Pan-Cellulosomics of Mesophilic Clostridia: Variations on a Theme
Abstract
:1. Introduction
2. Materials and Methods
2.1. Genomes Sequences
2.2. Bioinformatic Identification of Cellulosomal Components
3. Results
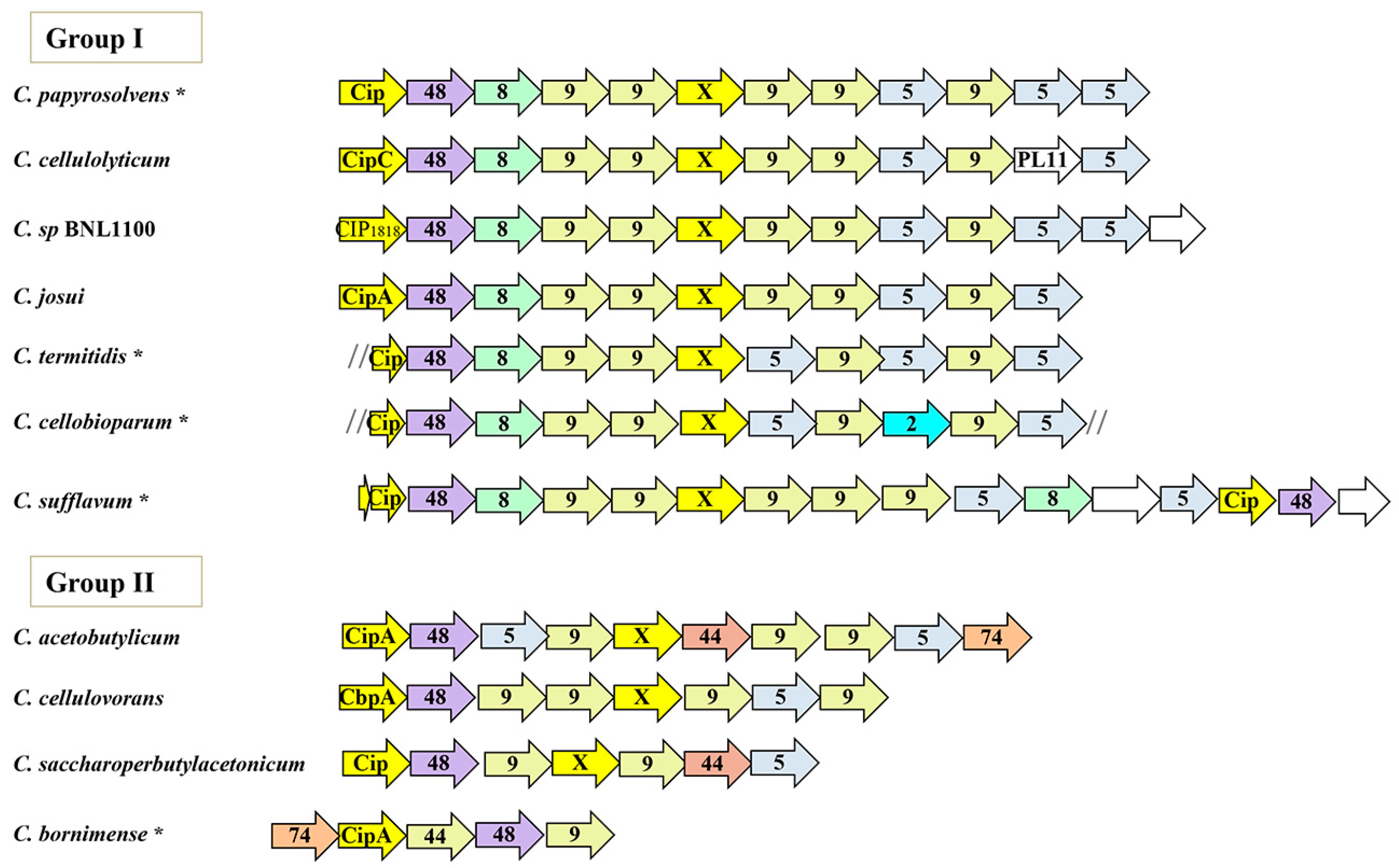
3.1. Comparative Characteristics of the Mesophilic Clostridia Cellulosomal Systems
3.2. Conserved Patterns in the Orthologous Sca Gene Cluster
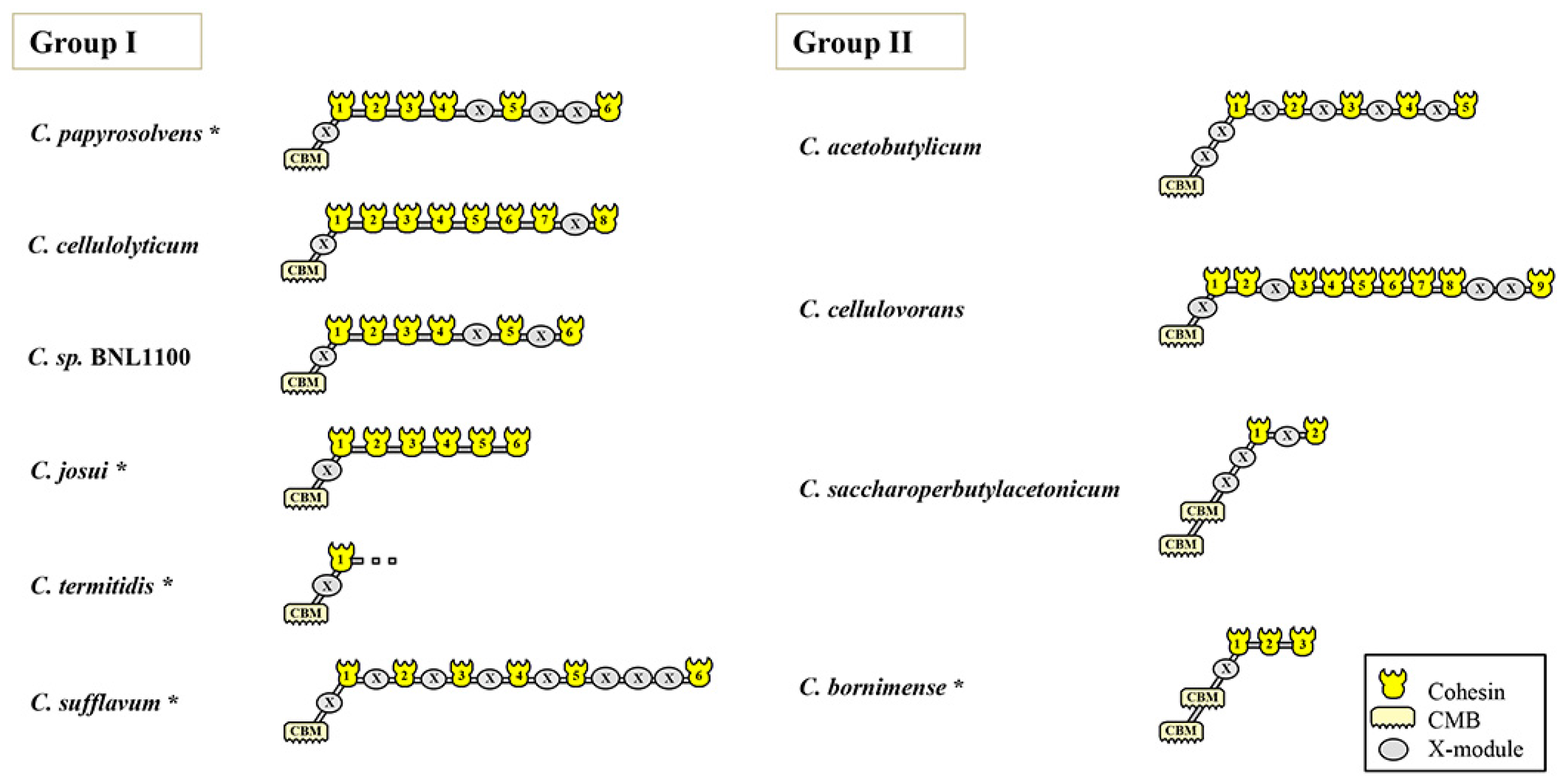
3.3. Modular Organization of the Major Scaffoldin Gene
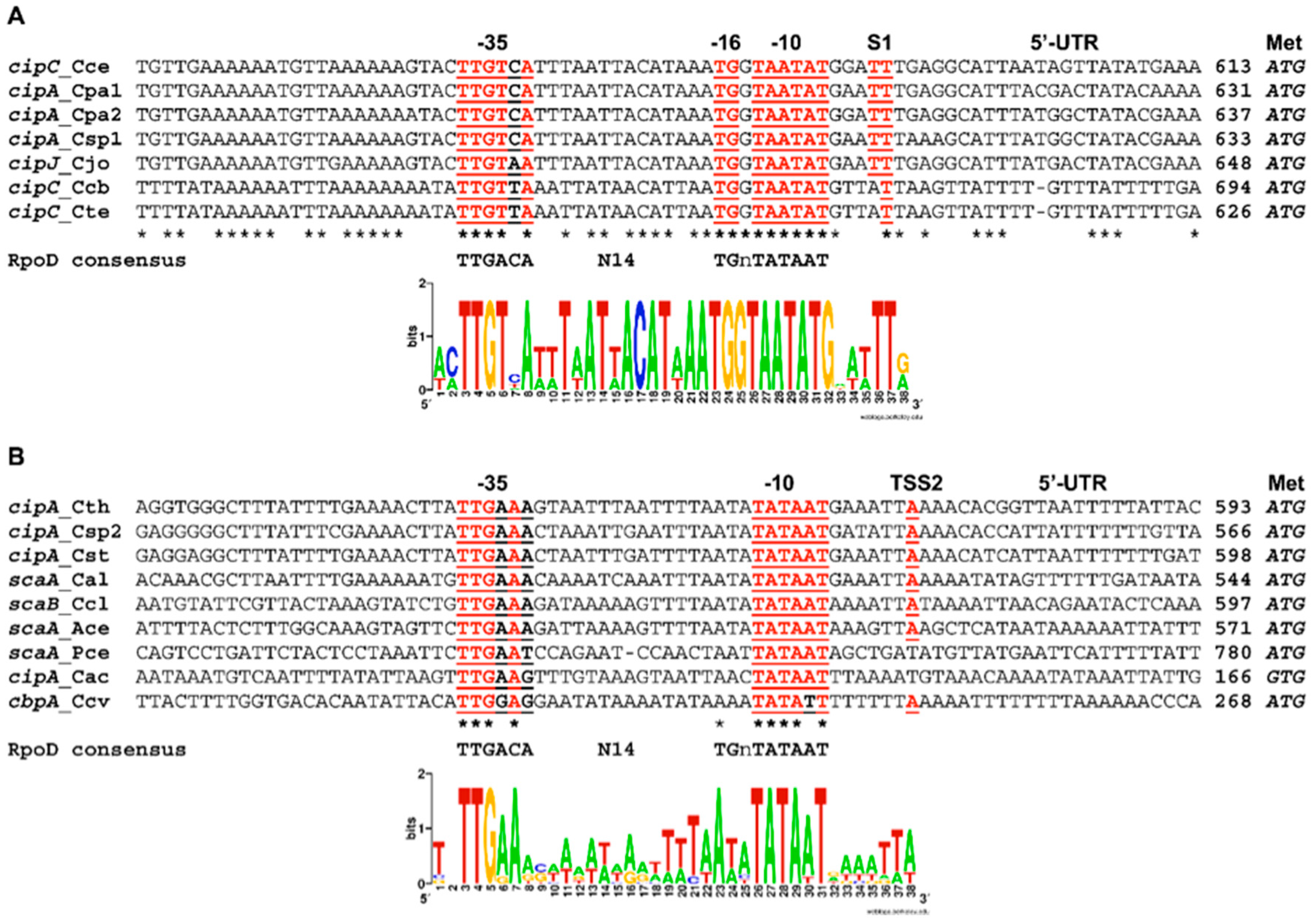
3.4. Regulation of the Sca Gene Cluster by a Conserved σA-Dependent Promoter
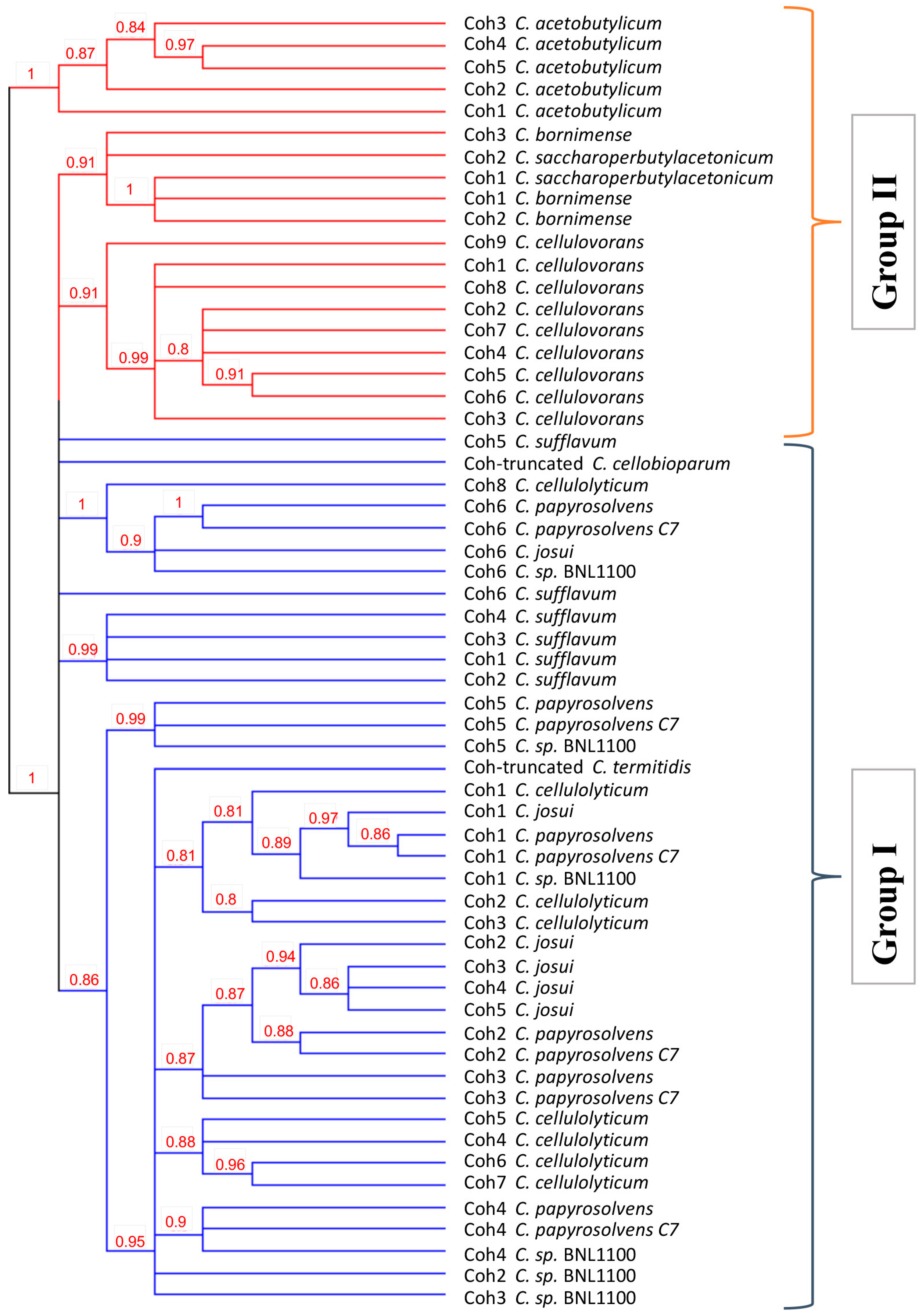
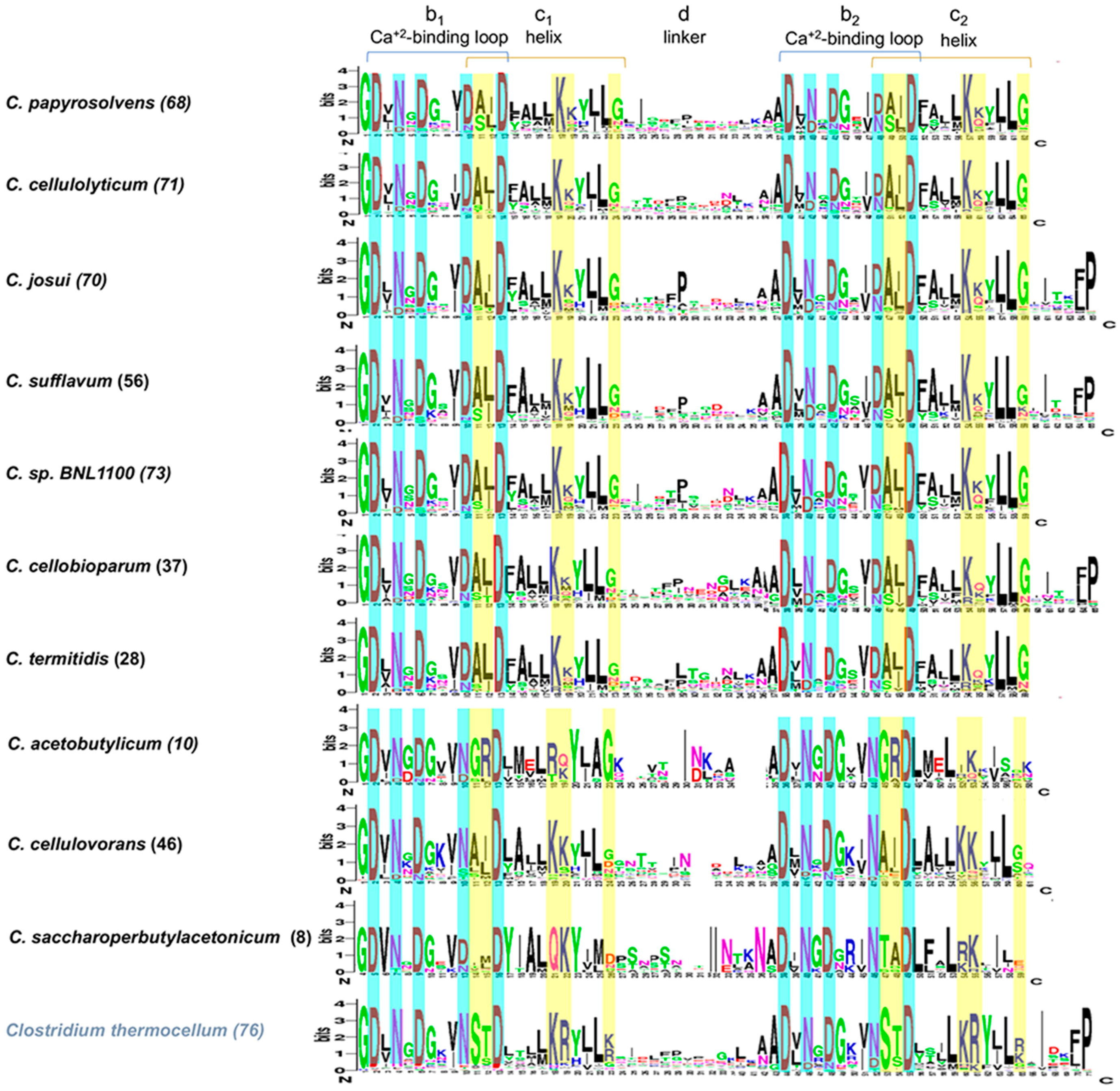
3.5. Sequence Conservation in Cohesins and Dockerins Suggest Cross-Species Recognition
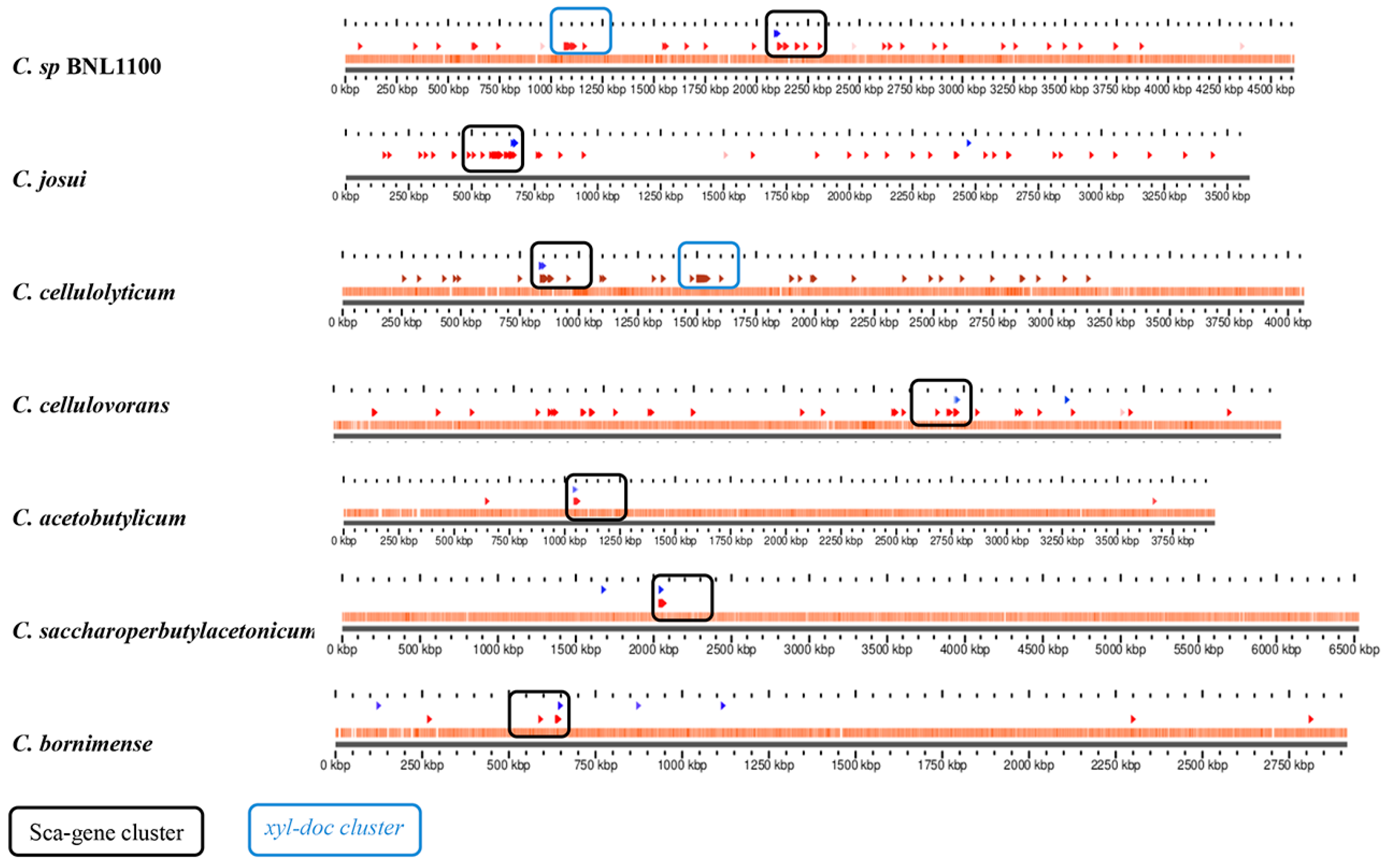
3.6. Sporadic Spatial Organization of the Cellulosomal Genes along the Bacterial Chromosome
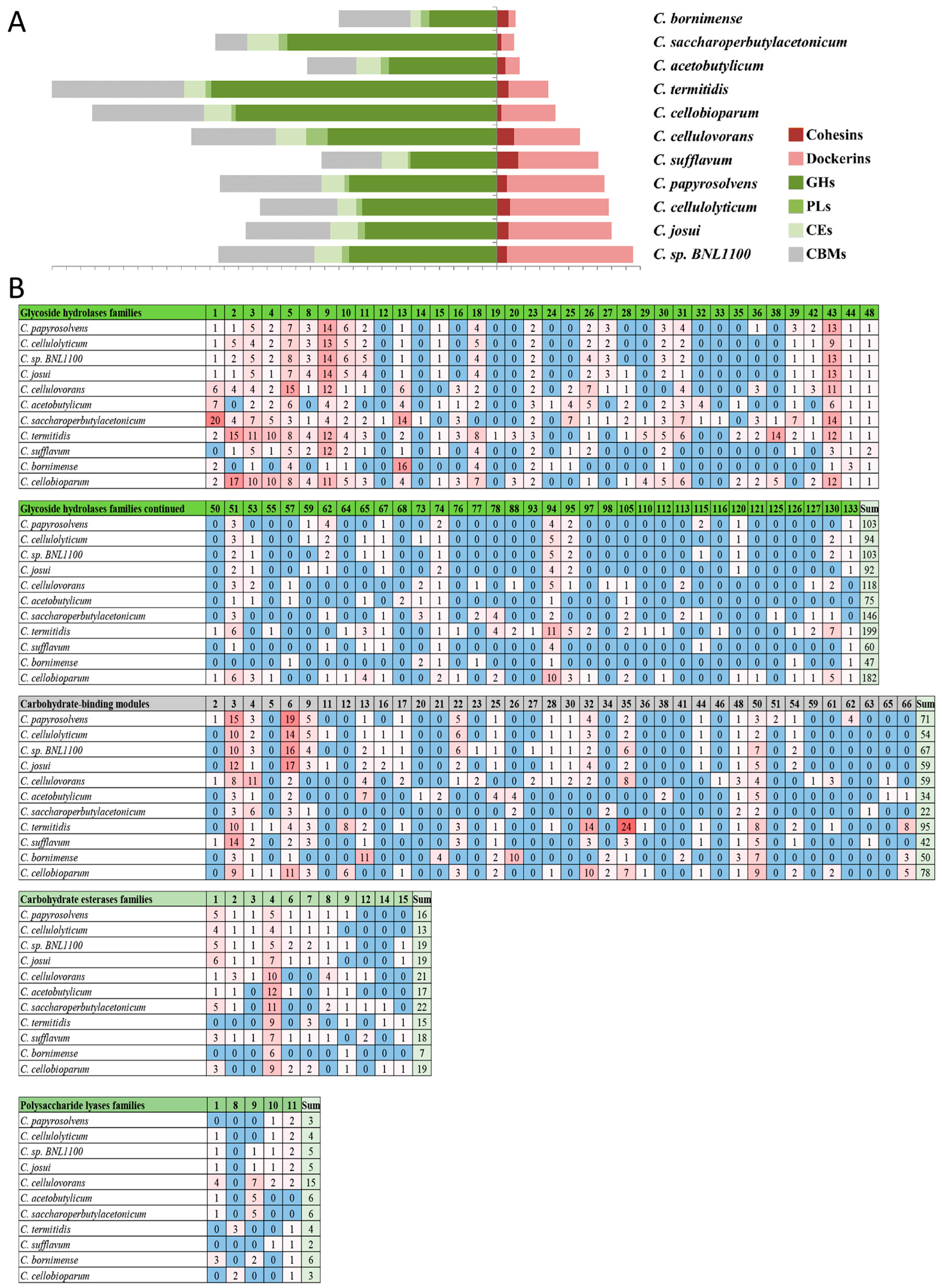
3.7. Profiling the Carbohydrate-Active Enzymes in the Cellulosome-Producing Mesophiles
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
References
- Albersheim, P.; Darvill, A.; Roberts, K.; Sederoff, R.; Staehelin, A. Plant Cell Walls; Garland Science: New York, NY, USA, 2010; p. 430. [Google Scholar]
- Artzi, L.; Bayer, E.A.; Moraïs, S. Cellulosomes: Bacterial nanomachines for dismantling plant polysaccharides. Nat. Rev. Microbiol. 2017, 15, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, W.H. The cellulosome and cellulose degradation by anaerobic bacteria. Appl. Microbiol. Biotechnol. 2001, 56, 634–649. [Google Scholar] [CrossRef] [PubMed]
- Himmel, M.E.; Xu, Q.; Luo, Y.; Ding, S.-Y.; Lamed, R.; Bayer, E.A. Microbial enzyme systems for biomass conversion: Emerging paradigms. Biofuels 2010, 1, 323–341. [Google Scholar] [CrossRef]
- Bayer, E.A.; Shoham, Y.; Lamed, R. Lignocellulose-decomposing bacteria and their enzyme systems. In The Prokaryotes, 4th ed.; Rosenberg, E., Ed.; Springer: Berlin, Germany, 2013; pp. 216–266. [Google Scholar]
- Doi, R.H. Cellulases of mesophilic microorganisms: Cellulosome and noncellulosome producers. Ann. N. Y. Acad. Sci. 2008, 1125, 267–279. [Google Scholar] [CrossRef] [PubMed]
- Bayer, E.A.; Belaich, J.-P.; Shoham, Y.; Lamed, R. The cellulosomes: Multi-enzyme machines for degradation of plant cell wall polysaccharides. Annu. Rev. Microbiol. 2004, 58, 521–554. [Google Scholar] [CrossRef] [PubMed]
- Belaich, J.-P.; Tardif, C.; Belaich, A.; Gaudin, C. The cellulolytic system of Clostridium cellulolyticum. J. Biotechnol. 1997, 57, 3–14. [Google Scholar] [CrossRef]
