Diversity and Control of Spoilage Fungi in Dairy Products: An Update
Abstract
1. Introduction
2. Diversity of Spoilage Fungi in Dairy Products
2.1. Sources of Fungal Contamination
2.2. Spoilage Yeasts
2.3. Molds in Dairy Products
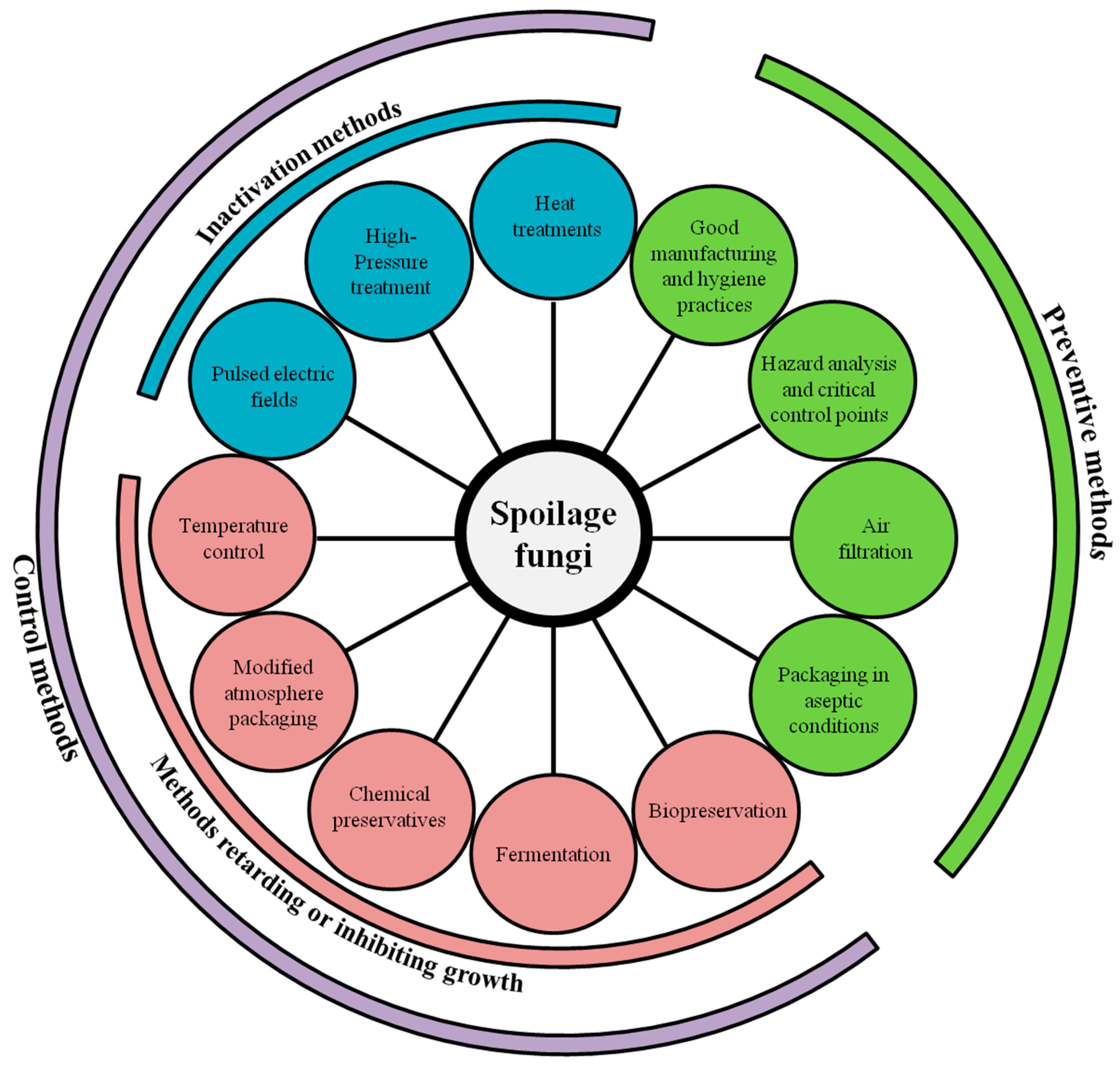
3. Prevention and Control of Fungal Spoilage in Dairy Products
3.1. Preventive Methods
3.1.1. Good Manufacturing and Distribution Practices
3.1.2. Air Filtration and Decontamination Systems
3.2. Control Methods
3.2.1. Inactivation Treatments
3.2.2. Temperature Control
3.2.3. Modified Atmosphere Packaging
3.2.4. Chemical Preservatives
3.2.5. Fermentation
4. Bioprotective Cultures
4.1. Lactic Acid Bacteria and Propionibacteria with Antifungal Activity
4.2. Fermentates
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- FAO. Global FOOD Losses and Food Waste—Extent, Causes and Prevention. Rome. 2011. Available online: http://www.fao.org/docrep/014/mb060e/mb060e00.pdf (accessed on 5 July 2017).
- Pitt, J.I.; Hocking, A.D. Fungi and Food Spoilage, 3rd ed.; Springer Science & Business Media: New York, NY, USA, 2009; pp. 1–2. ISBN 978-1-4615-6391-4. [Google Scholar]
- Shephard, G.S. Impact of mycotoxins on human health in developing countries. Food Addit. Contam. A 2008, 25, 146–151. [Google Scholar] [CrossRef] [PubMed]
- Filtenborg, O.; Frisvad, J.C.; Thrame, U. Moulds in food spoilage. Int. J. Food Microbiol. 1996, 33, 85–102. [Google Scholar] [CrossRef]
- Huis in’t Veld, J.H.J. Microbial and biochemical spoilage of foods: An overview. Int. J. Food Microbiol. 1996, 33, 1–18. [Google Scholar] [CrossRef]
- Prado, M.R.; Blandón, L.M.; Vandenberghe, L.P.S.; Rodrigues, C.; Castro, G.R.; Thomas-Soccol, V.; Soccol, C.R. Milk kefir: Composition, microbial cultures, biological activities and related products. Front. Microbiol. 2015, 6, 1177. [Google Scholar] [CrossRef] [PubMed]
- Desmasures, N. CHEESE-mold-ripened varieties. In Encyclopedia of Food Microbiology, 2nd ed.; Batt, C.A., Ed.; Elsevier Inc.: Atlanta, GA, USA, 2014; pp. 409–415. ISBN 9780123847331. [Google Scholar]
- Baroiller, C.; Schmidt, J.L. Contribution à l’étude de l’origine des levures du fromage de Camembert. Lait 1990, 70, 67–84. [Google Scholar] [CrossRef]
- Ledenbach, L.H.; Marshall, R.T. Microbiological spoilage of dairy products. In Compendium of the Microbiological Spoilage of Foods and Beverages, Food Microbiology and Food Safety; Sperber, W.H., Doyle, M.P., Eds.; Springer: New York, NY, USA, 2010; pp. 41–67. ISBN 1441908269, 9781441908261. [Google Scholar]
- Hymery, N.; Vasseur, V.; Coton, M.; Mounier, J.; Jany, J.L.; Barbier, G.; Coton, E. Filamentous fungi and mycotoxins in cheese: A review. Compr. Rev. Food Sci. Food Saf. 2014, 13, 437–456. [Google Scholar] [CrossRef]
- Westall, S.; Filtenborg, O. Spoilage yeasts of decorated soft cheese packed in modified atmosphere. Food Control 1998, 21, 136–142. [Google Scholar] [CrossRef]
- Ross, R.P.; Morgan, S.; Hill, C. Preservation and fermentation: Past, present and future. Int. J. Food Microbiol. 2002, 79, 3–16. [Google Scholar] [CrossRef]
- Stratford, M.; Steels, H.; Nebe-von-Caron, G.; Avery, S.V.; Novodvorska, M.; Archer, D.B. Population heterogeneity and dynamics in starter culture and lag phase adaptation of the spoilage yeast Zygosaccharomyces bailii to weak acid preservatives. Int. J. Food Microbiol. 2014, 181, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Brul, S.; Coote, P. Preservatives agents in foods: Mode of action and microbial resistance mechanisms. Int. J. Food Microbiol. 1999, 50, 1–17. [Google Scholar] [CrossRef]
- Delavenne, E.; Cliquet, S.; Trunet, C.; Barbier, G.; Mounier, J.; Le Blay, G. Characterization of the antifungal activity of Lactobacillus harbinensis K.V9.3.1.Np and Lactobacillus rhamnosus K. C8.3.1I in yoghurt. Food Microbiol. 2015, 45, 10–17. [Google Scholar] [CrossRef] [PubMed]
- Schoch, C.L.; Seifert, K.A.; Huhndorf, S.; Robert, V.; Spouge, J.L.; Levesque, C.A.; Chen, W. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for fungi. Proc. Natl. Acad. Sci. USA 2012, 109, 6241–6246. [Google Scholar] [CrossRef] [PubMed]
- Almeida, M.; Hébert, A.; Abraham, A.L.; Rasmussen, S.; Monnet, C.; Pons, N.; Delbès, S.; Loux, V.; Batta, J.M.; Leonard, P.; et al. Construction of a dairy microbial genome catalog opens new perspectives for the metagenomic analysis of dairy fermented products. BMC Genom. 2014, 15, 1257–1287. [Google Scholar] [CrossRef] [PubMed]
- Callon, C.; Retureau, E.; Didienne, R.; Montel, M.-C. Microbial biodiversity in cheese consortia and comparative Listeria growth on surfaces of uncookes pressed cheeses. Int. J. Food Microbiol. 2014, 174, 98–109. [Google Scholar] [CrossRef] [PubMed]
- Montel, M.-C.; Buchin, S.; Mallet, A.; Delbes-Paus, C.; Vuitton, D.A.; Desmasures, N.; Berthier, F. Traditional cheeses: Rich and diverse microbiota with associated benefits. Int. J. Food Microbiol. 2014, 177, 136–154. [Google Scholar] [CrossRef] [PubMed]
- Wolfe, B.E.; Button, J.E.; Santarelli, M.; Dutton, R.J. Cheese rind communities provide tractable systems for in situ and in vitro studies of microbial diversity. Cell 2014, 158, 422–433. [Google Scholar] [CrossRef] [PubMed]
- Quigley, L.; O’Sullivan, O.; Stanton, C.; Beresford, T.P.; Ross, R.P.; Fitzgerald, G.F.; Cotter, P.D. The complex microbiota of raw milk. FEMS Microbiol. Rev. 2013, 37, 664–698. [Google Scholar] [CrossRef] [PubMed]
- Dugat-Bony, E.; Garnier, L.; Denonfoux, J.; Ferreira, S.; Sarthou, A.S.; Bonnarme, P.; Irlinger, F. Highlighting the microbial diversity of 12 French cheese varieties. Int. J. Food Microbiol. 2016, 238, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Lavoie, K.; Touchette, M.; St-Gelais, D.; Labrie, S. Characterization of the fungal microflora in raw milk and specialty cheeses of the province of Quebec. Dairy Sci. Technol. 2012, 92, 455–468. [Google Scholar] [CrossRef] [PubMed]
- Marín, P.; Palmero, D.; Jurado, M. Occurrence of moulds associated with ovine raw milk and cheeses of the Spanish region of Castilla La Mancha. Int. J. Dairy Technol. 2015, 68, 565–572. [Google Scholar] [CrossRef]
- Vacheyrou, M.; Normand, A.C.; Guyot, P.; Cassagne, C.; Piarrous, R.; Bouton, Y. Cultivable microbial communities in raw cow milk and potential transfers from stables of sixteen French farms. Int. J. Food Microbiol. 2011, 146, 253–262. [Google Scholar] [CrossRef] [PubMed]
- Fleet, G.H. Yeasts in dairy products. J. Appl. Bacteriol. 1990, 68, 199–211. [Google Scholar] [CrossRef] [PubMed]
- Von Neubeck, M.; Baur, C.; Krewinkel, M.; Stoeckel, M.; Kranz, B.; Stressler, T.; Fisher, L.; Hinrichs, J.; Scherer, S.; Wenning, M. Biodiversity of refrigerated raw milk microbiota and their enzymatic spoilage potential. Int. J. Food Microbiol. 2015, 211, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Kure, C.F.; Skaar, I.; Brendehaug, J. Mould contamination in production of semi-hard cheese. Int. J. Food Microbiol. 2004, 93, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Beletsiotis, E.; Guikas, D.; Kalantzi, K. Incorporation of microbiological and molecular methods in HACCP monitoring scheme of molds and yeasts in a Greek dairy plant: A case study. Proc. Food Sci. 2011, 1, 1051–1059. [Google Scholar] [CrossRef]
- Temelli, S.; Anar, S.; Sen, C.; Akyuva, P. Determination of microbiological contamination sources during Turkish white cheese production. Food Control 2006, 17, 856–861. [Google Scholar] [CrossRef]
- Oyugi, E.; Buys, E.M. Microbiological quality of shredded Cheddar cheese packaged in modified atmospheres. Int. J. Dairy Technol. 2007, 60, 89–95. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Mills, D.A. Facility-specific “house” microbiome drives microbial landscape of artisan cheesemaking plants. Appl. Environ. Microbiol. 2013, 79, 5214–5223. [Google Scholar] [CrossRef] [PubMed]
- Radha, K.; Nath, L.S. Studies on the air quality in a dairy processing plant. Ind. J. Vet. Anim. Sci. Res. 2014, 43, 346–353. [Google Scholar] [CrossRef]
- Schön, K.; Schornsteiner, E.; Dzieciol, M.; Wagner, M.; Müller, M.; Schmitz-Esser, S. Microbial communities in dairy processing environment floor-drains are dominated by product-associated bacteria and yeasts. Food Control 2016, 70, 210–215. [Google Scholar] [CrossRef]
