Methods for Evaluating the Effects of 2D and 3D Culture Environment on Macrophage Response to Mycobacterium Infection
Abstract
1. Introduction
In Vitro Models for Characterizing Granuloma Formation
2. Materials and Methods
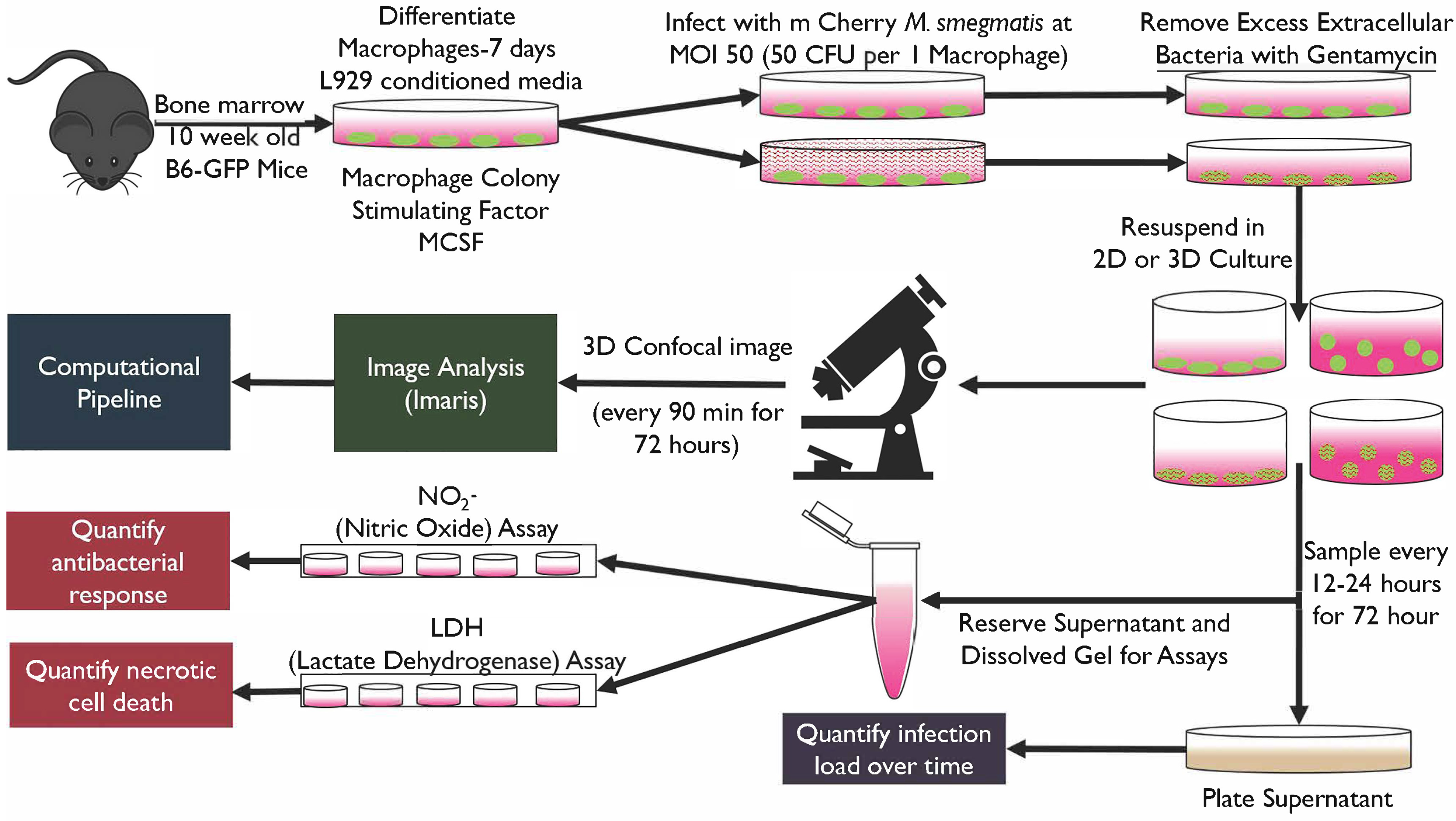
2.1. Developing and Characterizing System and Component Constraints for the Ex Vivo Infection Platform
2.1.1. Determining Optimal ECM Gel Height for 3D Cell Culture
2.1.2. mCherry M. smegmatis Fluorescence Homogeneity and Persistence in 2D/3D Culture
2.1.3. Impact of Macrophage EGFP and Bacterial mCherry on Infection Response
2.2. Development of 2D and 3D Ex Vivo Model of Mycobacteria Infection
2.2.1. Infection Studies in 2D/3D Environments
2.2.2. Quantifying Biochemical Response
2.3. Image-Based Quantification of Spatiotemporal Response
Image Processing and Analysis
2.4. Image and Data Analysis
2.4.1. Data Processing and Feature Analysis in MATLAB
2.4.2. Statistical Analysis
3. Results
3.1. Oxygen Diffusion in 3D Cell Culture Is Sufficient to Maintain Normoxia for Cell Populations
3.2. Fluorescent Strains Persist and Are Comparable to Wild Type in Growth and Infection Response
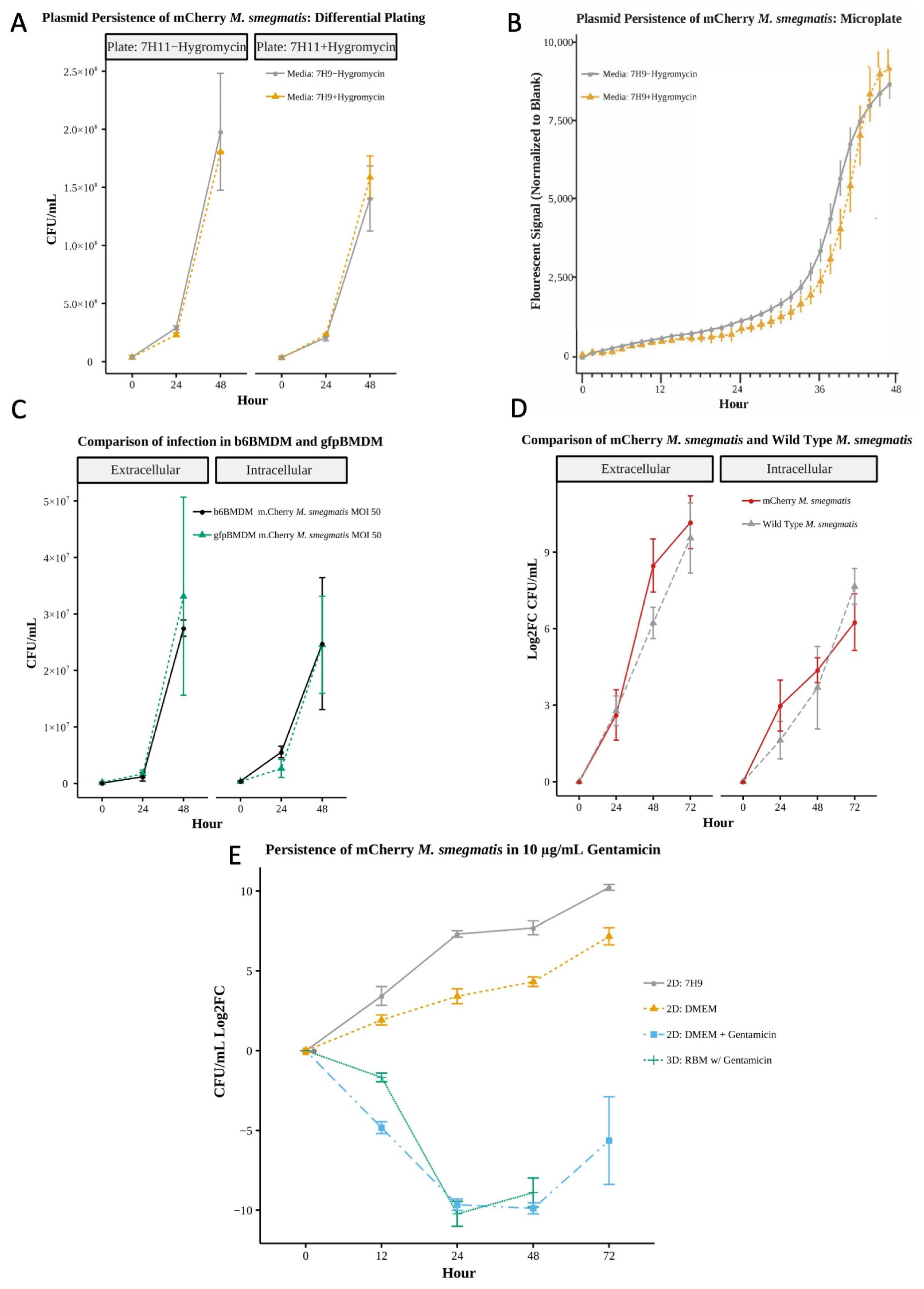
3.2.1. mCherry Growth and Fluorescence Maintained Absent Antibiotic Selection
3.2.2. Post-Infection Response Comparable for Fluorescent and Wild-Type Strains
3.2.3. Impact of Gentamicin on 3D Bacteria Culture Reproducible in 2D Culture
3.3. Extra-Matrix Sampling Sufficiently Captures Biochemical Response in 3D Infection in a Manner Comparable to Sample Collection in 2D Culture
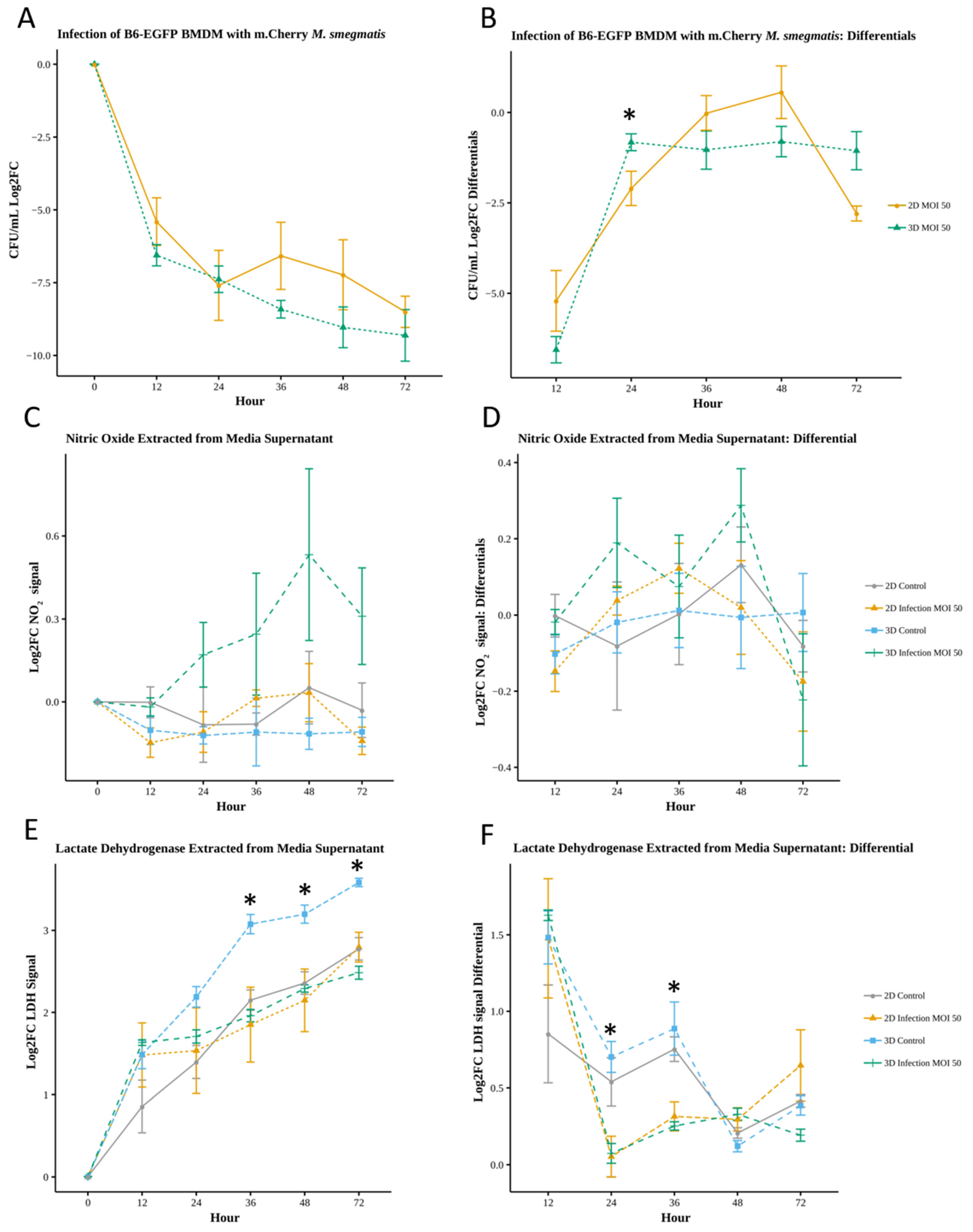
3.4. Macrophages in 3D Culture Exhibit Similar Infection Outcomes as in 2D Culture but Biochemical Response Dynamics Differ
3.4.1. Relative Increase in Oxidative Response During 3D Infection
3.4.2. Necrotic Cell Death Is Reduced for Infected Macrophages in 3D Culture
3.5. Imaging-Based Quantification Shows Differences in Macrophage 2D/3D Spatiotemporal Response
