Functional Profiling of Kiwifruit Phyllosphere Bacteria: Copper Resistance and Biocontrol Potential as a Foundation for Microbiome-Informed Strategies
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling and Processing
2.2. Bacterial Isolation and DNA Extraction
2.3. Fingerprinting Analysis
2.4. Phylogenetic Analysis
2.5. Bacterial Plant Growth-Promoting Potential
2.6. Antagonistic Activity Against Pseudomonas syringae pv. actinidiae
2.7. Copper Tolerance
3. Results
3.1. Bacterial Community Structure Based on Culturable Isolates
3.2. Phylogenetic Insights into the Culturable Bacteriome
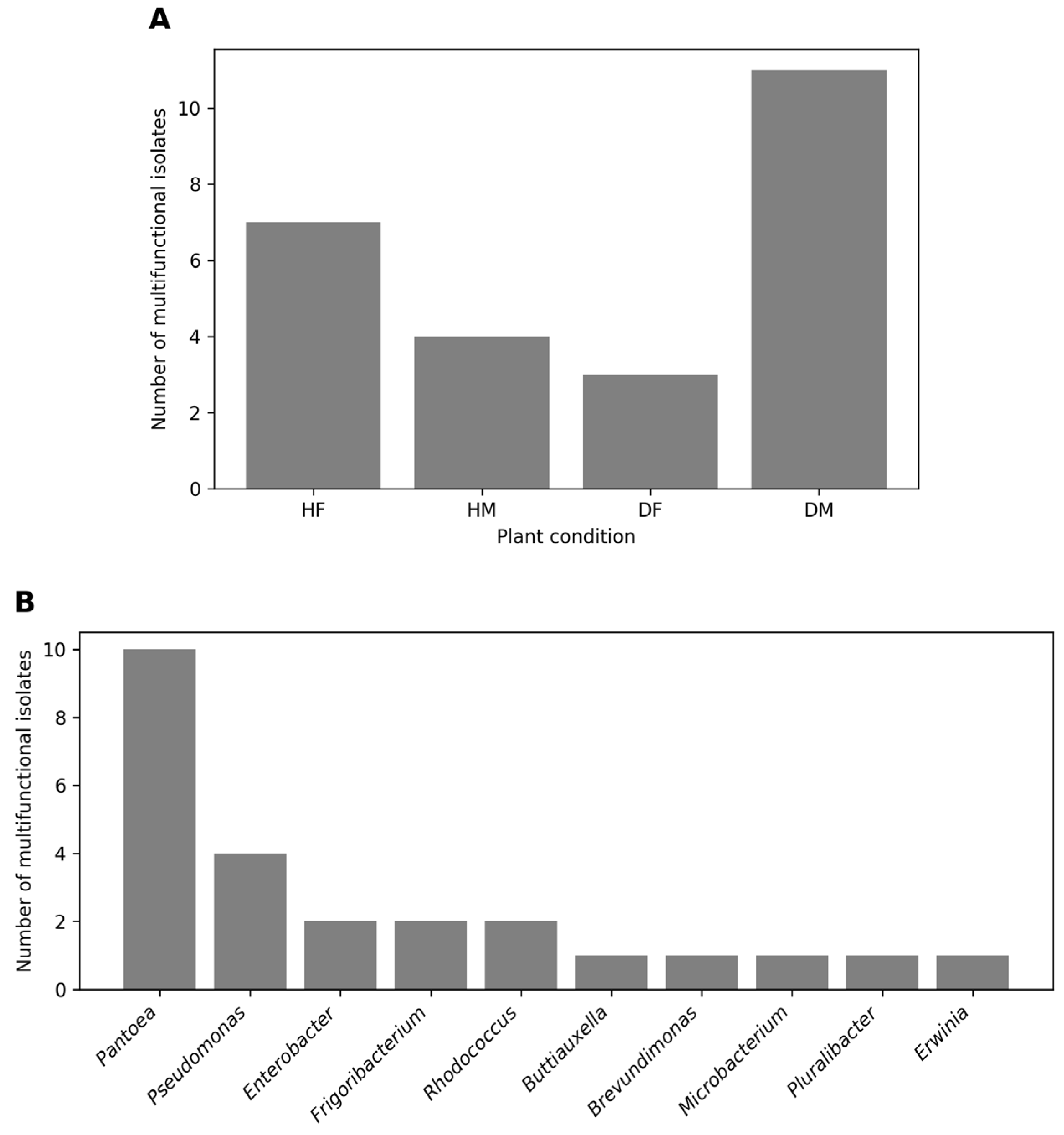
3.3. Distribution of Multifunctional Plant Growth Promoting Bacteria and Copper Tolerance
3.4. Antagonistic Activity Against Pseudomonas syringae pv. actinidiae and the Potential Application of Bacillus pumilus in Kiwifruit Disease Management
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ABM2 | Alkaliphilic buffered medium 2 |
| AIA | Average inhibition area |
| CAS | Chrome azurol S |
| CFU | Colony-forming unit |
| DF | Diseased female plants |
| DM | Diseased male plants |
| EPPO | European and Mediterranean Plant Protection Organization |
| HF | Healthy female plants |
| HM | Healthy male plants |
| IAA | Indole-3-acetic acid |
| ICE | Integrative conjugative element |
| MIC | Minimum inhibitory concentration |
| NJ | Neighbor-joining |
| PGP | Plant growth-promoting |
| PGPB | Plant growth-promoting bacteria |
| Psa | Pseudomonas syringae pv. actinidiae |
| R2A | Reasoner’s 2A agar |
| RAPD | Random amplified polymorphic DNA |
| SynCom | Synthetic microbial community |
| UNECE | United Nations Economic Commission for Europe |
References
- Antunes, M.D. Kiwi: Da Produção à Comercialização; Ciencias da Terra; Universidade do Algarve: Algarve, Portugal, 2008; 211p. [Google Scholar]
- FAOSTAT. Food and Agriculture Organization of the United Nations Statistics Division. 2017. Available online: https://www.fao.org/faostat/en/#data (accessed on 27 November 2025).