- Kaminarides, S.E.; Lakos, N.S. Yeasts in factory brine of feta cheese. Aust. J. Dairy Technol. 1992, 47, 68–71. [Google Scholar]
- Suriyarachchi, V.R.; Fleet, G.H. Occurrence and growth of yeasts in yoghurts. Appl. Environ. Microbiol. 1981, 42, 574–579. [Google Scholar] [CrossRef] [PubMed]
- Mayoral, M.B.; Martin, R.; Sanz, A.; Hernandez, P.E.; Gonzalez, I.; Garcia, T. Detection of Kluyveromyces marxianus and other spoilage yeasts in yoghurt using a PCR-culture technique. Int. J. Food. Microbiol. 2005, 105, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Penney, V.; Henderson, G.; Blum, C.; Johnson-Green, P. The potential phytopreservatives and nisin to control microbial spoilage of minimally processed fruit yoghurts. Innov. Food Sci. Emerg. Technol. 2004, 5, 369–375. [Google Scholar] [CrossRef]
- Tranquillini, R.; Scaramuzza, N.; Berni, E. Occurrence and ecological distribution of heat resistant moulds spores (HRMS) in raw materials used by food industry and thermal characterization of two Talaromyces isolates. Int. J. Food Microbiol. 2017, 242, 116–123. [Google Scholar] [CrossRef] [PubMed]
- Catellani, P.; Miotti Scapin, R.; Alberghini, L.; Radu, I.L.; Giaccone, V. Levels of microbial contamination of domestic refrigerators in Italy. Food Control 2014, 42, 257–262. [Google Scholar] [CrossRef]
- Carreira, A.; Ferreira, L.M.; Loureiro, V. Brown pigments produced by Yarrowia lipolytica result from extracellular accumulation of homogentisic acid. Appl. Environ. Microbiol. 2001, 67, 3463–3468. [Google Scholar] [CrossRef] [PubMed]
- Carreira, A.; Ferreira, L.M.; Loureiro, V. Production of brown tyrosine pigments by the yeast Yarrowia lipolytica. J. Appl. Microbiol. 2001, 90, 372–379. [Google Scholar] [CrossRef] [PubMed]
- Dalgaard, P.; Gram, L.; Huss, H.H. Spoilage and shelf life of cod fillets packed in vacuum or modified atmospheres. Int. J. Food Microbiol. 1993, 19, 283–294. [Google Scholar] [CrossRef]
- Gardini, F.; Tofalo, R.; Belleti, N.; Iucci, L.; Suzzi, G.; Torriani, S.; Guerzoni, M.E.; Lanciotti, R. Characterization of yeasts involved in the ripening of Pecorino Crotone cheese. Food Microbiol. 2006, 23, 641–648. [Google Scholar] [CrossRef] [PubMed]
- Roig-Sagués, A.X.; Molina, A.P.; Hernàndez-Herrero, M. Histamine and tyramine-forming microorganisms in Spanish traditional cheeses. Eur. Res. Technol. 2002, 215, 69–100. [Google Scholar] [CrossRef]
- Suzzi, G.; Schirone, M.; Martuscelli, M.; Gatti, M.; Fornasari, M.E.; Neviani, E. Yeasts associated with Manteca. FEMS Yeast Res. 2003, 3, 159–166. [Google Scholar] [CrossRef]
- Linares, D.M.; Martín, M.C.; Ladero, V.; Álvarez, M.A.; Fernández, M. Biogenic amines in dairy products. Crit. Rev. Food Sci. Nutr. 2011, 51, 691–703. [Google Scholar] [CrossRef] [PubMed]
- Leuschner, R.G.; Heidel, M.; Hammes, W.P. Histamine and tyramine degradation by food fermenting microorganisms. Int. J. Food Microbiol. 1998, 39, 1–10. [Google Scholar] [CrossRef]
- Garnier, L.; Valence, F.; Pawtowski, A.; Auhustsinava-Galerne, L.; Frotté, N.; Baroncelli, R.; Deniel, F.; Coton, E.; Mounier, J. Diversity of spoilage fungi associated with French dairy products. Int. J. Food Microbiol. 2017, 241, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Robinson, R.K. Dairy microbiology. In The Microbiology of Milk, 2nd ed.; Literary Cat Books: London, UK; New York, NY, USA, 1990; Volume 1, ISBN 978-0-471-38596-7. [Google Scholar]
- Minervini, F.; Montagna, M.T.; Spilotros, G.; Monaci, L.; Santacroce, M.P.; Visconti, A. Survey on mycoflora of cow and buffalo dairy products from Southern Italy. Int. J. Food Microbiol. 2001, 69, 141–146. [Google Scholar] [CrossRef]
- Jakobsen, M.; Narchus, J. Yeasts and their possible beneficial and negative effects on the quality of dairy products. Int. Dairy J. 1996, 6, 755–768. [Google Scholar] [CrossRef]
- Fleet, G.H.; Mian, M.A. The occurrence and growth of yeasts in dairy products. Int. J. Food Microbiol. 1987, 4, 145–155. [Google Scholar] [CrossRef]
- Hocking, A.D.; Faedo, M. Fungi causing thread mould spoilage of vacuum packaged Cheddar cheese during maturation. Int. J. Food Microbiol. 1992, 16, 123–130. [Google Scholar] [CrossRef]
- Fadda, M.E.; Cosentino, S.; Deplano, M.; Palmas, F. Yeasts populations in Sardinian feta cheese. Int. J. Food Microbiol. 2001, 69, 153–156. [Google Scholar] [CrossRef]
- Montagna, M.T.; Santacroce, M.P.; Spilotros, G.; Napoli, C.; Minervini, F.; Papa, A.; Dragoni, I. Investigation of fungal contamination in sheep and goat cheeses in southern Italy. Mycopathologia 2004, 158, 245–249. [Google Scholar] [CrossRef] [PubMed]
- Jodral, M.; Liñan, E.; Acosta, I.; Gallego, C.; Roja, F.; Bentabol, A. Mycoflora and toxigenic Aspergillus flavus in Spanish milks. Int. J. Food Microbiol. 1993, 36, 171–174. [Google Scholar] [CrossRef]
- Sensidoni, A.; Rondinini, G.; Peressini, D.; maifreni, M.; Bortolomeazzi, R. Presence of an off-flavor assiocated with the use of sorbates in cheese and margarine. Int. J. Food Sci. 1994, 6, 237–242. [Google Scholar]
- Lund, F.; Filtenborg, O.; Frisvad, J.C. Associated mycoflora of cheese. Food Microbiol. 1995, 12, 173–180. [Google Scholar] [CrossRef]
- Houbraken, J.; Samson, R.A. Phylogeny of Penicillium and the segregation of Trichocomoceae into three families. Stud. Mycol. 2011, 70, 1–51. [Google Scholar] [CrossRef] [PubMed]
- Frisvad, J.C.; Houbraken, J.; Popma, S.; Samson, R.A. Two new Penicillium species Penicillium buchwaldii and Penicillium spathulatum, producing the anticancer compound asperphenamate. FEMS Microbiol. Lett. 2013, 339, 77–92. [Google Scholar] [CrossRef] [PubMed]
- Samson, R.; Frisvad, J.; Hoekstra, E.; Filtenborg, O. Introduction to food and airborne fungi. In Mycopathologia, 6th ed.; Centraal Bureau Voor Schimmelcultures: Utrecht, The Netherlands, 2002; p. 609. ISBN 10 9070351528. [Google Scholar]
- Moubasher, A.H.; Abdel-Kader, M.I.A.; El-Kady, I.A. Toxigenic fungi isolated from Roquefort cheese. Mycopathologia 1978, 66, 187–190. [Google Scholar] [CrossRef]
- Kikoku, Y.; Tagashira, N.; Nakano, H. Heat resistance of fungi isolated from frozen blueberries. J. Food Prot. 2008, 71, 2030–2035. [Google Scholar] [CrossRef] [PubMed]
- Kure, C.F.; Wasteson, Y.; Brendehaug, J.; Skaar, I. Mould contaminants on Jarlsberg and Norvegia cheese blocks from four factories. Int. J. Food Microbiol. 2001, 70, 21–27. [Google Scholar] [CrossRef]
- Bullerman, L.B.; Olivigni, L.B. Mycotoxin producing-potential of molds isolated from cheddar cheese. J. Food Sci. 1974, 39, 1166–1168. [Google Scholar] [CrossRef]
- Hassanin, I.N. Detection of mycotoxigenic fungi and bacteria in processed cheese in Egypt. Int. Biodeterior. Biodegrad. 1993, 31, 15–23. [Google Scholar] [CrossRef]
- Aran, N.; Eke, D. Mould mycoflora of Kaşar cheese at the stage of consumption. Food Microbiol. 1987, 4, 101–104. [Google Scholar] [CrossRef]
- Kure, C.F.; Skaar, I. Mould growth on Norwegian semi-hard cheeses Norvegia and Jarlsberg. Int. J. Food Microbiol. 2000, 62, 133–137. [Google Scholar] [CrossRef]
- Basílico, J.C.; deBasílico, M.Z.; Chiericatti, C.; Vinderola, C.G. Characterizaion and control of thread mould in cheese. Lett. Appl. Microbiol. 2001, 32, 419–423. [Google Scholar] [CrossRef] [PubMed]
- Gould, G.W. Mechanisms of Action of Food Preservation Procedures; Gould, G.W., Ed.; Elsevier Applied Science: London, UK, 1989; ISBN 1851662936, 9781851662937. [Google Scholar]
- Rahman, M.S. Handbook of Food Preservation, 2nd ed.; CRC Press: Boca Raton, FL, USA, 2007; ISBN 9781574446067. [Google Scholar]
- Sofos, J.N. Current microbiological considerations in food preservation. Int. J. Food Microbiol. 1993, 19, 87–108. [Google Scholar] [CrossRef]
- Dalié, D.K.D.; Deschamps, A.M.; Richard-Forget, F. Lactic acid bacteria—Potential for control of mould growth and mycotoxins: A review. Food Control 2010, 21, 370–380. [Google Scholar] [CrossRef]
- Regulation (EC) No. 852/2004 of the European Parliament and of the Council of 29 April 2004 on the Hygiene and Foodstuffs. Available online: http://eur-lex.europa.eu/LexUriServ/LexUriServ.do?uri=OJ:L:2004:139:0001:0054:en:PDF (accessed on 27 July 2017).
