The Metabolism of Leuconostoc Genus Decoded by Comparative Genomics
Abstract
1. Introduction
2. Materials and Methods
3. Results and Discussion
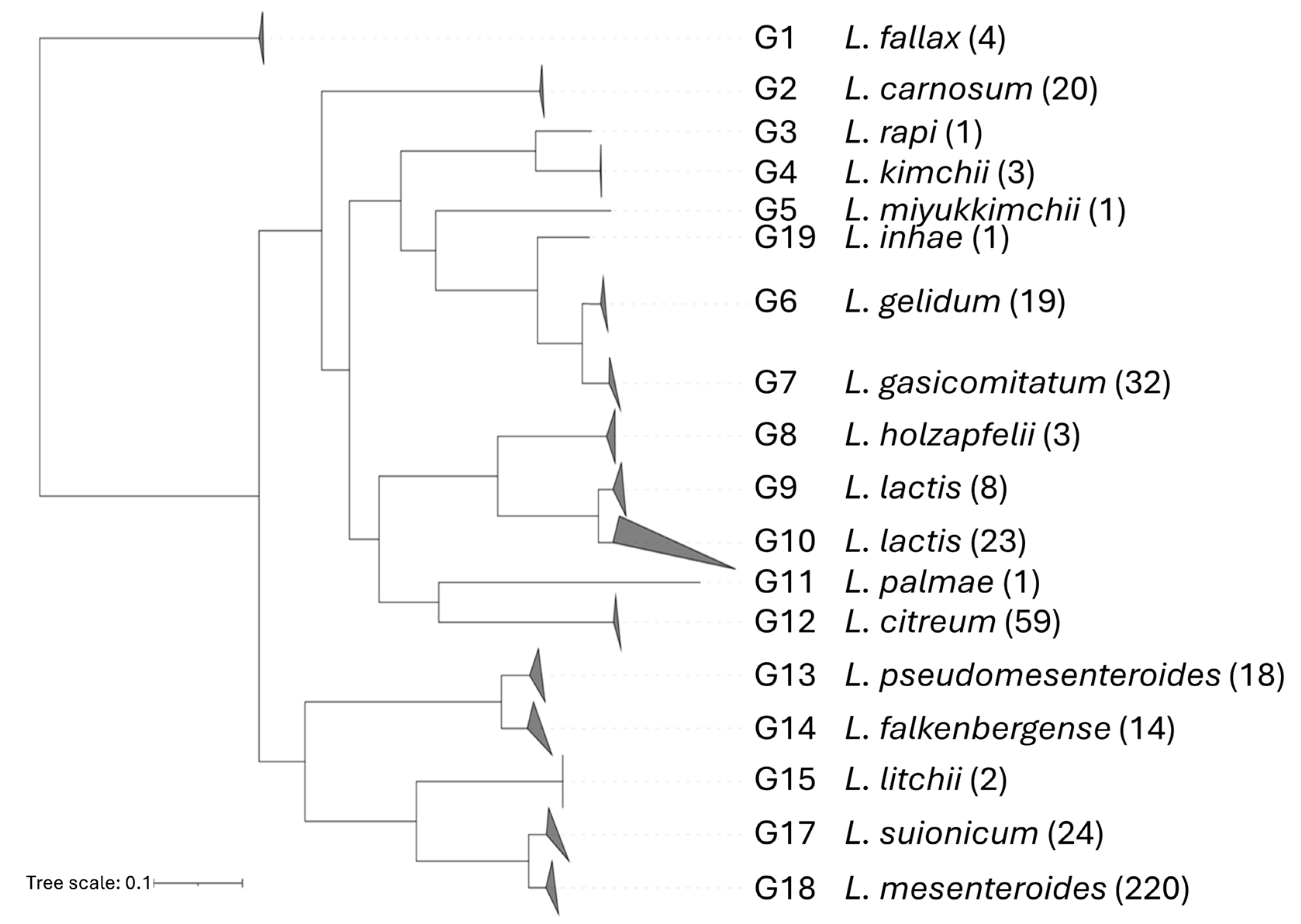
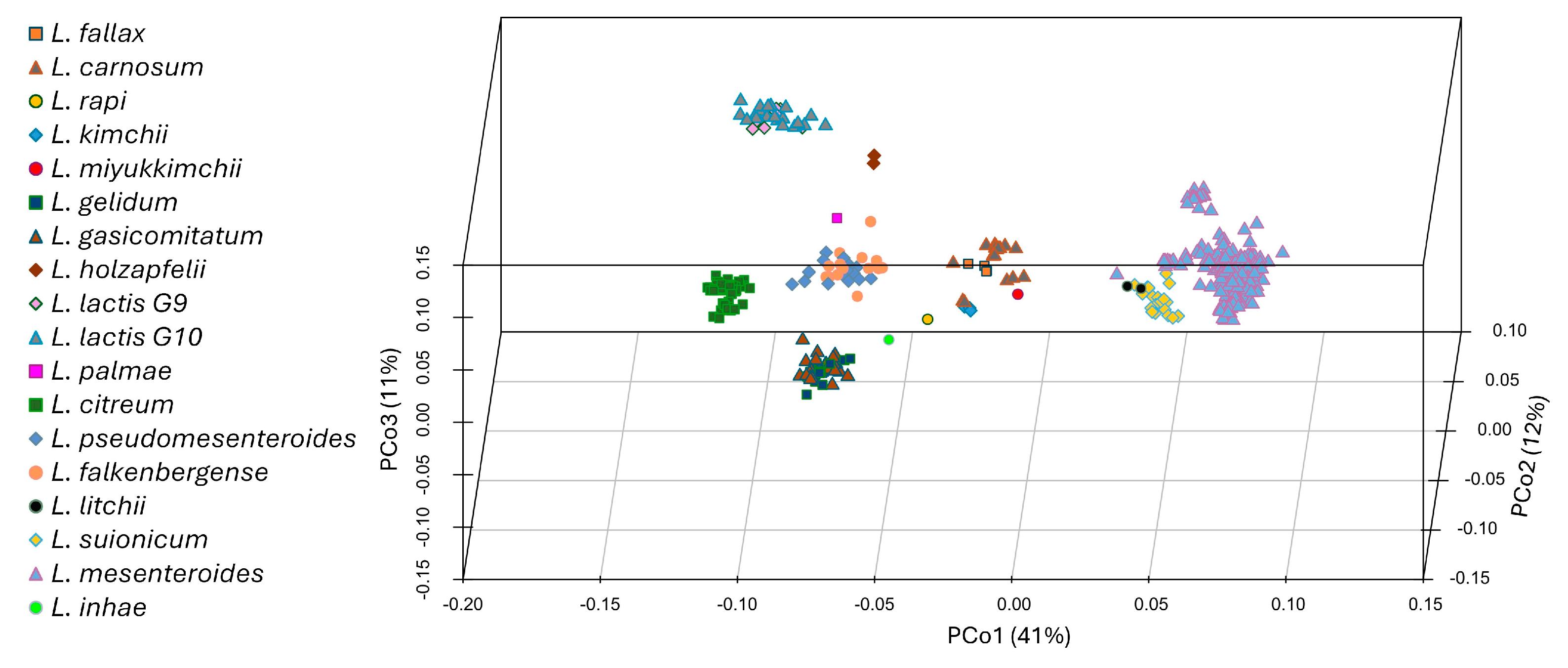
3.1. Phylogenomic Organization
3.2. Functional Analysis
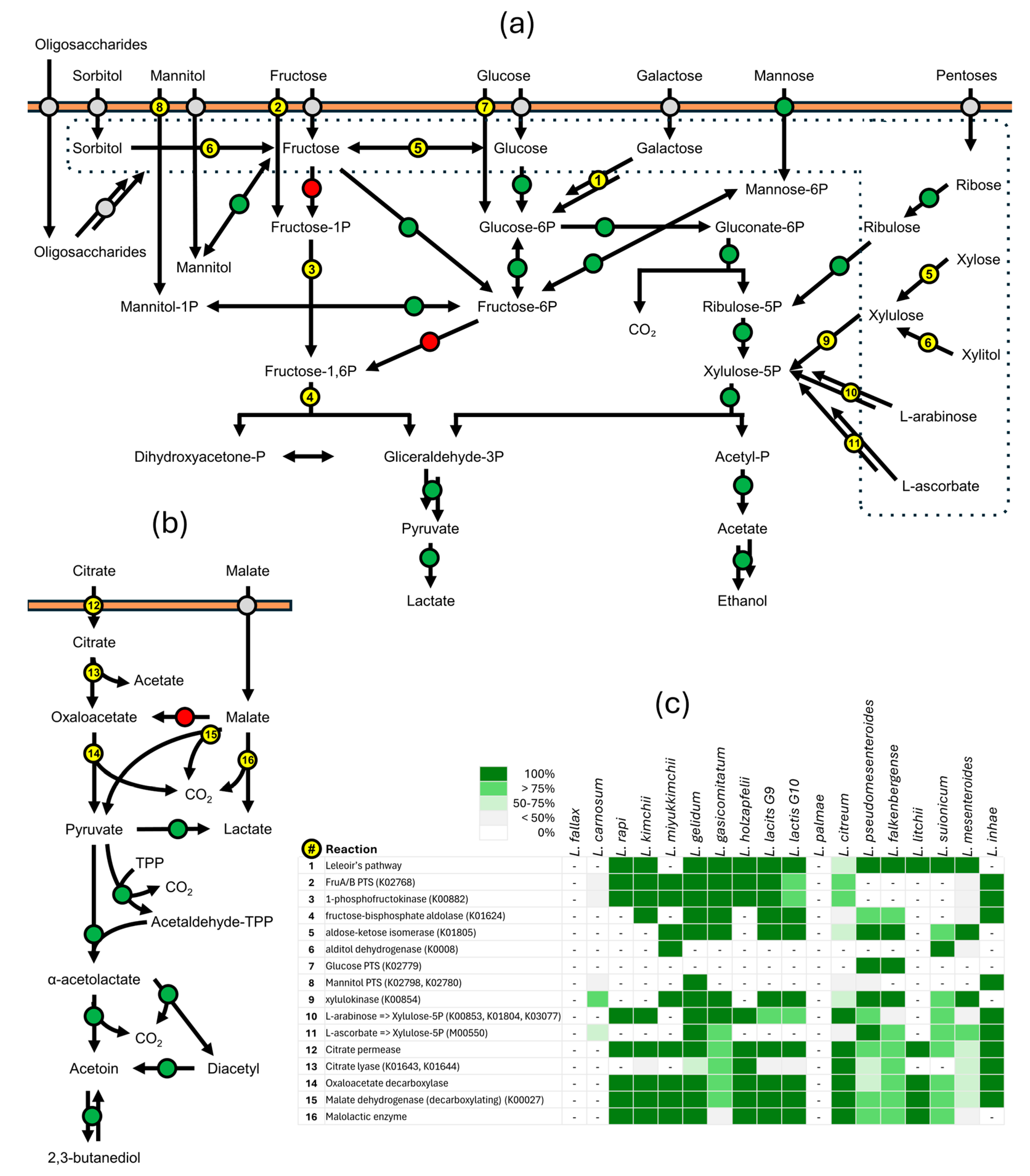
3.3. Metabolic Reconstruction
3.3.1. Central Catabolic Route
3.3.2. Uptake and Fermentation of Sugars
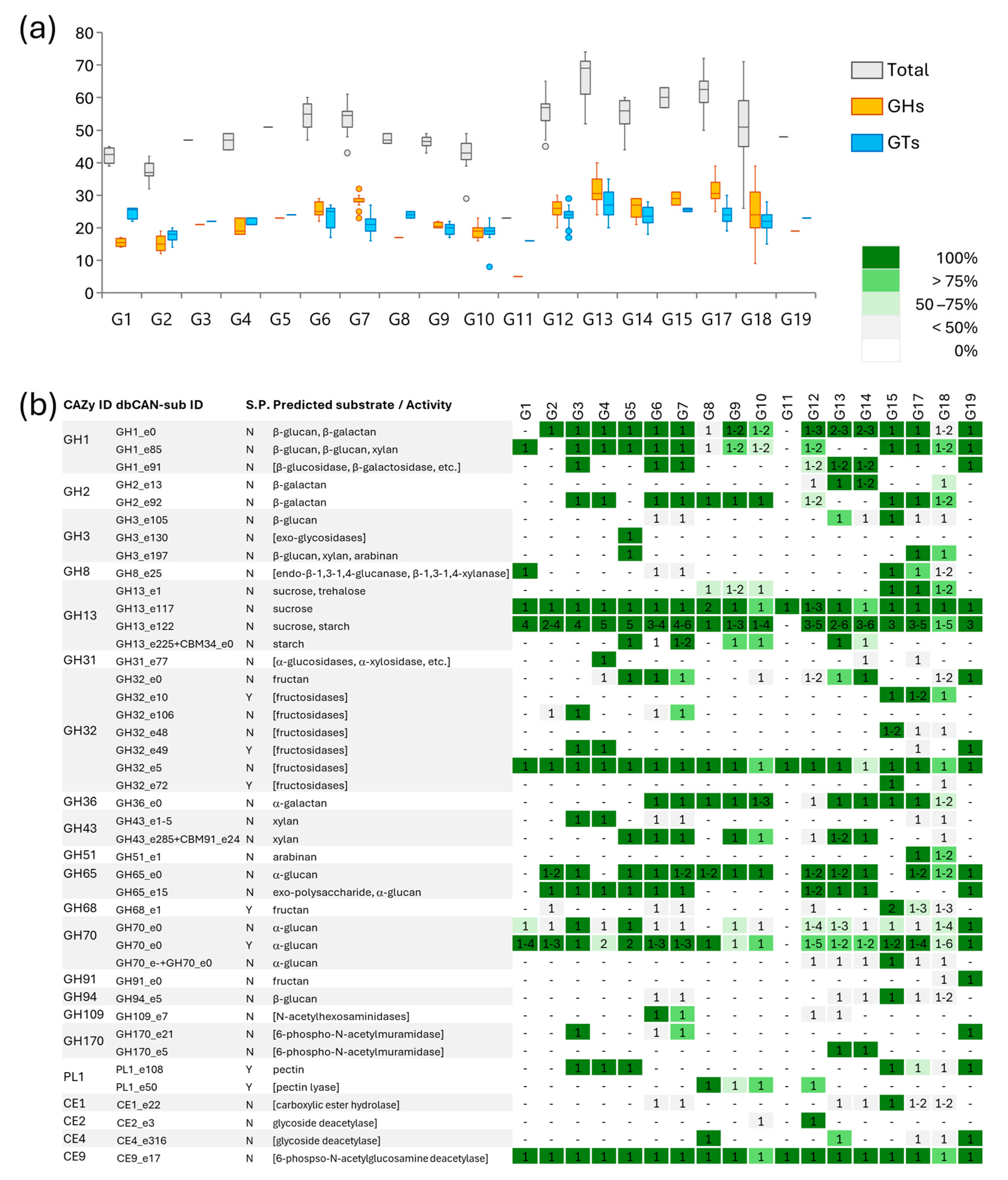
3.3.3. Carbohydrate-Active Enzymes (CAZymes)
3.3.4. Metabolism of Organic Acids
3.3.5. Metabolism of Amino Acids and Cofactors
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Nieminen, T.T.; Säde, E.; Endo, A.; Johansson, P.; Björkroth, J. The Family Leuconostocaceae. In The Prokaryotes, 4th ed.; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 215–240. [Google Scholar] [CrossRef]
- Bello, S.; Rudra, B.; Gupta, R.S. Phylogenomic and comparative genomic analyses of Leuconostocaceae species: Identification of molecular signatures specific for the genera Leuconostoc, Fructobacillus and Oenococcus and proposal for a novel genus Periweissella gen. nov. Int. J. Syst. Evol. Microbiol. 2022, 72, 005284. [Google Scholar] [CrossRef] [PubMed]
- Dellaglio, F.; Dicks, L.M.T.; Torriani, S. The Genus Leuconostoc. In The Genera of Lactic Acid Bacteria; Wood, B.J.B., Holzapfel, W.H., Eds.; Springer: Boston, MA, USA, 1995; Volume 2, pp. 235–278. [Google Scholar] [CrossRef]