3.5.1. Imaging Shows Structural and Morphological Differences Related to Infection and 3D Environment
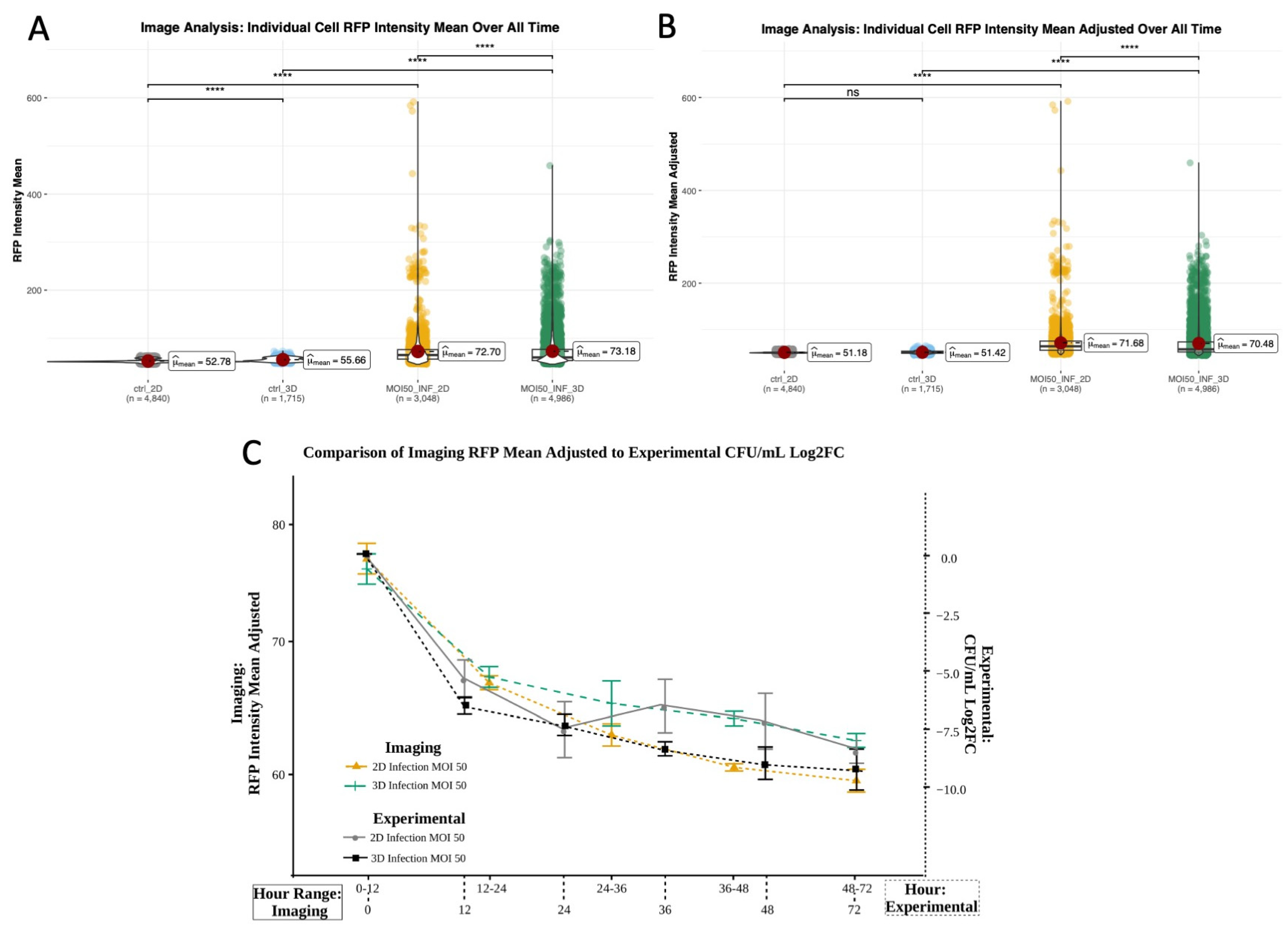
3.5.2. Image-Based Characterization of 2D and 3D Cytotoxicity and Bacterial Load Shows Concordance with Standard Downstream Assays
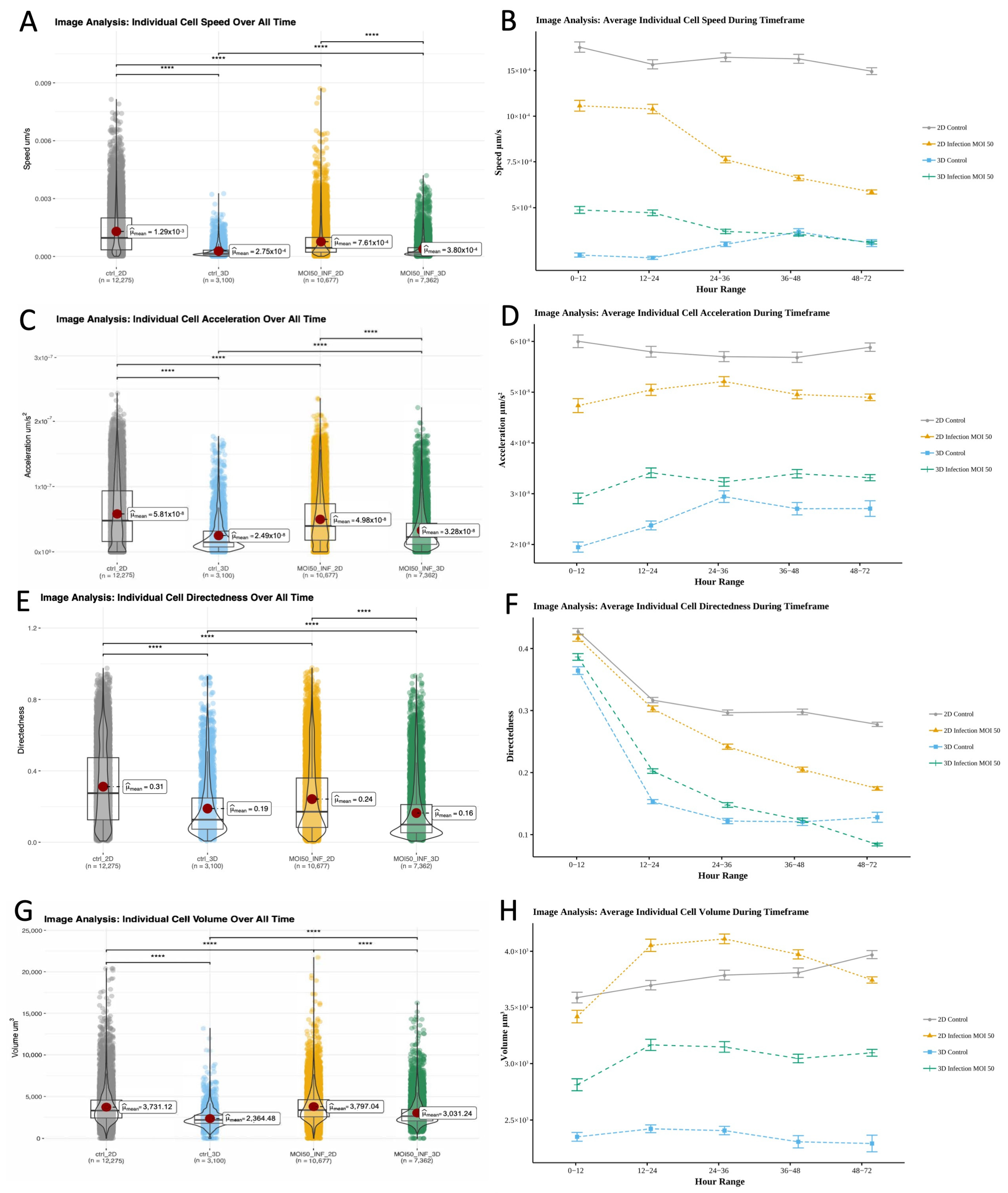
3.5.3. Environment Drives Differences in Cell Motility and Migration, While Volumetric Differences Driven by Infection State

3.6. Correlative Characteristics Vary Depending on Infection and Environment
3.6.1. Relationship Between Cytotoxicity, Speed, and Bacteria Load Concordant in 2D and 3D Cultures, but NO Negatively Correlates with Speed and Mycobacterial Load in 3D
3.6.2. Macrophage Speed and Directedness Positively Correlate with Bacteria Load with Significance in 3D Infection Conditions
4. Discussion
4.1. Effectiveness of Biochemical Response During Infection More Pronounced in 3D Environment
4.2. 3D Spatiotemporal Response Associates with Infection and NO and Is Consistent with Prior Observations of Virulence and Proinflammatory Modulation of Cell Speed and Morphology
4.3. BMDMs in 3D Environment Exhibit Significantly Increased Volume During Infection
4.4. 3D Environment Results in a Comparatively Distinct Biochemical and Structural Immune Response Signature
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Davis, J.M.; Ramakrishnan, L. The Role of the Granuloma in Expansion and Dissemination of Early Tuberculous Infection. Cell 2009, 136, 37–49. [Google Scholar] [CrossRef]
- Ernst, J.D. The immunological life cycle of tuberculosis. Nat. Rev. Immunol. 2012, 12, 581–591. [Google Scholar] [CrossRef]
- Batista, L.A.F.; Silva, K.J.S.; da Costa, E.S.L.M.; de Moura, Y.F.; Zucchi, F.C.R. Tuberculosis: A granulomatous disease mediated by epigenetic factors. Tuberculosis 2020, 123, 101943. [Google Scholar] [CrossRef]
