Quorum-Sensing C12-HSL Drives Antibiotic Resistance Plasmid Transfer via Membrane Remodeling, Oxidative Stress, and RpoS-RMF Crosstalk
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Culture Conditions
2.2. Reagents and Quorum Sensing Modulators
2.3. Minimum Inhibitory Concentration (MIC) Determination
2.4. Plasmid Conjugation Assay
2.5. Scanning Electron Microscopy (SEM)
2.6. Membrane Integrity Assessment
2.7. Intracellular Reactive Oxygen Species (ROS) Measurement
2.8. Construction of Isogenic Deletion Mutants and Complementation
2.9. Quantitative Reverse Transcription PCR (qRT-PCR)
2.10. Growth Curve Analysis
2.11. Statistical Analysis
3. Results
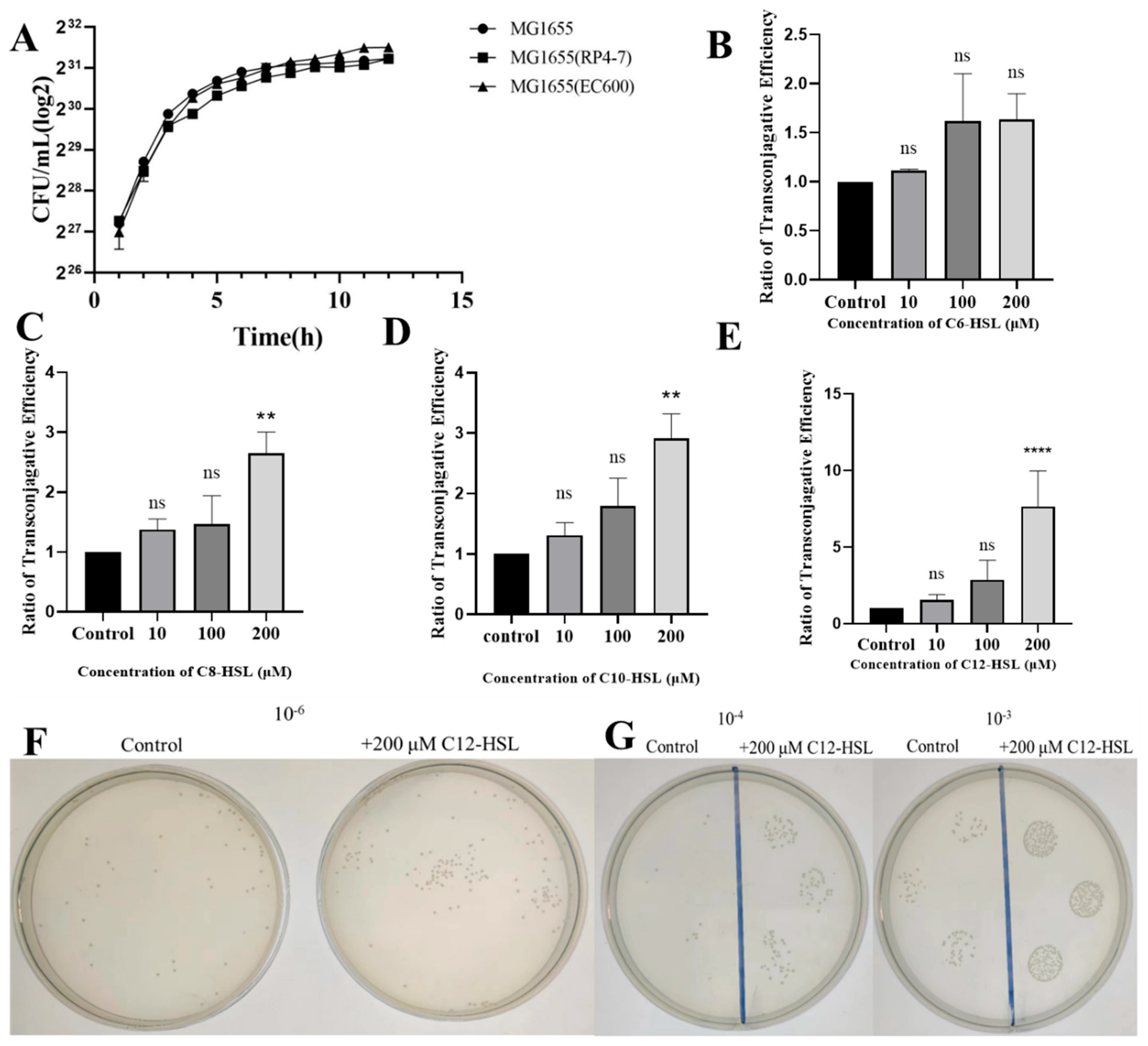
3.1. Ensuring Strain Growth Uniformity and Quantifying AHL-Specific Enhancement of Plasmid Conjugation in E. coli
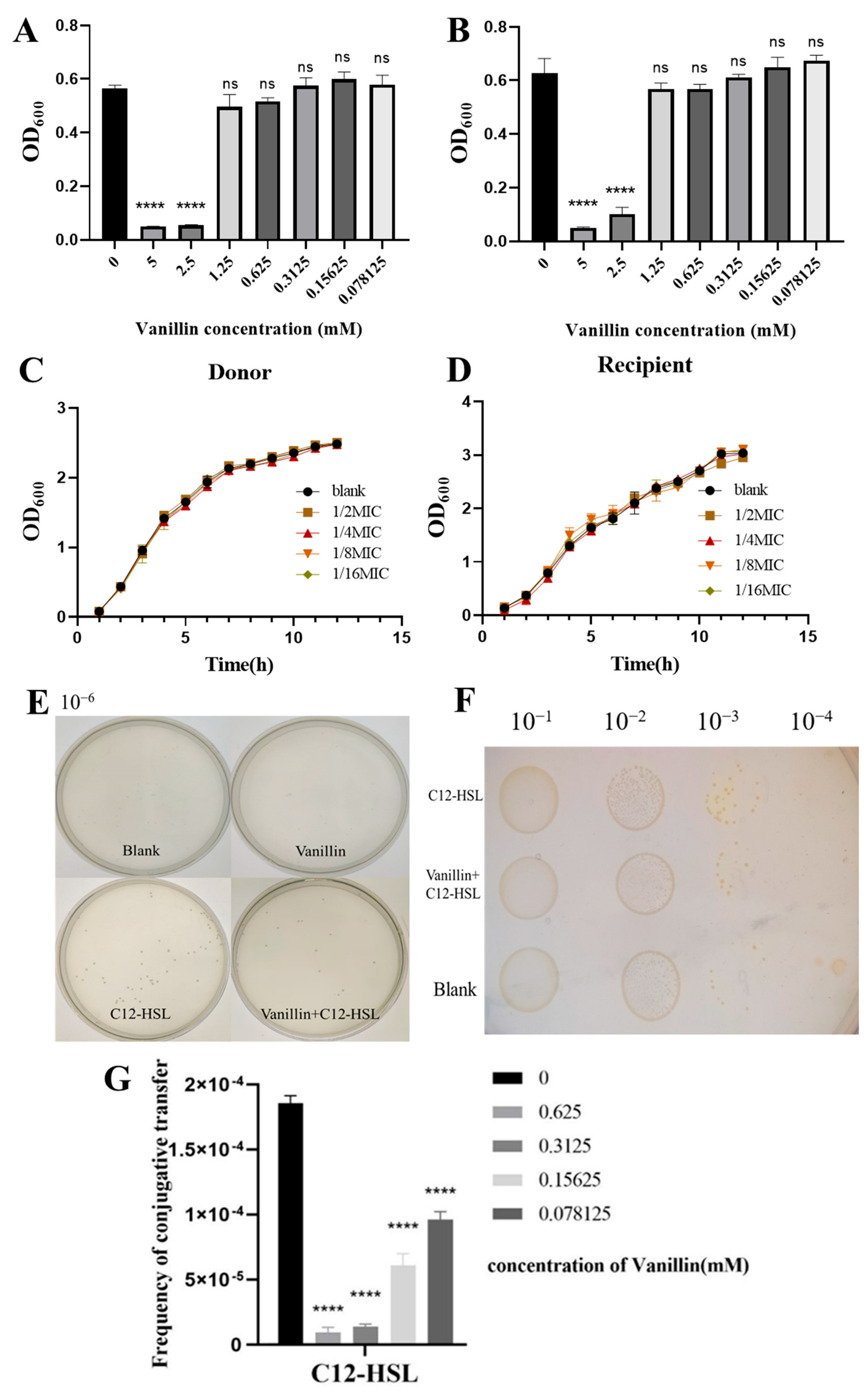
3.2. Vanillin Interferes with C12-HSL-Stimulated Plasmid Conjugation in E. coli
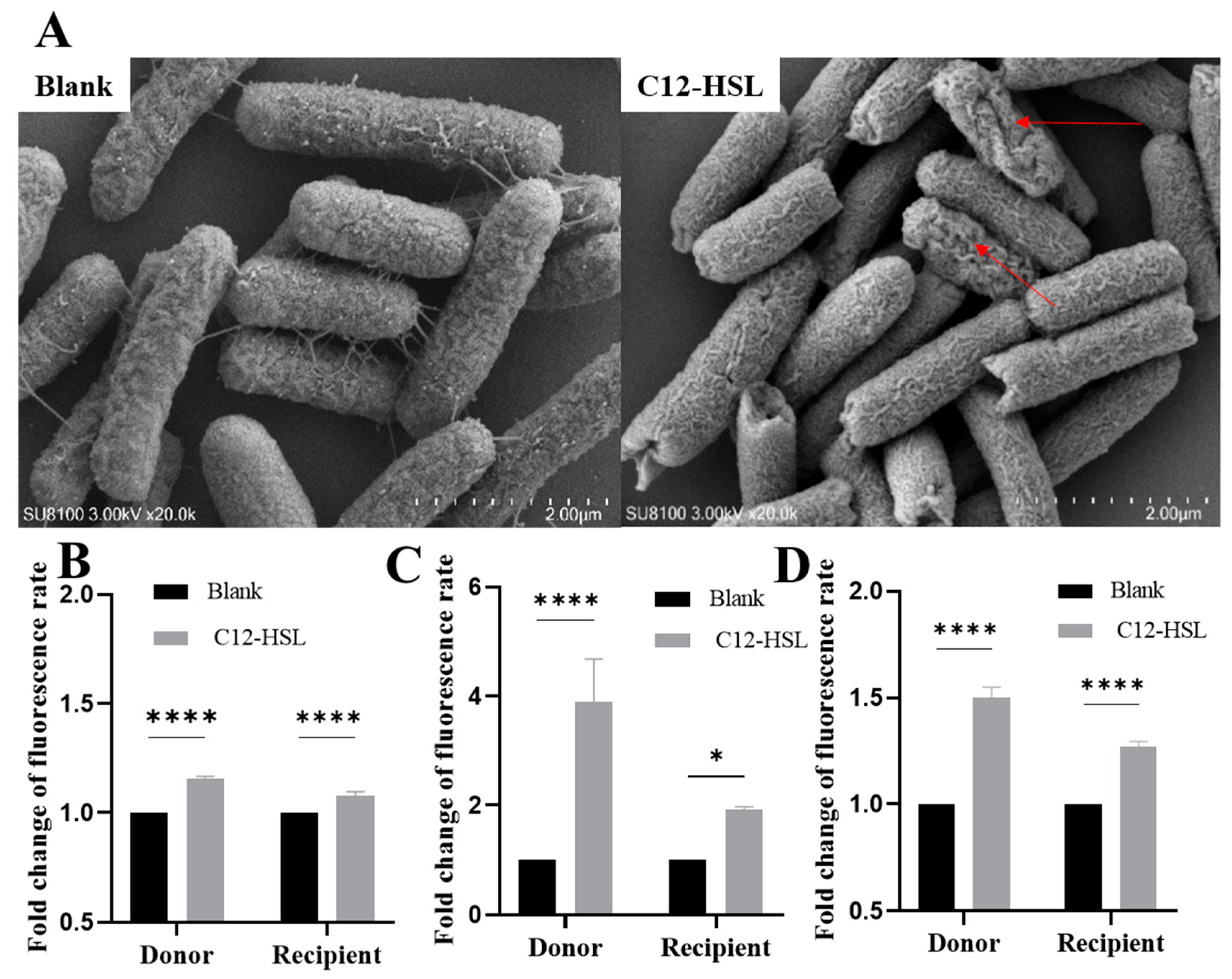
3.3. C12-HSL Disrupts Membrane Integrity and Induces Oxidative Stress to Facilitate Conjugation
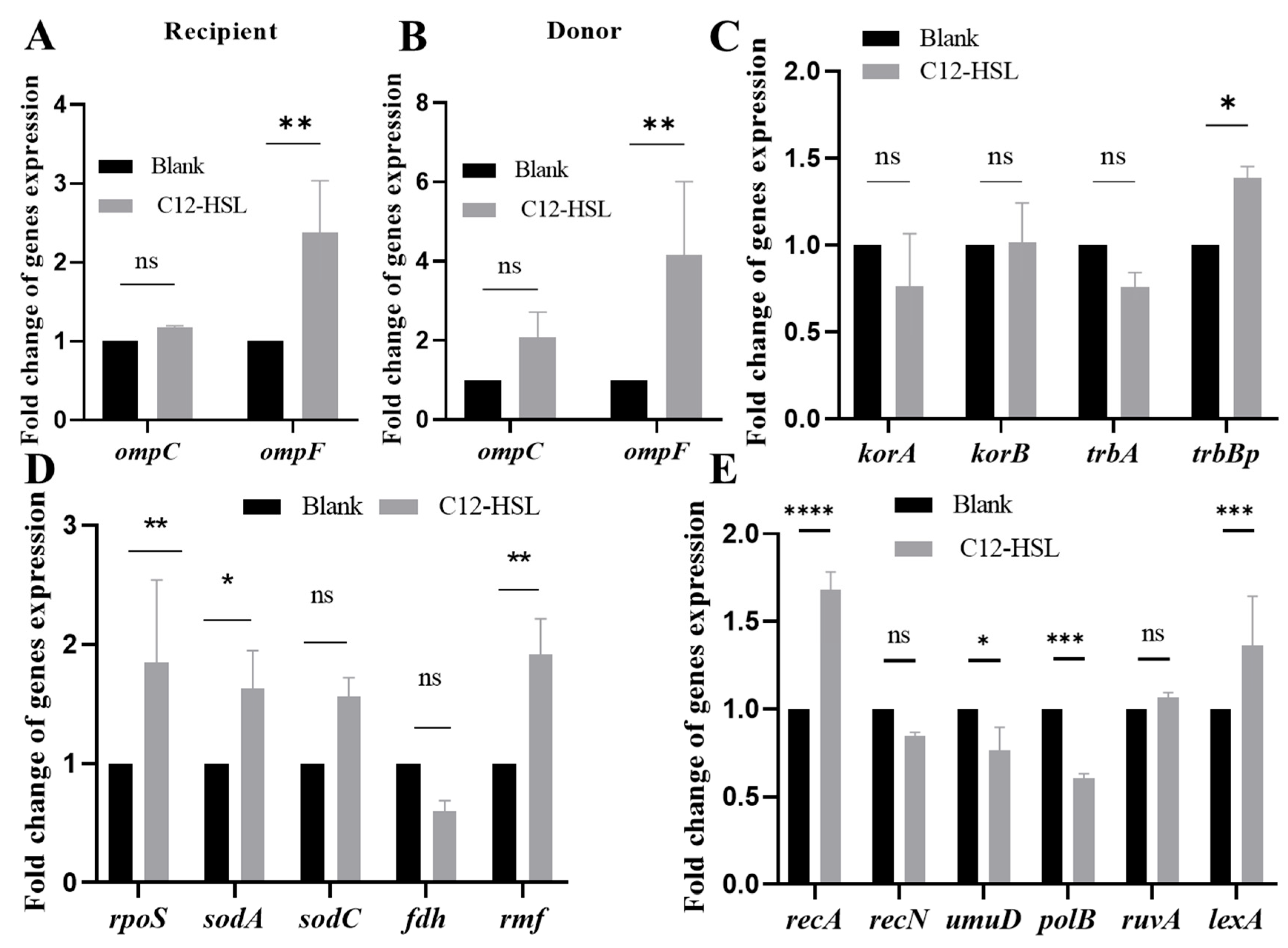
3.4. C12-HSL Transcriptional Reprogramming Optimizes Membrane Architecture and Genetic Plasticity for Conjugation
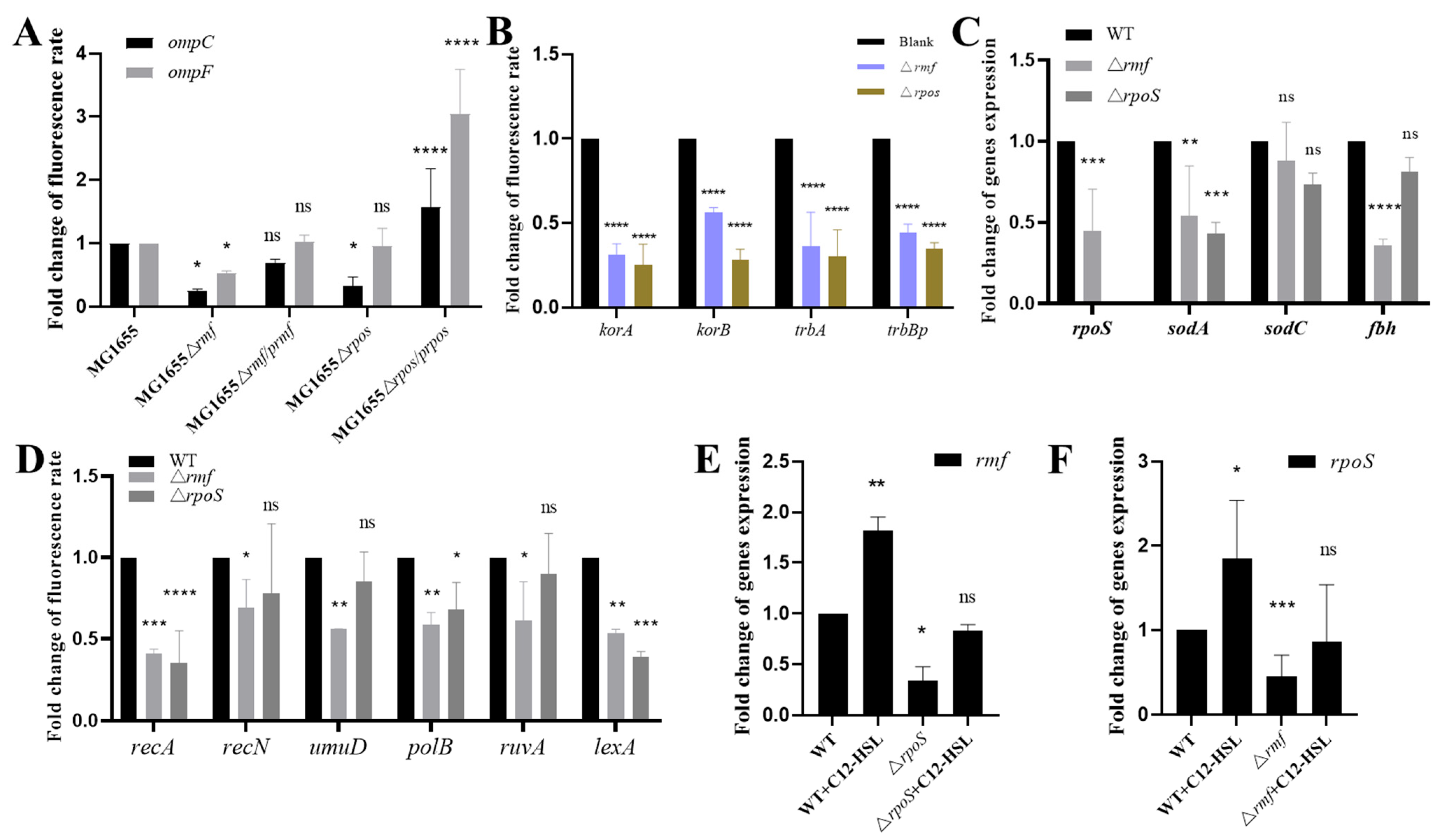
3.5. rmf and rpoS Govern Quorum Sensing-Driven Antibiotic Resistance Dissemination
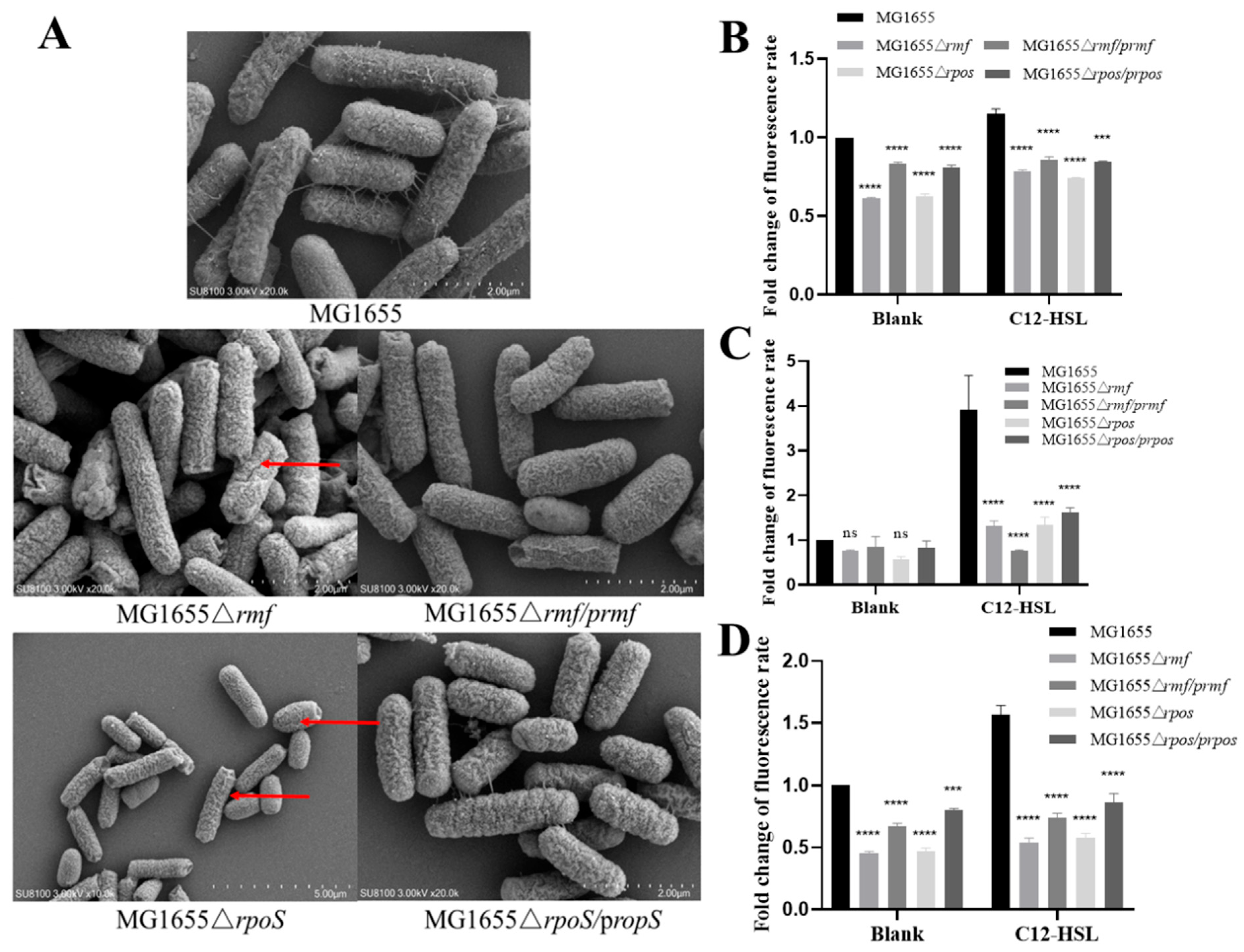
3.6. Genetic Control of Membrane Integrity and Oxidative Stress by rmf and rpoS