- United Nations Economic Commission for Europe. ECE/CTCS/WP.7/GE.1/2024/INF.02E: Economic and Social Council—Working Party on Agricultural Quality Standards—Specialized Section on Standardization of Fresh Fruit and Vegetables; UNCE: Geneva, Switzerland, 2024; Available online: https://unece.org/sites/default/files/2024-05/ECE_CTCS_WP.7_GE.1_2024_INF-02E.pdf (accessed on 15 October 2025).
- DGAV (Direção-Geral de Alimentação e Veterinária). Plano Nacional de Controlo do Cancro Bacteriano do Kiwi. 2015. Available online: https://www.dgav.pt/wp-content/uploads/2021/06/PNCPI-2015-17-final.pdf (accessed on 15 October 2025).
- Schroeder, C.A.; Fletcher, W.A. The Chinese Gooseberry (Actinidia chinensis) in New Zealand. Econ. Bot. 1967, 21, 81–92. [Google Scholar] [CrossRef]
- Vanneste, J.L. The scientific, economic, and social impacts of the New Zealand outbreak of bacterial canker of kiwifruit (Pseudomonas syringae pv. actinidiae). Annu. Rev. Phytopathol. 2017, 55, 377–399. [Google Scholar] [CrossRef] [PubMed]
- Balestra, G.M.; Renzi, M.; Mazzaglia, A. First report of bacterial canker of Actinidia deliciosa caused by Pseudomonas syringae pv. actinidiae in Portugal. New Dis. Rep. 2010, 22, 10. [Google Scholar] [CrossRef]
- DGAV (Direção-Geral de Alimentação e Veterinária). Comunicado sobre a bactéria Pseudomonas syringae pv. Actinidiae. 2010. Available online: https://www.dgav.pt/wp-content/uploads/2022/03/PSA_medidas_Kiwi_2022.pdf (accessed on 15 October 2025).
- Figueira, D.; Garcia, E.; Ares, A.; Tiago, I.; Veríssimo, A.; Costa, J. Genetic diversity of Pseudomonas syringae pv. actinidiae: Seasonal and spatial population dynamics. Microorganisms 2020, 8, 931. [Google Scholar] [CrossRef]
- Donati, I.; Buriani, G.; Cellini, A.; Mauri, S.; Costa, G.; Spinelli, F. New insights on the bacterial canker of kiwifruit (Pseudomonas syringae pv. actinidiae). J. Berry Res. 2014, 4, 53–67. [Google Scholar] [CrossRef]
- European Commission. Directive 2009/128/EC Establishing a Framework for Community Action to Achieve the Sustainable Use of Pesticides; Publication Office of EU: Luxembourg, 2009; Available online: https://eur-lex.europa.eu/legal-content/EN/ALL/?uri=celex%3A32009L0128 (accessed on 15 October 2025).