- Sabathe, F.; Belaich, A.; Soucaille, P. Characterization of the cellulolytic complex (cellulosome) of Clostridium acetobutylicum. FEMS Microbiol. Lett. 2002, 217, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Tamaru, Y.; Karita, S.; Ibrahim, A.; Chan, H.; Doi, R.H. A large gene cluster for the Clostridium cellulovorans cellulosome. J. Bacteriol. 2000, 182, 5906–5910. [Google Scholar] [CrossRef] [PubMed]
- Sakka, M.; Goto, M.; Fujino, T.; Fujino, E.; Karita, S.; Kimura, T.; Sakka, K. Analysis of a Clostridium josui cellulase gene cluster containing the man5a gene and characterization of recombinant Man5A. Biosci. Biotechnol. Biochem. 2010, 74, 2077–2082. [Google Scholar] [CrossRef] [PubMed]
- Poole, D.M.; Morag, E.; Lamed, R.; Bayer, E.A.; Hazlewood, G.P.; Gilbert, H.J. Identification of the cellulose binding domain of the cellulosome subunit S1 from Clostridium thermocellum. FEMS Microbiol. Lett. 1992, 99, 181–186. [Google Scholar] [CrossRef]
- Mosbah, A.; Belaich, A.; Bornet, O.; Belaich, J.P.; Henrissat, B.; Darbon, H. Solution structure of the module X2_1 of unknown function of the cellulosomal scaffolding protein CipC of Clostridium cellulolyticum. J. Mol. Biol. 2000, 304, 201–217. [Google Scholar] [CrossRef] [PubMed]
- Vazana, Y.; Barak, Y.; Unger, T.; Peleg, Y.; Ben Yehezkel, T.; Mazor, Y.; Shapiro, E.; Lamed, R.; Bayer, E.A. A synthetic biology approach for evaluating the functional contribution of designer cellulosome components to deconstruction of cellulosic substrates. Biotechnol. Biofuels 2013, 6, 182. [Google Scholar] [CrossRef] [PubMed]
- Bayer, E.A.; Smith, S.P.; Noach, I.; Alber, O.; Adams, J.J.; Lamed, R.; Shimon, L.J.W.; Frolow, F. Can we crystallize a cellulosome? In Biotechnology of Lignocellulose Degradation and Biomass Utilization; Sakka, K., Karita, S., Kimura, T., Sakka, M., Matsui, H., Miyake, H., Tanaka, A., Eds.; Ito Print Publishing Division: Tokyo, Japan, 2009; pp. 183–205. [Google Scholar]
- Fujino, T.; Béguin, P.; Aubert, J.-P. Organization of a Clostridium thermocellum gene cluster encoding the cellulosomal scaffolding protein cipa and a protein possibly involved in attachment of the cellulosome to the cell surface. J. Bacteriol. 1993, 175, 1891–1899. [Google Scholar] [CrossRef] [PubMed]
- Lamed, R.; Setter, E.; Bayer, E.A. Characterization of a cellulose-binding, cellulase-containing complex in Clostridium thermocellum. J. Bacteriol. 1983, 156, 828–836. [Google Scholar] [PubMed]