- Al Shammari, B.; Shiomi, T.; Tezera, L.; Bielecka, M.K.; Workman, V.; Sathyamoorthy, T.; Mauri, F.; Jayasinghe, S.N.; Robertson, B.D.; D’Armiento, J.; et al. The Extracellular Matrix Regulates Granuloma Necrosis in Tuberculosis. J. Infect. Dis. 2015, 212, 463–473. [Google Scholar] [CrossRef]
- Guirado, E.L.S.S.; Kaplan, G. Macrophages in Tuberculosis: Friend or Foe. Semin. Immunopathol. 2013, 35, 563–583. [Google Scholar] [CrossRef] [PubMed]
- Marakalala, M.J.; Raju, R.M.; Sharma, K.; Zhang, Y.J.; Eugenin, E.A.; Prideaux, B.; Daudelin, I.B.; Chen, P.Y.; Booty, M.G.; Kim, J.H. Inflammatory signaling in human Tuberculosis granulomas is spatially organized. Nat. Med. 2016, 22, 531–538. [Google Scholar] [CrossRef]
- Bold, T.D.; Ernst, J.D. Who Benefits from Granulomas, Mycobacteria or Host? Cell 2009, 136, 17–19. [Google Scholar] [CrossRef]
- Egen, J.G.; Rothfuchs, A.G.; Feng, C.G.; Winter, N.; Sher, A.; Germain, R.N. Macrophage and T Cell Dynamics during the Development and Disintegration of Mycobacterial Granulomas. Immunity 2008, 28, 271–284. [Google Scholar] [CrossRef]
- Flynn, J.L.; Gideon, H.P.; Mattila, J.T.; Lin, P.L. Immunology studies in non-human primate models of tuberculosis. Immunol. Rev. 2015, 264, 60–73. [Google Scholar] [CrossRef]
- Kramnik, I.; Beamer, G. Mouse models of human TB pathology: Roles in the analysis of necrosis and the development of host-directed therapies. Semin. Immunopathol. 2016, 38, 221–237. [Google Scholar] [CrossRef]
- Cutter, S.; Wright, M.D.; Reynolds, N.P.; Binger, K.J. Towards using 3D cellular cultures to model the activation and diverse functions of macrophages. Biochem. Soc. Trans. 2023, 51, 387–401. [Google Scholar] [CrossRef]
- Mukundan, S.; Bhatt, R.; Lucas, J.; Tereyek, M.; Chang, T.L.; Subbian, S.; Parekkadan, B. 3D host cell and pathogen-based bioassay development for testing anti-tuberculosis (TB) drug response and modeling immunodeficiency. Biomol. Concepts 2021, 12, 117–128. [Google Scholar] [CrossRef]
