GWAS Study Applied to Phenotypically Slow Growth Strains of Listeria monocytogenes Workflow and Outcome
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection and Preparation
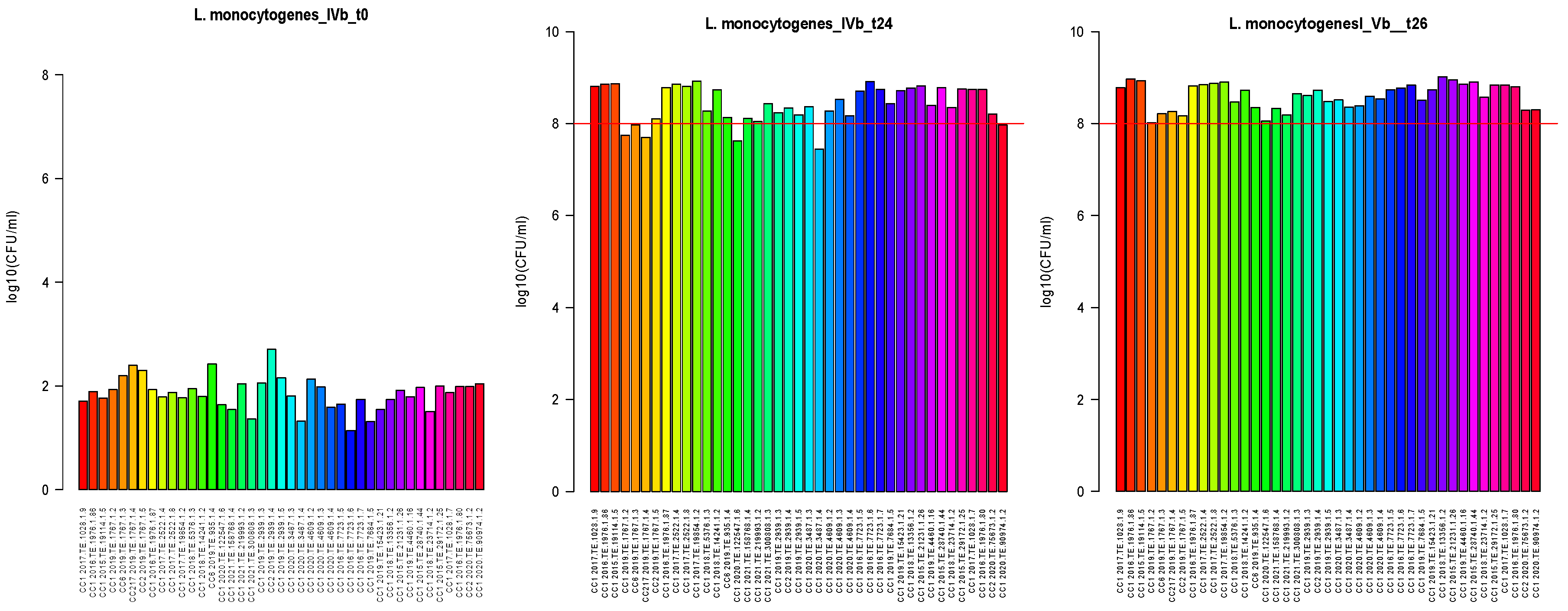
2.2. Growth Potential Evaluation
2.2.1. Stocks Preparation
2.2.2. Inoculum Preparation and Contamination of Half Fraser Broth
2.2.3. Enumeration and Growth Rate
2.2.4. Graphics Analysis
2.3. DNA Extraction, Sequencing and Bioinformatic Analysis
2.4. Genome Annotation and GWAS
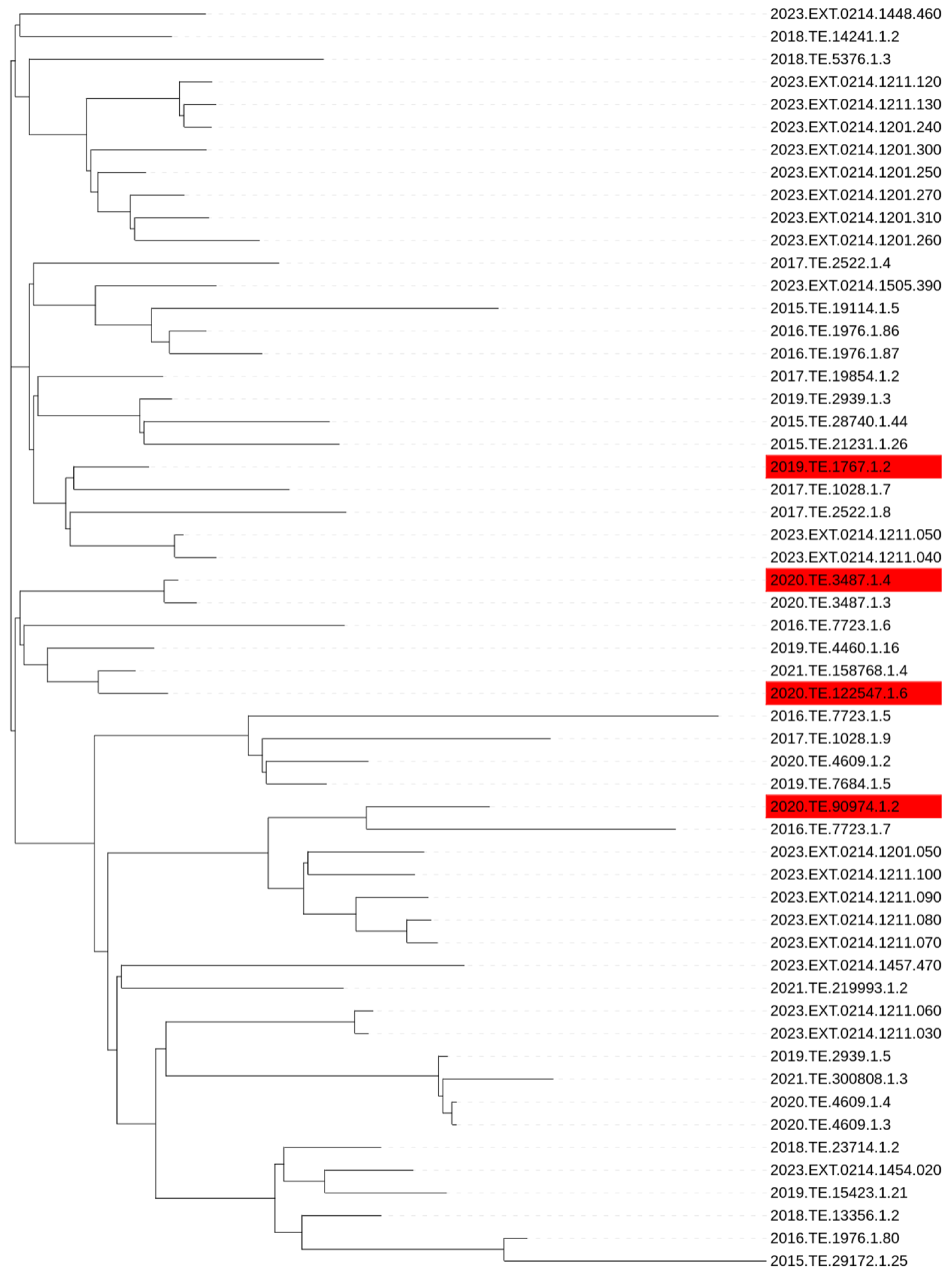
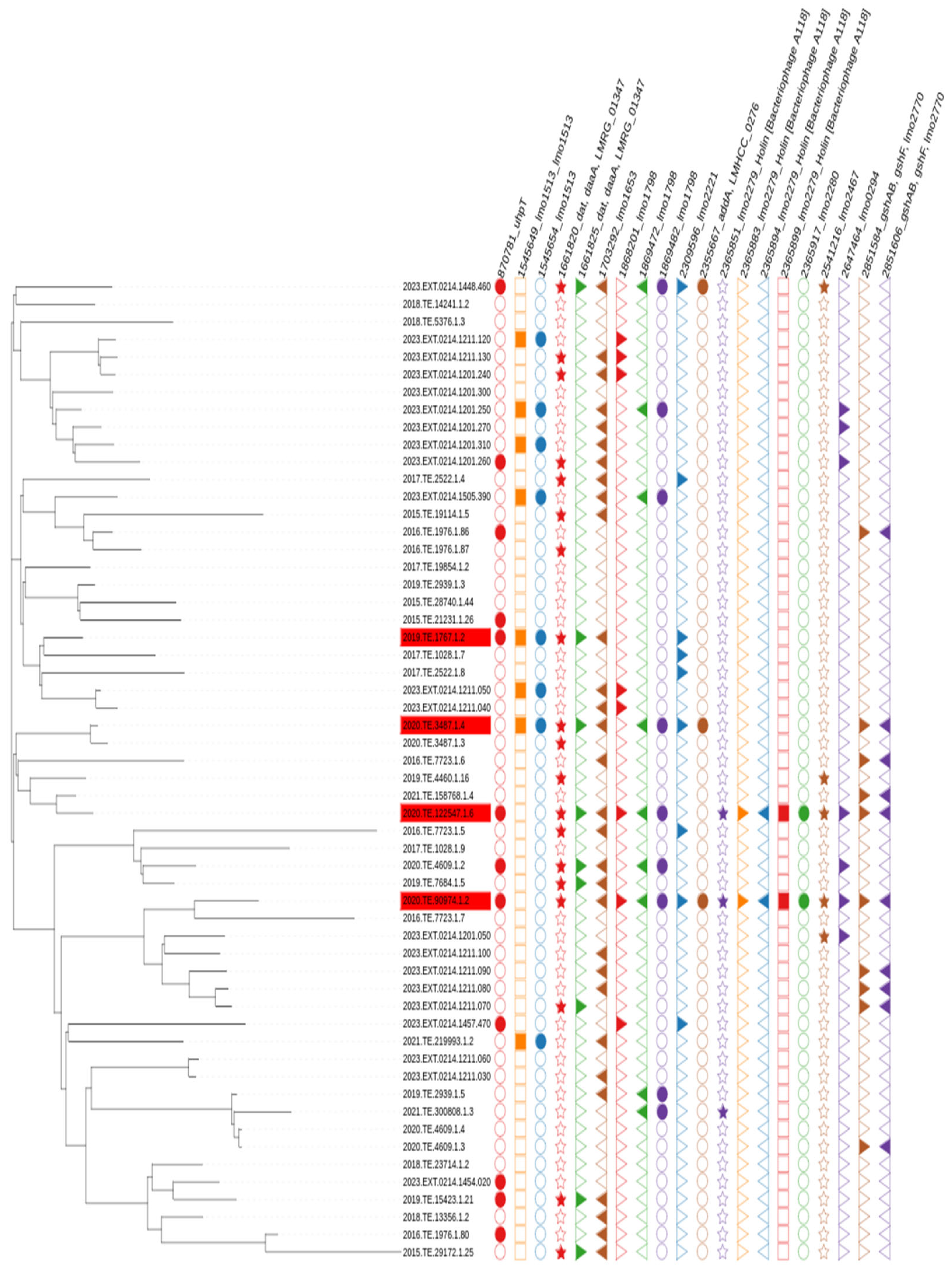
2.5. Phylogenetic Analysis
2.6. Gene Alignment and Protein Similarity Visualization
2.7. Protein Translation and Function
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Lm | Listeria monocytogenes |