- Marcelletti, S.; Ferrante, P.; Petriccione, M.; Firrao, G.; Scortichini, M. Pseudomonas syringae pv. actinidiae draft genomes comparison reveal strain-specific features involved in adaptation and virulence to Actinidia species. PLoS ONE 2011, 6, e27297. [Google Scholar] [CrossRef]
- Thomidis, T.; Pagoulatou, M.G.; Alexandridis, E.; Mpalantinaki, E.; Goumas, D.E. Copper resistance in Pseudomonas syringae pv. actinidiae strains from the prefecture of Pella, Northern Greece, and a comparison of the effectiveness of several commercial products to control. Agriculture 2025, 15, 157. [Google Scholar] [CrossRef]
- Colombi, E.; Straub, C.; Künzel, S.; Templeton, M.D.; McCann, H.C.; Rainey, P.B. Evolution of copper resistance in the kiwifruit pathogen Pseudomonas syringae pv. actinidiae through acquisition of integrative conjugative elements and plasmids. Environ. Microbiol. 2017, 19, 819–832. [Google Scholar] [CrossRef]
- Michelotti, V.; Lamontanara, A.; Buriani, G.; Orrù, L.; Cellini, A.; Donati, I.; Vanneste, J.L.; Cattivelli, L.; Tacconi, G.; Spinelli, F. Comparative transcriptome analysis of the interaction between Actinidia chinensis var. chinensis and Pseudomonas syringae pv. actinidiae in absence and presence of acibenzolar-S-methyl. BMC Genom. 2018, 19, 585–607. [Google Scholar] [CrossRef]
- de Jong, H.; Reglinski, T.; Elmer, P.A.G.; Wurms, K.; Vanneste, J.L.; Guo, L.F.; Alavi, M. Integrated use of Aureobasidium pullulans strain CG163 and acibenzolar-S-methyl for management of bacterial canker in kiwifruit. Plants 2019, 15, 287. [Google Scholar] [CrossRef]
- Reglinski, T.; Wurms, K.V.; Vanneste, J.L.; Ah Chee, A.; Elmer, P.A.G. Transient changes in defence gene expression and phytohormone content induced by acibenzolar-S-methyl in glasshouse and orchard grown kiwifruit. Front. Agron. 2022, 3, 831172. [Google Scholar] [CrossRef]
- Reglinski, T.; Anderson, C.R.; Vanneste, J.L.; Wurms, K.V.; Ah Chee, A.; Elmer, P.A.G. Postharvest application of acibenzolar-S-methyl activates salicylic acid pathway genes in kiwifruit vines. Plants 2023, 12, 833. [Google Scholar] [CrossRef] [PubMed]
- Reglinski, T.; Wurms, K.V.; Vanneste, J.L.; Ah Chee, A.; Schipper, M.; Cornish, D.; Yu, J.; McAlinden, J.; Hedderley, D. Kiwifruit resistance to Sclerotinia sclerotiorum and Pseudomonas syringae pv. actinidiae and defence induction by acibenzolar S methyl and methyl jasmonate are cultivar dependent. Int. J. Mol. Sci. 2023, 24, 15952. [Google Scholar] [CrossRef] [PubMed]
- European Commission. The European Green Deal. COM 640 Final; Publication Office of EU: Luxembourg, 2019; Available online: https://ec.europa.eu/info/publications/communication-european-green-deal_en (accessed on 15 October 2025).
- European Commission. Farm to Fork Strategy: For a Fair, Healthy and Environmentally Friendly Food System; Publication Office of EU: Luxembourg, 2020; Available online: https://ec.europa.eu/food/farm2fork_en (accessed on 15 October 2025).