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zheng, J.; Chen, T.; Chen, H. Antibiotic resistome promotion in drinking water during biological activated carbon treatment: Is it influenced by quorum sensing? Sci. Total Environ. 2018, 612, 1–8. [Google Scholar] [CrossRef]
- Chen, C.-Y.; Fuqua, C.; Jackson, C.R.; Kadlec, K.; Top, E.M. Editorial: Plasmid transfer-mechanisms, ecology, evolution and applications. Front. Microbiol. 2022, 13, 993628. [Google Scholar] [CrossRef]
- Wang, M.; Lian, Y.; Wang, Y.; Zhu, L. The role and mechanism of quorum sensing on environmental antimicrobial resistance. Environ. Pollut. 2023, 322, 121238. [Google Scholar] [CrossRef]
- Ahmed, S.K.; Hussein, S.; Qurbani, K.; Ibrahim, R.H.; Fareeq, A.; Mahmood, K.A.; Mohamed, M.G. Antimicrobial resistance: Impacts, challenges, and future prospects. J. Med. Surg. Public Health 2024, 2, 100081. [Google Scholar] [CrossRef]
- Brito, I.L. Examining horizontal gene transfer in microbial communities. Nat. Rev. Microbiol. 2021, 19, 442–453. [Google Scholar] [CrossRef]
- Liu, G.; Thomsen, L.E.; Olsen, J.E. Antimicrobial-induced horizontal transfer of antimicrobial resistance genes in bacteria: A mini-review. J. Antimicrob. Chemother. 2022, 77, 556–567. [Google Scholar] [CrossRef]
- Waksman, G. Molecular basis of conjugation-mediated DNA transfer by gram-negative bacteria. Curr. Opin. Struct. Biol. 2025, 90, 102978. [Google Scholar] [CrossRef]
- Virolle, C.; Goldlust, K.; Djermoun, S.; Bigot, S.; Lesterlin, C. Plasmid Transfer by Conjugation in Gram-Negative Bacteria: From the Cellular to the Community Level. Genes 2020, 11, 1239. [Google Scholar] [CrossRef]