- Rysstad, G.; Kolstad, J. Extended shelf life milk—Advances in technology. Int. J. Dairy Technol. 2006, 85–96. [Google Scholar] [CrossRef]
- Loureiro, V.; Malfeito-Ferreira, M. Spoilage yeasts in foods and beverages: Characterization and ecology for improved diagnosis and control. Food Res. Int. 2000, 33, 247–256. [Google Scholar] [CrossRef]
- Loureiro, V.; Querol, A. The prevalence and control of spoilage yeasts in foods and beverages. Trends Food Sci. Technol. 1999, 10, 356–365. [Google Scholar] [CrossRef]
- Odet, G. Qualité bactériologique des fromages au lait cru. Cahiers de Nutrition et de Diététique 1999, 34, 47–53. [Google Scholar]
- Faye, B.; Loiseau, G. Sources of Contamination in Dairy Supply Chains and Approaches to Quality Control; Paper Read at International Cooperation in International Research for Agricultural Development (CIRAD)-FAO: Montpellier, France, 2002; ISBN 92-5-004787-8. [Google Scholar]
- Sandrou, D.K.; Arvanitoyannis, I.S. Implementation of Hazard Analysis Critical Control Point (HACCP) to the dairy industry: Current status and perspectives. Food Rev. Int. 2000, 16, 77–111. [Google Scholar] [CrossRef]
- De Steur, H.; Wesana, J.; Dora, M.K.; Pearce, D.; Gellynck, X. Applying value stream mapping to reduce food losses and wastes in supply chains: A systematic review. Waste Manag. 2016, 58, 359–368. [Google Scholar] [CrossRef] [PubMed]
- Pinto, A.T.; Schmidt, V.; Raimundo, S.A.; Raihmer, F. Use of ozone to control fungi in a cheese ripening room. Acta Sci. Vet. 2007, 35, 333–337. [Google Scholar] [CrossRef]
- Serra, R.; Abrunhosa, L.; Kozakiewicz, Z.; Venâncio, A.; Lima, N. Use of ozone to reduce molds in a cheese ripening room. J. Food Prot. 2003, 66, 2355–2358. [Google Scholar] [CrossRef] [PubMed]
- Varga, L.; Szigeti, J. Use of ozone in the dairy industry: A review. Int. J. Dairy Technol. 2016, 69, 157–168. [Google Scholar] [CrossRef]
- Korukluoglu, M.; Sahan, Y.; Yigit, A. The fungicidal efficacy of various commercial disinfectants used in the food industry. Ann. Microbiol. 2006, 56, 325–330. [Google Scholar] [CrossRef]
- Bundgaard-Nielsen, K.; Nielsen, P.V. Fungicidal effect of 15 disinfectants against 25 fungal contaminants commonly found in bread and cheese manufacturing. J. Food Prot. 1995, 59, 268–275. [Google Scholar] [CrossRef]
- Martin, H.; Maris, P. Synergism between hydrogen peroxide and seventeen acids against five agri-food-borne fungi and one yeast strain. J. Appl. Microbiol. 2012, 113, 1451–1460. [Google Scholar] [CrossRef] [PubMed]
- Lado, B.H.; Yousef, A.E. Alternative food-preservation technologies: Efficacy and mechanisms. Microbes Infect. 2002, 4, 433–440. [Google Scholar] [CrossRef]
- European Union Council Directive 92/46/EEC of 16 June 1992 Laying Down the Health Rules for the Production and Placing on the Market or Raw Milk, Heat-Treated Mil and Milk-Based Products. Available online: https://www.fsvps.ru/fsvps-docs/ru/usefulinf/files/es9246.pdf (accessed on 27 July 2017).
- Sakkas, L.; Moutafi, A.; Moschopoulou, E.; Moatsou, G. Assessment of treatment of various types of milk. Food Chem. 2014, 15, 293–301. [Google Scholar] [CrossRef] [PubMed]
- Houbraken, J.; Samson, R.A.; Frisvad, J.C. Byssochlamys: Significance of heat resistance and mycotoxin production. In Advances in Food Mycology; Hocking, A.D., Pitt, J.I., Samson, R.A., Thrane, U., Eds.; Springer: New York, NY, USA, 2005; pp. 211–224. [Google Scholar]
- Scaramuzza, N.; Berni, E. Heat-resistance of Hamigera avellanea and Thermoascus crustaceus isolated from pasteurized acid products. Int. J. Food Microbiol. 2014, 169, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Salomão, B.C.M.; Slongo, A.P.; Aragão, G.M.F. Heat resistance of Neosartorya fischeri in various juices. LWT Food Sci. Technol. 2007, 40, 676–680. [Google Scholar] [CrossRef]
- Trujillo, A.J.; Capellas, M.; Saldo, J.; Gervilla, R.; Guamis, B. Applications of high-hydrostatic pressure on milk and dairy products: A review. Innov. Food Sci. Emerg. Technol. 2002, 3, 295–307. [Google Scholar] [CrossRef]
- Mainville, I.; Montpetit, D.; Durand, N.; Farnworth, E.R. Deactivating the bacteria and yeast in kefir using heat treatment, irradiation and high pressure. Int. Dairy J. 2001, 11, 45–49. [Google Scholar] [CrossRef]
- Reps, A.; Krzyzewska, A.; Lanievska-Moroz, L.; Iwanczak, M. Effect of high pressure on microflora of kefir. In Proceedings of the First International Conference on High Pressure Bioscience and Biotechnology HPBB-2000, Kyoto, Japan, 26–30 November 2000; Volume 19, pp. 1–652. [Google Scholar] [CrossRef]
- Evert-Arriagada, K.; Hernández-Herrero, M.M.; Guamis, B.; Trujillo, A.J. Commercial application of high-pressure processing for increasing starter-free fresh cheese shelf-life. Food Sci. Technol. 2014, 55, 498–505. [Google Scholar] [CrossRef]
- Ravishankar, S.; Zhang, H.; Kempkes, M.L. Pulsed electric fields. Food Sci. Technol. Int. 2008, 14, 429–432. [Google Scholar] [CrossRef]
- Buckow, R.; Scott Chandry, P.; Ng, S.Y.; McAuley, C.M.; Swanson, B.G. Opportunities and challenges in pulse electric field processing of dairy products. Int. Dairy J. 2014, 34, 199–212. [Google Scholar] [CrossRef]
- Wan, J.; Coventry, J.; Swiergon, P.; Sanguansri, P.; Viersteeg, C. Advances in innovative processing technologies for microbial inactivation and enhancement of food safety—Pulsed electric field and low temperature plasma. Trends Food Sci. Technol. 2009, 20, 414–424. [Google Scholar] [CrossRef]
- Gougouli, M.; Kalantzi, K.; Beletsiotis, E.; Koutsoumanis, K.P. Development and application of predictive models for fungal growth as tools to improve quality control in yoghurt production. Food Microbiol. 2011, 28, 1453–1462. [Google Scholar] [CrossRef] [PubMed]
- Sandhya. Modified atmosphere packaging of fresh produce: Currentstatus and future needs. Food Sci. Technol. 2010, 43, 381–392. [Google Scholar] [CrossRef]
- Ooraikul, B.; Stiles, M.E. Modified Atmosphere Packaging of Food; Ellis Horwood Publishing: New York, NY, USA, 1991; ISBN 978-0-7476-0064-0. [Google Scholar]
- Nguyen Van Long, N.; Joly, C.; Dantigny, P. Active packaging with antifungal activities. Int. J. Food Microbiol. 2016, 220, 73–90. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Aguilera, R.; Oliveira, J.C.; Montanez, J.C.; Mahajan, P.V. Effect of modified atmosphere packaging on quality factors and shelf-life of surface mould ripened cheese: Part II varying storage temperature. LWT Food Sci. Technol. 2011, 44, 330–336. [Google Scholar] [CrossRef]
- Sing, P.; Wani, A.A.; Karim, A.A.; Langowski, H.-C. The use of carbon dioxide in the processing and packaging of milk and dairy products: A review. Int. J. Dairy Technol. 2012, 65, 161–177. [Google Scholar] [CrossRef]
- Hotchkiss, J.H.; Werner, B.G.; Lee, E.Y.C. Addition of carbon dioxide to dairy products to improve quality: A comprehensive review. Compr. Rev. Food Sci. Food Saf. 2006, 5, 158–168. [Google Scholar] [CrossRef]
- Gonzalez-Fandos, E.; Sanz, S.; Olarte, C. Microbiological, physicochemical and sensory characteristics of Cameros cheese packaged under modified atmospheres. Food Microbiol. 2000, 17, 407–414. [Google Scholar] [CrossRef]
- Taniwaki, M.H.; Hocking, A.D.; Pitt, J.I.; Fleet, G.H. Growth and mycotoxin production by fungi in atmosphere containing 80% carbon dioxide and 20% oxygen. Int. J. Food Microbiol. 2010, 143, 218–225. [Google Scholar] [CrossRef] [PubMed]
- Khoshgozaran, S.; Azizi, M.H.; Bagheripoor-Fallah, N. Evaluating the effect of modified atmosphere packaging on cheese characteristics: A review. Dairy Sci. Technol. 2012, 92, 1–24. [Google Scholar] [CrossRef]
- Taniwaki, M.H.; Hocking, A.D.; Pitt, J.I.; Fleet, G.H. Growth and mycotoxin production by food spoilage under high carbon dioxide and low oxygen atmospheres. Int. J. Food Microbiol. 2009, 132, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Dermiki, M.; Ntzimani, A.; Badeka, A.; Savvaidis, I.N.; Kontominas, M.G. Shelf-life extension and quality attributes of the whey cheese “Myzithra Kalathaki” using modified atmosphere packaging. LWT Food Sci. Technol. 2008, 41, 284–294. [Google Scholar] [CrossRef]
- Eliot, S.C.; Vuillemard, J.-C.; Edmond, J.-P. Stability of shredded Mozzarella cheese under modified atmospheres. J. Food Sci. 1998, 63, 1075–1080. [Google Scholar] [CrossRef]
- Fedio, W.M.; Macleod, A.; Ozimek, L. The effect of modified atmosphere packaging on the growth of microorganisms in cottage cheese. Milchwissenschaft 1994, 49, 622–629. [Google Scholar]
- Genigeorgis, C. Microbial and safety implications of the use of modified atmospheres to extend the storage life of fresh meat and fish. Int. J. Food Microbiol. 1985, 1, 237–251. [Google Scholar] [CrossRef]
- Phillips, C.A. Review: Modified atmosphere packaging and its effects on the microbiological quality and safety of produce. Int. J. Food Sci. Technol. 1996, 31, 463–479. [Google Scholar] [CrossRef]
- Rodriguez-Aguilera, R.; Oliveira, J.C.; Montanez, J.C.; Mahajan, P.V. Effect of modified atmosphere packaging on quality factors and shelf-life of surface mould ripened cheese: Part I constant temperature. LWT Food Sci. Technol. 2011, 44, 337–342. [Google Scholar] [CrossRef]
- Blocher, J.C.; Busta, F.F. Multiple modes of inhibition of spore germination and outgrowth by reduced pH and sorbate. J. Appl. Bacteriol. 1985, 59, 469–478. [Google Scholar] [CrossRef] [PubMed]
- Guynot, M.E.; Ramos, A.J.; Sanchis, V.; Marín, S. Study of benzoate, propionate, and sorbate salts as mould spoilage inhibitors on intermediate moisture bakery products of low pH (4.5–5.5). Int. J. Food Microbiol. 2005, 101, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Wilson, M.; Chapman, B.; Hocking, A.D. Evaluation of the efficacy of four weak acids as antifungal preservatives in low-acid intermediate moisture model food systems. Food Microbiol. 2009, 27, 6–33. [Google Scholar] [CrossRef] [PubMed]
- Stratford, M.; Nebe-von-Caron, G.; Steels, H.; Novodvorska, M.; Ueckert, J.; Archer, D.B. Weak-acid preservatives: pH and proton movements in the yeast Saccharomyces cerevisiae. Int. J. Food Microbiol. 2013, 161, 164–171. [Google Scholar] [CrossRef] [PubMed]
- Fajardo, P.; Martins, J.T.; Fuciños, C.; Pastrana, L.; Teixera, J.A.; Vicente, A.A. Evaluation of a chitosan-based edible film as carrier of natamycin to improve the storability of Saloio cheese. J. Food Eng. 2010, 101, 349–356. [Google Scholar] [CrossRef]
- EFSA. Scientific Opinion on the use of Natamycin (E 235) as a food additive. EFSA Panel on Food Additives and Nutrient Sources added to Food (ANS). EFSA J. 2009, 7, 1412. [Google Scholar] [CrossRef]
- FDA (2015). ICFSAN/Office of Food Additive Safety. GRAS Notice Natamycin. Published: 6 November 2015. Response Letter GRAS Notice No. GRN 000578. Available online: https://www.fda.gov/Food/IngredientsPackagingLabeling/GRAS/NoticeInventory/ucm484505.htm (accessed on 27 July 2017).