- Yu, A.O.; Leveau, J.H.J.; Marco, M.L. Abundance, diversity and plant-specific adaptations of plant-associated lactic acid bacteria. Environ. Microbiol. Rep. 2020, 12, 16–29. [Google Scholar] [CrossRef] [PubMed]
- Candeliere, F.; Raimondi, S.; Spampinato, G.; Tay, M.Y.F.; Amaretti, A.; Schlundt, J.; Rossi, M. Comparative Genomics of Leuconostoc carnosum. Front. Microbiol. 2021, 11, 605127. [Google Scholar] [CrossRef] [PubMed]
- Vedamuthu, E.R. The dairy Leuconostoc: Use in dairy products. J. Dairy Sci. 1994, 77, 2725–2737. [Google Scholar] [CrossRef]
- Raimondi, S.; Spampinato, G.; Candeliere, F.; Amaretti, A.; Brun, P.; Castagliuolo, I.; Rossi, M. Phenotypic Traits and Immunomodulatory Properties of Leuconostoc carnosum Isolated From Meat Products. Front. Microbiol. 2021, 12, 730827. [Google Scholar] [CrossRef] [PubMed]
- Alegria, A.; Delgado, S.; Florez, A.B.; Mayo, B. Identification, typing, and functional characterization of Leuconostoc spp. strains from traditional, starter-free cheeses. Dairy Sci. Technol. 2013, 93, 657–673. [Google Scholar] [CrossRef]
- Samet-Bali, O.; Bellila, A.; Ayadi, M.-A.; Marzouk, B.; Attia, H. A comparison of the physicochemical, microbiological and aromatic composition of Traditional and Industrial Leben in Tunisia. Int. J. Dairy Technol. 2020, 63, 98–104. [Google Scholar] [CrossRef]