- Boto, L.; Pineda, M.; Pineda, R. Potential impacts of horizontal gene transfer on human health and physiology and how anthropogenic activity can affect it. FEBS J. 2019, 286, 3959–3967. [Google Scholar] [CrossRef]
- Zhang, Y.; Ma, Q.; Su, B.; Chen, R.; Lin, J.; Lin, Z.; Wang, D.; Yu, Y. A study on the role that quorum sensing play in antibiotic-resistant plasmid conjugative transfer in Escherichia coli. Ecotoxicology 2018, 27, 209–216. [Google Scholar] [CrossRef]
- Lu, Y.; Zeng, J.; Wu, B.; E, S.; Wang, L.; Cai, R.; Zhang, N.; Li, Y.; Huang, X.; Huang, B.; et al. Quorum Sensing N-acyl Homoserine Lactones-SdiA Suppresses Escherichia coli-Pseudomonas aeruginosa Conjugation through Inhibiting traI Expression. Front. Cell. Infect. Microbiol. 2017, 7, 7. [Google Scholar] [CrossRef]
- van Gestel, J.; Bareia, T.; Tenennbaum, B.; Dal Co, A.; Guler, P.; Aframian, N.; Puyesky, S.; Grinberg, I.; D’Souza, G.G.; Erez, Z.; et al. Short-range quorum sensing controls horizontal gene transfer at micron scale in bacterial communities. Nat. Commun. 2021, 12, 2324. [Google Scholar] [CrossRef]
- Striednig, B.; Hilbi, H. Bacterial quorum sensing and phenotypic heterogeneity: How the collective shapes the individual. Trends Microbiol. 2022, 30, 379–389. [Google Scholar] [CrossRef]
- Zhu, L.; Chen, T.; Xu, L.; Zhou, Z.; Feng, W.; Liu, Y.; Chen, H. Effect and mechanism of quorum sensing on horizontal transfer of multidrug plasmid RP4 in BAC biofilm. Sci. Total Environ. 2020, 698, 134236. [Google Scholar] [CrossRef]
- Zhao, X.; Yu, Z.; Ding, T. Quorum-Sensing Regulation of Antimicrobial Resistance in Bacteria. Microorganisms 2020, 8, 425. [Google Scholar] [CrossRef]