| RTE | Ready-to-eat food |
| WGS | Whole Genome Sequencing |
| GWAS | Genome Wide Association Studies |
| C&D | Cleaning and disinfection |
| CCs | Clonal complexes |
| GENPAT | National Reference Centre for whole genome sequencing of microbial pathogens: database and bioinformatic analysis |
| NCBI | National Center for Biotechnology Information |
| MLST | Multilocus sequence typing |
| ST | Sequence type |
| cgMLST | Core genome MLST |
| NA | Not applicable |
References
- Dhama, K.; Karthik, K.; Tiwari, R.; Shabbir, M.Z.; Barbuddhe, S.; Malik, S.V.S.; Singh, R.K. Listeriosis in Animals, Its Public Health Significance (Food-Borne Zoonosis) and Advances in Diagnosis and Control: A Comprehensive Review. Vet. Q. 2015, 35, 211–235. [Google Scholar] [CrossRef]
- European Food Safety Authority; European Centre for Disease Prevention and Control. Prolonged multi-country outbreak of Listeria monocytogenes ST173 linked to consumption of fish products. EFSA Support. Publ. 2024, 21, 8885E. [Google Scholar] [CrossRef]
- Maury, M.M.; Tsai, Y.-H.; Charlier, C.; Touchon, M.; Chenal-Francisque, V.; Leclercq, A.; Criscuolo, A.; Gaultier, C.; Roussel, S.; Brisabois, A.; et al. Uncovering Listeria monocytogenes Hypervirulence by Harnessing Its Biodiversity. Nat. Genet. 2016, 48, 308–313. [Google Scholar] [CrossRef] [PubMed]
- ISO 11290-1:2017; Microbiology of the Food Chain—Horizontal Method for the Detection and Enumeration of Listeria monocytogenes and of Listeria spp.—Part 1: Detection Method. International Organization for Standardization: Geneva, Switzerland, 2017.