- Tsitko, I.; Manninen, J.; Smart, K.; James, S.; Laitila, A. Management of barley-associated bacterial biofilms: A key to improving wort separation. J. Inst. Brew. 2018, 124, 325–335. [Google Scholar] [CrossRef]
- Shin, S.Y.; Han, N.S. Leuconostoc spp. as starters and their beneficial roles in fermented foods. In Beneficial Microorganisms in Food and Nutraceuticals. Microbiology Monographs; Liong, M.T., Ed.; Springer: Cham, Switzerland, 2015; Volume 27, pp. 111–132. [Google Scholar] [CrossRef]
- Venegas-Ortega, M.G.; Flores-Gallegos, A.C.; Martínez-Hernández, J.L.; Aguilar, C.N.; Nevárez-Moorillón, G.V. Production of bioactive peptides from lactic acid bacteria: A sustainable approach for healthier foods. Compr. Rev. Food Sci. Food Saf. 2019, 18, 1039–1051. [Google Scholar] [CrossRef]
- Ahmadi-Ashtiani, H.-R.; Baldisserotto, A.; Cesa, E.; Manfredini, S.; Sedghi Zadeh, H.; Ghafori Gorab, M.; Khanahmadi, M.; Zakizadeh, S.; Buso, P.; Vertuani, S. Microbial Biosurfactants as Key Multifunctional Ingredients for Sustainable Cosmetics. Cosmetics 2020, 7, 46. [Google Scholar] [CrossRef]
- Costa, S.; Summa, D.; Semeraro, B.; Zappaterra, F.; Rugiero, I.; Tamburini, E. Fermentation as a strategy for bio-transforming waste into resources: Lactic acid production from agri-food residues. Fermentation 2021, 7, 3. [Google Scholar] [CrossRef]
