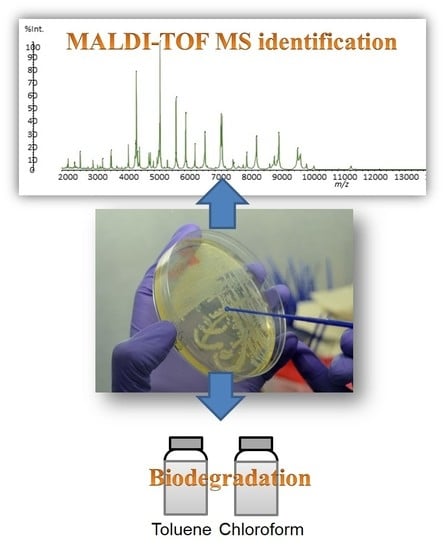
MALDI-TOF MS for the Identification of Cultivable Organic-Degrading Bacteria in Contaminated Groundwater near Unconventional Natural Gas Extraction Sites
Abstract
:1. Introduction
2. Materials and Methods
2.1. Reagents
2.2. Groundwater Isolates
2.3. MALDI-TOF MS
2.4. Identification of the Isolated Strains
2.5. Biodegradation Studies of Toluene and Chloroform by the Isolated Bacteria
2.6. Antibiotic Susceptibility Tests
3. Results and Discussion
3.1. Water Quality
3.2. Bacteria Identification by MALDI-MS
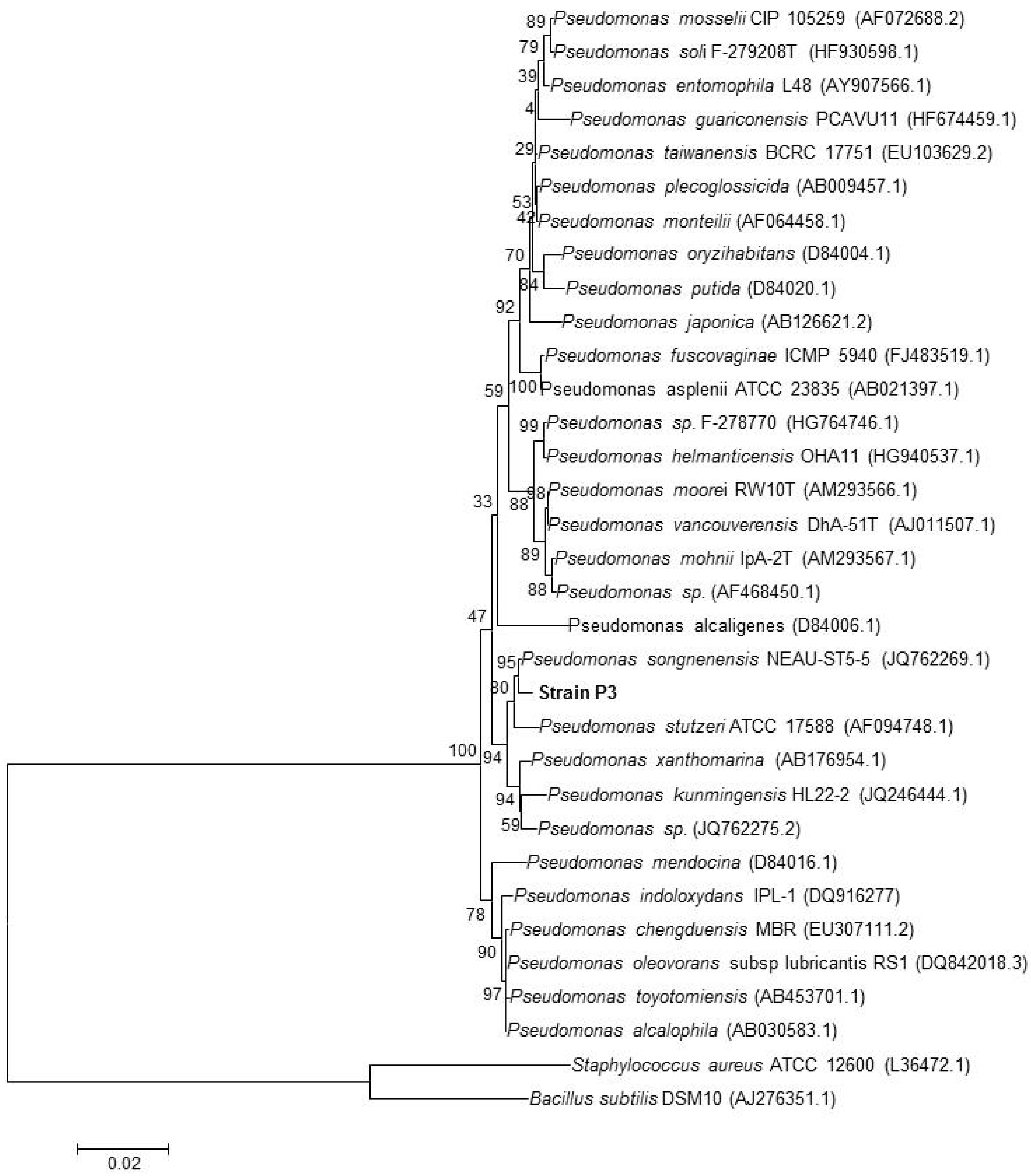