- Pacheco, T.; Gomes, A.É.; Siqueira, N.M.; Assoni, L.; Darrieux, M.; Venter, H.; Ferraz, L.F. SdiA, a Quorum-Sensing Regulator, Suppresses Fimbriae Expression, Biofilm Formation, and Quorum-Sensing Signaling Molecules Production in Klebsiella pneumoniae. Front. Microbiol. 2021, 12, 597735. [Google Scholar] [CrossRef]
- Bouyahya, A.; Abrini, J.; Dakka, N.; Bakri, Y. Essential oils of Origanum compactum increase membrane permeability, disturb cell membrane integrity, and suppress quorum-sensing phenotype in bacteria. J. Pharm. Anal. 2019, 9, 301–311. [Google Scholar] [CrossRef]
- Zhou, J.-W.; Ruan, L.-Y.; Chen, H.-J.; Luo, H.-Z.; Jiang, H.; Wang, J.-S.; Jia, A.-Q. Correction to Inhibition of Quorum Sensing and Virulence in Serratia marcescens by Hordenine. J. Agric. Food Chem. 2019, 67, 2420. [Google Scholar] [CrossRef]
- Zhou, J.-W.; Muhammad, J.; Sun, B.; Yang, R.; Wadood, A.; Wang, J.-S.; Jia, A.-Q. Metabolomic analysis of quorum sensing inhibitor hordenine on Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2019, 103, 6271–6285. [Google Scholar] [CrossRef]
- Wang, H.; Shang, F.; Shen, J.; Xu, J.; Chen, X.; Ni, J.; Yu, L.; Xue, T. LsrR, the effector of AI-2 quorum sensing, is vital for the H2O2 stress response in mammary pathogenic Escherichia coli. Vet. Res. 2021, 52, 127. [Google Scholar] [CrossRef]
- Xiao, X.; Zeng, F.; Li, R.; Liu, Y.; Wang, Z. Subinhibitory Concentration of Colistin Promotes the Conjugation Frequencies of Mcr-1- and blaNDM-5-Positive Plasmids. Microbiol. Spectr. 2022, 10, e02160-21. [Google Scholar] [CrossRef]