- Berg, G.; Rybakova, D.; Grube, M.; Köberl, M. The plant microbiome explored: Implications for experimental botany. J. Exp. Bot. 2016, 67, 995–1002. [Google Scholar] [CrossRef] [PubMed]
- Berg, G.; Rybakova, D.; Fischer, D.; Cernava, T.; Champomier Vergès, M.-C.; Charles, T.; Chen, X.; Cocolin, L.; Eversole, K.; Herrero Corral, G.; et al. Microbiome definition re-visited: Old concepts and new challenges. Microbiome 2020, 8, 103, Correction in Microbiome 2020, 8, 119. [Google Scholar] [CrossRef]
- Kostic, T.; Schloter, M.; Arruda, P.; Berg, G.; Charles, T.C.; Cotter, P.D.; Kiran, G.S.; Lange, L.; Maguin, E.; Meisner, A.; et al. Concepts and criteria defining emerging microbiome applications. Microb. Biotechnol. 2024, 17, e14550. [Google Scholar] [CrossRef]
- Purahong, W.; Orru, L.; Donati, I.; Perpetuini, G.; Cellini, A.; Lamontanara, A.; Michelotti, V.; Tacconi, G.; Spinelli, F. Plant microbiome and its link to plant health: Host species, organs and Pseudomonas syringae pv. actinidiae infection shaping bacterial phyllosphere communities of kiwifruit plants. Front. Plant Sci. 2018, 9, 1563. [Google Scholar] [CrossRef]
- Fu, M.; Chen, Y.; Liu, Y.-X.; Chang, X.; Zhang, L.; Yang, X.; Li, L.; Zhang, L. Genotype-associated core bacteria enhance host resistance against kiwifruit bacterial canker. Hortic. Res. 2024, 11, uhae236. [Google Scholar] [CrossRef]
- Correia, C.; Cellini, A.; Donati, I.; Voulgaris, P.; Obafemi, A.E.; Soriato, E.; Vandelle, E.; Santos, C.; Spinelli, F. Pseudomonas azotoformans and Pseudomonas putida: Novel kiwifruit-native biological control agents against Pseudomonas syringae pv. actinidiae. Biocontrol 2025, 105, 105706. [Google Scholar] [CrossRef]
- Patterson, H.R.; Hemara, L.M.; Templeton, M.D.; Jayaraman, J. Diverse Pseudomonas species engage in beneficial and suppressive interactions with the kiwifruit pathogen Psa across Actinidia germplasm. bioRxiv 2025. bioRxiv:2025.01.31.635981. [Google Scholar] [CrossRef]
- Vorholt, J.A. Microbial life in the phyllosphere. Nat. Rev. Microbiol. 2012, 10, 828–840. [Google Scholar] [CrossRef] [PubMed]
- Vandenkoornhuyse, P.; Quaiser, A.; Duhamel, M.; Le Van, A.; Dufresne, A. The importance of the microbiome of the plant holobiont. New Phytol. 2015, 206, 1196–1206. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Nomura, K.; Wang, X.; Sohrabi, R.; Xu, J.; Yao, L.; Paasch, B.C.; Ma, L.; Kremer, J.; Cheng, Y.; et al. Plant genetic network for preventing dysbiosis in the phyllosphere. Nature 2020, 580, 653–657. [Google Scholar] [CrossRef] [PubMed]
- Arnault, G.; Mony, C.; Vandenkoornhuyse, P. Plant microbiota dysbiosis and the Anna Karenina Principle. Trends Plant Sci. 2023, 28, 18–30. [Google Scholar] [CrossRef]
- Ares, A.; Pereira, J.; Garcia, E.; Costa, J.; Tiago, I. The leaf bacterial microbiota of female and male kiwifruit plants in distinct seasons: Assessing the impact of Pseudomonas syringae pv. actinidiae. Phytobiomes J. 2021, 5, 275–287. [Google Scholar] [CrossRef]
- EPPO (European and Mediterranean Plant Protection Organization). Pseudomonas syringae pv. actinidiae. EPPO Bull. 2021, 51, 549–567. [Google Scholar] [CrossRef]
- Tiago, I.; Chung, A.P.; Veríssimo, A. Bacterial diversity in a nonsaline alkaline environment: Heterotrophic aerobic populations. App. Environ. Microb. 2004, 70, 7378–7387. [Google Scholar] [CrossRef]
- Reasoner, D.J.; Geldreich, E.E. A new medium for the enumeration and subculture of bacteria from potable water. App. Environ. Microbiol. 1985, 49, 1–7. [Google Scholar] [CrossRef]
- Costa, J.; Tiago, I.; da Costa, M.S.; Veríssimo, A. Presence and persistence of Legionella spp. in groundwater. Appl. Environ. Microbiol. 2005, 71, 663–671. [Google Scholar] [CrossRef]