- Xu, Q.; Bayer, E.A.; Goldman, M.; Kenig, R.; Shoham, Y.; Lamed, R. Architecture of the Bacteroides cellulosolvens cellulosome: Description of a cell-surface anchoring scaffoldin and a family-48 cellulase. J. Bacteriol. 2004, 186, 968–977. [Google Scholar] [CrossRef] [PubMed]
- Zhivin, O.; Dassa, B.; Moraïs, S.; Uttukar, S.M.; Brown, S.D.; Henrissat, B.; Lamed, R.; Bayer, E.A. Unique organization and unprecedented diversity of the Bacteroides (Pseudobacteroides) cellulosolvens cellulosome system. Biotechnol. Biofuels 2017, 10, 211. [Google Scholar] [CrossRef] [PubMed]
- Dassa, B.; Borovok, I.; Lamed, R.; Henrissat, B.; Coutinho, P.; Hemme, C.L.; Huang, Y.; Zhou, Z.; Bayer, E.A. Genome-wide analysis of Acetivibrio cellulolyticus provides a blueprint of an elaborate cellulosome system. BMC Genom. 2012, 13, 210. [Google Scholar] [CrossRef] [PubMed]
- Hamberg, Y.; Ruimy-Israeli, V.; Dassa, B.; Barak, Y.; Lamed, R.; Cameron, K.; Fontes, C.M.; Bayer, E.A.; Fried, D.B. Elaborate cellulosome architecture of Acetivibrio cellulolyticus revealed by selective screening of cohesin-dockerin interactions. PeerJ 2014, 2, e636. [Google Scholar] [CrossRef] [PubMed]
- Israeli-Ruimy, V.; Bule, P.; Jindou, S.; Dassa, B.; Barak, Y.; Slutzki, M.; Hamberg, Y.; Cardoso, V.; Alves, V.D.; Najmudin, S.; et al. Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions. Sci. Rep. 2017, 7, 42355. [Google Scholar] [CrossRef] [PubMed]
- Rincon, M.T.; Cepeljnik, T.; Martin, J.C.; Barak, Y.; Lamed, R.; Bayer, E.A.; Flint, H.J. A novel cell surface-anchored cellulose-binding protein encoded by the sca gene cluster of Ruminococcus flavefaciens. J. Bacteriol. 2007, 189, 4774–7283. [Google Scholar] [CrossRef] [PubMed]
- Artzi, L.; Dassa, B.; Borovok, I.; Shamshoum, M.; Lamed, R.; Bayer, E.A. Cellulosomics of the cellulolytic thermophile Clostridium clariflavum. Biotechnol. Biofuels 2014, 7, 100. [Google Scholar] [CrossRef] [PubMed]
- Artzi, L.; Morag, E.; Barak, Y.; Lamed, R.; Bayer, E.A. Clostridium clariflavum: Key cellulosome players are revealed by proteomic analysis. mBio 2015, 6, e00411–00415. [Google Scholar] [CrossRef] [PubMed]
- Tamaru, Y.; Miyake, H.; Kuroda, K.; Nakanishi, A.; Matsushima, C.; Doi, R.H.; Ueda, M. Comparison of the mesophilic cellulosome-producing Clostridium cellulovorans genome with other cellulosome-related clostridial genomes. Microb. Biotechnol. 2011, 4, 64–73. [Google Scholar] [CrossRef] [PubMed]