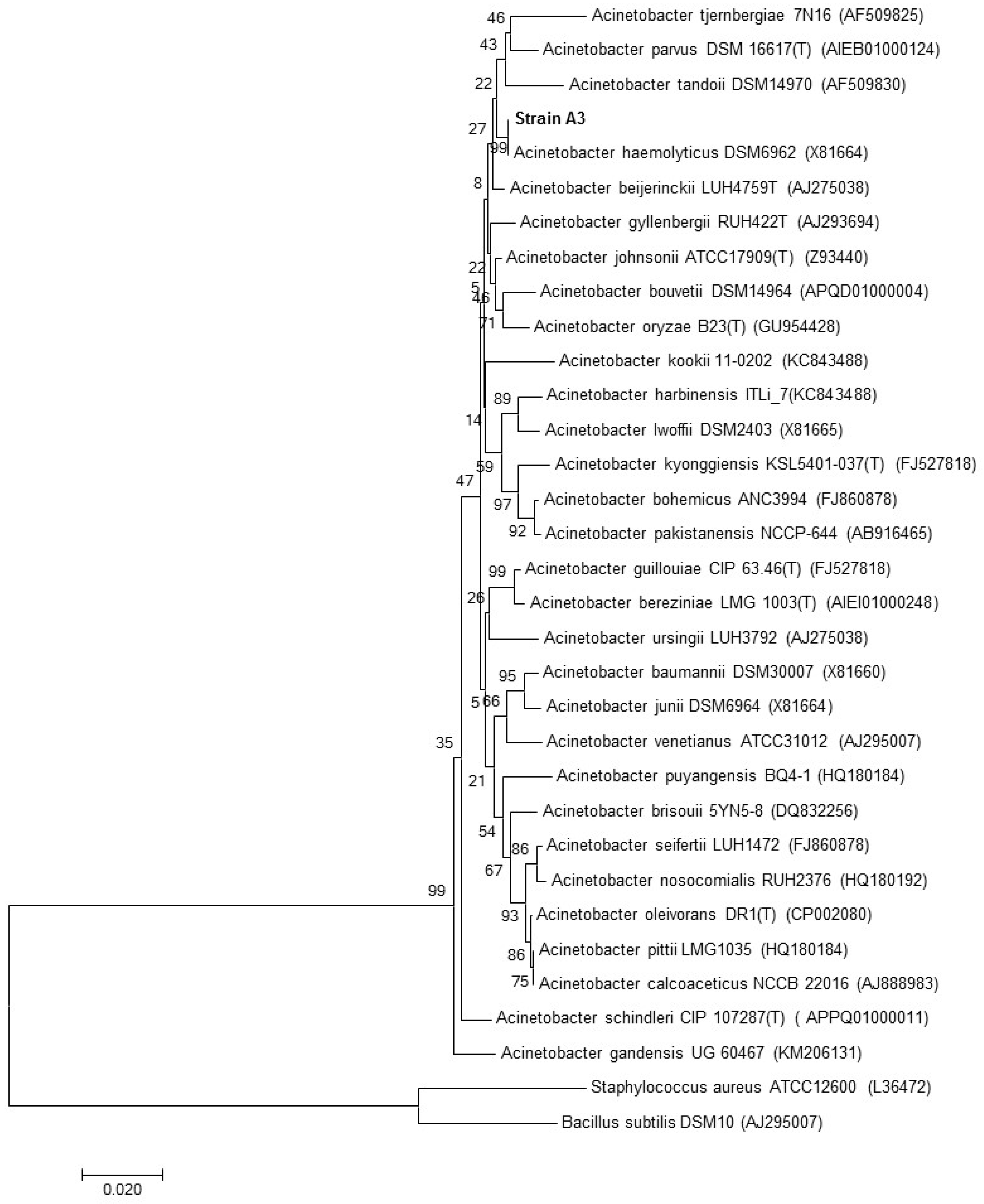
3.3. Identification of the Strains by 16S rRNA Gene Sequencing
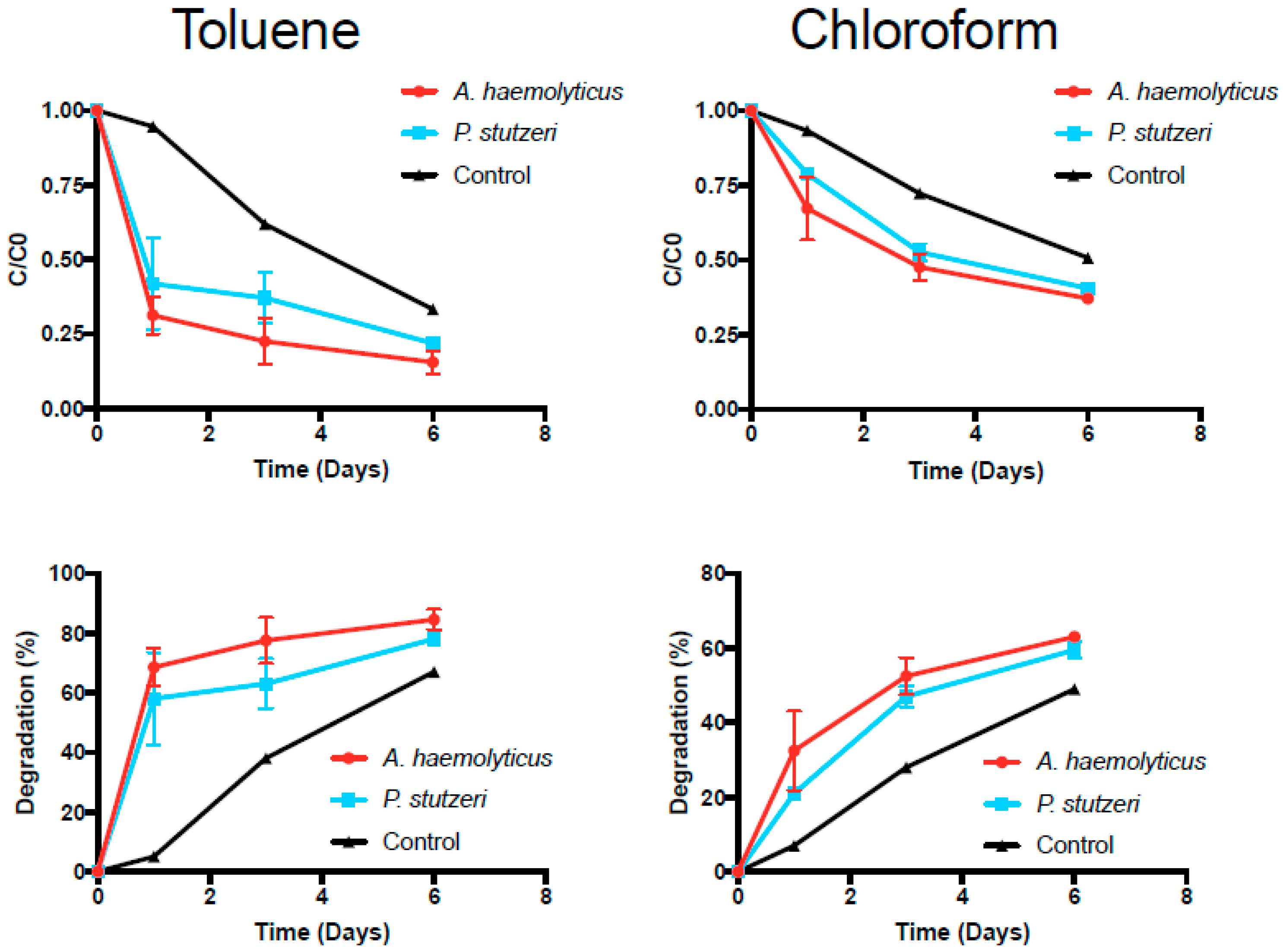
3.4. Toluene and Chloroform Degradation Studies
3.5. Antibiotic Susceptibility
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Centers for Disease Control and Prevention. Private Ground Water Wells. Available online: http://www.cdc.gov/healthywater/drinking/private/wells/ (accessed on 5 January 2015).
- United States Environmental Protection Agency. Ground Water. Available online: http://water.epa.gov/type/groundwater/ (accessed on 5 January 2015).
- United States Environmental Protection Agency. Human Health. Available online: http://water.epa.gov/drink/info/well/health.cfm (accessed on 5 January 2015).
- Sovacool, B.K. Cornucopia or curse? Reviewing the costs and benefits of shale gas hydraulic fracturing (fracking). Renew. Sustain. Energ. Rev. 2014, 37, 249–269. [Google Scholar] [CrossRef]
- FracFocus. Hydraulic Fracturing: The Process. Available online: http://fracfocus.org/hydraulic-fracturing-how-it-works/hydraulic-fracturing-process (accessed on 6 January 2015).