- Brandão, L.R.; Rosa, J.C.C.; Nicoli, J.R.; Almeida, M.V.S.; do Carmo, A.P.; Queiros, H.T.; Castro, I.M. Investigating acid stress response in different Saccharomyces strains. J. Mycol. 2014, 12, 1–9. [Google Scholar] [CrossRef]
- Ullah, A.; Chandrasekaran, G.; Brul, S.; Smits, J.G. Yeast adaptation to weak acids prevents futile energy expenditure. Front. Microbiol. 2013, 4, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Casas, E.; Ancos, B.; Valderrama, M.J.; Cano, P.; Peinado, J.M. Pentadiene production from potassium sorbate by osmotolerant yeasts. Int. J. Food Microbiol. 2004, 94, 93–96. [Google Scholar] [CrossRef] [PubMed]
- Plumridge, A.; Stratford, M.; Lowe, K.C.; Archer, D.B. The weak-acid preservative sorbic acid is decarboxylated and detoxified by a phynylacrylic acid decarboxylase, PadA1, in the spoilage mold Aspergillus niger. Appl. Environ. Microbiol. 2008, 74, 550–552. [Google Scholar] [CrossRef] [PubMed]
- Kallinteri, L.D.; Kostoula, O.K.; Savvaidis, I.N. Efficacy of nisin and/or natamycin to improve the shelf-life of Galotyri cheese. Food Microbiol. 2013, 36, 176–181. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, T.M.; De Fátima Ferreira Soares, N.; Pereira, R.M.; De Freitas Fraga, K. Development and evalutation of antimicrobial natamycin-incorporated film in Gorgonzola cheese conservation. Packag. Technol. Sci. 2007, 20, 147–153. [Google Scholar] [CrossRef]
- Ollé Resa, C.P.; Gerschenson, L.N.; Jagus, R.J. Natamycin and nisin supported on starch edible films for controlling mixed growth on model systems and Port Salut cheese. Food Control 2014, 44, 146–151. [Google Scholar] [CrossRef]
- Ollé Resa, C.P.; Jagus, R.J.; Gerschenson, L.N. Natamycin effiency for controlling yeast growth in models systems and on cheese surfaces. Food Control 2014, 35, 101–108. [Google Scholar] [CrossRef]
- WHO. Safety Evaluation of Certain Food Additives and Contaminants; Natamycin (Pimaricin); WHO Food Additives Series 48; WHO: Geneva, Switzerland, 2002; Available online: http://apps.who.int/iris/bitstream/10665/42578/1/WHO_TRS_909.pdf (accessed on 27 July 2017).
- Van Leeuwen, M.R.; De Boer, W.; Smant, W.; Dijksterhuis, J. Filipin is a reliable in situ marker of ergosterol in the plasma membrane of germinating conidia (spores) of Penicillium discolor and stains intensively at the site of germ tube formation. J. Microbiol. Methods 2008, 74, 64–73. [Google Scholar] [CrossRef] [PubMed]
- Van Leeuwen, M.R.; Van Doorn, T.; Golovina, E.A.; Stark, J.; Dijksterhuis, J. Waterand air-distributed conidia exhibit differences in sterol content and cytoplasmic microviscosity. Appl. Environ. Microbiol. 2010, 67, 366–369. [Google Scholar] [CrossRef] [PubMed]
- Stern, G.A. In vitro antibiotic synergism against ocular fungal isolates. Am. J. Ophthalmol. 1978, 86, 359–367. [Google Scholar] [CrossRef]
- Streekstra, H.; Verkennis, A.E.E.; Jacobs, R.; Dekker, A.; Stark, J.; Dijksterhuis, J. Fungal strain and the development of tolerance against natamycin. Int. J. Food Microbiol. 2016, 238, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Joint FAO/WHO Expert Committee on Food Additives. Evaluation of Certain Food Additives and Contaminants: 67th Report of the Joint FAO/WHO Expert Committee on Food Additives; WHO Technical Report Series; WHO: Geneva, Switzerland, 2007; p. 25. Available online: http://www.who.int/ipcs/publications/jecfa/reports/trs940.pdf (accessed on 27 July 2017).
- Bourdichon, F.; Casaregola, S.; Farrokh, C.; Frisvad, J.C.; Gerds, M.L.; Hammes, W.P.; Harnett, J.; Huys, G.; Laulund, S.; Ouwehand, A.; et al. Food fermentations: Microorganisms with technological beneficial use. Int. J. Food Microbiol. 2012, 154, 87–97. [Google Scholar] [CrossRef] [PubMed]
- Caplice, E.; Fitzgerald, G.F. Food fermentations: Role of microorganisms in food production and preservation. Int. J. Food Microbiol. 1999, 50, 131–149. [Google Scholar] [CrossRef]
- Brimelow, C.J.B.; Collins-Thompson, D. Food preservatives. Food Control 1992, 3, 218–219. [Google Scholar] [CrossRef]
- Elsser-Gravesen, D.; Elsser-Gravesen, A. Biopreservatives. In Biotechnology of Food and feed Additives; Zorn, H., Czermak, P., Eds.; Springer: Berlin, Germany, 2014; ISBN 978-3-662-43761-2. [Google Scholar]
- Stiles, M.E. Biopreservation by lactic acid bacteria. Antonie van Leeuwenhoek 1996, 70, 331–345. [Google Scholar] [CrossRef] [PubMed]
- Holzapfel, W.H.; Geisen, R.; Schillinger, U. Biological preservation of foods with reference to protective cultures, bacteriocins and food-grade enzymes. Int. J. Food Microbiol. 1995, 24, 343–362. [Google Scholar] [CrossRef]
- Magnusson, J.; Schnurer, J. Lactobacillus coryniformis subs. coryniformis strain Si3 produces a broad-spectrum proteinaceous antfungal compound. Appl. Environ. Microbiol. 2001, 67, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Lassois, L.; de Lapeyre de Bellaire, L.; Jijakli, M.H. Biological control of crown rot of bananas with Pichia anomala strain K and Candida oleophila strain O. Biol. Control 2008, 45, 410–418. [Google Scholar] [CrossRef]
- Zhang, H.; Wang, L.; Ma, L.; Dong, Y.; Jiang, S.; Xu, B.; Zheng, X. Biocontrol of major postharvest pathogens on apple using Rhodotorula glutinis and its effects on postharvest quality parameters. Biol. Control 2009, 48, 79–83. [Google Scholar] [CrossRef]
- Carr, F.J.; Chill, D.; Maida, N. The lactic acid bacteria: A literature survey. Crit. Rev. Microbiol. 2002, 28, 281–370. [Google Scholar] [CrossRef] [PubMed]
- Schwenninger, S.M.; Meile, L. A mixed culture of Propionibacterium jensenii and Lactobacillus paracasei subs. paracasei inhibits food spoilage yeasts. Syst. Appl. Microbiol. 2004, 27, 229–237. [Google Scholar] [CrossRef] [PubMed]
- Aljewicz, M.; Cichosz, G. Protective effects of Lactobacillus cultures in Dutch-type cheese like products. LWT Food Sci. Technol. 2015, 63, 52–56. [Google Scholar] [CrossRef]
- Aunsbjerg, S.D.; Honoré, A.H.; Marcussen, J.; Ebrahimi, P.; Vogensen, F.K.; Benfeldt, C.; Skov, T.; KnØchel, S. Contribution of volatiles to the antifungal effect of Lactobacillus paracasei in defined medium and yoghurt. Int. J. Food Microbiol. 2015, 194, 46–53. [Google Scholar] [CrossRef] [PubMed]
- Crowley, S.; Mahony, J.; van Sinderen, D. Curent perspectives on antifungal lactic acid bacteria as natural bio-preservatives. Trends Food Sci. Technol. 2013, 33, 93–109. [Google Scholar] [CrossRef]
- Inglin, R.C.; Stevens, M.J.A.; Meile, L.; Lacroix, C.; Meile, L. High-throughput screening assays for antibacterial and antifungal activities of Lactobacillus species. J. Microbiol. Methods 2015, 114, 26–29. [Google Scholar] [CrossRef] [PubMed]
- Stiles, J.; Penkar, S.; Plocková, M.; Chumchalová, J.; Bullerman, L.B. Antifungal activity of sodium acetate and Lactobacillus rhamnosus. J. Food Prot. 2002, 65, 1188–1191. [Google Scholar] [CrossRef] [PubMed]
- Le Lay, C.; Mounier, J.; Vasseur, V.; Weill, A.; Le Blay, G.; Barbier, G.; Coton, E. In vitro and in situ screening of lactic acid bacteria and propionibacteria antifungal activities against bakery product spoilage molds. Food Control 2016, 60, 247–255. [Google Scholar] [CrossRef]
- Delavenne, E.; Mounier, J.; Deniel, F.; Barbier, G.; Le Blay, G. Biodiversity of antifungal lactic acid bacteria isolated from raw milk samples from cow, ewe and goat over one-year period. Int. J. Food. Microbiol. 2012, 155, 185–190. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, B.; Vimon, A.; Desfossés-Foucault, E.; Daga, M.; Arora, G.; Fliss, I. Antifungal activity of lactic and propionic acid bacteria and their potential as protective culture in cottage cheese. Food Control 2017, 78, 350–356. [Google Scholar] [CrossRef]
- Axelsson, L.; Chung, T.C.; Debrogosz, W.J.; Lindgren, S. Production of a broad spectrum antimicrobial substance by Lactobacillus reuteri. Microb. Ecol. Health Dis. 1989, 2, 131–136. [Google Scholar] [CrossRef]
- Dagnas, S.; Gauvry, E.; Onno, B.; Membré, J.M. Quantifying effect of lactic, acetic, and propionic acids on growth of molds isolated from spoiled bakery products. J. Food Prot. 2015, 78, 1689–1698. [Google Scholar] [CrossRef] [PubMed]
- Coallier-Ascah, J.; Idziak, E.S. Interaction between Streptococcus lactis and Aspergillus flavus on production of aflatoxin. Appl. Environ. Microbiol. 1985, 49, 163–167. [Google Scholar] [CrossRef] [PubMed]
- Luchese, R.H.; Harrigan, W.F. Growth of, and aflatoxin production by Aspergillus paraciticus when in the presence of either Lactococcus lactis or lactic acid and at different initial pH values. J. Bacteriol. 1990, 69, 512–519. [Google Scholar] [CrossRef]
- Batish, V.K.; Roy, U.; Lal, R.; Grower, S. Antifungal attributes of lactic acid bacteria—A review. Crit. Rev. Biotechnol. 1997, 17, 209–225. [Google Scholar] [CrossRef] [PubMed]
- Roy, U.; Batish, V.K.; Grover, S.; Neelakantan, S. Production of antifungal substance by Lactococcus lactis subsp. lactis CHD-28.3. Int. J. Food Microbiol. 1996, 32, 27–34. [Google Scholar] [CrossRef]
- Florianowicz, T. Antifungal activity of some microorganisms against Penicillium expansum. Eur. Food Res. Technol. 2001, 212, 282–286. [Google Scholar] [CrossRef]
- Voulgari, K.; Hatzikamari, M.; Delepoglou, A.; Georgakipoulos, P.; Litopoulou-Tzanetaki, E.; Tzanetakis, N. Antifungal activity of non-starter lactic acid bacteria isolates from dairy products. Food Control 2010, 21, 136–142. [Google Scholar] [CrossRef]
- Garofalo, C.; Zannini, E.; Aquilanti, L.; Silvestri, G.; Fierro, O.; Picariello, G.; Clementi, F. Selection of sourdough lactobacilli with antifungal activity for use as biopreservatives in bakery products. J. Agric. Food Chem. 2012, 60, 7719–7728. [Google Scholar] [CrossRef] [PubMed]