- Udaondo, Z.; Duque, E.; Ramos, J.L. The pangenome of the genus Clostridium. Environ. Microbiol. 2017, 19, 2588–2603. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Huang, R.; Teng, L.; Jing, X.; Hu, J.; Cui, G.; Wang, Y.; Cui, Q.; Xu, J. Cellulosome stoichiometry in Clostridium cellulolyticum is regulated by selective RNA processing and stabilization. Nat. Commun. 2015, 6, 6900. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schaffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped blast and psi-blast: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Lombard, V.; Golaconda Ramulu, H.; Drula, E.; Coutinho, P.M.; Henrissat, B. The carbohydrate-active enzymes database (CAZy). Nucleic Acids Res. 2014, 42, D490–D495. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer, A.; Bo, Y.; Han, L.; He, J.; Lanczycki, C.J.; Lu, S.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; et al. Cdd/sparcle: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017, 45, D200–D203. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Soding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef] [PubMed]
- Crooks, G.E.; Hon, G.; Chandonia, J.M.; Brenner, S.E. Weblogo: A sequence logo generator. Genome Res. 2004, 14, 1188–1190. [Google Scholar] [CrossRef] [PubMed]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The rast server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef] [PubMed]
- Lefort, V.; Longueville, J.E.; Gascuel, O. SMS: Smart model selection in PhyML. Mol. Biol. Evol. 2017, 34, 2422–2424. [Google Scholar] [CrossRef] [PubMed]
- Nolling, J.; Breton, G.; Omelchenko, M.V.; Makarova, K.S.; Zeng, Q.; Gibson, R.; Lee, H.M.; Dubois, J.; Qiu, D.; Hitti, J.; et al. Genome sequence and comparative analysis of the solvent-producing bacterium Clostridium acetobutylicum. J. Bacteriol. 2001, 183, 4823–4838. [Google Scholar] [CrossRef] [PubMed]
- Kakiuchi, M.; Isui, A.; Suzuki, K.; Fujino, T.; Fujino, E.; Kimura, T.; Karita, S.; Sakka, K.; Ohmiya, K. Cloning and DNA sequencing of the genes encoding Clostridium josui scaffolding protein CipA and cellulase CelD and identification of their gene products as major components of the cellulosome. J. Bacteriol. 1998, 180, 4303–4308. [Google Scholar] [PubMed]
- Pohlschröder, M.; Leschine, S.B.; Canale-Parola, E. Regulation of the multicomplex cellulase-xylanase system of Clostridium papyrosolvens. In Genetics, Biochemistry and Ecology of Lignocellulose Degradation; Shimada, K., Hoshino, S., Ohmiya, K., Sakka, K., Kobayashi, Y., Karita, S., Eds.; Uni Publishers Co., Ltd.: Tokyo, Japan, 1993; pp. 86–94. [Google Scholar]
- Pagès, S.; Belaich, A.; Belaich, J.-P.; Morag, E.; Lamed, R.; Shoham, Y.; Bayer, E.A. Species-specificity of the cohesin-dockerin interaction between Clostridium thermocellum and Clostridium cellulolyticum: Prediction of specificity determinants of the dockerin domain. Proteins 1997, 29, 517–527. [Google Scholar] [CrossRef]