- Melikoglu, M. Shale gas: Analysis of its role in the global energy market. Renew. Sustain. Energ. Rev. 2014, 37, 460–468. [Google Scholar] [CrossRef]
- Vidic, R.D.; Brantley, S.L.; Vandenbossche, J.M.; Yoxtheimer, D.; Abad, J.D. Impact of shale gas development on regional water quality. Science 2013, 340, 1235009. [Google Scholar] [CrossRef] [PubMed]
- Fontenot, B.E.; Hunt, L.R.; Hildenbrand, Z.L.; Carlton, D.D., Jr.; Oka, H.; Walton, J.L.; Hopkins, D.; Osorio, A.; Bjorndal, B.; Hu, Q.H.; et al. An evaluation of water quality in private drinking water wells near natural gas extraction sites in the Barnett shale formation. Environ. Sci. Technol. 2013, 47, 10032–10040. [Google Scholar] [CrossRef] [PubMed]
- Rozell, D.J.; Reaven, S.J. Water pollution risk associated with natural gas extraction from the Marcellus shale. Risk Anal. 2012, 32, 1382–1393. [Google Scholar] [CrossRef] [PubMed]
- Osborn, S.G.; Vengosh, A.; Warner, N.R.; Jackson, R.B. Methane contamination of drinking water accompanying gas-well drilling and hydraulic fracturing. Proc. Natl. Acad. Sci. USA 2011, 108, 8172–8176. [Google Scholar] [CrossRef] [PubMed]
- Warner, N.R.; Jackson, R.B.; Darrah, T.H.; Osborn, S.G.; Down, A.; Zhao, K.; White, A.; Vengosh, A. Geochemical evidence for possible natural migration of Marcellus Formation brine to shallow aquifers in Pennsylvania. Proc. Natl. Acad. Sci. USA 2012, 109, 11961–11966. [Google Scholar] [CrossRef] [PubMed]
- Hildenbrand, Z.L.; Carlton, D.D., Jr.; Fontenot, B.E.; Meik, J.M.; Walton, J.L.; Taylor, J.T.; Thacker, J.B.; Korlie, S.; Shelor, C.P.; Henderson, D.; et al. A Comprehensive Analysis of Groundwater Quality in The Barnett Shale Region. Environ. Sci. Technol. 2015, 49, 8254–8262. [Google Scholar] [CrossRef] [PubMed]
- Simon, J.A.; Fleming, M.E. Editor’s perspective-Shale gas development: Environmental issues and opportunities. Remediation 2011, 21, 1–10. [Google Scholar] [CrossRef]
- Cluff, M.A.; Hartsock, A.; MacRae, J.D.; Carter, K.; Mouser, P.J. Temporal changes in microbial ecology and geochemistry in produced water from hydraulically fractured Marcellus shale gas wells. Environ. Sci. Technol. 2014, 48, 6508–6517. [Google Scholar] [CrossRef] [PubMed]
- Davis, J.P.; Struchtemeyer, C.G.; Elshahed, M.S. Bacterial communities associated with production facilities of two newly drilled thermogenic natural gas wells in the Barnett shale (Texas, USA). Microb. Ecol. 2012, 64, 942–954. [Google Scholar] [CrossRef] [PubMed]
- Kahrilas, G.A.; Blotevogel, J.; Stewart, P.S.; Borch, T. Biocides in hydraulic fluids: A critical review of their usage, mobility, degradation, and toxicity. Environ. Sci. Technol. 2015, 49, 16–32. [Google Scholar] [CrossRef] [PubMed]
- Ohio State University. DNA Analysis of Microbes in A Fracking Site Yields Surprises. Available online: www.sciencedaily.com/releases/2012/12/121203145834.htm (accessed on 3 December 2012).
