Comparison of Microbiological and Probiotic Characteristics of Lactobacilli Isolates from Dairy Food Products and Animal Rumen Contents
Abstract
:1. Introduction
2. Experimental Section
2.1. Isolation, Phenotypic Characterization and Carbohydrate Fermentation Profile of Strains
2.2. Identification of LAB
2.3. Acid Tolerance
2.4. Bile Salt Tolerance
2.5. Haemolytic Activity
2.6. Antibiotic Resistance
2.7. Antimicrobial Activity
2.8. Assessment of Bacterial Hydrophobicity
2.9. Statistical Analysis
3. Results
3.1. Isolation, Phenotypic Characterization and Carbohydrate Fermentation Profile of Strains
3.2. Identification of LAB
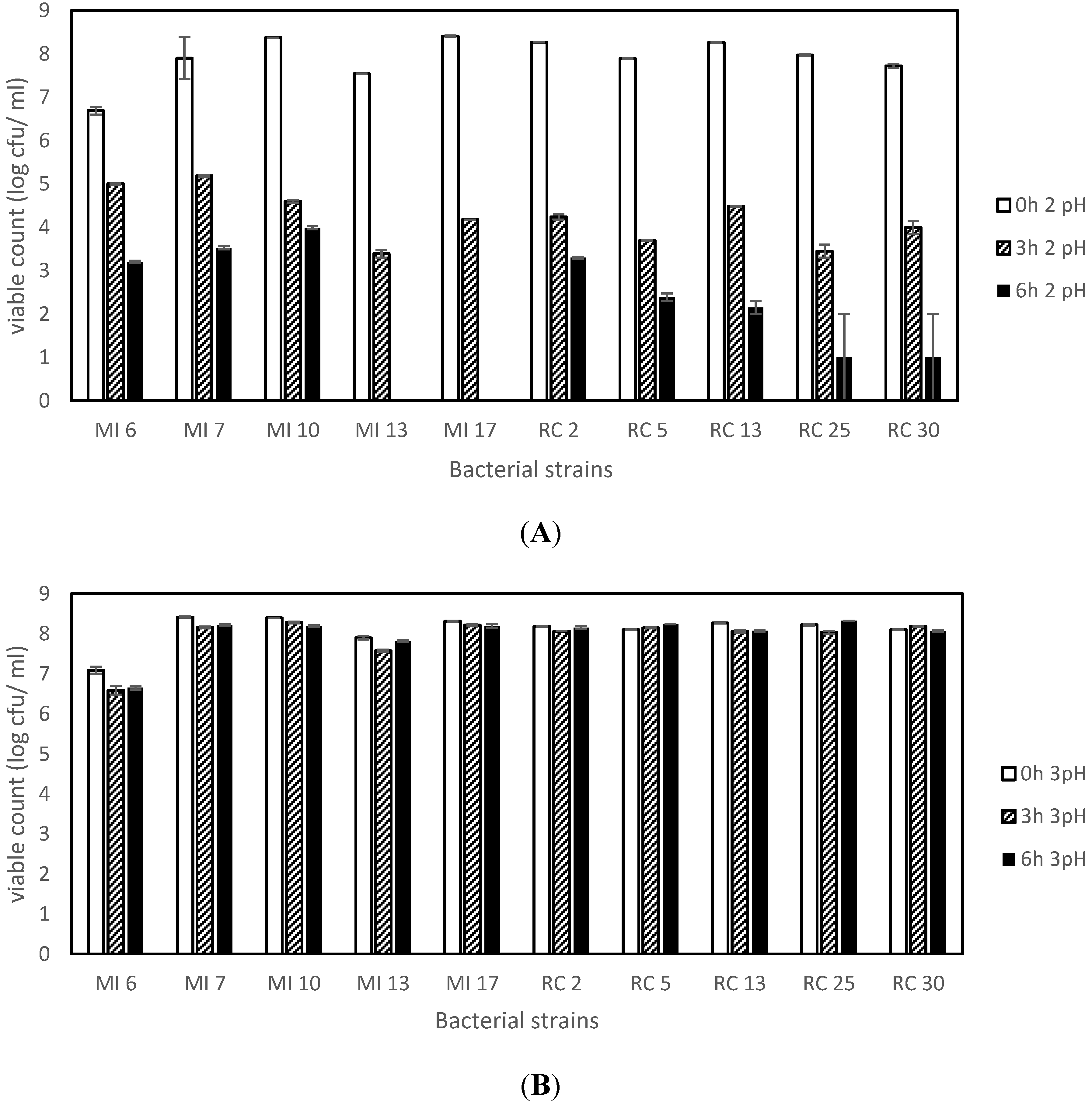
3.3. Acid Tolerance
| Strain | Source | Arabinose | Cellobiose | Fructose | Glucose | Galactose | Glycerol | Lactose | Mannose | Mannitol | Melibiose | Maltose | Raffinose | Ribose | Sorbitol | Sucrose |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MI 6 | dairy food | ++ | − | − | ++ | ++ | − | ++ | − | − | ++ | ++ | ++ | + | − | ++ |
| MI 7 | dairy food | ++ | − | − | ++ | ++ | − | ++ | − | − | ++ | ++ | ++ | + | − | ++ |
| MI 10 | dairy food | ++ | − | + | ++ | ++ | − | ++ | − | − | ++ | ++ | ++ | + | − | ++ |
| MI 13 | dairy food | ++ | ++ | ++ | ++ | ++ | − | ++ | ++ | ++ | ++ | ++ | ++ | ++ | + | ++ |
| MI 17 | dairy food | ++ | ++ | ++ | ++ | ++ | − | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ |
| RC 2 | animal rumen | ++ | ++ | ++ | ++ | ++ | − | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ |
| RC 5 | animal rumen | ++ | ++ | ++ | ++ | ++ | − | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ |
| RC 13 | animal rumen | ++ | ++ | ++ | ++ | ++ | − | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ |
| RC 25 | animal rumen | − | ++ | ++ | ++ | ++ | − | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ |
| RC 30 | animal rumen | ++ | ++ | ++ | ++ | ++ | − | ++ | ++ | ++ | ++ | ++ | + | + | ++ | ++ |
| Sl. No | Microbial ID | Source | Genetic Identification |
|---|---|---|---|
| 1 | MI 6 | dairy food (yoghurt) | Lactobacillus reuteri TD1, complete genome |
| 2 | MI 7 | dairy food (yoghurt) | Lactobacillus reuteri JCM 1112 DNA, complete genome |
| 3 | MI 10 | dairy food (yoghurt) | Lactobacillus reuteri strain C16 |
| 4 | MI 13 | dairy food (cheese) | Lactobacillus rhamnosus LOCK 908, complete genome |
| 5 | MI 17 | dairy food (cheese) | Lactobacillus rhamnosus LOCK 908, complete genome |
| 6 | RC 2 | animal rumen (cow) | Lactobacillus plantarum 16, complete genome |
| 7 | RC 5 | animal rumen (cow) | Lactobacillus plantarum 16, complete genome |
| 8 | RC 13 | animal rumen (cow) | Lactobacillus plantarum subsp. plantarum ST-III, complete genome |
| 9 | RC 25 | animal rumen (cow) | Lactobacillus plantarum 16, complete genome |
| 10 | RC 30 | animal rumen (cow) | Lactobacillus plantarum subsp. plantarum ST-III, complete genome |
3.4. Bile Salt Tolerance
| Strain | Source | 0.3% Bile | 2% Bile | ||||
|---|---|---|---|---|---|---|---|
| 0 h | 3 h | 6 h | 0 h | 3 h | 6 h | ||