- Ogier, J.C.; Casalta, E.; Farrokh, C.; Saihi, A. Safety assessment of dairy microorganisms: The Leuconostoc genus. Int. J. Food Microbiol. 2008, 126, 286–290. [Google Scholar] [CrossRef] [PubMed]
- Ghobrial, M.; Ibrahim, M.; Streit, S.G.; Staiano, P.P.; Seeram, V. A Rare Case of Leuconostoc pseudomesenteroides Bacteremia and Refractory Septic Shock. Cureus 2023, 15, e38312. [Google Scholar] [CrossRef] [PubMed]
- Modaweb, A.; Mansoor, Z.; Alsarhan, A.; Abuhammour, W. A Case of Successfully Treated Central Line-Associated Bloodstream Infection Due to Vancomycin-Resistant Leuconostoc citreum in a Child With Biliary Atresia. Cureus 2022, 14, e21227. [Google Scholar] [CrossRef] [PubMed]
- Hosoya, S.; Kutsuna, S.; Shiojiri, D.; Tamura, S.; Isaka, E.; Wakimoto, Y.; Nomoto, H.; Ohmagari, N. Leuconostoc lactis and Staphylococcus nepalensis Bacteremia, Japan. Emerg. Infect. Dis. 2020, 26, 2283–2285. [Google Scholar] [CrossRef] [PubMed]
- Antunes, A.; Rainey, F.A.; Nobre, M.F.; Schumann, P.; Ferreira, A.M.; Ramos, A.; Santos, H.; da Costa, M.S. Leuconostoc ficulneum sp. nov.; a novel lactic acid bacterium isolated from a ripe fig, and reclassification of Lactobacillus fructosus as Leuconostoc fructosum comb. nov. Int. J. Syst. Evol. Microbiol. 2002, 52 Pt 2, 647–655. [Google Scholar] [CrossRef] [PubMed]