- ISO 11290-2:2017; Microbiology of the Food Chain—Horizontal Method for the Detection and Enumeration of Listeria monocytogenes and of Listeria spp.—Part 2: Enumeration Method. International Organization for Standardization: Geneva, Switzerland, 2017.
- Zilelidou, E.A.; Skandamis, P.N. Growth, Detection and Virulence of Listeria monocytogenes in the Presence of Other Microorganisms: Microbial Interactions from Species to Strain Level. Int. J. Food Microbiol. 2018, 277, 10–25. [Google Scholar] [CrossRef] [PubMed]
- Disson, O.; Moura, A.; Lecuit, M. Making Sense of the Biodiversity and Virulence of Listeria monocytogenes. Trends Microbiol. 2021, 29, 811–822. [Google Scholar] [CrossRef]
- Bannenberg, J.W.; Abee, T.; Zwietering, M.H.; Den Besten, H.M.W. Variability in Lag Duration of Listeria monocytogenes Strains in Half Fraser Enrichment Broth after Stress Affects the Detection Efficacy Using the ISO 11290-1 Method. Int. J. Food Microbiol. 2021, 337, 108914. [Google Scholar] [CrossRef]
- Allende, A.; Alvarez-Ordóñez, A.; Bolton, D.; Bover-Cid, S.; Chemaly, M.; Davies, R.; De Cesare, A.; Herman, L.; Hilbert, F.; Koutsoumanis, K.; et al. Whole Genome Sequencing and Metagenomics for Outbreak Investigation, Source Attribution and Risk Assessment of Food-Borne Microorganisms. EFSA J. 2018, 16, e00058. [Google Scholar] [CrossRef]
- Chiaverini, A.; Guidi, F.; Torresi, M.; Acciari, V.A.; Centorotola, G.; Cornacchia, A.; Centorame, P.; Marfoglia, C.; Blasi, G.; Di Domenico, M.; et al. Phylogenetic Analysis and Genome-Wide Association Study Applied to an Italian Listeria monocytogenes Outbreak. Front. Microbiol. 2021, 12, 750065. [Google Scholar] [CrossRef]
- Guidi, F.; Centorotola, G.; Chiaverini, A.; Iannetti, L.; Schirone, M.; Visciano, P.; Cornacchia, A.; Scattolini, S.; Pomilio, F.; D’Alterio, N.; et al. The Slaughterhouse as Hotspot of CC1 and CC6 Listeria monocytogenes Strains with Hypervirulent Profiles in an Integrated Poultry Chain of Italy. Microorganisms 2023, 11, 1543. [Google Scholar] [CrossRef]
- Portmann, A.-C.; Fournier, C.; Gimonet, J.; Ngom-Bru, C.; Barretto, C.; Baert, L. A Validation Approach of an End-to-End Whole Genome Sequencing Workflow for Source Tracking of Listeria monocytogenes and Salmonella enterica. Front. Microbiol. 2018, 9, 446. [Google Scholar] [CrossRef]