- Sleat, R.; Mah, R.A.; Robinson, R. Isolation and characterization of an anaerobic, cellulolytic bacterium, Clostridium cellulovorans, sp. nov. Appl. Environ. Microbiol. 1984, 48, 88–93. [Google Scholar] [PubMed]
- Lamed, R.; Naimark, J.; Morgenstern, E.; Bayer, E.A. Specialized cell surface structures in cellulolytic bacteria. J. Bacteriol. 1987, 169, 3792–3800. [Google Scholar] [CrossRef] [PubMed]
- Li, L.L.; Taghavi, S.; Izquierdo, J.A.; van der Lelie, D. Complete genome sequence of Clostridium sp. Strain BNL1100, a cellulolytic mesophile isolated from corn stover. J. Bacteriol. 2012, 194, 6982–6983. [Google Scholar] [CrossRef] [PubMed]
- Hahnke, S.; Striesow, J.; Elvert, M.; Mollar, X.P.; Klocke, M. Clostridium bornimense sp. nov., isolated from a mesophilic, two-phase, laboratory-scale biogas reactor. Int. J. Syst. Evol. Microbiol. 2014, 64, 2792–2797. [Google Scholar] [CrossRef] [PubMed]
- Hahnke, S.; Wibberg, D.; Tomazetto, G.; Puhler, A.; Klocke, M.; Schluter, A. Whole genome sequence of Clostridium bornimense strain M2/40 isolated from a lab-scale mesophilic two-phase biogas reactor digesting maize silage and wheat straw. J. Biotechnol. 2014, 184, 199–200. [Google Scholar] [CrossRef] [PubMed]
- Poehlein, A.; Krabben, P.; Durre, P.; Daniel, R. Complete genome sequence of the solvent producer Clostridium saccharoperbutylacetonicum strain DSM 14923. Genome Announc. 2014, 2, e01056-14. [Google Scholar] [CrossRef] [PubMed]
- Lal, S.; Ramachandran, U.; Zhang, X.; Munir, R.; Sparling, R.; Levin, D.B. Draft genome sequence of the cellulolytic, mesophilic, anaerobic bacterium Clostridium termitidis strain CT1112 (DSM 5398). Genome Announc. 2013, 1, e00281-13. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, T.; Ueki, A.; Kaku, N.; Ueki, K. Clostridium sufflavum sp. nov., isolated from a methanogenic reactor treating cattle waste. Int. J. Syst. Evol. Microbiol. 2009, 59, 981–986. [Google Scholar] [CrossRef] [PubMed]
- Zverlov, V.V.; Kellermann, J.; Schwarz, W.H. Functional subgenomics of Clostridium thermocellum cellulosomal genes: Identification of the major catalytic components in the extracellular complex and detection of three new enzymes. Proteomics 2005, 5, 3646–3653. [Google Scholar] [CrossRef] [PubMed]
- Morag, E.; Halevy, I.; Bayer, E.A.; Lamed, R. Isolation and properties of a major cellobiohydrolase from the cellulosome of Clostridium thermocellum. J. Bacteriol. 1991, 173, 4155–4162. [Google Scholar] [CrossRef] [PubMed]