- Bouillet, S.; Hamdallah, I.; Majdalani, N.; Tripathi, A.; Gottesman, S.; Papenfort, K. A negative feedback loop is critical for recovery of RpoS after stress in Escherichia coli. PLoS Genet. 2024, 20, e1011059. [Google Scholar] [CrossRef]
- Zhu, Z.; Zhang, Y.; Li, J.; Dong, H. Insight into quorum sensing and microbial community of an anammox consortium in response to salt stress: From “Candaditus brocadia” to “Candaditus scalindua”. Sci. Total Environ. 2021, 796, 148979. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, J.; Hu, Z.; Li, J.; Lu, H. Oxytetracycline stress stimulates antibiotic resistance gene proliferation and quorum sensing response of marine anammox bacteria in seawater-based wastewater treatment. Chem. Eng. J. 2022, 447, 137539. [Google Scholar] [CrossRef]
- Schellhorn, H.E. Function, Evolution, and Composition of the RpoS Regulon in Escherichia coli. Front. Microbiol. 2020, 11, 560099. [Google Scholar] [CrossRef]
- Zhang, D.; Li, S.H.-J.; King, C.G.; Wingreen, N.S.; Gitai, Z.; Li, Z.; Sinha, S. Global and gene-specific translational regulation in Escherichia coli across different conditions. PLoS Comput. Biol. 2022, 18, e1010641. [Google Scholar] [CrossRef]
- Dapa, T.; Fleurier, S.; Bredeche, M.-F.; Matic, I. The SOS and RpoS Regulons Contribute to Bacterial Cell Robustness to Genotoxic Stress by Synergistically Regulating DNA Polymerase Pol II. Genetics 2017, 206, 1349–1360. [Google Scholar] [CrossRef]
- Zhang, Y.; Guo, M.; Shen, P.; Xie, Z. The stress response factor RpoS is required for the natural transformation of Escherichia coli. Chin. Sci. Bull. 2014, 59, 521–527. [Google Scholar] [CrossRef]