- Cito, F.; Di Pasquale, A.; Cammà, C.; Cito, P. The Italian Information System for the Collection and Analysis of Complete Genome Sequence of Pathogens Isolated from Animal, Food and Environment. Int. J. Infect. Dis. 2018, 73, 296–297. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Timme, R.E.; Lafon, P.C.; Balkey, M.; Adams, J.K.; Wagner, D.; Carleton, H.; Strain, E.; Hoffmann, M.; Sabol, A.; Rand, H. Gen-FS Coordinated Proficiency Test Data for Genomic Foodborne Pathogen Surveillance, 2017 and 2018 Exercises. Sci. Data 2020, 7, 402. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.; Machado, M.P.; Silva, D.N.; Rossi, M.; Moran-Gilad, J.; Santos, S.; Ramirez, M.; Carriço, J.A. chewBBACA: A Complete Suite for Gene-by-Gene Schema Creation and Strain Identification. Microb. Genom. 2018, 4, e000166. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid Prokaryotic Genome Annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef] [PubMed]
- Tonkin-Hill, G.; MacAlasdair, N.; Ruis, C.; Weimann, A.; Horesh, G.; Lees, J.A.; Gladstone, R.A.; Lo, S.; Beaudoin, C.; Floto, R.A.; et al. Producing Polished Prokaryotic Pangenomes with the Panaroo Pipeline. Genome Biol. 2020, 21, 180. [Google Scholar] [CrossRef] [PubMed]
- Brynildsrud, O.; Bohlin, J.; Scheffer, L.; Eldholm, V. Rapid Scoring of Genes in Microbial Pan-Genome-Wide Association Studies with Scoary. Genome Biol. 2016, 17, 238. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An Online Tool for Phylogenetic Tree Display and Annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Seemann, T. Snippy: Fast Bacterial Variant Calling from NGS Reads. GitHub. 2015. Available online: https://github.com/tseemann/snippy (accessed on 9 August 2025).
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve Years of SAMtools and BCFtools. GigaScience 2021, 10, giab008. [Google Scholar] [CrossRef]
- Kaas, R.S.; Leekitcharoenphon, P.; Aarestrup, F.M.; Lund, O. Solving the Problem of Comparing Whole Bacterial Genomes across Different Sequencing Platforms. PLoS ONE 2014, 9, e104984. [Google Scholar] [CrossRef] [PubMed]
- Gilchrist, C.L.M.; Chooi, Y.-H. Clinker & Clustermap.Js: Automatic Generation of Gene Cluster Comparison Figures. Bioinformatics 2021, 37, 2473–2475. [Google Scholar] [CrossRef]