- Vidali, M. Bioremediation. An overview. Pure Appl. Chem. 2001, 73, 1163–1172. [Google Scholar] [CrossRef]
- Farhadian, M.; Duchez, D.; Vachelard, C.; Larroche, C. Monoaromatics removal from polluted water through bioreactors-A review. Water Res. 2008, 42, 1325–1341. [Google Scholar] [CrossRef] [PubMed]
- Karigar, C.S.; Rao, S.S. Role of microbial enzymes in the bioremediation of pollutants: A review. Enzym. Res. 2011. [Google Scholar] [CrossRef] [PubMed]
- Dingle, T.C.; Butler-Wu, S.M. MALDI-TOF mass spectrometry for microorganism identification. Clin. Lab. Med. 2013, 33, 589–609. [Google Scholar] [CrossRef] [PubMed]
- Dalluge, J.J. Mass spectrometry for direct determination of proteins in cells: Applications in biotechnology and microbiology. Fresenius J. Anal. Chem. 2000, 366, 701–711. [Google Scholar] [CrossRef] [PubMed]
- Fenselau, C.C. Rapid characterization of microorganisms by mass spectrometry–What can be learned and how? J. Am. Soc. Mass Spectrom. 2013, 24, 1161–1166. [Google Scholar] [CrossRef] [PubMed]
- Fenselau, C.; Demirev, P.A. Characterization of intact microorganisms by MALDI mass spectrometry. Mass Spectrom. Rev. 2001, 20, 157–171. [Google Scholar] [CrossRef] [PubMed]
- Kliem, M.; Sauer, S. The essence on mass spectrometry based microbial diagnostics. Curr. Opin. Microbiol. 2012, 15, 397–402. [Google Scholar] [CrossRef] [PubMed]
- Krásný, L.; Hynek, R.; Hochel, I. Identification of bacteria using mass spectrometry techniques. Int. J. Mass Spectrom. 2013, 353, 67–79. [Google Scholar] [CrossRef]
- Lay, J.O., Jr. MALDI-TOF mass spectrometry of bacteria. Mass Spectrom. Rev. 2001, 20, 172–194. [Google Scholar] [CrossRef] [PubMed]
- Havlicek, V.; Lemr, K.; Schug, K.A. Current trends in microbial diagnostics based on mass spectrometry. Anal. Chem. 2013, 85, 790–797. [Google Scholar] [CrossRef] [PubMed]
- Amann, R.I.; Ludwig, W.; Schleifer, K.H. Phylogenetic identification and in situ detection of individual cells without cultivation. Microbiol. Rev. 1995, 59, 143–169. [Google Scholar] [PubMed]
- APHA-AWWA-WPCF. Standard Methods for the Examination of Water and Wastewater, 20th ed.; American Public Health Association: Washington, DC, USA, 1998; Chapter 9; pp. 67–78. [Google Scholar]
- Freiwald, A.; Sauer, S. Phylogenetic classification and identification of bacteria by mass spectrometry. Nat. Protoc. 2009, 4, 732–742. [Google Scholar] [CrossRef] [PubMed]
- Ryzhov, V.; Fenselau, C. Characterization of the protein subset desorbed by MALDI from whole bacterial cells. Anal. Chem. 2001, 73, 746–750. [Google Scholar] [CrossRef] [PubMed]
- Amorim, C.L.; Ferreira, A.C.S.; Carvalho, M.F.; Afonso, C.M.M.; Castro, P.M.L. Mineralization of 4-fluorocinnamic acid by a Rhodococcus strain. Appl. Microbiol. Biotechnol. 2014, 98, 1893–1905. [Google Scholar] [CrossRef] [PubMed]
- Amorim, C.L.; Moreira, I.S.; Maia, A.S.; Tiritan, M.E.; Castro, P.M.L. Biodegradation of ofloxacin, norfloxacin, and ciprofloxacin as single and mixed substrates by Labrys portucalensis F11. Appl. Microbiol. Biotechnol. 2014, 98, 3181–3190. [Google Scholar] [CrossRef] [PubMed]
- Andrews, J.M. Determination of minimum inhibitory concentrations. J. Antimicrob. Chemother. 2001, 48 (Suppl. 1), S5–S16. [Google Scholar] [CrossRef]
- Chemicals Used in Hydraulic Fracturing. United States House of Representatives Committee on Energy and Commerce. Washington, DC, USA. Available online: http://www.conservation.ca.gov/dog/general_information/Documents/Hydraulic%20Fracturing%20Report%204%2018%2011.pdf (accessed on 8 August 2017).
- WHO. Benzene in Drinking-Water. Background Document for Development of WHO Guidelines for Drinking-Water Quality. World Health Organization: Geneva, Switzerland, 2003. Available online: http://www.who.int/water_sanitation_health/dwq/benzene.pdf (accessed on 5 January 2015).
- United States Environmental Protection Agency. Chloroform. Available online: http://www3.epa.gov/airtoxics/hlthef/chlorofo.html (accessed on 5 January 2015).
- United States Environmental Protection Agency. Drinking Water Contaminants. Available online: http://water.epa.gov/drink/contaminants/ (accessed on 5 January 2015).