- Mukundan, S.; Singh, P.; Shah, A.; Kumar, R.; O’Neill, K.C.; Carter, C.L.; Russell, D.G.; Subbian, S.; Parekkadan, B. In Vitro Miniaturized Tuberculosis Spheroid Model. Biomedicines 2021, 9, 1209. [Google Scholar] [CrossRef]
- Puissegur, M.-P.; Botanch, C.; Duteyrat, J.-L.; Delsol, G.; Caratero, C.; Altare, F. An in vitro dual model of mycobacterial granulomas to investigate the molecular interactions between mycobacteria and human host cells. Cell. Microbiol. 2004, 6, 423–433. [Google Scholar] [CrossRef] [PubMed]
- Parasa, V.R.; Rahman, M.J.; Hoang, A.T.N.; Svensson, M.; Brighenti, S.; Lerm, M. Modeling Mycobacterium tuberculosis early granuloma formation in experimental human lung tissue. Dis. Model. Mech. 2014, 7, 281–288. [Google Scholar] [PubMed]
- Bielecka, M.K.; Tezera, L.B.; Zmijan, R.; Drobniewski, F.; Zhang, X.; Jayasinghe, S.; Elkington, P. A Bioengineered Three-Dimensional Cell Culture Platform Integrated with Microfluidics to Address Antimicrobial Resistance in Tuberculosis. mBio 2017, 8, e02073-16. [Google Scholar] [CrossRef] [PubMed]
- Berry, S.B.; Gower, M.S.; Su, X.; Seshadri, C.; Theberge, A.B. A Modular Microscale Granuloma Model for Immune-Microenvironment Signaling Studies in vitro. Front. Bioeng. Biotechnol. 2020, 8, 931. [Google Scholar] [CrossRef]
- Guirado, E.; Mbawuike, U.; Keiser, T.L.; Arcos, J.; Azad, A.K.; Wang, S.H.; Schlesinger, L.S. Characterization of host and microbial determinants in individuals with latent tuberculosis infection using a human granuloma model. mBio 2015, 6, e02537-14. [Google Scholar] [CrossRef]
- Kapoor, N.; Pawar, S.; Sirakova, T.D.; Deb, C.; Warren, W.L.; Kolattukudy, P.E. Human Granuloma In Vitro Model, for TB Dormancy and Resuscitation. PLoS ONE 2013, 8, e53657. [Google Scholar] [CrossRef]
- Griffith, L.G.; Swartz, M.A. Capturing complex 3D tissue physiology in vitro. Nat. Rev. Mol. Cell Biol. 2006, 7, 211–224. [Google Scholar] [CrossRef]
- Kim, J.; Koo, B.K.; Knoblich, J.A. Human organoids: Model systems for human biology and medicine. Nat Rev Mol Cell Biol 2020, 21, 571–584. [Google Scholar] [CrossRef]