- Coda, R.; Rizello, C.G.; Nigri, F.; De Angelis, M.; Arnault, P.; Gobbetti, M. Long-term fungi inhibitory activity of water-soluble extract from Phaseolus vulgaris cv. pinto and sourdough lactic acid bacteria during bread storage. Appl. Environ. Microbiol. 2008, 74, 7391–7398. [Google Scholar] [CrossRef] [PubMed]
- Gerez, C.L.; Torres, M.J.; Font de Valdez, G.; Rollán, G. Control of spoilage fungi by lactic acid bacteria. Biol. Control 2013, 64, 231–237. [Google Scholar] [CrossRef]
- Mauch, A.; Dal Bello, F.; Coffey, A.; Arendt, E.K. The use of Lactobacillus brevis PS1 to in vitro inhibit the outgrowth of Fusarium culmorum and other common Fusarium species found on barley. Int. J. Food Microbiol. 2010, 141, 116–121. [Google Scholar] [CrossRef] [PubMed]
- Falguni, P.; Shilpa, V.; Mann, B. Production of proteinaceous antifungal substances from Lactobacillus brevis NCDC. Int. J. Dairy Technol. 2010, 63, 70–76. [Google Scholar] [CrossRef]
- Gourama, H. Inhibition of growth and mycotoxin production of Penicillium by Lactobacillus species. LWT Food Sci. Technol. 1997, 30, 279–283. [Google Scholar] [CrossRef]
- Li, H.; Liu, L.; Zhang, S.; Uluko, H.; Cui, W.; Lv, J. Potential use of Lactobacillus casei AST18 as bioprotective culture in yoghurt. Food Control 2013, 34, 675–680. [Google Scholar] [CrossRef]
- Atanassova, M.R.; Fernández-Otero, C.; Rodríguez-Alonso, P.; Fernández-No, I.C.; Garabal, J.I.; Centeno, J.A. Characterization of yeasts isolated from artisanal short-ripened cows’ cheeses produced in Galicia (NW Spain). Food Microbiol. 2016, 53, 172–181. [Google Scholar] [CrossRef] [PubMed]
- Ryan, L.A.; Zannini, E.; Dal Bello, F.; Pawlowska, A.; Koehler, P.; Arendt, E.K. Lactobacillus amylovorus DSM 19280 as a novel food-grade antifungal agent for bakery products. Int. J. Food Microbiol. 2011, 146, 276–283. [Google Scholar] [CrossRef]
- Lynch, K.M.; Pawlowska, A.M.; Brosnan, B.; Coffey, A.; Zannini, E.; Furey, A.; McSweeney, P.L.H.; Waters, D.M.; Arendt, E.K. Application of Lactobacillus amylovorus as an antifungal adjunct to extend the shelf-life of Cheddar cheese. Int. J. Food Microbiol. 2014, 34, 167–173. [Google Scholar] [CrossRef]
- Corsetti, A.; Gobetti, M.; Rossi, J.; Damiani, P. Antimould activity of sourdough lactic acid bacteria: Identification of a mixture of organic acids produced by Lactobacillus sanfrancisco CB1. Appl. Microbiol. Biotechnol. 1998, 50, 253–256. [Google Scholar] [CrossRef] [PubMed]
- Corsetti, A.; Settani, L.; van Sinderen, D.; Felis, G.; Dellaglio, F.; Gobetti, M. Lactobacillus rossii sp. nov., isolated from wheat sourdough. Int. J. Syst. Evol. Microbiol. 2004, 55, 35–40. [Google Scholar] [CrossRef]
- Muhialdin, B.J.; Hassan, Z.; Sadon, S.K.; Zulkifli, N.A.; Azfar, A.A. Effect of pH and heat treatment on antifungal activity of Lactobacillus fermentum TE007, Lactobacillus pentosus G004 and Pediococcus pentosaceus TE010. Innov. Romanian Food Biotechnol. 2011, 8, 41–53. [Google Scholar] [CrossRef]
- Coloretti, F.; Carri, S.; Armaforte, E.; Chiavari, C.; Grazia, L.; Zambonelli, C. Antifungal activity of lactobacilli isolated from salami. FEMS Microbiol. Lett. 2007, 271, 245–250. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Mauch, A.; Galle, S.; Murphy, P.; Arendt, E.K.; Coffey, A. Inhibition of growth of Trichophyton tonsurans by Lactobacillus reuteri. J. Appl. Microbiol. 2011, 111, 474–483. [Google Scholar] [CrossRef] [PubMed]
- Niku-Paavola, M.L.; Laitila, A.; Mattila-Sandholm, T.; Haikara, A. New types of antimicrobial compounds produced by Lactobacillus plantarum. J. Appl. Microbiol. 1999, 86, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Dal Bello, F.; Clarke, C.; Ryan, L.; Ulmer, H.; Schober, T.; Strom, K.; Sjögren, J.; van Sinderen, D.; Schnürer, J.; Arendt, E.K. Improvement of the quality and shelf life of wheat bread by fermentation with the antifungal strain Lactobacillus plantarum FST 1.7. J. Cereal Sci. 2007, 45, 309–318. [Google Scholar] [CrossRef]
- Lavermicocca, P.; Valerio, F.; Visconti, A. Antifungal activity of phenyllactic acid against molds isolated from Bakery products. Appl. Environ. Microbiol. 2003, 69, 634–640. [Google Scholar] [CrossRef] [PubMed]
- Ström, K.; Sjogren, J.; Broberg, A.; Schnürer, J. Lactobacillus plantarum MiLAB 393 produces the antifungal cyclic dipeptides cyclo (l-Phe-l-Pro) and cyclo (l-Phe-trans-4-OH-l-Pro) and 3-phenyllactic acid. Appl. Environ. Microbiol. 2002, 68, 4322–4327. [Google Scholar] [CrossRef] [PubMed]
- Sjogren, J.; Magnusson, J.; Broberg, A.; Schnu rer, J.; Kenne, L. Antifungal 3- hydroxy fatty acids from Lactobacillus plantarum MiLAB 14. Appl. Environ. Microbiol. 2003, 69, 7554–7557. [Google Scholar] [CrossRef] [PubMed]
- Sathe, S.J.; Nawani, N.N.; Dhakephalkar, P.K.; Kapadnis, B.P. Antifungal lactic acid bacteria with potential to prolong shelf-life of fresh vegetables. J. Appl. Microbiol. 2007, 103, 2622–2628. [Google Scholar] [CrossRef] [PubMed]
- Yang, E.J.; Chang, H.C. Purification of a new antifungal compound produced by Lactobacillus plantarum AF1 isolated from kimchi. Int. J. Food Microbiol. 2010, 139, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Prema, P.; Smila, D.; Palavesan, A.; Immanuel, G. Production and characterization of an antifungal compound (3-phenyllactic acid) produced by Lactobacillus plantarum strain. Food Bioprocess Technol. 2008, 3, 379–386. [Google Scholar] [CrossRef]
- Rouse, S.; Harnett, D.; Vaughan, A.; Sinderen, D.V. Lactic acid bacteria with potential to eliminate fungal spoilage in foods. J. Appl. Microbiol. 2008, 104, 915–923. [Google Scholar] [CrossRef] [PubMed]
- Ndagano, D.; Lamoureux, T.; Dortu, C.; Vandermoten, S.; Thonart, P. Antifungal activity of 2 lactic acid bacteria of the Weissella Genus Isolated from Food. J. Food Sci. 2011, 76, 305–311. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Sun, Y.; Chen, C.; Sun, Z.; Zhou, Y.; Shen, F. Genome shuffling of Lactobacillus plantarum for improving antifungal activity. Food Control 2013, 32, 341–347. [Google Scholar] [CrossRef]
- Laitila, A.; Alakomi, H.L.; Raaska, L.; Mattila-Sandholm, T.; Haikara, A. Antifungal activities of two Lactobacillus plantarum strains against Fusarium moulds in vitro and in malting of barley. J. App. Microbiol. 2002, 93, 566–576. [Google Scholar] [CrossRef]
- Digaitiene, A.; Hansen, A.; Juodeikiene, G.; Eidukonyte, D.; Josephsen, J. Lactic acid bacteria isolated from rye sourdoughs produce bacteriocin-like inhibitory substances active against Bacillus subtilis and fungi. J. Appl. Microbiol. 2012, 112, 732–742. [Google Scholar] [CrossRef] [PubMed]
- Lind, H.; Jonsson, H.; Schnürer, J. Antifungal effect of dairy propionibacteria–contribution of organic acids. Int. J. Food Microbiol. 2003, 98, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Mieszkin, S.; Hymery, N.; Debaets, S.; Coton, E.; Le Blay, G.; Valence, F.; Mounier, J. Action mechanism involved in the bioprotective effect of Lactobacillus harbinensis K.V9.3.1Np against Yarrowia lipolytica in fermented milk. Int. J. Food Microbiol. 2017, 248, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Al-Zoreky, N.; Ayres, J.W.; Sandine, W.E. Antimicrobial activity of Microgard against food spoilage and pathogenic microorganisms. J. Dairy Sci. 1991, 74, 758–763. [Google Scholar] [CrossRef]
- Daeschel, A.M. Antimicrobial substances from lactic acid bacteria for use as food preservatives. Food Technol. 1989, 43, 91–94. [Google Scholar] [CrossRef]
| Phylum | Species | Product Types | References |
|---|---|---|---|
| Ascomycota | Atelosaccharomyces pseudotropicalis | Yoghurt | [26] |
| Blastodendrion arztii | Fresh unripened cheese | [51] | |
| Candida acidothermophilum | Yoghurt | [26,36,52] | |
| Candida apis | Hard or semi-hard cheese | [53] | |
| Candida beverwijkiae | Hard or semi-hard cheese | [51] | |
| Yoghurt | [26] | ||
| Candida blankii | Heat-treated milk and dairy product | [53] | |
| Candida glabrata | Soft and semi-soft cheese | [51] | |
| Candida glaebosa | Hard or semi-hard cheese | [54] | |
| Candida inconspicua | Buffalo, goat, or sheep cheese | [51] | |
| Fresh unripened cheese | [49,51] | ||
| Soft and semi-soft cheese | [51] | ||
| Candida intermedia | Hard or semi-hard cheese | [2,54] | |
| Other dairy products | [49] | ||
| Yoghurt | [26] | ||
| Candida krusei | Soft and semi-soft cheese | [26] | |
| Candida lusitaniae | Buffalo, goat, or sheep cheese | [51] | |
| Fresh unripened cheese | [51] | ||
| Soft and semi-soft cheese | [51] | ||
| Yoghurt | [26,52] | ||
| Candida multigemmis | Hard or semi-hard cheese | [2,54] | |
| Candida mycoderma | Buffalo, goat, or sheep cheese | [51] | |
| Candida norvegica | Soft and semi-soft cheese | [11] | |
| Candida parapsilosis | Fresh unripened cheese | [51] | |
| Hard or semi-hard cheese | [2,51,54] | ||
| Soft and semi-soft cheese | [11,49,51] | ||
| Yoghurt | [2,49] | ||
| Candida pseudoglaebosa | Raw milk | [27] | |
| Candida rugosa | Soft and semi-soft cheese | [11] | |
| Candida saitoana | Yoghurt | [26] | |
| Candida sake | Fresh unripened cheese | [55] | |
| Hard or semi-hard cheese | [54] | ||
| Candida sphaerica | Fresh unripened cheese | [26] | |
| Candida stellata | Hard or semi-hard cheese | [53] | |
| Yoghurt | [26,53] | ||
| Candida versatilis | Yoghurt | [26,36] | |
| Candida zeylanoides | Fresh unripened cheese | [51] | |
| Hard or semi-hard cheese | [51] | ||
| Soft and semi-soft cheese | [51] | ||
| Candida sp. | Mold ripened cheese | [26] | |
| Cyberlindnera jadinii | Soft and semi-soft cheese | [11] | |
| Debaryomyces hansenii | Blue-veined cheese | [26] | |
| Buffalo, goat, or sheep cheese | [51] | ||
| Fresh unripened cheese | [51,53,55] | ||
| Hard or semi-hard cheese | [26,49,51] | ||
| Heat-treated milk and dairy product | [26,52,53] | ||