- Lalucat, J.; Bennasar, A.; Bosch, R.; García-Valdés, E.; Palleroni, N.J. Biology of Pseudomonas stutzeri. Microbiol. Mol. Biol. Rev. 2006, 70, 510–547. [Google Scholar] [CrossRef] [PubMed]
- Potvliege, C.; Jonckheer, J.; Lenclud, C.; Hansen, W. Pseudomonas stutzeri pneumonia and septicemia in a patient with multiple myeloma. J. Clin. Microbiol. 1987, 25, 458–459. [Google Scholar] [PubMed]
- Goetz, A.; Yu, V.L.; Hanchett, J.E.; Rihs, J.D. Pseudomonas stutzeri bacteremia associated with hemodialysis. Arch. Intern. Med. 1983, 143, 1909–1912. [Google Scholar] [CrossRef] [PubMed]
- Vanbroekhoven, K.; Ryngaert, A.; Wattiau, P.; De Mot, R.; Springael, D. Acinetobacter diversity in environmental samples assessed by 16S rRNA gene PCR-DGGE fingerprinting. FEMS Microbiol. Ecol. 2004, 50, 37–50. [Google Scholar] [CrossRef] [PubMed]
- Bélanger, S.D.; Boissinot, M.; Ménard, C.; Picard, F.J.; Bergeron, M.G. Rapid detection of shiga toxin-producing bacteria in feces by multiplex PCR with molecular beacons on the smart cycler. J. Clin. Microbiol. 2002, 40, 1436–1440. [Google Scholar] [CrossRef] [PubMed]
- Grotiuz, G.; Sirok, A.; Gadea, P.; Varela, G.; Schelotto, F. Shiga toxin 2-Producing Acinetobacter haemolyticus associated with a case of bloody diarrhea. J. Clin. Microbiol. 2006, 44, 3838–3841. [Google Scholar] [CrossRef] [PubMed]
- Wasi, S.; Tabrez, S.; Ahmad, M. Use of Pseudomonas spp. or the bioremediation of environmental pollutants: A review. Environ. Monit. Assess. 2013, 185, 8147–8155. [Google Scholar] [CrossRef] [PubMed]
- Fritsche, W.; Hofrichter, M. Aerobic Degradation by Microorganisms. In Biotechnology: Environmental Processes II, 2nd ed.; Rehm, H.-J., Reed, G., Eds.; Wiley-VCH Verlag GmbH: Weinheim, Germany, 2000; Volume 11b, Chapter 6; pp. 146–155. [Google Scholar]
- McClay, K.; Fox, B.G.; Steffan, R.J. Chloroform mineralization by toluene-oxidizing bacteria. Appl. Environ. Microbiol. 1996, 62, 2716–2722. [Google Scholar] [PubMed]
- Chauhan, S.; Barbieri, P.; Wood, T.K. Oxidation of trichloroethylene, 1,1-dichloroethylene, and chloroform by toluene/o-xylene monooxygenase from Pseudomonas stutzeri OX1. Appl. Environ. Microbiol. 1998, 64, 3023–3024. [Google Scholar] [PubMed]
- Cappelletti, M.; Frascari, D.; Zannoni, D.; Fedi, S. Microbial degradation of chloroform. Appl. Microbiol. Biotechnol. 2012, 96, 1395–1409. [Google Scholar] [CrossRef] [PubMed]
- Jindrova, E.; Chocová, M.; Demnerová, K.; Brenner, V. Bacterial aerobic degradation of benzene, toluene, ethylbenzene and xylene. Folia Microbiol. 2002, 47, 83–93. [Google Scholar] [CrossRef]
- Vaz-Moreira, I.; Nunes, O.C.; Manaia, C.M. Diversity and antibiotic resistance in Pseudomonas spp. from drinking water. Sci. Total Environ. 2012, 426, 366–374. [Google Scholar] [CrossRef] [PubMed]
- Gellatly, S.L.; Hancock, R.E.W. Pseudomonas aeruginosa: New insights into pathogenesis and host defenses. Pathog. Dis. 2013, 67, 159–173. [Google Scholar] [CrossRef] [PubMed]
- Galindo, E.; Ramírez, O.R. Advances in Bioprocess Engineering; Springer Science & Business Media: Dordrecht, The Netherlands, 2013; Volume 2, pp. 219–222. [Google Scholar]