- Kong, Y.; Yang, D.; Cirillo, S.L.; Li, S.; Akin, A.; Francis, K.P.; Maloney, T.; Cirillo, J.D. Application of fluorescent protein expressing strains to evaluation of anti-tuberculosis therapeutic efficacy in vitro and in vivo. PLoS ONE 2016, 11, e0149972. [Google Scholar] [CrossRef] [PubMed]
- Colom, A.; Galgoczy, R.; Almendros, I.; Xaubet, A.; Farré, R.; Alcaraz, J. Oxygen diffusion and consumption in extracellular matrix gels: Implications for designing three-dimensional cultures. J. Biomed. Mater. Res. A 2014, 102, 2776–2784. [Google Scholar] [CrossRef] [PubMed]
- Pavlou, S.; Lindsay, J.; Ingram, R.; Xu, H.; Chen, M. Sustained high glucose exposure sensitizes macrophage responses to cytokine stimuli but reduces their phagocytic activity. BMC Immunol. 2018, 19, 24. [Google Scholar] [CrossRef] [PubMed]
- Gough, M.E.; Graviss, E.A.; May, E.E. The dynamic immunomodulatory effects of vitamin D3 during Mycobacterium infection. Innate Immun. 2017, 23, 506–523. [Google Scholar] [CrossRef]
- Lagier, B.; Pelicic, V.; Lecossier, D.; Prod’hom, G.; Rauzier, J.; Guilhot, C.; Gicquel, B.; Hance, A.J. Identification of genetic loci implicated in the survival of Mycobacterium smegmatis in human mononuclear phagocytes. Mol. Microbiol. 1998, 29, 465–475. [Google Scholar] [CrossRef]
- Xu, C.; Inokuma, M.S.; Denham, J.; Golds, K.; Kundu, P.; Gold, J.D.; Carpenter, M.K. Feeder-free growth of undifferentiated human embryonic stem cells. Nat. Biotechnol. 2001, 19, 971–974. [Google Scholar] [CrossRef]
- Krombach, F.; Münzing, S.; Allmeling, A.-M.; Gerlach, J.T.; Behr, J.; Dörger, M. Cell Size of Alveolar Macrophages: An Interspecies Comparison. Environ. Health Perspect. 1997, 105, 1261–1263. [Google Scholar]