- BLAST® Command Line Applications User Manual [Internet]. National Center for Biotechnology Information (US): Bethesda, MD, USA, 2008. Available online: https://www.ncbi.nlm.nih.gov/books/NBK279690/ (accessed on 9 August 2025).
- Denes, T.; Vongkamjan, K.; Ackermann, H.-W.; Moreno Switt, A.I.; Wiedmann, M.; den Bakker, H.C. Comparative Genomic and Morphological Analyses of Listeria Phages Isolated from Farm Environments. Appl. Environ. Microbiol. 2014, 80, 4616–4625. [Google Scholar] [CrossRef] [PubMed]
- Read, T.D.; Massey, R.C. Characterizing the Genetic Basis of Bacterial Phenotypes Using Genome-Wide Association Studies: A New Direction for Bacteriology. Genome Med. 2014, 6, 109. [Google Scholar] [CrossRef] [PubMed]
- Loessner, M.J.; Inman, R.B.; Lauer, P.; Calendar, R. Complete Nucleotide Sequence, Molecular Analysis and Genome Structure of Bacteriophage A118 of Listeria monocytogenes: Implications for Phage Evolution. Mol. Microbiol. 2000, 35, 324–340. [Google Scholar] [CrossRef]
- Pasechnek, A.; Rabinovich, L.; Stadnyuk, O.; Azulay, G.; Mioduser, J.; Argov, T.; Borovok, I.; Sigal, N.; Herskovits, A.A. Active Lysogeny in Listeria monocytogenes Is a Bacteria-Phage Adaptive Response in the Mammalian Environment. Cell Rep. 2020, 32, 107956. [Google Scholar] [CrossRef]
- Karlsmose, A.K.; Ivanova, M.; Kragh, M.L.; Kjeldgaard, J.S.; Otani, S.; Svendsen, C.A.; Papić, B.; Zdovc, I.; Tasara, T.; Stephan, R.; et al. A Novel Metagenomic Approach Uncovers Phage Genes as Markers for Increased Disinfectant Tolerance in Mixed Listeria monocytogenes Communities. Infect. Genet. Evol. 2024, 119, 105582. [Google Scholar] [CrossRef]
- Kawacka, I.; Olejnik-Schmidt, A.; Schmidt, M.; Sip, A. Effectiveness of Phage-Based Inhibition of Listeria monocytogenes in Food Products and Food Processing Environments. Microorganisms 2020, 8, 1764. [Google Scholar] [CrossRef]




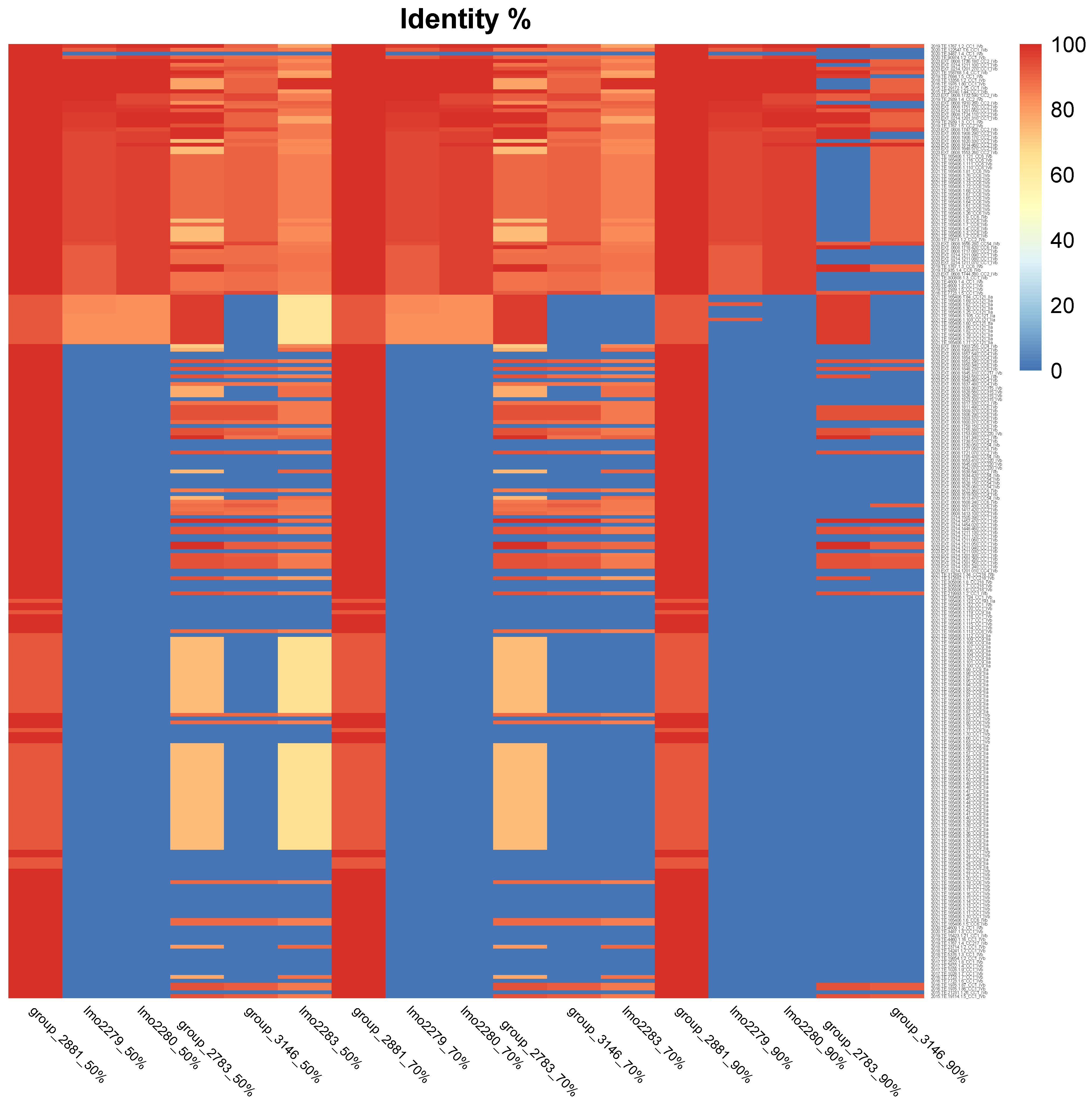
| Gene | Threshold (%) | n. Absent | Absent % | n. Similarity 50–90% | Similarity 50–90 % | n. Similarity > 90% | Similarity > 90% |
|---|---|---|---|---|---|---|---|
| Group_2881 | 50.00% | 0 | NA | 0 | NA | 251 | 100% |
| Group_2881 | 70.00% | 0 | NA | 0 | NA | 251 | 100% |
| Group_2881 | 90.00% | 0 | NA | 0 | NA | 251 | 100% |
| lmo2279 | 50.00% | 173 | 69% | 13 | 5% | 65 | 26% |