- Espinal, P.; Roca, I.; Vila, J. Clinical impact and molecular basis of antimicrobial resistance in non-baumannii Acinetobacter. Future Microbiol. 2011, 6, 495–511. [Google Scholar] [CrossRef] [PubMed]


| Sample ID | Depth (ft) | Temp (°C) | DO (mg/L) | Specific Conductance (mS/cm) | TDS (mg/L) | Salinity (mg/L) | pH | Water Use |
|---|---|---|---|---|---|---|---|---|
| 314 | 500 | 21.1 | 29.2 | 1.01 | 657 | 0.50 | 9.04 | Residential |
| 316 | 350–400 | 17.4 | 3.66 | 0.77 | 501 | 0.38 | 8.78 | Residential |
| 346 | - | 19.8 | 1.23 | 1.01 | 657 | 0.50 | 8.94 | Residential |
| 369 | 300 | 22.3 | 1.70 | 0.97 | 631 | 0.48 | 8.98 | Residential |
| 377 | 1580 | 30.1 | 1.56 | 1.11 | 722 | 0.55 | 8.58 | Municipal |
| 380 | 3370 | 31 | 1.57 | 1.52 | 988 | 0.74 | 8.08 | Municipal |
| 526 | 735 | 25.8 | 1.26 | 0.88 | 572 | 0.43 | 9.42 | Residential |
| (mg/L) | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample ID | TOC | IC | TN | MeOH | EtOH | DCM | Propargyl Alcohol | Acetone | ACN | Ethyl Acetate | Chloroform | Cyclo-hexane | Benzene | Toluene | Ethyl Benzene | m,p-Xylene | o-Xylene |
| 314 | 0.1 | 91.2 | 0.5 | 44.6 | 0.874 | 0.000 | 0.000 | 0.740 | 23.0 | 1.14 | 0.000 | 0.017 | 0.000 | 0.000 | 0.007 | 0.009 | 0.017 |
| 316 | 0.2 | 94.6 | 0.5 | 2.17 | 7.03 | 0.000 | 22.1 | 0.331 | 2.57 | 1.19 | 0.000 | 0.014 | 0.083 | 0.000 | 0.002 | 0.000 | 0.023 |
| 346 | 0.1 | 0.1 | 0.5 | 0.00 | 0.000 | 0.216 | 0.000 | 0.000 | 0.037 | 0.009 | 2.14 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.009 |
| 369 | 0.1 | 0.1 | 0.5 | 0.00 | 0.000 | 0.136 | 0.000 | 0.000 | 0.028 | 0.000 | 0.000 | 0.014 | 0.000 | 0.000 | 0.000 | 0.003 | 0.010 |
| 377 | 0.4 | 85.8 | 0.5 | 0.00 | 3.71 | 0.203 | 0.000 | 0.178 | 1.90 | 6.41 | 0.000 | 0.000 | 0.000 | 0.000 | 0.012 | 0.005 | 0.006 |
| 380 | 2.2 | 119 | 0.7 | 0.00 | 0.413 | 0.144 | 0.000 | 0.026 | 0.019 | 0.000 | 0.000 | 0.013 | 0.048 | 0.051 | 0.007 | 0.000 | 0.010 |
| 526 | 0.0 | 89.4 | 0.6 | 0.00 | 0.000 | 0.000 | 0.000 | 0.000 | 0.031 | 0.039 | 0.000 | 0.000 | 0.000 | 0.001 | 0.000 | 0.000 | 0.005 |
| A. haemolyticus | P. stutzeri | E. coli | |||
|---|---|---|---|---|---|
| m/z | Intensity | m/z | Intensity | m/z | Intensity |
| 2131.99 | 4,269,529 | 2420.83 | 2,147,092 | 2547.83 | 2,131,223 |
| 2482.59 | 1,506,536 | 2788.75 | 4,242,038 | 3127.48 | 5,397,132 |
| 2806.23 | 10,347,161 | 2984.35 | 3,109,296 | 3157.67 | 2,467,099 |
| 3052.56 | 3,607,775 | 3229.38 | 6,287,977 | 3163.06 | 1,425,940 |
| 3725.27 | 15,720,611 | 3571.07 | 12,904,611 | 4364.47 | 26,901,980 |
| 3738.74 | 1,824,080 | 3584.09 | 3,649,719 | 4768.65 | 5,157,224 |
| 3763.72 | 4,437,623 | 3635.36 | 11,766,971 | 4777.67 | 4,935,549 |
| 4265.64 | 30,881,306 | 3934.45 | 3,198,564 | 4870.50 | 3,468,297 |
| 4281.13 | 3,399,151 | 4095.20 | 6,913,595 | 5381.71 | 14,711,287 |
| 4648.25 | 27,630,984 | 4536.57 | 12,977,799 | 5613.05 | 2,307,677 |
| 4690.28 | 5,806,014 | 4851.46 | 6,236,720 | 6255.81 | 24,465,422 |
| 5417.54 | 6,592,444 | 5001.59 | 4,327,399 | 6267.09 | 5,618,275 |
| 5526.04 | 3,239,353 | 5585.99 | 9,931,623 | 6294.67 | 6,707,362 |
| 6106.49 | 14,694,100 | 5976.90 | 6,644,226 | 6316.83 | 13,191,460 |
| 6643.96 | 2,894,466 | 6162.98 | 3,457,942 | 6354.56 | 1,996,031 |
| 7409.36 | 2,194,172 | 6466.64 | 10,736,716 | 7275.17 | 11,564,301 |
| 7451.24 | 42,138,924 | 7151.01 | 30,722,710 | 7721.46 | 89,658.7891 |