- Leisner, J.J.; Vancanneyt, M.; Van der Meulen, R.; Lefebvre, K.; Engelbeen, K.; Hoste, B.; Laursen, B.G.; Bay, L.; Rusul, G.; De Vuyst, L.; et al. Leuconostoc durionis sp. nov.; a heterofermenter with no detectable gas production from glucose. Int. J. Syst. Evol. Microbiol. 2005, 55 Pt 3, 1267–1270. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Wittouck, S.; Salvetti, E.; Franz, C.M.A.P.; Harris, H.M.B.; Mattarelli, P.; O’Toole, P.W.; Pot, B.; Vandamme, P.; Walter, J.; et al. A taxonomic note on the genus Lactobacillus: Description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int. J. Syst. Evol. Microbiol. 2020, 70, 2782–2858. [Google Scholar] [CrossRef] [PubMed]
- Parte, A.C.; Sardà Carbasse, J.; Meier-Kolthoff, J.P.; Reimer, L.C.; Göker, M. List of prokaryotic names with standing in nomenclature (LPSN) moves to the DSMZ. Int. J. Syst. Evol. Microbiol. 2020, 70, 5607–5612. [Google Scholar] [CrossRef]
- Vandamme, P.; Pot, B.; Gillis, M.; de Vos, P.; Kersters, K.; Swings, J. Polyphasic taxonomy, a consensus approach to bacterial systematics. Microbiol. Rev. 1996, 60, 407–438. [Google Scholar] [CrossRef]
- Stackebrandt, E.; Goebel, B.M. Taxonomic note: A place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in Bacteriology. Int. J. Syst. Bacteriol. 1994, 44, 846–849. [Google Scholar] [CrossRef]
- Parks, D.H.; Imelfort, M.; Skennerton, C.T.; Hugenholtz, P.; Tyson, G.W. CheckM: Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015, 25, 1043–1055. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Page, A.J.; Cummins, C.A.; Hunt, M.; Wong, V.K.; Reuter, S.; Holden, M.T.; Fookes, M.; Falush, D.; Keane, J.A.; Parkhill, J. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 2015, 31, 3691–3693. [Google Scholar] [CrossRef]