| MI 6 | dairy food | 9.794 ± 0.054 | 8.016 ± 0.088 | 8.777 ± 0.087 | 9.658 ± 0.007 | 9.078 ± 0.051 | 9.078 ± 0.051 |
| MI 7 | dairy food | 10.021 ± 0.012 | 8.574 ± 0.008 | 8.562 ± 0.008 | 10.158 ± 0.013 | 8.889 ± 0.157 | 9.230 ± 0 |
| MI 10 | dairy food | 9.602 ± 0 | 8.984 ± 0.003 | 8.469 ± 0.031 | 9.511 ± 0.113 | 9.102 ± 0.144 | 8.778 ± 0.249 |
| MI 13 | dairy food | 9.451 ± 0.213 | 8.067 ± 0.021 | 9.434 ± 0.025 | 7.827 ± 0.181 | 8.661 ± 0.260 | 7.389 ± 0.125 |
| MI 17 | dairy food | 8.866 ± 0.125 | 9.075 ± 0.010 | 8.396 ± 0.049 | 8.651 ± 0.069 | 8.540 ± 0.088 | 7.500 ± 0.281 |
| RC 2 | animal rumen | 9.217 ± 0.019 | 9.428 ± 0.017 | 9.477 ± 0 | 8.923 ± 0.110 | 9.289 ± 0.047 | 9.755 ± 0.043 |
| RC 5 | animal rumen | 10.069 ± 0.013 | 9.041 ± 0.017 | 9.477 ± 0 | 10.234 ± 0.009 | 9.136 ± 0.134 | 9.581 ± 0.088 |
| RC 13 | animal rumen | 9.413 ± 0.047 | 9.118 ± 0.012 | 9.346 ± 0.008 | 9.802 ± 0.044 | 8.690 ± 0.125 | 9.711 ± 0.042 |
| RC 25 | animal rumen | 8.922 ± 0.110 | 8.841 ± 0.013 | 9.477 ± 0 | 9.589 ± 0.063 | 9.300 ± 0.031 | 9.920 ± 0.048 |
| RC 30 | animal rumen | 9.096 ± 0.025 | 8.679 ± 0.051 | 9.229 ± 0.033 | 9.871 ± 0.037 | 9.918 ± 0.052 | 9.590 ± 0.032 |
3.5. Haemolytic Activity
3.6. Antibiotic Resistance
| Strain | Source | Antibiotic Resistance * | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| TE | ST | NA | CN | FA | VA | K | C | E | CIP | AMP | ||
| MI 6 | dairy food | R | R | R | R | R | R | R | R | R | R | R |
| MI 7 | dairy food | R | R | R | R | R | R | R | R | R | R | R |