- Vijay, S.; Nair, R.R.; Sharan, D.; Jakkala, K.; Mukkayyan, N.; Swaminath, S.; Pradhan, A.; Joshi, N.V.; Ajitkumar, P. Mycobacterial Cultures Contain Cell Size and Density Specific Sub-populations of Cells with Significant Differential Susceptibility to Antibiotics, Oxidative and Nitrite Stress. Front. Microbiol. 2017, 8, 463. [Google Scholar] [CrossRef]
- Yamane, T. Elementary Sampling Theory; Prentice-Hall, Inc.: Englewood Cliffs, NJ, USA, 1967. [Google Scholar]
- Stolley, D.L.; May, E.E. Spatiotemporal Analysis of Mycobacterium-Dependent Macrophage Response. In Proceedings of the 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, USA, 18–21 July 2018; pp. 2390–2393. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- McWhorter, F.Y.; Wang, T.; Nguyen, P.; Chung, T.; Liu, W.F. Modulation of macrophage phenotype by cell shape. Proc. Natl. Acad. Sci. USA 2013, 110, 17253–17258. [Google Scholar] [CrossRef]
- Webb, S.E.; Pollard, J.W.; Jones, G.E. Direct observation and quantification of macrophage chemoattraction to the growth factor CSF-1. J. Cell Sci. 1996, 109, 793–803. [Google Scholar] [CrossRef]
- Cui, K.; Ardell, C.L.; Podolnikova, N.P.; Yakubenko, V.P. Distinct Migratory Properties of M1, M2, and Resident Macrophages Are Regulated by αDβ2 and αMβ2 Integrin-Mediated Adhesion. Front. Immunol. 2018, 9, 2650. [Google Scholar] [CrossRef]
- Rothe, G.; Valet, G. Phagocytosis, intracellular pH, and cell volume in the multifunctional analysis of granulocytes by flow cytometry. Cytometry 1988, 9, 316–324. [Google Scholar] [CrossRef]
- Gough, M.E.; Graviss, E.A.; Chen, T.A.; Obasi, E.M.; May, E.E. Compounding effect of vitamin D3 diet, supplementation, and alcohol exposure on macrophage response to mycobacterium infection. Tuberculosis 2019, 116S, S42–S58. [Google Scholar] [CrossRef] [PubMed]
- Marim, F.M.; Silveira, T.N.; Lima, D.S., Jr.; Zamboni, D.S. A Method for Generation of Bone Marrow-Derived Macrophages from Cryopreserved Mouse Bone Marrow Cells. PLoS ONE 2010, 5, e15263. [Google Scholar] [CrossRef] [PubMed]
- Loose, L.D. Characterization of Macrophage Dysfunction in Rodent Malaria. J. Leukoc. Biol. 1984, 36, 703–718. [Google Scholar] [CrossRef] [PubMed]