- Cantalapiedra, C.P.; Hernández-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. eggNOG-mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Mol. Biol. Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef]
- Huerta-Cepas, J.; Szklarczyk, D.; Heller, D.; Hernández-Plaza, A.; Forslund, S.K.; Cook, H.; Mende, D.R.; Letunic, I.; Rattei, T.; Jensen, L.J.; et al. eggNOG 5.0: A hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 2019, 47, D309–D314. [Google Scholar] [CrossRef] [PubMed]
- Palù, M.; Basile, A.; Zampieri, G.; Treu, L.; Rossi, A.; Morlino, M.S.; Campanaro, S. KEMET—A python tool for KEGG Module evaluation and microbial genome annotation expansion. Comput. Struct. Biotechnol. J. 2022, 20, 1481–1486. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Ge, Q.; Yan, Y.; Zhang, X.; Huang, L.; Yin, Y. dbCAN3: Automated carbohydrate-active enzyme and substrate annotation. Nucleic Acids Res. 2023, 51, W115–W121. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; O’hara, R.B.; Simpson, G.L.; Solymos, P.; Stevens, M.H.H.; Wagner, H.; Barbour, M.; et al. vegan: Community Ecology Package. R Package Version 2.5-6. 2019. Available online: https://CRAN.R-project.org/package=vegan (accessed on 1 December 2023).