- Sebastian, H.; Finkel, S.E.; Yap, M.-N.F. Absence of Ribosome Modulation Factor Alters Growth and Competitive Fitness of Escherichia coli. Microbiol. Spectr. 2022, 10, e0223921. [Google Scholar] [CrossRef]
- Usachev, K.S.; Yusupov, M.M.; Validov, S.Z. Hibernation as a Stage of Ribosome Functioning. Biochemistry 2020, 85, 1434–1442. [Google Scholar] [CrossRef]
- Polikanov, Y.S.; Blaha, G.M.; Steitz, T.A. How hibernation factors RMF, HPF, and YfiA turn off protein synthesis. Science 2012, 336, 915–918. [Google Scholar] [CrossRef]
- Maki, Y.; Yoshida, H. Ribosomal Hibernation-Associated Factors in Escherichia coli. Microorganisms 2021, 10, 33. [Google Scholar] [CrossRef]
- Shastry, R.P.; Ghate, S.D.; Kumar, B.S.; Srinath, B.S.; Kumar, V. Vanillin derivative inhibits quorum sensing and biofilm formation in Pseudomonas aeruginosa: A study in a Caenorhabditis elegans infection model. Nat. Prod. Res. 2021, 36, 1610–1615. [Google Scholar] [CrossRef]
- Li, T.; He, B.; Mei, Y.; Wang, D.; Sun, X.; Li, J. Inhibitory effect of vanillin on the virulence factors and biofilm formation of Hafnia alvei. Food Sci. Technol. 2019, 102, 223–229. [Google Scholar] [CrossRef]
- Yang, Y.; Yao, F.; Zhou, M.; Zhu, J.; Zhang, X.; Bao, W.; Wu, S.; Hardwidge, P.R.; Zhu, G. F18ab Escherichia coli flagella expression is regulated by acyl-homoserine lactone and contributes to bacterial virulence. Vet. Microbiol. 2013, 165, 378–383. [Google Scholar] [CrossRef]
- Yang, Y.; Zhou, M.; Hou, H.; Zhu, J.; Yao, F.; Zhang, X.; Zhu, X.; Hardwidge, P.R.; Zhu, G. Quorum-sensing gene luxS regulates flagella expression and Shiga-like toxin production in F18ab Escherichia coli. Can. J. Microbiol. 2014, 60, 355–361. [Google Scholar] [CrossRef]