- Ravachol, J.; Borne, R.; Tardif, C.; de Philip, P.; Fierobe, H.P. Characterization of all family-9 glycoside hydrolases synthesized by the cellulosome-producing bacterium Clostridium cellulolyticum. J. Biol. Chem. 2014, 289, 7335–7348. [Google Scholar] [CrossRef] [PubMed]
- Tomazetto, G.; Hahnke, S.; Koeck, D.E.; Wibberg, D.; Maus, I.; Pühler, A.; Klocke, M.; Schluter, A. Complete genome analysis of Clostridium bornimense strain M2/40(T): A new acidogenic Clostridium species isolated from a mesophilic two-phase laboratory-scale biogas reactor. J. Biotechnol. 2016, 232, 38–49. [Google Scholar] [CrossRef] [PubMed]
- Abdou, L.; Boileau, C.; de Philip, P.; Pages, S.; Fierobe, H.P.; Tardif, C. Transcriptional regulation of the Clostridium cellulolyticum cip-cel operon: A complex mechanism involving a catabolite-responsive element. J. Bacteriol. 2008, 190, 1499–1506. [Google Scholar] [CrossRef] [PubMed]
- Asai, K.; Ootsuji, T.; Obata, K.; Matsumoto, T.; Fujita, Y.; Sadaie, Y. Regulatory role of RsgI in sigI expression in Bacillus subtilis. Microbiology 2007, 153, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Ortiz de Ora, L.; Munoz-Gutierrez, I.; Bayer, E.A.; Shoham, Y.; Lamed, R.; Borovok, I. Revisiting the regulation of the primary scaffoldin gene in Clostridium thermocellum. Appl. Environ. Microbiol. 2017, 83, e03088-16. [Google Scholar] [CrossRef] [PubMed]
- Rainey, F.A.; Stackebrandt, E. 16S rDNA analysis reveals phylogenetic diversity among the polysaccharolytic clostridia. FEMS Microbiol. Lett. 1993, 113, 125–128. [Google Scholar] [CrossRef] [PubMed]
- Gal, L.; Pagès, S.; Gaudin, C.; Belaich, A.; Reverbel-Leroy, C.; Tardif, C.; Belaich, J.-P. Characterization of the cellulolytic complex (cellulosome) produced by Clostridium cellulolyticum. Appl. Environ. Microbiol. 1997, 63, 903–909. [Google Scholar] [PubMed]
- Mechaly, A.; Yaron, S.; Lamed, R.; Fierobe, H.P.; Belaich, A.; Belaich, J.P.; Shoham, Y.; Bayer, E.A. Cohesin-dockerin recognition in cellulosome assembly: Experiment versus hypothesis. Proteins 2000, 39, 170–177. [Google Scholar] [CrossRef]
- Schaeffer, F.; Matuschek, M.; Guglielmi, G.; Miras, I.; Alzari, P.M.; Béguin, P. Duplicated dockerin subdomains of Clostridium thermocellum endoglucanase CelD bind to a cohesin domain of the scaffolding protein CipA with distinct thermodynamic parameters and a negative cooperativity. Biochemistry 2002, 41, 2106–2114. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, B.A.; Proctor, M.R.; Martinez-Fleites, C.; Prates, J.A.; Money, V.A.; Davies, G.J.; Bayer, E.A.; Fontesm, C.M.; Fierobe, H.P.; Gilbert, H.J. The Clostridium cellulolyticum dockerin displays a dual binding mode for its cohesin partner. J. Biol. Chem. 2008, 283, 18422–18430. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Huang, R.; Teng, L.; Wang, D.; Hemme, C.L.; Borovok, I.; He, Q.; Lamed, R.; Bayer, E.A.; Zhou, J.; et al. Structure and regulation of the cellulose degradome in Clostridium cellulolyticum. Biotechnol. Biofuels 2013, 6, 73. [Google Scholar] [CrossRef] [PubMed]
- Lamed, R.; Bayer, E.A. The cellulosome concept: Exocellular/extracellular enzyme reactor centers for efficient binding and cellulolysis. In Biochemistry and Genetics of Cellulose Degradation; Aubert, J.-P., Beguin, P., Millet, J., Eds.; Academic Press: London, UK, 1988; pp. 101–116. [Google Scholar]
- Haimovitz, R.; Barak, Y.; Morag, E.; Voronov-Goldman, M.; Lamed, R.; Bayer, E.A. Cohesin-dockerin microarray: Diverse specificities between two complementary families of interacting protein modules. Proteomics 2008, 8, 968–979. [Google Scholar] [CrossRef] [PubMed]
- Bayer, E.A.; Shoham, Y.; Lamed, R. The cellulosome—An exocellular organelle for degrading plant cell wall polysaccharides. In Glycomicrobiology; Doyle, R.J., Ed.; Kluwer Academic/Plenum Publishers: New York, NY, USA, 2000; pp. 387–439. [Google Scholar]
- Bayer, E.A.; Morag, E.; Lamed, R. The cellulosome—A treasure-trove for biotechnology. Trends Biotechnol. 1994, 12, 379–386. [Google Scholar] [CrossRef]
- Davidi, L.; Moraïs, S.; Artzi, L.; Knop, D.; Hadar, Y.; Arfi, Y.; Bayer, E.A. Toward combined delignification and saccharification of wheat straw by a laccase-containing designer cellulosome. Proc. Natl. Acad. Sci. USA 2016, 113, 10854–10859. [Google Scholar] [CrossRef] [PubMed]
- Fierobe, H.-P.; Mingardon, F.; Mechaly, A.; Belaich, A.; Rincon, M.T.; Lamed, R.; Tardif, C.; Belaich, J.-P.; Bayer, E.A. Action of designer cellulosomes on homogeneous versus complex substrates: Controlled incorporation of three distinct enzymes into a defined tri-functional scaffoldin. J. Biol. Chem. 2005, 280, 16325–16334. [Google Scholar] [CrossRef] [PubMed]
- Moraïs, S.; Morag, E.; Barak, Y.; Goldman, D.; Hadar, Y.; Lamed, R.; Shoham, Y.; Wilson, D.B.; Bayer, E.A. Deconstruction of lignocellulose into soluble sugars by native and designer cellulosomes. mBio 2012, 3, e00508-12. [Google Scholar] [CrossRef] [PubMed]
- Stern, J.; Moraïs, S.; Lamed, R.; Bayer, E.A. Adaptor scaffoldins: An original strategy for extended designer cellulosomes, inspired from nature. mBio 2016, 7, e00083-16. [Google Scholar] [CrossRef] [PubMed]
