sequencing and beyond: Integrating molecular ‘omics’ for microbial community profiling. Nat. Rev. Microbiol. 2015, 13, 360–372. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; de Vos, W.M. Next-Generation Beneficial Microbes: The Case of Akkermansia muciniphila. Front. Microbiol. 2017, 8, 1765. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Siles, M.; Duncan, S.H.; Garcia-Gil, L.J.; Martinez-Medina, M. Faecalibacterium prausnitzii: From microbiology to diagnostics and prognostics. ISME J. 2017, 11, 841–852. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).