| MI 10 | dairy food | R | R | R | R | R | R | R | R | R | R | R |
| MI 13 | dairy food | S | R | R | R | R | R | R | S | S | R | S |
| MI 17 | dairy food | R | R | R | R | R | R | R | S | I | R | S |
| RC 2 | animal rumen | R | R | R | R | R | R | R | S | I | R | S |
| RC 5 | animal rumen | I | R | R | R | R | R | R | S | I | R | S |
| RC 13 | animal rumen | I | R | R | R | R | R | R | S | I | R | S |
| RC 25 | animal rumen | I | R | R | R | R | R | R | S | I | R | S |
| RC 30 | animal rumen | I | R | R | R | R | R | R | S | I | R | S |
3.7. Antimicrobial Activity
| Strain | Source | Escherichia coli | Listeria monocytogenes | Enterobacter aerogenes | Staphylococcus aureus | Salmonella menston |
|---|---|---|---|---|---|---|
| MI 6 | dairy food | − | − | ± | ± | + |
| MI 7 | dairy food | − | − | ± | + | + |
| MI 10 | dairy food | − | − | ± | ± | ± |
| MI 13 | dairy food | − | + | − | + | ± |
| MI 17 | dairy food | − | +++ | ± | ± | + |
| RC 2 | animal rumen | − | + | ± | + | + |
| RC 5 | animal rumen | − | ++ | + | − | + |
| RC 13 | animal rumen | − | +++ | + | ± | + |
| RC 25 | animal rumen | − | +++ | + | + | + |
| RC 30 | animal rumen | − | +++ | + | + | + |
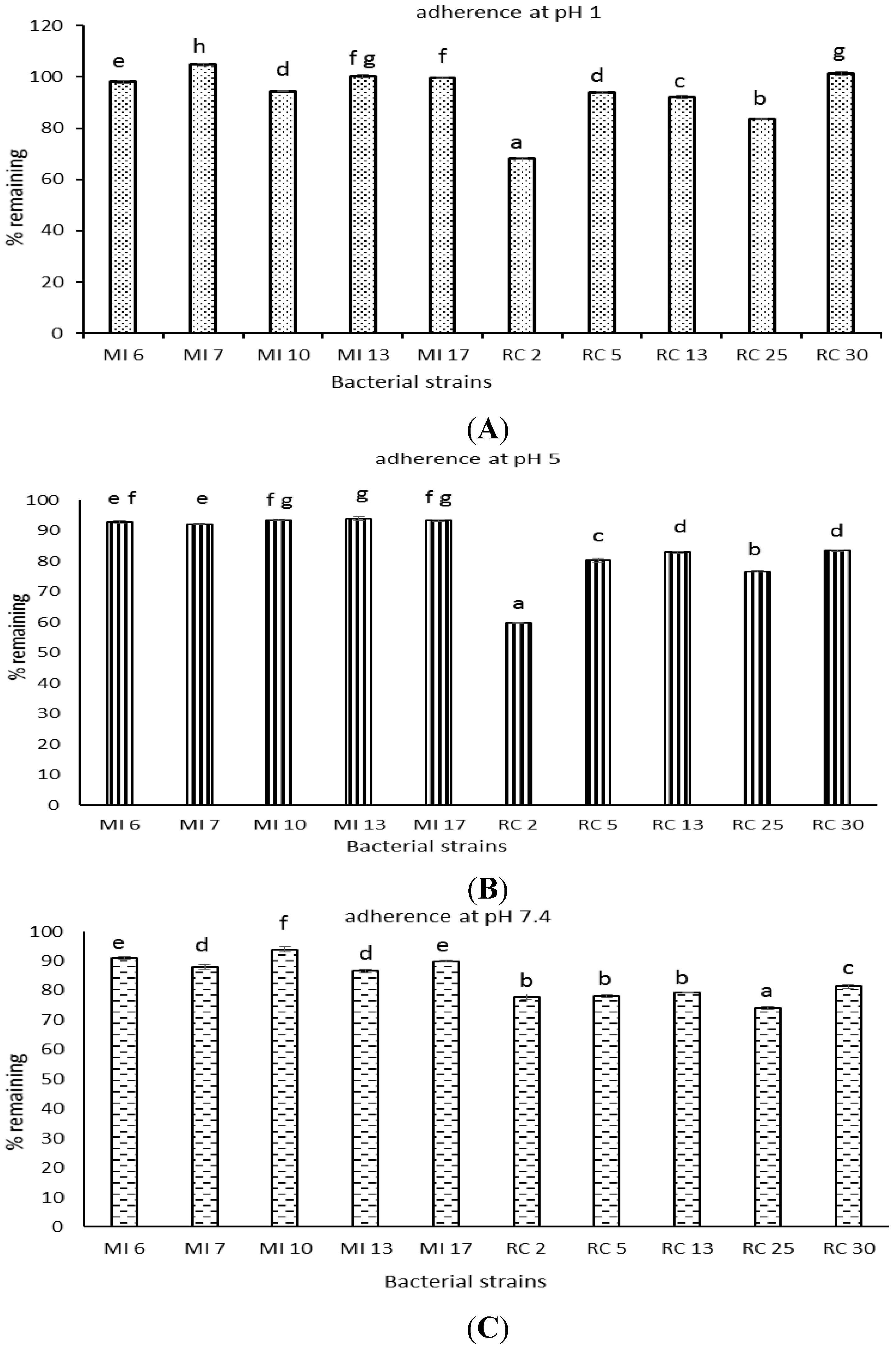
3.8. Bacterial Adherence to Hydrocarbons
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Probiotics Market—Global Industry Analysis, Market Size, Share, Trends, Analysis, Growth and Forecast. 2012—2018. Available online: http://www.transparencymarketresearch.com/probioticsmarket.html2013 (accessed on 24 October 2014).