- Yang, Y.; Zhang, X.; Zhang, B.; Zhou, M.; Duan, Q.; Li, Z.; Zhang, X.; Zhu, G. Quorum sensing-1 signaling of N-hexanoyl-l-homoserine lactone contributes to virulence in avian pathogenic Escherichia coli. Arch. Microbiol. 2021, 203, 6079–6089. [Google Scholar] [CrossRef]
- Yang, Y.; Zhang, X.; Yang, J.; Wu, Z.; Li, J.; Song, R.; Meng, C.; Zhu, G. N-(3-Oxododecanoyl)-Homoserine Lactone Induces Intestinal Barrier Damage in Piglets via the Lipid Raft-Mediated Apoptosis Pathway. Vet. Sci. 2025, 12, 233. [Google Scholar] [CrossRef]
- Yang, Y.; Shao, J.; Zhou, M.; Duan, Q.; Zhang, X.; Zhu, G. Effects of Quorum Sensing AHL Signaling on the Biological Characteristics of Porcine Derived F4ac+ Enterotoxigenic Escherichia coli. Pak. J. Zool. 2022, 54, 701–708. [Google Scholar] [CrossRef]
- Ramsay, J.P.; Bastholm, T.R.; Verdonk, C.J.; Tambalo, D.D.; Sullivan, J.T.; Harold, L.K.; Panganiban, B.A.; Colombi, E.; Perry, B.J.; Jowsey, W.; et al. An epigenetic switch activates bacterial quorum sensing and horizontal transfer of an integrative and conjugative element. Nucleic Acids Res. 2021, 50, 975–988. [Google Scholar] [CrossRef]
- Qiu, X.; Wang, B.; Ren, S.; Liu, X.; Wang, Y. Regulation of quorum sensing for the manipulation of conjugative transfer of antibiotic resistance genes in wastewater treatment system. Water Res. 2024, 253, 121222. [Google Scholar] [CrossRef]
- Wang, T.; Liang, C.; Zheng, M.; Liu, L.; An, Y.; Xu, H.; Xiao, S.; Nie, L. Ribosome Hibernation as a Stress Response of Bacteria. Protein Pept. Lett. 2020, 27, 1082–1091. [Google Scholar] [CrossRef]