| 7478.33 | 6,692,804 | 7178.22 | 11,632,331 | 8327.55 | 2,868,725 |
| 8187.53 | 5,622,694 | 7279.12 | 23,694,890 | 8370.46 | 7,628,921 |
| 8200.57 | 2,720,580 | 7306.05 | 3,114,205 | 9065.06 | 5,520,206 |
| 8905.78 | 1,443,282 | 7853.02 | 3,044,757 | 9227.75 | 9,474,368 |
| 9296.67 | 45,184,156 | 8199.13 | 28,981,870 | 9537.25 | 14,085,582 |
| 9324.89 | 10,356,339 | 8228.94 | 12,505,299 | 9555.28 | 15,322,127 |
| 10,080.44 | 1,686,673 | 8434.89 | 5,943,598 | 9741.23 | 15,633,452 |
| 10,834.33 | 5,093,954 | 9081.59 | 14,799,761 | 10,140.12 | 1,406,545 |
| 10,862.14 | 769,789 | 9711.42 | 3,061,340 | 10,653.57 | 863,826 |
| 12,050.36 | 14,516,812 | 10,569.06 | 527,712 | 11,225.28 | 2,137,546 |
| 12,078.37 | 3,561,288 | 11,838.82 | 1,172,665 | 11,452.91 | 484,301 |
| 12,473.59 | 939,518 | ||||
| Sample ID | Strain P3 cfu/mL ± SD | Strain A3 cfu/mL ± SD |
|---|---|---|
| 314 | 9.2 (± 0.6) × 103 | - |
| 316 | 2.0 (± 0.2) × 102 | - |
| 346 | 1.68 (± 0.009) × 103 | 21 ± 5 |
| 369 | 20 ± 3 | 3 ± 1 |
| 377 | 2.1 (± 0.1) × 103 | - |
| 380 | 55 ± 9 | - |
| 526 | 110 ± 1 | - |
| Bacteria | Toluene | Chloroform | ||
|---|---|---|---|---|
| k (day−1) | t1/2 (day) | k (day−1) | t1/2 (day) | |
| Acinetobacter sp. A3 | 0.67 ± 0.12 | 1.41 ± 0.24 | 0.27 ± 0.04 | 2.96 ± 0.12 |
| Pseudomonas sp. P3 | 0.50 ± 0.15 | 1.90 ± 0.31 | 0.20 ± 0.00 | 3.66 ± 0.03 |
| Family | Antibiotic | Pseudomonas sp. P3 | Acinetobacter sp. A3 |
|---|---|---|---|
| Aminoglycoside | Streptomycin | R | R |
| β-lactam, extended spectrum penicillin | Piperacillin | S | S |
| β-lactam, semisynthetic penicillin | Oxacillin | S | S |
| Glycopeptide | Vancomycin | R | R |
| β-lactam, extended spectrum penicillin | Ampicillin | S | R |
| Macrolide | Erythromycin | R | R |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Santos, I.C.; Martin, M.S.; Carlton, D.D.; Amorim, C.L.; Castro, P.M.L.; Hildenbrand, Z.L.; Schug, K.A. MALDI-TOF MS for the Identification of Cultivable Organic-Degrading Bacteria in Contaminated Groundwater near Unconventional Natural Gas Extraction Sites. Microorganisms 2017, 5, 47. https://doi.org/10.3390/microorganisms5030047
Santos IC, Martin MS, Carlton DD, Amorim CL, Castro PML, Hildenbrand ZL, Schug KA. MALDI-TOF MS for the Identification of Cultivable Organic-Degrading Bacteria in Contaminated Groundwater near Unconventional Natural Gas Extraction Sites. Microorganisms. 2017; 5(3):47. https://doi.org/10.3390/microorganisms5030047
Chicago/Turabian StyleSantos, Inês C., Misty S. Martin, Doug D. Carlton, Catarina L. Amorim, Paula M. L. Castro, Zacariah L. Hildenbrand, and Kevin A. Schug. 2017. "MALDI-TOF MS for the Identification of Cultivable Organic-Degrading Bacteria in Contaminated Groundwater near Unconventional Natural Gas Extraction Sites" Microorganisms 5, no. 3: 47. https://doi.org/10.3390/microorganisms5030047
APA StyleSantos, I. C., Martin, M. S., Carlton, D. D., Amorim, C. L., Castro, P. M. L., Hildenbrand, Z. L., & Schug, K. A. (2017). MALDI-TOF MS for the Identification of Cultivable Organic-Degrading Bacteria in Contaminated Groundwater near Unconventional Natural Gas Extraction Sites. Microorganisms, 5(3), 47. https://doi.org/10.3390/microorganisms5030047