- Felix, G.E.; Dellaglio, F. Taxonomy of lactobacilli and bifidobacteria. Curr. Issues Intest. Microbiol. 2007, 8, 44–61. [Google Scholar] [PubMed]
- Isolauri, E. The role of probiotics in paediatrics. Curr. Pediatr. 2004, 14, 104–109. [Google Scholar] [CrossRef]
- Czerucka, D.; Piche, T.; Rampal, P. Yeast as probiotics-Saccharomyces boulardii. Aliment. Pharmacol. Ther. 2007, 26, 767–778. [Google Scholar] [CrossRef] [PubMed]
- Gupta, V.; Garg, R. Probiotics. Indian J. Med. Microbiol. 2009, 27, 202–209. [Google Scholar] [CrossRef] [PubMed]
- Holzapfel, W.H.; Haberer, P.; Geisen, R.; Bjorkroth, J.; Schillinger, U. Taxonomy and important features of probiotic microorganisms in food and nutrition. Am. J. Clin. Nutr. 2001, 73, 365S–373S. [Google Scholar] [PubMed]
- Tambekar, D.H.; Bhutada, S.A. An evaluation of probiotic potential of Lactobacillus sp. from milk of domestic animals and commercial available probiotic preparations in prevention of enteric bacterial infections. Recent Res. Sci. Technol. 2010, 2, 82–88. [Google Scholar]
- Parvez, S.; Malik, K.A.; Ah Kang, S.; Kim, H.Y. Probiotics and their fermented food products are beneficial for health. J. Appl. Microbiol. 2006, 100, 1171–1185. [Google Scholar] [CrossRef] [PubMed]
- Mombelli, B.; Gismondo, M.R. The use of probiotics in medical practice. Int. J. Antimicrob. Agents 2000, 16, 531–536. [Google Scholar] [CrossRef] [PubMed]
- Kailasapathy, K.; Chin, J. Survival and therapeutic potential of probiotic organisms with reference to Lactobacillus acidophilus and Bifidobacterium spp. Immunol. Cell Biol. 2000, 78, 80–88. [Google Scholar] [CrossRef] [PubMed]
- Kechagia, M.; Basoulis, D.; Konstantopoulou, S.; Dimitriadi, D.; Gyftopoulou, K.; Skarmoutsou, N.; Fakiri, E.M. Health benefits of probiotics: A review. ISRN Nutr. 2013, 2013. [Google Scholar] [CrossRef]
- Musikasang, H.; Tani, A.; H-kittikun, A. Probiotic potential of lactic acid bacteria isolated from chicken gastrointestinal digestive tract. World J. Microbiol. Biotechnol. 2009, 25, 1337–1345. [Google Scholar] [CrossRef]
- Shah, N.P. Probiotic bacteria: Selective enumeration and survival in dairy foods. J. Dairy Sci. 2000, 83, 894–907. [Google Scholar] [CrossRef] [PubMed]
- Chou, L.S.; Weimer, B. Isolation and characterization of acid and bile tolerant isolates from strains of Lactobacillus acidophilus. J. Dairy Sci. 1999, 82, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Ranadheera, C.S.; Evans, C.A.; Adams, M.C.; Baines, S.K. In vitro analysis of gastrointestinal tolerance and intestinal cell ashesion of probiotics in goat’s milk ice cream and yogurt. Food Res. Int. 2012, 49, 619–625. [Google Scholar] [CrossRef]
- Jose, N.M.; Bunt, C.R.; Hussain, M.A. Implications of antibiotic resistance in probiotics. Food Rev. Int. 2015, 31, 52–62. [Google Scholar] [CrossRef]
- Alvarez-Olmos, M.I.; Oberhelman, R.A. Probiotic agents and infectious diseases: A modern perspective on a traditional therapy. Clin. Infect. Dis. 2001, 32, 1567–1576. [Google Scholar] [CrossRef] [PubMed]