- Feaga, H.A.; Dworkin, J. Transcription regulates ribosome hibernation. Mol. Microbiol. 2021, 116, 663–673. [Google Scholar] [CrossRef]
- Khaova, E.A.; Kashevarova, N.M.; Tkachenko, A.G. Ribosome Hibernation: Molecular Strategy of Bacterial Survival. Appl. Biochem. Microbiol. 2022, 58, 213–231. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Y.; Wu, Z.; Zhu, L.; Han, Z.; Li, J.; Fang, Q.; Zhu, G. Quorum-Sensing C12-HSL Drives Antibiotic Resistance Plasmid Transfer via Membrane Remodeling, Oxidative Stress, and RpoS-RMF Crosstalk. Microorganisms 2025, 13, 1837. https://doi.org/10.3390/microorganisms13081837
Yang Y, Wu Z, Zhu L, Han Z, Li J, Fang Q, Zhu G. Quorum-Sensing C12-HSL Drives Antibiotic Resistance Plasmid Transfer via Membrane Remodeling, Oxidative Stress, and RpoS-RMF Crosstalk. Microorganisms. 2025; 13(8):1837. https://doi.org/10.3390/microorganisms13081837
Chicago/Turabian StyleYang, Yang, Ziyan Wu, Li’e Zhu, Zixin Han, Junpeng Li, Qiaoqiao Fang, and Guoqiang Zhu. 2025. "Quorum-Sensing C12-HSL Drives Antibiotic Resistance Plasmid Transfer via Membrane Remodeling, Oxidative Stress, and RpoS-RMF Crosstalk" Microorganisms 13, no. 8: 1837. https://doi.org/10.3390/microorganisms13081837
APA StyleYang, Y., Wu, Z., Zhu, L., Han, Z., Li, J., Fang, Q., & Zhu, G. (2025). Quorum-Sensing C12-HSL Drives Antibiotic Resistance Plasmid Transfer via Membrane Remodeling, Oxidative Stress, and RpoS-RMF Crosstalk. Microorganisms, 13(8), 1837. https://doi.org/10.3390/microorganisms13081837