- Martins, J.; Ares, A.; Casais, V.; Costa, J.; Canhoto, J. Identification and characterization of Arbutus unedo L. endophytic bacteria isolated from wild and cultivated trees for the biological control of Phytophthora cinnamomi. Plants 2021, 10, 1569. [Google Scholar] [CrossRef]
- Turner, S.; Pryer, K.M.; Miao, V.P.; Palmer, J.D. Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis. J. Eukaryot. Microbiol. 1999, 46, 327–338. [Google Scholar] [CrossRef]
- Okonechnikov, K.; Golosova, O.; Fursov, M.; The UGENE team. Unipro UGENE: A unified bioinformatics toolkit. Bioinformatics 2012, 28, 1166–1167. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Niu, B.; Gao, Y.; Fu, L.; Li, W. CD-HIT Suite: A web server for clustering and comparing biological sequences. Bioinformatics 2010, 26, 680–682. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, W.; Strunk, O.; Westram, R.; Richter, L.; Meier, H.; Yadhukumar; Buchner, A.; Lai, T.; Steppi, S.; Jobb, G.; et al. ARB: A software environment for sequence data. Nucleic Acids Res. 2004, 32, 1363–1371. [Google Scholar] [CrossRef] [PubMed]
- Almoneafy, A.A.; Xie, G.L.; Tian, W.X.; Xu, L.H.; Zhang, G.Q.; Ibrahim, M. Characterization and evaluation of Bacillus isolates for their potential plant growth and biocontrol activities against tomato bacterial wilt. Afr. J. Biotechnol. 2012, 11, 7193–7201. [Google Scholar]
- Singh, D.; Kumar Yadav, D. Potential of Bacillus amyloliquefaciens for biocontrol of bacterial wilt of tomato incited by Ralstonia solanacearum. J. Plant Pathol. Microbiol. 2016, 7, 1000327. [Google Scholar] [CrossRef]
- Lertcanawanichakul, M.; Sawangnop, S. A comparison of two methods used for measuring the antagonistic activity of Bacillus species. Walailak J. Sci. Technol. 2008, 5, 161–171. [Google Scholar]
- Tontou, R.; Giovanardi, D.; Ferrari, M.; Stefani, E. Isolation of bacterial endophytes from Actinidia chinensis and preliminary studies on their possible use as antagonists against Pseudomonas syringae pv. actinidiae. J. Berry Res. 2016, 6, 395–406. [Google Scholar] [CrossRef]
- Beltrán, M.F.; Osorio, V.; Lemus, G.; Millas, P.; France, A.; Correa, F.; Sagredo, B. High incidence of copper resistance in Pseudomonas and Pantoea strains from stone fruit phyllosphere in central Chile. Chil. J. Agric. Res. 2021, 81, 378–389. [Google Scholar] [CrossRef]
- Zaneveld, J.R.; McMinds, R.; Vega Thurber, R.L. Stress and stability: Applying the Anna Karenina principle to animal microbiomes. Nat. Microbiol. 2017, 2, 17121. [Google Scholar] [CrossRef]
- Wicaksono, W.A.; Jones, E.E.; Casonato, S.; Monk, J.; Ridgway, H.J. Biological control of Pseudomonas syringae pv. actinidiae (Psa), the causal agent of bacterial canker of kiwifruit, using endophytic bacteria recovered from a medicinal plant. Biol. Control 2018, 116, 103–112. [Google Scholar] [CrossRef]
- Wang, F.; Wei, Y.; Yan, T.; Wang, C.; Chao, Y.; Jia, M.; An, L.; Sheng, H. Sphingomonas sp. Hbc-6 alters physiological metabolism and recruits beneficial rhizosphere bacteria to improve plant growth and drought tolerance. Front. Plant Sci. 2022, 13, 1002772. [Google Scholar] [CrossRef] [PubMed]
- Mazoyon, C.; Firmin, S.; Bensaddek, L.; Pecourt, A.; Chabot, A.; Faucon, M.P.; Duclercq, J. Optimizing crop production with bacterial inputs: Insights into chemical dialogue between Sphingomonas sediminicola and Pisum sativum. Microorganisms 2023, 11, 1847. [Google Scholar] [CrossRef] [PubMed]
- Palberg, D.; Kisiała, A.; Jorge, G.L.; Emery, R.J.N.; Przybył, K. A survey of Methylobacterium species and strains reveals widespread production and varying profiles of cytokinin phytohormones. BMC Microbiol. 2022, 22, 49. [Google Scholar] [CrossRef]