- Gueimonde, M.; Salminen, S. New methods for selecting and evaluating probiotics. Dig. Liver Dis. 2006, 38, S242–S247. [Google Scholar] [CrossRef] [PubMed]
- Suvarna, V.C.; Boby, V.U. Probiotics in human health: A current assessment. Curr. Sci. 2005, 88, 1744–1748. [Google Scholar]
- Bao, Y.; Zhang, Y.; Zhang, Y.; Liu, Y.; Wang, S.; Dong, X.; Wang, Y.; Zhang, H. Screening of potential probiotic properties of Lactobacillus fermentum isolated from traditional dairy products. Food Control 2010, 21, 695–701. [Google Scholar] [CrossRef]
- Guo, X.H.; Kim, J.M.; Nam, H.M.; Park, S.Y.; Kim, J.M. Screening lactic acid bacteria from swine origins for multistrain probiotics based on in vitro functional properties. Anaerobe 2010, 16, 321–326. [Google Scholar] [CrossRef] [PubMed]
- Kirtzalidou, E.; Pramateftaki, P.; Kotsou, M.; Kyriacou, A. Screening for lactobacilli with probiotic properties in the infant gut microbiota. Anaerobe 2011, 17, 440–443. [Google Scholar] [CrossRef] [PubMed]
- Zago, M.; Fornasari, M.E.; Carminati, D.; Burns, P.; Suarez, V.; Vinderola, G.; Reinheimer, J.; Giraffa, G. Characterization and probiotic potential of Lactobacillus plantarum strains isolated from cheeses. Food Microbiol. 2011, 28, 1033–1040. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.K.; Mital, B.K.; Garg, S.K. Characterization of Lactobacillus acidophilus strains for use as dietary adjunct. Int. J. Food Microbiol. 1996, 29, 105–109. [Google Scholar] [CrossRef] [PubMed]
- Maragkoudakis, P.A.; Zoumpopoulou, G.; Miaris, C.; Kalantzopoulos, G.; Pot, B.; Tsakalidou, E. Probiotic potential of Lactobacillus strains isolated from dairy products. Int. Dairy J. 2006, 16, 189–199. [Google Scholar] [CrossRef]
- Thirabunyanon, M.; Boonprasom, P.; Niamsup, P. Probiotic potential of lactic acid bacteria isolated from fermented dairy milks on antiproliferation of colon cancer cells. Biotechnol. Lett. 2009, 31, 571–576. [Google Scholar] [CrossRef] [PubMed]
- Vinderola, G.; Capellini, B.; Villarreal, F.; Suarez, V.; Quiberoni, A.; Reinheimer, J. Usefulness of a simple set of in vitro tests for the screening and identification of probiotic candidate strains for dairy use. LWT—Food Sci. Technol. 2008, 41, 1678–1688. [Google Scholar] [CrossRef]
- Abe, F.; Ishibashi, N.; Shimamura, S. Effect of administration of bifidobacteria and lactic acid bacteria to new born calves and piglets. J. Dairy Sci. 1995, 78, 2838–2846. [Google Scholar] [CrossRef] [PubMed]
- Otero, M.C.; Morelli, L.; Nader-Macias, M.E. Probiotic properties of vaginal lactic acid bacteria to prevent metritis in cattle. Lett. Appl. Microbiol. 2006, 43, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.K.; Salminen, S. The coming of age of probiotics. Trends Food Sci. Technol. 1995, 6, 241–245. [Google Scholar] [CrossRef]
- Liu, X.; Liu, W.; Zhang, Q.; Tian, F.; Wang, G.; Zhang, H.; Chen, W. Screening of lactobacilli with antagonistic activity against enteroinvasive Escherichia coli. Food Control 2013, 30, 563–568. [Google Scholar] [CrossRef]