- Sershen, C.L.; Plimpton, S.J.; May, E.E. Oxygen Modulates the Effectiveness of Granuloma Mediated Host Response to Mycobacterium tuberculosis: A Multiscale Computational Biology Approach. Front. Cell. Infect. Microbiol. 2016, 6, 6. [Google Scholar] [CrossRef]

| Variable | Description and Values Used |
|---|---|
| (Maximum) gel thickness | |
| Thiele modulus for normoxic environment (19 in for half gel depth, 16.45 for full gel depth (estimate)) assuming an oxygen tension limit of 50 mmHg [23]; normoxic oxygen tension is 20 to 50 mm Hg in most tissues (excluding high oxygenated environments such as arteries and alveoli) | |
| Oxygen concentration at which OCR is half of Vmax; (0.45 mmHg = 0.58 × 10−3 mol m−3) [23] | |
| Oxygen diffusivity in the gel in the presence of cells (1.7 × 10−5 cm2/s, graphical estimation) [23] | |
| Initial cell seeding density (2.5 × 106 cells/mL) | |
| Maximum OCR per cell Maximum OCR for A549 in gel is ~3/10ths of 2D OCR (graphical estimation) [23] Maximum OCR for BMDMs in 2D culture is ~1.7 × 10−16 moles/(cell (second)) [24], which equates to an estimate of 5 × 10−17 moles/(cell (second)) for BMDMs in 3D RBM culture | |
| Time constant describing cell proliferation rate (54 h for A549 cells in gel) [23]; however, primary BMDMs do not proliferate well and attempts to extract cells from gel after 24 h produce fewer cells than seeded, so the exponential term will trend towards 1, changing the cell growth term from , an assumption of exponential growth, to an assumption of no cell growth | |
| t | Time in cell culture (hours) |
| Hour-Range | 2D Control | 3D Control | 2D Infection MOI 50 | 3D Infection MOI 50 |
|---|---|---|---|---|
| 0–12 | 1586 | 740 | 1205 | 959 |
| 12–24 | 2171 | 855 | 1779 | 1309 |
| 24–36 | 2502 | 718 | 2044 | 1341 |
| 36–48 | 2337 | 453 | 2107 | 1452 |
| 48–72 | 3679 | 334 | 3542 | 2301 |
| Total | 12,275 | 3100 | 10,677 | 7362 |