- Paradis, E.; Claude, J.; Strimmer, K. APE: Analyses of Phylogenetics and Evolution in R language. Bioinformatics 2004, 20, 289–290. [Google Scholar] [CrossRef]
- Gosselin, S.; Fullmer, M.S.; Feng, Y.; Gogarten, J.P. Improving Phylogenies Based on Average Nucleotide Identity, Incorporating Saturation Correction and Nonparametric Bootstrap Support. Syst. Biol. 2022, 71, 396–409. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Desper, R.; Gascuel, O. Fast and accurate phylogeny reconstruction algorithms based on the minimum-evolution principle. J. Comput. Biol. 2002, 9, 687–705. [Google Scholar] [CrossRef] [PubMed]
- Schliep, K.P. Phangorn: Phylogenetic analysis in R. Bioinformatics 2011, 27, 592–593. [Google Scholar] [CrossRef] [PubMed]
- Huson, D.H.; Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 2006, 23, 254–267. [Google Scholar] [CrossRef] [PubMed]
- Bandelt, H.J.; Dress, A.W. Split decomposition: A new and useful approach to phylogenetic analysis of distance data. Mol. Phylogenet. Evol. 1992, 1, 242–252. [Google Scholar] [CrossRef] [PubMed]
- Raimondi, S.; Candeliere, F.; Amaretti, A.; Costa, S.; Vertuani, S.; Spampinato, G.; Rossi, M. Phylogenomic analysis of the genus Leuconostoc. Front. Microbiol. 2022, 13, 897656. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wu, J.; Lv, M.; Shao, Z.; Hungwe, M.; Wang, J.; Bai, X.; Xie, J.; Wang, Y.; Geng, W. Metabolism Characteristics of Lactic Acid Bacteria and the Expanding Applications in Food Industry. Front. Bioeng. Biotechnol. 2021, 9, 612285. [Google Scholar] [CrossRef] [PubMed]
- Gänzle, M.G. Lactic metabolism revisited: Metabolism of lactic acid bacteria in food fermentations and food spoilage. Curr. Opinin. Food Sci. 2015, 2, 106–117. [Google Scholar] [CrossRef]
- Cogan, T.M.; Jordan, K.N. Metabolism of Leuconostoc bacteria. J. Dairy Sci. 1994, 77, 2704–2717. [Google Scholar] [CrossRef]
- Ren, Q.; Kang, K.H.; Paulsen, I.T. TransportDB: A relational database of cellular membrane transport systems. Nucleic Acids Res. 2004, 32 (Suppl. S1), D284–D288. [Google Scholar] [CrossRef]
- Reque, P.M.; Pinilla, C.M.B.; Tinello, F.; Corich, V.; Lante, A.; Giacomini, A.; Brandelli, A. Biochemical and functional properties of wheat middlings bioprocessed by lactic acid bacteria. J. Food Biochem. 2020, 44, e13262. [Google Scholar] [CrossRef] [PubMed]
- Zafar, H.; Saier, M.H., Jr. Comparative Genomics of the Transport Proteins of Ten Lactobacillus Strains. Genes 2020, 11, 1234. [Google Scholar] [CrossRef] [PubMed]
- Mende, S.; Rohm, H.; Jaros, D. Influence of exopolysaccharides on the structure, texture, stability and sensory properties of yoghurt and related products. Int. Dairy J. 2016, 52, 57–71. [Google Scholar] [CrossRef]
- Kumar, S.; Bansal, K.; Sethi, S.K. Comparative genomics analysis of genus Leuconostoc resolves its taxonomy and elucidates its biotechnological importance. Food Microbiol. 2022, 106, 104039. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Li, N.; Zhao, S.; Zhang, C.; Qiao, N.; Duan, H.; Xiao, Y.; Yan, B.; Zhao, J.; Tian, F.; et al. Integrated phenotypic–genotypic analysis of Latilactobacillus sakei from different niches. Foods 2021, 10, 1717. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Sharma, N.; Gupta, D.; Lee, H.J.; Park, Y.S. Comparative genome analysis of four Leuconostoc strains with a focus on carbohydrate-active enzymes and oligosaccharide utilization pathways. Comput. Struct. Biotechnol. J. 2022, 20, 4771–4785. [Google Scholar] [CrossRef] [PubMed]