- Mishra, V.; Prasad, D.N. Application of in vitro methods for selection of Lactobacillus casei strains as potential probiotics. Int. J. Food Microbiol. 2005, 103, 109–115. [Google Scholar] [CrossRef] [PubMed]
- Hoque, M.Z.; Akter, F.; Hossain, K.M.; Rahman, M.S.M.; Billah, M.M.; Islam, K.M.D. Isolation, identification and analysis of probiotic properties of Lactobacillus spp. From selective regional yoghurts. World J. Dairy Food Sci. 2010, 5, 39–46. [Google Scholar]
- Santini, C.; Baffoni, L.; Gaggia, F.; Granata, M.; Gasbarri, R.; di Gioia, D.; Biavati, B. Characterization of probiotic strains: An application as feed additives in poultry against Campylobacter jejuni. Int. J. Food Microbiol. 2010, 141, S98–S108. [Google Scholar] [CrossRef] [PubMed]
- DʼAimmo, M.R.; Modesto, M.; Biavati, B. Antibiotic resistance of lactic acid bacteria and Bifidobacterium spp. isolated from dairy and pharmaceutical products. Int. J. Food Microbiol. 2007, 115, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Fukao, M.; Yajima, N. Assessment of antibiotic resistance in probiotic lactobacilli. In Antibiotic Resistant Bacteria—A Continuous Challenge in the New Millennium; InTech: Rijeka, Croatia, 2012; pp. 503–512. [Google Scholar]
- Ammor, M.S.; Florez, A.B.; Mayo, B. Antibiotic resistance in non-enterococcal lactic acid bacteria and bifidobacteria. Food Microbiol. 2007, 24, 559–570. [Google Scholar] [CrossRef] [PubMed]
- Gueimonde, M.; Sanchez, B.; de los Reyes-Gavilan, C.G.; Margolles, A. Antibiotic resistance in probiotic bacteria. Front. Microbiol. 2013, 202, 1–6. [Google Scholar]
- Resta-Lenert, S.; Barrett, K.E. Live probiotics protect intestinal epithelial cells from the effects of infection with enteroinvasive Escherichia coli. Gut 2003, 52, 988–997. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jose, N.M.; Bunt, C.R.; Hussain, M.A. Comparison of Microbiological and Probiotic Characteristics of Lactobacilli Isolates from Dairy Food Products and Animal Rumen Contents. Microorganisms 2015, 3, 198-212. https://doi.org/10.3390/microorganisms3020198
Jose NM, Bunt CR, Hussain MA. Comparison of Microbiological and Probiotic Characteristics of Lactobacilli Isolates from Dairy Food Products and Animal Rumen Contents. Microorganisms. 2015; 3(2):198-212. https://doi.org/10.3390/microorganisms3020198
Chicago/Turabian StyleJose, Neethu Maria, Craig R. Bunt, and Malik Altaf Hussain. 2015. "Comparison of Microbiological and Probiotic Characteristics of Lactobacilli Isolates from Dairy Food Products and Animal Rumen Contents" Microorganisms 3, no. 2: 198-212. https://doi.org/10.3390/microorganisms3020198
APA StyleJose, N. M., Bunt, C. R., & Hussain, M. A. (2015). Comparison of Microbiological and Probiotic Characteristics of Lactobacilli Isolates from Dairy Food Products and Animal Rumen Contents. Microorganisms, 3(2), 198-212. https://doi.org/10.3390/microorganisms3020198