| lmo2279 | 70.00% | 173 | 69% | 13 | 5% | 65 | 26% |
| lmo2279 | 90.00% | 184 | 73% | 0 | 0 | 67 | 27% |
| lmo2280 | 50.00% | 173 | 69% | 13 | 5% | 65 | 26% |
| lmo2280 | 70.00% | 173 | 69% | 13 | 5% | 65 | 26% |
| lmo2280 | 90.00% | 186 | 74% | 0 | 0 | 65 | 26% |
| group_2783 | 50.00% | 77 | 31% | 113 | 45% | 61 | 24% |
| group_2783 | 70.00% | 77 | 31% | 113 | 45% | 61 | 24% |
| group_2783 | 90.00% | 190 | 76% | 0 | 0 | 61 | 24% |
| group_3146 | 50.00% | 147 | 59% | 34 | 14% | 70 | 28% |
| group_3146 | 70.00% | 147 | 59% | 34 | 14% | 70 | 28% |
| group_3146 | 90.00% | 180 | 72% | 0 | 0 | 71 | 28% |
| lmo2283 | 50.00% | 77 | 31% | 168 | 67% | 6 | 2% |
| lmo2283 | 70.00% | 138 | 55% | 107 | 43% | 6 | 2% |
| lmo2283 | 90.00% | 240 | 96% | 0 | 0 | 11 | 4% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Angelis, M.E.; Robbe, G.A.; Cammà, C.; Ancora, M.; Bosica, S.; Ferrara, M.; Torresi, M.; Aarestrup, F.M.; Gmeiner, A.; Nuanmuang, N.; et al. GWAS Study Applied to Phenotypically Slow Growth Strains of Listeria monocytogenes Workflow and Outcome. Microorganisms 2025, 13, 2011. https://doi.org/10.3390/microorganisms13092011
De Angelis ME, Robbe GA, Cammà C, Ancora M, Bosica S, Ferrara M, Torresi M, Aarestrup FM, Gmeiner A, Nuanmuang N, et al. GWAS Study Applied to Phenotypically Slow Growth Strains of Listeria monocytogenes Workflow and Outcome. Microorganisms. 2025; 13(9):2011. https://doi.org/10.3390/microorganisms13092011
Chicago/Turabian StyleDe Angelis, Maria Elisabetta, Giovanna Alessia Robbe, Cesare Cammà, Massimo Ancora, Serena Bosica, Mattia Ferrara, Marina Torresi, Frank M. Aarestrup, Alexander Gmeiner, Narong Nuanmuang, and et al. 2025. "GWAS Study Applied to Phenotypically Slow Growth Strains of Listeria monocytogenes Workflow and Outcome" Microorganisms 13, no. 9: 2011. https://doi.org/10.3390/microorganisms13092011
APA StyleDe Angelis, M. E., Robbe, G. A., Cammà, C., Ancora, M., Bosica, S., Ferrara, M., Torresi, M., Aarestrup, F. M., Gmeiner, A., Nuanmuang, N., Njage, P. M. K., Bravaccini, C., Belardo, V., Di Iorio, C., Di Zacomo, S., Fazii, P., Pomilio, F., & Leekitcharoenphon, P. (2025). GWAS Study Applied to Phenotypically Slow Growth Strains of Listeria monocytogenes Workflow and Outcome. Microorganisms, 13(9), 2011. https://doi.org/10.3390/microorganisms13092011