| Other dairy products | [52,53] | ||
| Soft and semi-soft cheese | [11,51] | ||
| Mold ripened cheese | [26] | ||
| Yoghurt | [2,26,36,52,53] | ||
| Dekkera anomala | Fresh unripened cheese | [55] | |
| Yoghurt | [26] | ||
| Exophiala sp. | Fresh unripened cheese | [49] | |
| Galactomyces geotrichum | Butter and margarine | [2] | |
| Fresh unripened cheese | [2,49,55] | ||
| Heat-treated milk and dairy product | [2] | ||
| Other dairy products | [49] | ||
| Geotrichum capitatum | Soft and semi-soft cheese | [51] | |
| Geotrichum klebahnii | Hard or semi-hard cheese | [51] | |
| Geotrichum sp. | Buffalo, goat, or sheep cheese | [56] | |
| Raw milk | [57] | ||
| Kazachstania unispora | Soft and semi-soft cheese | [11] | |
| Kluyveromyces lactis | Fresh unripened cheese | [49,55] | |
| Kluyveromyces marxianus | Blue-veined cheese | [26] | |
| Fresh unripened cheese | [26,53] | ||
| Hard or semi-hard cheese | [26,53,54] | ||
| Heat-treated milk and dairy product | [26,53] | ||
| Mold ripened cheese | [26] | ||
| Other dairy products | [52] | ||
| Raw milk | [27] | ||
| Soft and semi-soft cheese | [49] | ||
| Yoghurt | [2,26,36,53] | ||
| Meyerozyma guilliermondii | Buffalo, goat, or sheep cheese | [51] | |
| Fresh unripened cheese | [49] | ||
| Soft and semi-soft cheese | [11,51] | ||
| Yoghurt | [49] | ||
| Naumovia dairenensis | Soft and semi-soft cheese | [11] | |
| Peterozyma toletana | Yoghurt | [36] | |
| Pichia fermentans | Soft and semi-soft cheese | [11,49] | |
| Pichia membranifaciens | Fresh unripened cheese | [26] | |
| Soft and semi-soft cheese | [11] | ||
| Pichia norvegensis | Soft and semi-soft cheese | [11] | |
| Priceomyces carsonii | Hard or semi-hard cheese | [54] | |
| Saccharomyces cerevisiae | Blue-veined cheese | [26] | |
| Buffalo, goat, or sheep cheese | [51] | ||
| Fresh unripened cheese | [51,53] | ||
| Hard or semi-hard cheese | [53] | ||
| Heat-treated milk and dairy product | [26] | ||
| Other dairy products | [52] | ||
| Soft and semi-soft cheese | [11,51] | ||
| Yoghurt | [26,36,53] | ||
| Torulaspora delbrueckii | Fresh unripened cheese | [51] | |
| Soft and semi-soft cheese | [11] | ||
| Torulopsis sp. | Yoghurt | [26] | |
| Wickerhomomyces anomalus | Soft and semi-soft cheese | [11] | |
| Yoghurt | [2] | ||
| Yarrowia lipolytica | Blue-veined cheese | [26] | |
| Buffalo, goat, or sheep cheese | [51] | ||
| Butter and margarine | [53] | ||
| Fresh unripened cheese | [26,55] | ||
| Hard or semi-hard cheese | [2,49,51,54] | ||
| Mold ripened cheese | [2,26] | ||
| Soft and semi-soft cheese | [11,49] | ||
| Yoghurt | [2] | ||
| Basidiomycota | Cryptococcus humicola | Butter and margarine | [2] |
| Hard or semi-hard cheese | [2] | ||
| Cryptococcus laurentii | Butter and margarine | [53] | |
| Fresh unripened cheese | [26,53] | ||
| Cryptococcus pseudolongus | Fresh unripened cheese | [49] | |
| Cryptococcus sp. | Heat-treated milk and dairy products | [52] | |
| Soft and semi-soft cheese | [11] | ||
| Cystobasidium minuta | Soft and semi-soft cheese | [11] | |
| Cutaneotrichosporon curvatus | Raw milk | [26] | |
| Cutaneotrichosporon cutaneum | Hard or semi-hard cheese | [51] | |
| Fresh unripened cheese | [51] | ||
| Naganishia albida | Butter and margarine | [53] | |
| Fresh unripened cheese | [53] | ||
| Other dairy products | [53] | ||
| Yoghurt | [53] | ||
| Rhodotorula diffluens | Butter and margarine | [53] | |
| Fresh unripened cheese | [53] | ||
| Hard or semi-hard cheese | [26,53] | ||
| Heat-treated milk and dairy products | [26,52,53] | ||
| Yoghurt | [2,26,53] | ||
| Rhodotorula glutinis | Butter and margarine | [53] | |
| Fresh unripened cheese | [53] | ||
| Other dairy products | [52,53] | ||
| Soft and semi-soft cheese | [11] | ||
| Rhodotorula mucilaginosa | Butter and margarine | [53] | |
| Fresh unripened cheese | [51,53] | ||
| Other dairy products | [52,53] | ||
| Soft and semi-soft cheese | [49] | ||
| Yoghurt | [2,36] | ||
| Rhodotorula sp. | Heat-treated milk and dairy products | [26] | |
| Saitozyma flava | Fresh unripened cheese | [53] | |
| Hard or semi-hard cheese | [53] | ||
| Heat-treated milk and dairy products | [26,53] | ||
| Other dairy products | [53] | ||
| Sporobolomyces roseus | Fresh unripened cheese | [26] | |
| Sporodiobolus salmonicolor | Hard or semi-hard cheese | [49] | |
| Trichosporon asahii | Soft and semi-soft cheese | [49] | |
| Vanrija humicola | Fresh unripened cheese | [51] |
| Phylum | Genera | Species | Product Types | References |
|---|---|---|---|---|
| Ascomycota | Acremonium | Nd * | Buffalo, goat, or sheep cheese | [56] |
| Alternaria | Alternaria alternata | Blue-veined cheese | [63] | |
| Hard or semi-hard cheese | [54,65] | |||
| Nd * | Hard or semi-hard cheese | [66] | ||
| Raw milk | [25] | |||
| Aspergillus section Aspergillus | Aspergillus glaucus | Heat-treated milk | [57] | |
| Raw milk | [57] | |||
| Aspergillus section Candidi | Aspergillus candidus | Buffalo, goat, or sheep cheese | [56] | |
| Aspergillus section Circumdati | Aspergillus ochraceus | Buffalo, goat, or sheep cheese | [56] | |
| Aspergillus section Flavi | Aspergillus flavus | Hard or semi-hard cheese | [67] | |
| Heat-treated milk | [57] | |||
| Raw milk | [57] | |||
| Aspergillus tamarii | Blue-veined cheese | [63] | ||
| Aspergillus section Fumigati | Aspergillus fumigatus | Blue-veined cheese | [63] | |
| Hard or semi-hard cheese | [65] | |||
| Heat-treated milk | [57] | |||
| Raw milk | [57] | |||
| Aspergillus section Nidulantes | Aspergillus sydowii | Yoghurt | [2] | |
| Aspergillus versicolor | Blue-veined cheese | [63] | ||
| Hard or semi-hard cheese | [59,68] | |||
| Semi-soft cheese | [4,59] | |||
| Buffalo, goat, or sheep cheese | [56] | |||
| Aspergillus section Nigri | Aspergillus niger | Blue-veined cheese | [63,67] | |
| Aspergillus | Nd * | Hard or semi-hard cheese | [54,66] | |
| Raw milk | [25] | |||
| Aureobasisium | Aureobasidium pullulans | Hard or semi-hard cheese | [69] | |
| Bipolaris | Bipolaris australiensis | Blue-veined cheese | [63] | |
| Botrytis | Botrytis cinerea | Hard or semi-hard cheese | [54] | |
| Byssochlamys | Byssochlamys nivea | Heat-treated milk | [2] | |
| Cladosporium cladosporioides complex | Cladosporium | Blue-veined cheese | [63] | |
| cladosporioides | Buffalo, goat, or sheep cheese | [56] | ||
| Butter and margarine | [2] | |||
| Hard or semi-hard cheese | [2,54,65] | |||
| Buffalo, goat, or sheep cheese | [56] | |||
| Cladosporium phyllophilum | Butter and margarine | [49] | ||
| Cladosporium herbarum complex | Cladosporium butyri | Butter and margarine | [2] | |
| Cladosporium herbarum | Hard or semi-hard cheese | [2,54] | ||
| Cladosporium macrocarpum | Hard or semi-hard cheese | [54] | ||
| Cladosporium sphaerospermum complex | Cladosporium halotolerans | Heat-treated milk | [49] | |
| Cladosporium sphaerospermum | Hard or semi-hard cheese | [49,54] | ||
| Cladosporium | Nd * | Hard or semi-hard cheese | [59,66] | |
| Semi-soft cheese | [59] | |||
| Didymella | Didymella pinodella | Fresh unripened cheese | [49] | |
| Other dairy product | [49] | |||
| Epicoccum | Epicoccum nigrum | Hard or semi-hard cheese | [54,69] | |
| Eupenicillium | Eupenicillium brefeldianum | Heat-treated milk | [2] | |
| Eurotium | Eurotium herbariorum | Hard or semi-hard cheese | [65] | |
| Nd * | Raw milk | [25,68] | ||
| Exophiala | Nd * | Cream cheese | [49] | |
| Fusarium | Fusarium avenaceum | Buffalo, goat, or sheep cheese | [56] | |
| Fusarium oxysporum | Hard or semi-hard cheese | [54,59] | ||
| Semi-soft cheese | [59] | |||
| Heat-treated milk | [2] | |||
| Fusarium solani | Hard or semi-hard cheese | [59] | ||
| Semi-soft cheese | [59] | |||
| Fusarium verticillioides | Hard or semi-hard cheese | [67] | ||
| Nd * | Hard or semi-hard cheese | [66] | ||
| Heat-treated milk | [57] | |||
| Raw milk | [57] | |||
| Gliocladium | Gliocladium roseum | Hard or semi-hard cheese | [54] | |
| Hamigera | Hamigera avellanea | Heat-treated milk | [2] | |
| Lecanicillium | Lecanicillium lecanii | Hard or semi-hard cheese | [59] | |
| Semi-soft cheese | [59] | |||
| Naumovia | Naumovia dairenensis | Soft cheese | [11] | |
| Neosartorya | Neosartorya fischeri | Heat-treated milk | [2] | |
| Penicillium section | Penicillium glabrum | Butter and margarine | [2] | |
| Aspergilloides | Hard or semi-hard cheese | [2,54,70] | ||
| Fresh unripened cheese | [2] | |||
| Yoghurt | [2] | |||
| Penicillium spinulosum | Hard or semi-hard cheese | [65] | ||
| Penicillium section | Penicillium funiculosum | Buffalo, goat, or sheep cheese | [56] | |
| Biverticillium | Penicillium minioluteum | Hard or semi-hard cheese | [69] | |
| Penicillium purpureogenum | Hard or semi-hard cheese | [68] | ||
| Penicillium rugulosum | Hard or semi-hard cheese | [54,59] | ||
| Semi-soft cheese | [59] | |||
| Penicillium section | Penicillium bialowiezense | Hard or semi-hard cheese | [49] | |
| Brevicompacta | Yoghurt | [49] | ||
| Penicillium brevicompactum | Hard or semi-hard cheese | [2,49,54,59,65,69,70] | ||
| Semi-soft cheese | [59] | |||
| Buffalo, goat, or sheep cheese | [56] | |||
| Penicillium section Sclerotiora | Penicillium adametzioides | Fresh unripened cheese | [49] | |
| Penicillium section | Penicillium antarcticum | Hard or semi-hard cheese | [49] | |
| Canescentia | Penicillium arenicola | Buffalo, goat, or sheep cheese | [56] | |
| Penicillium canescens | Hard or semi-hard cheese | [68] | ||
| Penicillium novae-zeelandiae | Hard or semi-hard cheese | [54] | ||
| Penicillium section Charlesii | Penicillium charlesii | Fresh unripened cheese | [49] | |
| Heat-treated milk | [49] | |||
| Penicillium dierckxii | Fresh unripened cheese | [49] | ||