- Frantzen, C.; Kot, W.; Pedersen, T.B.; Ardö, Y.; Broadbent, J.R.; Neve, H.; Hansen, L.H.; Dal Bello, F.; Østlie, H.M.; Kleppen, H.P.; et al. Genomic characterization of dairy associated Leuconostoc species and diversity of leuconostocs in undefined mixed mesophilic starter cultures. Front. Microbiol. 2017, 8, 132. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Feng, F.; Zhou, Q.; Zhao, F.; Du, R.; Zhou, Z.; Han, Y. Isolation, purification and characterization of exopolysaccharide produced by Leuconostoc pseudomesenteroides YF32 from soybean paste. Int. J. Biol. Macromol. 2018, 114, 529–535. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Yan, D.; Liu, Y.; Luo, X.; Li, Y.; Cao, C.; Li, M.; Han, Q.; Wang, C.; Wu, R.; et al. Purification, Structural Characteristics, and Biological Activities of Exopolysaccharide Isolated from Leuconostoc mesenteroides SN-8. Front. Microbiol. 2021, 12, 644226. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Du, R.; Qiao, X.; Zhao, B.; Zhou, Z.; Han, Y. Optimization and characterization of exopolysaccharides with a highly branched structure extracted from Leuconostoc citreum B-2. Int. J. Biol. Macromol. 2020, 142, 73–84. [Google Scholar] [CrossRef]
- Wu, Y.; Gu, C.T. Leuconostoc falkenbergense sp. nov.; isolated from a lactic culture, fermentating string beans and traditional yogurt. Int. J. Syst. Evol. Microbiol. 2021, 71. [Google Scholar] [CrossRef] [PubMed]
- Jay, J.M.; Rivers, G.M.; Boisvert, W.E. Antimicrobial Properties of α-Dicarbonyl and Related Compounds. J. Food. Prot. 1983, 46, 325–329. [Google Scholar] [CrossRef] [PubMed]
- García Quintans, N.; Blancato, V.; Repizo, G.; Magni, C.; López, P. Citrate metabolism and aroma compound production in lactic acid bacteria. In Molecular Aspects of Lactic Acid Bacteria for Traditional and New Applications; Mayo, B., López, P., Pérez-Martínez, G., Eds.; Research Signpost: Kerala, India, 2008; Chapter 3; pp. 65–88. [Google Scholar]
- Laëtitia, G.; Pascal, D.; Yann, D. The Citrate Metabolism in Homo- and Heterofermentative LAB: A Selective Means of Becoming Dominant over Other Microorganisms in Complex Ecosystems. Food Nutr. Sci. 2014, 5, 953–969. [Google Scholar] [CrossRef]
- Prusova, B.; Licek, J.; Kumsta, M.; Baron, M.; Sochor, J. Effect of new methods for inhibiting malolactic fermentation on the analytical and sensory parameters of wines. Fermentation 2024, 10, 122. [Google Scholar] [CrossRef]
- Schümann, C.; Michlmayr, H.; Del Hierro, A.M.; Kulbe, K.D.; Jiranek, V.; Eder, R.; Nguyen, T.H. Malolactic enzyme from Oenococcus oeni: Heterologous expression in Escherichia coli and biochemical characterization. Bioengineered 2013, 4, 147–152. [Google Scholar] [CrossRef]
- Montaño, A.; Sánchez, A.H.; Casado, F.J.; Beato, V.M.; de Castro, A. Degradation of ascorbic acid and potassium sorbate by different Lactobacillus species isolated from packed green olives. Food Microbiol. 2013, 34, 7–11. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Candeliere, F.; Sola, L.; Busi, E.; Rossi, M.; Amaretti, A.; Raimondi, S. The Metabolism of Leuconostoc Genus Decoded by Comparative Genomics. Microorganisms 2024, 12, 1487. https://doi.org/10.3390/microorganisms12071487
Candeliere F, Sola L, Busi E, Rossi M, Amaretti A, Raimondi S. The Metabolism of Leuconostoc Genus Decoded by Comparative Genomics. Microorganisms. 2024; 12(7):1487. https://doi.org/10.3390/microorganisms12071487
Chicago/Turabian StyleCandeliere, Francesco, Laura Sola, Enrico Busi, Maddalena Rossi, Alberto Amaretti, and Stefano Raimondi. 2024. "The Metabolism of Leuconostoc Genus Decoded by Comparative Genomics" Microorganisms 12, no. 7: 1487. https://doi.org/10.3390/microorganisms12071487
APA StyleCandeliere, F., Sola, L., Busi, E., Rossi, M., Amaretti, A., & Raimondi, S. (2024). The Metabolism of Leuconostoc Genus Decoded by Comparative Genomics. Microorganisms, 12(7), 1487. https://doi.org/10.3390/microorganisms12071487