| Condition 1 | Condition 2 | Log2 Fold Change |
|---|---|---|
| 2D control | 3D control | 2.235 |
| 2D infection | 3D infection | 1.0015 |
| 2D control | 2D infection | 0.766 |
| 3D control | 3D infection | −0.467 |
| Condition 1 | Condition 2 | Log2 Fold Change |
|---|---|---|
| 2D control | 3D control | 1.223 |
| 2D infection | 3D infection | 0.601 |
| 2D control | 2D infection | 0.224 |
| 3D control | 3D infection | −0.389 |
| Condition 1 | Condition 2 | Log2 Fold Change |
|---|---|---|
| 2D control | 3D control | 0.723 |
| 2D infection | 3D infection | 0.564 |
| 2D control | 2D infection | 0.365 |
| 3D control | 3D infection | 0.206 |
| Condition 1 | Condition 2 | Log2 Fold Change |
|---|---|---|
| 2D control | 3D control | 0.658 |
| 2D infection | 3D infection | 0.325 |
| 2D control | 2D infection | −0.025 |
| 3D control | 3D infection | −0.357 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stolley, D.L.; Rasaputra, K.S.; May, E.E. Methods for Evaluating the Effects of 2D and 3D Culture Environment on Macrophage Response to Mycobacterium Infection. Microorganisms 2025, 13, 2026. https://doi.org/10.3390/microorganisms13092026
Stolley DL, Rasaputra KS, May EE. Methods for Evaluating the Effects of 2D and 3D Culture Environment on Macrophage Response to Mycobacterium Infection. Microorganisms. 2025; 13(9):2026. https://doi.org/10.3390/microorganisms13092026
Chicago/Turabian StyleStolley, Danielle L., Komal S. Rasaputra, and Elebeoba E. May. 2025. "Methods for Evaluating the Effects of 2D and 3D Culture Environment on Macrophage Response to Mycobacterium Infection" Microorganisms 13, no. 9: 2026. https://doi.org/10.3390/microorganisms13092026
APA StyleStolley, D. L., Rasaputra, K. S., & May, E. E. (2025). Methods for Evaluating the Effects of 2D and 3D Culture Environment on Macrophage Response to Mycobacterium Infection. Microorganisms, 13(9), 2026. https://doi.org/10.3390/microorganisms13092026