| Penicillium section Chrysogena | Penicillium chrysogenum | Butter and margarine | [2] | |
| Blue-veined cheese | [63] | |||
| Hard or semi-hard cheese | [54,59,62,65,69,70] | |||
| Semi-soft cheese | [59] | |||
| Buffalo, goat, or sheep cheese | [56] | |||
| Fresh unripened cheese | [2] | |||
| Yoghurt | [2] | |||
| Penicillium griseofulvum | Hard or semi-hard cheese | [68] | ||
| Penicillium nalgiovense | Hard or semi-hard cheese | [4,49,59] | ||
| Semi-soft cheese | [4,59] | |||
| Buffalo, goat, or sheep cheese | [56] | |||
| Penicillium section Citrina | Penicillium steckii | Hard or semi-hard cheese | [69] | |
| Penicillium section Exilicaulis | Penicillium corylophitum | Hard or semi-hard cheese | [65] | |
| Penicillium section Fasciculata | Penicillium aurantiogriseum | Hard or semi-hard cheese | [65,69] | |
| Buffalo, goat, or sheep cheese | [56] | |||
| Yoghurt | [2] | |||
| Penicillium camembertii | Hard or semi-hard cheese | [68] | ||
| Penicillium commune | Other dairy products | [49] | ||
| Hard or semi-hard cheese | [2,4,49,54,59,65,69,70] | |||
| Mold ripened cheese | [2] | |||
| Semi-soft cheese | [4,59] | |||
| Buffalo, goat, or sheep cheese | [56] | |||
| Fresh unripened cheese | [2] | |||
| Penicillium crustosum | Buffalo, goat, or sheep cheese | [56] | ||
| Hard or semi-hard cheese | [54,59,62,65,69,70] | |||
| Semi-soft cheese | [59] | |||
| Buffalo, goat, or sheep cheese | [56] | |||
| Penicillium discolor | Hard or semi-hard cheese | [49,59] | ||
| Semi-soft cheese | [59] | |||
| Penicillium echinulatum | Hard or semi-hard cheese | [59,65,68,69] | ||
| Semi-soft cheese | [59] | |||
| Penicillium hirsutum | Buffalo, goat, or sheep cheese | [56] | ||
| Penicillium nordicum | Hard or semi-hard cheese | [49] | ||
| Penicillium palitans | Fresh unripened cheese | [49] | ||
| Hard or semi-hard cheese | [49,65,69] | |||
| Penicillium solitum | Buffalo, goat, or sheep cheese | [56] | ||
| Hard or semi-hard cheese | [2,49,54,59,65,69] | |||
| Other dairy products | [49] | |||
| Semi-soft cheese | [4,59] | |||
| Yoghurt | [49] | |||
| Penicillium verrucosum | Buffalo, goat, or sheep cheese | [56] | ||
| Hard or semi-hard cheese | [54,59,68,69] | |||
| Mold ripened cheese | [2] | |||
| Semi-soft cheese | [4,59] | |||
| Penicillium viridicatum | Hard or semi-hard cheese | [2,54,65,67,68,69] | ||
| Penicillium section Paradoxa | Penicillium atramentosum | Blue-veined cheese | [63] | |
| Hard or semi-hard cheese | [59] | |||
| Semi-soft cheese | [59] | |||
| Penicillium section Penicillium | Penicillium expansum | Butter and margarine | [2] | |
| Hard or semi-hard cheese | [2,65,68,69] | |||
| Penicillium section Lanata- | Penicillium oxalicum | Buffalo, goat, or sheep cheese | [56] | |
| Divaricata | Penicillium simplicissimum | Hard or semi-hard cheese | [57] | |
| Penicillium section | Penicillium roquefortii | Buffalo, goat, or sheep cheese | [56] | |
| Roquefortum | Hard or semi-hard cheese | [2,54,59,65,68,69] | ||
| Mold ripened cheese | [2] | |||
| Semi-soft cheese | [4,59] | |||
| Penicllium | Nd * | Buffalo, goat, or sheep cheese | [56] | |
| Hard or semi-hard cheese | [54,66] | |||
| Heat-treated milk | [57] | |||
| Raw milk | [25,57] | |||
| Phaeramularia | Nd * | Hard or semi-hard cheese | [54] | |
| Phoma | Phoma glomerata | Hard or semi-hard cheese | [2,70] | |
| Phoma nebulosa | Hard or semi-hard cheese | [69] | ||
| Nd * | Buffalo, goat, or sheep cheese | [56] | ||
| Hard or semi-hard cheese | [2,54,65,69] | |||
| Rhinocladiella | Nd * | Blue-veined cheese | [63] | |
| Sarocladium | Sarocladium strictum | Hard or semi-hard cheese | [69] | |
| Scopulariopsis | Scopulariopsis brevicaulis | Buffalo, goat, or sheep cheese | [56] | |
| Hard or semi-hard cheese | [51,59] | |||
| Semi-soft cheese | [4,59] | |||
| Nd * | Hard or semi-hard cheese | [54,67] | ||
| Talaromyces | Talaromyces macrosporus | Heat-treated milk | [2] | |
| Trichoderma | Trichoderma harzianum | Hard or semi-hard cheese | [65] | |
| Ulocladium | Ulocladium chartarum | Hard or semi-hard cheese | [69] | |
| Basidiomycota | Wallemia | Wallemia sebi | Raw milk | [25] |
| Zygomycota | Mucor | Mucor circinelloides | Hard or semi-hard cheese | [49,69] |
| Yoghurt | [2] | |||
| Mucor hiemalis | Buffalo, goat, or sheep cheese | [56] | ||
| Hard or semi-hard cheese | [2,65,70] | |||
| Yoghurt | [2] | |||
| Mucor plumbeus | Hard or semi-hard cheese | [65,69] | ||
| Mucor racemosus | Hard or semi-hard cheese | [49,59,65,68,69] | ||
| Semi-soft cheese | [59] | |||
| Yoghurt | [2] | |||
| Rhizopus | Rhizopus stolonifer | Blue-veined cheese | [63] | |
| Buffalo, goat, or sheep cheese | [56] | |||
| Thamnidium | Thamnidium elegans | Hard or semi-hard cheese | [49] |
| Preservatives | Dairy Product | USA Regulation | Food and Drug Administration (FDA) Code | EU Legislation | EU Code |
|---|---|---|---|---|---|
| Natamycin (pimaricin) | Cheese | 20 mg/kg | 172.155 | - | - |
| Uncut hard, semi-hard, and semi-soft cheese | 20 mg/kg | 172.155 | 1 mg/dm2, surface (not present at a depth of 5 mm) | E 235 | |
| Sorbic acid | Margarine | 1000 mg/kg | 182.3089 | - | E 200 |
| Flavoured fermented milk | 2000 mg/kg | 1500 mg/kg | |||
| Non-heat-treated dairy-based desserts | 2000 mg/kg | 300 mg/kg | |||
| Whey cheeses, cheese products, processed cheeses | 2000 mg/kg | 2000 mg/kg | |||
| Curdled milk, unripened products, ripened products, pre-packed, sliced; layered ripened products | 2000 mg/kg | 1000 mg/kg | |||
| Potassium sorbate and calcium sorbate | Cold-pack cheese, cream cheese, pasteurized process cheese food, pasteurized process cheese spread, semi-soft part-skim cheeses | 3000 mg/kg | 182.3640 and 182.3225 | - | E 202 and E 203 |
| Flavoured fermented milk products including heat-treated products | 3000 mg/kg | 1500 mg/kg | |||
| Non-heat-treated dairy-based desserts | 3000 mg/kg | 300 mg/kg | |||
| Whey cheeses, cheese products, processed cheeses | 3000 mg/kg | 2000 mg/kg | |||
| Curdled milk, unripened products, ripened products, pre-packed, sliced; layered ripened products | 3000 mg/kg | 1000 mg/kg | |||
| Flavoured fermented milk products including heat-treated products | 3000 mg/kg | 1500 mg/kg | |||
| Non-heat-treated dairy-based desserts | 3000 mg/kg | 300 mg/kg | |||
| Whey cheeses, cheese products, processed cheeses | 3000 mg/kg | 2000 mg/kg | |||
| Curdled milk, unripened products, ripened products, pre-packed, sliced; layered ripened products | 3000 mg/kg | 1000 mg/kg | |||
| Sodium benzoate | Margarine | 1000 mg/kg | 184.1733 | - | E 211 |
| Flavoured fermented milk products including heat-treated products | - | 1500 mg/kg | |||
| Non-heat-treated dairy-based desserts | - | 300 mg/kg | |||
| Whey cheeses, cheese products, processed cheeses | - | 2000 mg/kg | |||
| Curdled milk, unripened products, ripened products, pre-packed, sliced; layered ripened products | - | 1000 mg/kg | |||
| Potassium benzoate and calcium benzoate | Flavoured fermented milk products including heat-treated products | - | 184.1081 | 1500 mg/kg | E 212 and E 213 |
| Non-heat-treated dairy-based desserts | - | 300 mg/kg | |||
| Whey cheeses, cheese products, processed cheeses | - | 2000 mg/kg | |||
| Curdled milk, unripened products, ripened products, pre-packed, sliced; layered ripened products | - | 1000 mg/kg | |||
| Propionic acid | Gruyere cheese, swiss cheese, and emmentaler cheese | surface | - | E 280 | |
| Ripened cheese | - | surface treatment | |||
| Sodium propionate | Cheeses and related cheese products | surface | 184.1784 | - | E 281 |
| Ripened cheese | - | surface treatment |
| Lactic Acid Bacteria (LAB) and Propionibacteria | Matrix | Metabolites | Targets | References |
|---|---|---|---|---|
| Genus Lactococcus | ||||
| Lc. lactis | Lab-Lemco tryptone broth (LTB) | Nd * | Aspergillus flavus | [162] |
| LTB | Nisin | Aspergillus parasiticus | [163] | |
| Potatoe Dextrose Agar (PDA) + 0.1% Triton X-100 | Nd * | Aspergillus fumigatus, A. parasiticus, A. flavus | [164] | |
| PDA + 0.1% Triton X-100 | Possibly proteinaceous compound(s) | Fusarium spp., A. flavus, A. parasiticus | [165] | |
| LTB and PDA | Possibly proteinaceous compound(s), lactic acid | Penicillium expansum | [166] | |
| Genus Lactobacillus | ||||
| Lactobacillus spp. | De Man, Rogosa and Sharpe (MRS) agar | Acetic acid, propionic acid, lactic acid, peptides | Penicillium candidum, Debaryomyces hansenii | [167] |
| Group Lb. alimentarius/Lb. farciminis | ||||
| Lb. paralimentarus | Modified Sabouraud Dextrose Broth (mSDB) medium | Lactic acid, phenyllactic acid, acetic acid, peptides | Aspergillus japonicus, Eurotium repens, Penicillium roseopurpureum | [168] |
| Group Lb. brevis | ||||
| Lb. brevis | PDA | Peptide | Penicillium roqueforti | [169] |
| wheat flour hydrolysate (WFH) broth | Acetic acid, phenyllactic acid, lactic acid | Fusarium graminearum | [170] | |
| mMRS agar | Organic acids and proteinaceous compounds | Fusarium spp. | [171] | |
| MRS agar or PDA | Peptide | Penicillium camemberti, P. roqueforti, Aspergillus niger, Rhizopus oryzae, Kluyveromyces marxianus, Torulopsis candida, Meyerozyma guillermondii, Saccharomyces cerevisiae | [172] | |
| Group Lb. casei | ||||
| Lb. casei | PDA + 0.1% Triton X-100 | Possibly proteinaceous compound(s) | A. flavus, A. parasiticus, Fusarium sp. | [165] |
| PDA | Peptide | Penicillium citrinum, P. expansum, A. flavus | [173] | |
| Yoghurt | Lactic acid and cyclo-(Leu-Pro) | Penicillium sp. | [174] | |
| LTB and PDA | Possibly proteinaceous compound(s), lactic acid | P. expansum | [166] | |
| Lb. paracasei | Yoghurt and cheese surface | Propionic acid, acetic acid, lactic acid, succinic acid, 2-pyrrolidone-5-carboxylic acid, 3-phynyllactic acid, hydroxyphenyllactic acid | Candida pulcherrima, Candida magnoliae, Candida parapsilosis, Zygosaccharomyces bailii | [151] |