- Berg, G.; Koskella, B. Nutrient- and dose-dependent microbiome-mediated protection against a plant pathogen. Curr. Biol. 2018, 28, 2487–2492.e3. [Google Scholar] [CrossRef]
- Grossi, C.E.M.; Fantino, E.; Serral, F.; Zawoznik, M.S.; Fernandez Do Porto, D.A.; Ulloa, R.M. Methylobacterium sp. 2A is a plant growth-promoting rhizobacteria that has the potential to improve potato crop yield under adverse conditions. Front. Plant Sci. 2020, 11, 71. [Google Scholar] [CrossRef]
- Hernández-Huerta, J.; Tamez-Guerra, P.; Gomez-Flores, R.; Delgado-Gardea, M.C.E.; García-Madrid, M.S.; Robles-Hernández, L.; Infante-Ramirez, R. Prevalence of Xanthomonas euvesicatoria (formally X. perforans) associated with bacterial spot severity in Capsicum annuum crops in South Central Chihuahua, Mexico. Peer J. 2021, 9, e10913. [Google Scholar] [CrossRef]
- Berg, J.; Tom-Petersen, A.; Nybroe, O. Copper amendment of agricultural soil selects for bacterial antibiotic resistance in the field. Lett. Appl. Microbiol. 2005, 40, 146–151. [Google Scholar] [CrossRef]
- Hu, H.W.; Wang, J.T.; Li, J.; Li, J.J.; Ma, Y.B.; Chen, D.; He, J.Z. Field-based evidence for copper contamination induced changes of antibiotic resistance in agricultural soils. Environ. Microbiol. 2016, 18, 3896–3909. [Google Scholar] [CrossRef]
- Kang, Y.; Lee, W.; Kim, S.; Jang, G.; Kim, B.G.; Yoon, Y. Enhancing the copper-sensing capability of Escherichia coli-based whole-cell bioreporters by genetic engineering. Appl. Microbiol. Biotechnol. 2018, 102, 1513–1521. [Google Scholar] [CrossRef]
- Lindow, S.E. Horizontal gene transfer gone wild: Promiscuity in a kiwifruit pathogen leads to resistance to chemical control. Environ. Microbiol. 2017, 19, 1363–1365. [Google Scholar] [CrossRef]
- Lorenzi, A.S.; Bonatelli, M.L.; Chia, M.A.; Peressim, L.; Quecine, M.C. Opposite sides of Pantoea agglomerans and its associated commercial outlook. Microorganisms 2022, 10, 2072. [Google Scholar] [CrossRef]
- Cornish, D.A.; Schipper, M.M.; Oldham, J.M.; Yu, J.; Vanneste, J.L. The biological control agent for bacterial canker of kiwifruit, in Aureo® Gold, is a strain of Aureobasidium pullulans identifiable by novel SCAR marker primers. Biol. Control 2025, 202, 105709. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Casais, V.; Pereira, J.; Garcia, E.; Coelho, C.; Figueira, D.; Ares, A.; Tiago, I.; Costa, J. Functional Profiling of Kiwifruit Phyllosphere Bacteria: Copper Resistance and Biocontrol Potential as a Foundation for Microbiome-Informed Strategies. Microorganisms 2026, 14, 321. https://doi.org/10.3390/microorganisms14020321
Casais V, Pereira J, Garcia E, Coelho C, Figueira D, Ares A, Tiago I, Costa J. Functional Profiling of Kiwifruit Phyllosphere Bacteria: Copper Resistance and Biocontrol Potential as a Foundation for Microbiome-Informed Strategies. Microorganisms. 2026; 14(2):321. https://doi.org/10.3390/microorganisms14020321
Chicago/Turabian StyleCasais, Vinicius, Joana Pereira, Eva Garcia, Catarina Coelho, Daniela Figueira, Aitana Ares, Igor Tiago, and Joana Costa. 2026. "Functional Profiling of Kiwifruit Phyllosphere Bacteria: Copper Resistance and Biocontrol Potential as a Foundation for Microbiome-Informed Strategies" Microorganisms 14, no. 2: 321. https://doi.org/10.3390/microorganisms14020321
APA StyleCasais, V., Pereira, J., Garcia, E., Coelho, C., Figueira, D., Ares, A., Tiago, I., & Costa, J. (2026). Functional Profiling of Kiwifruit Phyllosphere Bacteria: Copper Resistance and Biocontrol Potential as a Foundation for Microbiome-Informed Strategies. Microorganisms, 14(2), 321. https://doi.org/10.3390/microorganisms14020321