| Species | Genome Accession | Genome Sequencing Level | No. of Contigs | Scaffoldins | Cohesins | Dockerins | GHs | PLs | CEs | CBMs | Total CAZYmes | Source | References |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Clostridium sp. BNL1100 | CP003259.1 | Complete | 1 | 2 | 7 | 88 | 103 | 5 | 19 | 67 | 127 | Corn stover | [42] |
| C. josui JCM17888 | JAGE00000000.1 | Draft | 2 | 3 | 8 | 72 | 92 | 5 | 19 | 59 | 116 | Compost | [37] |
| C. cellulolyticum H10 | CP001348.1 | Complete | 1 | 2 | 9 | 69 | 94 | 4 | 13 | 54 | 111 | Compost | [39] |
| C. papyrosolvens DSM 2782 | ACXX00000000.2 | Draft | 31 | 2 | 7 | 68 | 103 | 3 | 16 | 71 | 122 | Paper mill | [38] |
| C. sufflavum DSM 19573 | PRJNA262320 | Draft | 57 | 5 (6) | 15 | 56 | 60 | 2 | 18 | 42 | 80 | Methanogenic reactor | [47] |
| C. cellulovorans 743B | CP002160.1 | Complete | 1 | 5 | 12 | 46 | 118 | 15 | 21 | 59 | 154 | Wood fermenter | [40] |
| C. cellobioparum DSM 1351 | JHYD01000000.1 | Draft | 80 | 3 | 3 | 38 | 182 | 3 | 19 | 78 | 204 | Rumen of cattle | [41] |
| C. termitidis CT1112 | AORV00000000.1 | Draft | 78 | 7 | 8 | 28 | 199 | 4 | 15 | 95 | 218 | Gut of termite | [46] |
| C. acetobutylicum DSM 1731 | CP002660.1 | Complete | 1 | 2 | 6 | 10 | 75 | 6 | 17 | 34 | 98 | Soil | [36] |
| C. saccharoperbutylacetonicum N1-4 (HMT) | CP004121.1 | Complete | 1 | 2 | 3 | 9 | 146 | 6 | 22 | 22 | 174 | Soil | [45] |
| C. bornimense (=Clostridium sp. M2/40) | HG917868.1, HG917869.1 | Draft | 2 | 2 | 8 | 5 | 47 | 6 | 7 | 50 | 60 | Biogas reactor | [43] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dassa, B.; Borovok, I.; Lombard, V.; Henrissat, B.; Lamed, R.; Bayer, E.A.; Moraïs, S. Pan-Cellulosomics of Mesophilic Clostridia: Variations on a Theme. Microorganisms 2017, 5, 74. https://doi.org/10.3390/microorganisms5040074
Dassa B, Borovok I, Lombard V, Henrissat B, Lamed R, Bayer EA, Moraïs S. Pan-Cellulosomics of Mesophilic Clostridia: Variations on a Theme. Microorganisms. 2017; 5(4):74. https://doi.org/10.3390/microorganisms5040074
Chicago/Turabian StyleDassa, Bareket, Ilya Borovok, Vincent Lombard, Bernard Henrissat, Raphael Lamed, Edward A. Bayer, and Sarah Moraïs. 2017. "Pan-Cellulosomics of Mesophilic Clostridia: Variations on a Theme" Microorganisms 5, no. 4: 74. https://doi.org/10.3390/microorganisms5040074
APA StyleDassa, B., Borovok, I., Lombard, V., Henrissat, B., Lamed, R., Bayer, E. A., & Moraïs, S. (2017). Pan-Cellulosomics of Mesophilic Clostridia: Variations on a Theme. Microorganisms, 5(4), 74. https://doi.org/10.3390/microorganisms5040074