| MRS agar | Peptide | Candida albicans, Candida blankii, Candida pseudointermedia | [175] | |
| Chemically defined interaction medium | Diacetyl | Penicillium solitum, Penicillium sp. | [153] | |
| Yoghurt and acidified milk | Diacetyl, acetic acid, butanoic acid, 2,3-pentadione | P. solitum, Penicillium sp. | [153] | |
| Lb. rhamnosus | Yoghurt | Acetic acid, lactic acid | Rhodotorula mucilaginosa | [15,158] |
| Group Lb. coryniformis | ||||
| Lb. coryniformis | MRS agar or PDA | Peptide +/−3 KDa, phenyllactic acid, cyclo(Phe-Pro), cyclo(Phe-OH-Pro), reuterin | Broad spectrum | [147] |
| Group Lb. delbrueckii | ||||
| Lb. acidophilus | PDA + 0.1% Triton X | Nd * | A. fumigatus | [164] |
| Lb. amylovorus | MRS broth | 3-phenylpropanoic acid, p-coumaric, (E)-2-methylcinnamic acid, 3-phenyllactic acid, 3-(4-hydroxyphenyl)lactic acid, lactic acid, acetic acid, d-glucuronic acid, salicylic acid, cytidine and 2′-deoxycytidine, sodium decanoate, | A. fumigatus, Fusarium culmorum | [175] |
| Cheese | Cyclo(l-Pro-l-Pro), cyclo(l-Leu-l-Pro), cyclo(l-Tyr-l-Pro) cyclo(l-Met-l-Pro) and cyclo(l-His-l-Pro) | P. roqueforti, P. expansum | [176] | |
| Milk agar and cheese | dl-ρ-hydroxyphenyllactic acid, 4-hydroxybenzoic acid, (S)-(−)-2-hydroxyisocapric acid, azelaic acid, phenyllactic acid, benzoic acid, hydrocinnacmic acid, 3-hydroxydecanoic acid, dl-β-hydroxylauric acid, decanoic acid, salicylic acid, 4-hydroxybenzoic, vanillic acid, (S)-(−)-2-hydroxyisocapric acid | P. expansum | [177] | |
| Lb. delbrueckii | LTB and PDA | Possibly proteinaceous compound(s), lactic acid | P. expansum | [166] |
| Group Lb. fructivorans | ||||
| Lb. sanfranciscencis | Malt-agar medium | Caproic acid, propionic acid, butyrix acid, acetic acid, valeric acid | F. graminearum | [178,179] |
| Group Lb. perolens | ||||
| Lb. harbinensis | Yoghurt | Acetic acid, lactic acid | Yarrowia lipolytica, P. expansum, Penicillium brevicompactum, D. hansenii, R. mucilaginosa, Kluyveromyces lactis | [15,158] |
| Group Lb. plantarum | ||||
| Lb. pentosus | MRS | Peptide | A. niger | [180] |
| MRS agar | Peptide, phenyllactic and hydroxyphenyllactic acid | Penicillium nalgiovense, Aspergillus candidus | [181] | |
| Lb. plantarum | mMRS agar | (S)-(−)-2-hydroxyisocapric acid, hydrocinnamic acid, phenyllactic acid, decanoic acid, azelaic acid, 4-hydroxybenzoic acid, p-coumaric acid, vanillic acid, dl-Þ-hydroxyphenyllactic acid, 3-hydroxydecanoic acid | Microsporum canis, Microsporum gypseum, Epidermophyton floccosum | [182] |
| MRS broth | Benzoic acid, 5-methyl-2,4-imidazolidinedione, tetrahydro-4-hydroxy-4-methyl-2H-pyran-2-one, 3-(2-methylpropyl)-2,5-piperazinedione, cyclo(glycyl-l-leucyl)). | Fusarium avenaceum | [183] | |
| MRS agar | Lactic acid, PLA, cyclo(l-Leu-l-Pro), cyclo(l-Phe-l-Pro) | A. niger, F. graminearum, F. culmorum, Fusarium oxysporum | [184] | |
| Wheat flour hydrolysate (WFH) | Phenyllactic acid, 4-hydroxy-phenillactic acid | E. repens, Eurotium rubrum, Penicillium corylophilum, P. roqueforti, P. expansum, Endomyces fibuliger, A. niger, A. flavus, Monilia sitophila, F. graminearum | [185] | |
| MRS broth | 3-phenyllactic acid, cyclo(Phe-Pro), cyclo(Phe-OH-Pro), cyclo(l-Phe-LPro) and cyclo(l-Phe-trans-4-OH-l-Pro) dipeptides | Fusarium sporotrichioides, A. fumigatus, K. marxianus | [186] | |
| Wheat flour hydrolysate (WFH) | Acetic acid, phenyllactic acid, lactic acid | F. graminearum, A. niger | [170] | |
| MRS agar | 3-(R)-hydroxydecanoic acid, 3-hydroxy-5-cis-dodecenoic acid, 3-(R)-hydroxydodecanoic acid and 3-(R)-hydroxytetradecanoic acid | A. fumigatus, Aspergillus nidulans, K. marxianus, P. roqueforti, Penicillium commune, Penicillium anomala, R. mucilaginosa | [187] | |
| MRS agar plates | Acetic acid | A. flavus, F. graminearum, Rhizopus stolonifer, Sclerotium oryzae, Rhizoctonia solani, Botrytis cinerea, Sclerotinia minor | [188] | |
| MRS agar | Peptide, phenyl-lactic and hydroxy-phenyllactic acid | P. nalgiovense, P. camemberti, Penicillium verrucosum, Penicillium chrysogenum, A. candidus, A. flavus, A. ochraceus, A. fumigatus, Galactomyces geotrichum, Moniliella spp., Mucor racemosus, Wallemia sebi, Eurotium herbariorum | [181] | |
| Soybean | 3,6-bis(2-methylpropyl)-2,5-piperazinedion | A. flavus | [189] | |
| MRS agar plates | 3-3-phenyllactic acid (PLA), lactic acid, acetic acid | A. fumigatus, Rhizopus stolonifer | [190] | |
| MRS agar medium, apple-based agar growth medium | Lactic acid, acetic acid | P. expansum, Penicillium notatum | [191] | |
| MRS agar medium | 2-hydroxy-4-methylpentanoic acid | A. niger, Aspergillus tubingensis, Penicillium crustosum | [192] | |
| PDA plates | 3-PLA, benzeneacetic acid, 2-propenyl ester | B. cinerea, Glomerella cingulate, Phytophthora drechsleri Tucker, P. citrinum, Penicillium digitatum, F. oxysporum | [193] | |
| Chopped Meat Carbohydrate (CMC) broth | Lactic acid | F. avenaceum, F. culmorum, F. graminearum, F. oxyporum | [194] | |
| Group Lb. reuteri | ||||
| Lb. fermentum | MRS | Peptide | A. niger | [180] |
| Lb. reuteri | Wheat flour hydrolysate (WFH) broth | Acetic acid, phenyllactic acid, lactic acid | F. graminearum, A. niger | [170] |
| mMRS agar | Acid, vanillic acid, dl-Þ-hydroxyphenyllactic acid, 3-hydroxydecanoic acid, (S)-(-)-2–hydroxyisocapric acid, hydrocinnamic acid, phenyllactic acid, decanoic acid, azelaic acid, 4-hydroxybenzoic acid, p-coumaric | M. canis, M. gypseum, E. floccosum | [182] | |
| Lb. rossiae | mSDB agar medium | Lactic acid, phenyllactic acid, acetic acid, peptides | A. japonicus, E. repens, P. roseopurpureum | [168] |
| Group Lb. sakei | ||||
| Lb. sakei | MRS | Sakacin KTU05-6 | A. flavus, A. fumigatus, A. niger, Aspergillus versicolor, F. culmorum, Fusarium poae, Mucor spp., P. chrysogenum, P. expansum, Penicillium spp. | [195] |
| Genus Pediococcus | ||||
| Pc. acidilactici | MRS | Pediocin KTU05-08 | A. flavus, A. fumigatus, A. niger, A. versicolor, F. culmorum, F. poae, Mucor spp., P. chrysogenum, P. expansum, Penicillium spp. | [195] |
| Pc. pentosaceous | MRS | Peptide | A. niger | [180] |
| MRS agar plates | Acetic acid | A. flavus, F. graminearum, R. stolonifer, S. oryzae, R. solani, B. cinerea, S. minor | [188] | |
| MRS | Pediocin KTU05-09, KTU05-10 and AcKTU05-67 | A. flavus, A. fumigatus, A. niger, A. versicolor, F. culmorum, F. poae, Mucor spp., P. chrysogenum, P. expansum, Penicillium spp. | [195] | |
| MRS agar medium | Possibly cyclic dipeptide | P. expansum, Penicillium notatum | [191] | |
| Genus Weissella | ||||
| W. confusa | MRS agar medium | Lactic acid, acetic acid | P. expansum, P. notatum | [191] |
| W. cibaria | MRS agar medium | Lactic acid, acetic acid | P. expansum, P. notatum | [191] |
| W. paramesenteroides | MRS agar medium | 2-hydroxy-4-methylpentanoic acid, lactic acid, acetic acid | A. niger, A. tubingensis, P. crustosum | [192] |
| MRS agar plates | Acetic acid | A. flavus, F. graminearum, R. stolonifer, S. oryzae, B. cinerea, S. minor | [188] | |
| Genus Propionibacteria | ||||
| P. acidipropionici | Sodium lactate (SL) medium | Lactic acid, propionic acid, acetic acid | A. fumigatus | [196] |
| MRS | Propionic acid, acetic acid | R. mucilaginosa, P. roqueforti, A. fumigatus | [196] | |
| MRS-acetate | Propionic acid, acetic acid | R. mucilaginosa, P. roqueforti, A. fumigatus | [196] | |
| SL broth | PLA | A. fumigatus | [196] | |
| P. freudenreichii subsp. shermanii | Skim milk | propionic acid, 3-phenyllactic acid | P. chrysogenum | [159] |
| MRS | Acetic acid, propionic acid, 3-phenyllactic acid, 4-hydroxy-phenyllactic acid | P. chrysogenum | [159] | |
| MRS | Propionic acid, acetic acid | P. roqueforti, A. fumigatus | [196] | |
| MRS-acetate | Propionic acid, acetic acid | R. mucilaginosa, P. roqueforti, A. fumigatus | [196] | |
| P. freudenreichii subsp. freudenreichii | MRS | Propionic acid, acetic acid | R. mucilaginosa, P. roqueforti, A. fumigatus | [196] |
| MRS-acetate | Propionic acid, acetic acid | R. mucilaginosa, P. roqueforti, A. fumigatus | [196] | |
| P. jensenii | Yoghurt and cheese surface | Propionic acid, acetic acid, lactic acid, succinic acid, 2-pyrrolidone-5-carboxylic acid, 3-phynyllactic acid, hydroxyphenyllactic acid | C. pulcherrima, C. magnollae, C. parapsilosis, Z. bailii | [151] |
| MRS | Propionic acid, acetic acid | R. mucilaginosa, P. roqueforti, A. fumigatus | [196] | |
| MRS-acetate | Nd * | R. mucilaginosa, P. roqueforti, A. fumigatus | [196] | |
| P. thoenii | Sodium Lactate (SL) medium | Lactic acid, propionic acid | A. fumigatus | [196] |
| SL broth | PLA | A. fumigatus | [196] | |
| MRS | Propionic acid, acetic acid | R. mucilaginosa, P. roqueforti, A. fumigatus, K. marxianus | [196] | |
| MRS-acetate | Propionic acid, acetic acid | R. mucilaginosa, P. roqueforti, A. fumigatus | [196] | |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Garnier, L.; Valence, F.; Mounier, J. Diversity and Control of Spoilage Fungi in Dairy Products: An Update. Microorganisms 2017, 5, 42. https://doi.org/10.3390/microorganisms5030042
Garnier L, Valence F, Mounier J. Diversity and Control of Spoilage Fungi in Dairy Products: An Update. Microorganisms. 2017; 5(3):42. https://doi.org/10.3390/microorganisms5030042
Chicago/Turabian StyleGarnier, Lucille, Florence Valence, and Jérôme Mounier. 2017. "Diversity and Control of Spoilage Fungi in Dairy Products: An Update" Microorganisms 5, no. 3: 42. https://doi.org/10.3390/microorganisms5030042
APA StyleGarnier, L., Valence, F., & Mounier, J. (2017). Diversity and Control of Spoilage Fungi in Dairy Products: An Update. Microorganisms, 5(3), 42. https://doi.org/10.3390/microorganisms5030042






