Antimicrobial Resistance and Genomic Characteristics of Escherichia coli Strains Isolated from the Poultry Industry in Henan Province, China
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Bacterial Isolation
2.2. Antimicrobial Susceptibility Testing
2.3. DNA Extraction, WGS, and Bioinformatic Analysis
2.4. Statistical Analysis
3. Results
3.1. Prevalence of E. coli
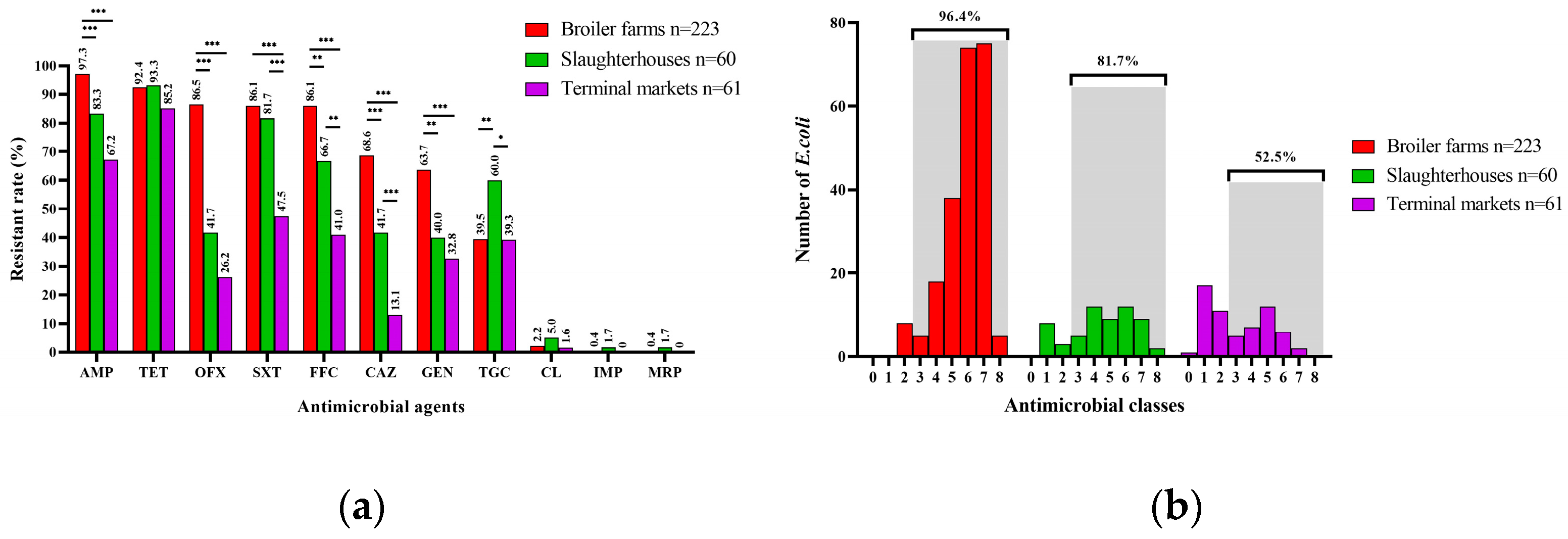
3.2. Antimicrobial Resistance Phenotypes of E. coli
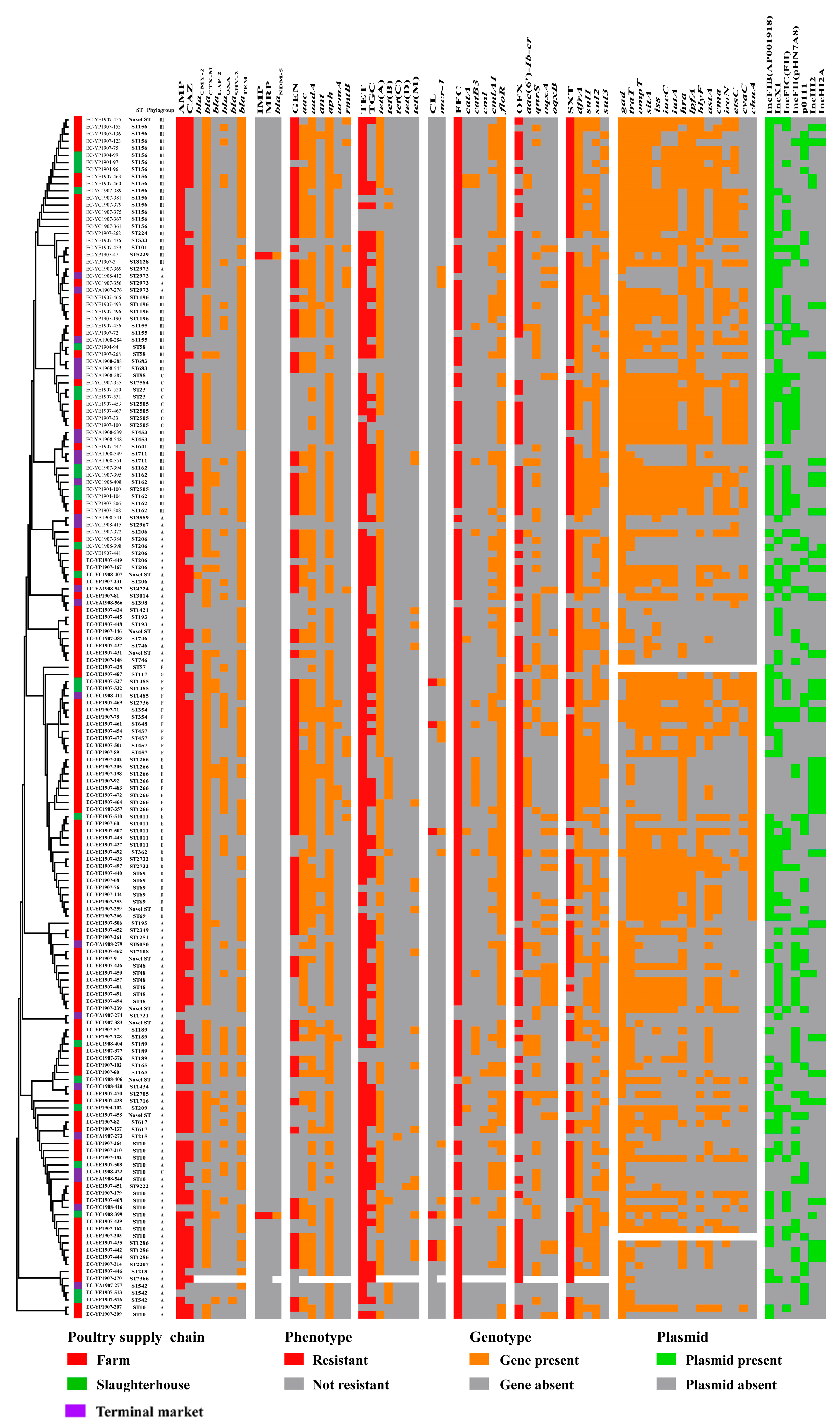
3.3. Sequence Type, Phylogenetic Group and O-Serogroup
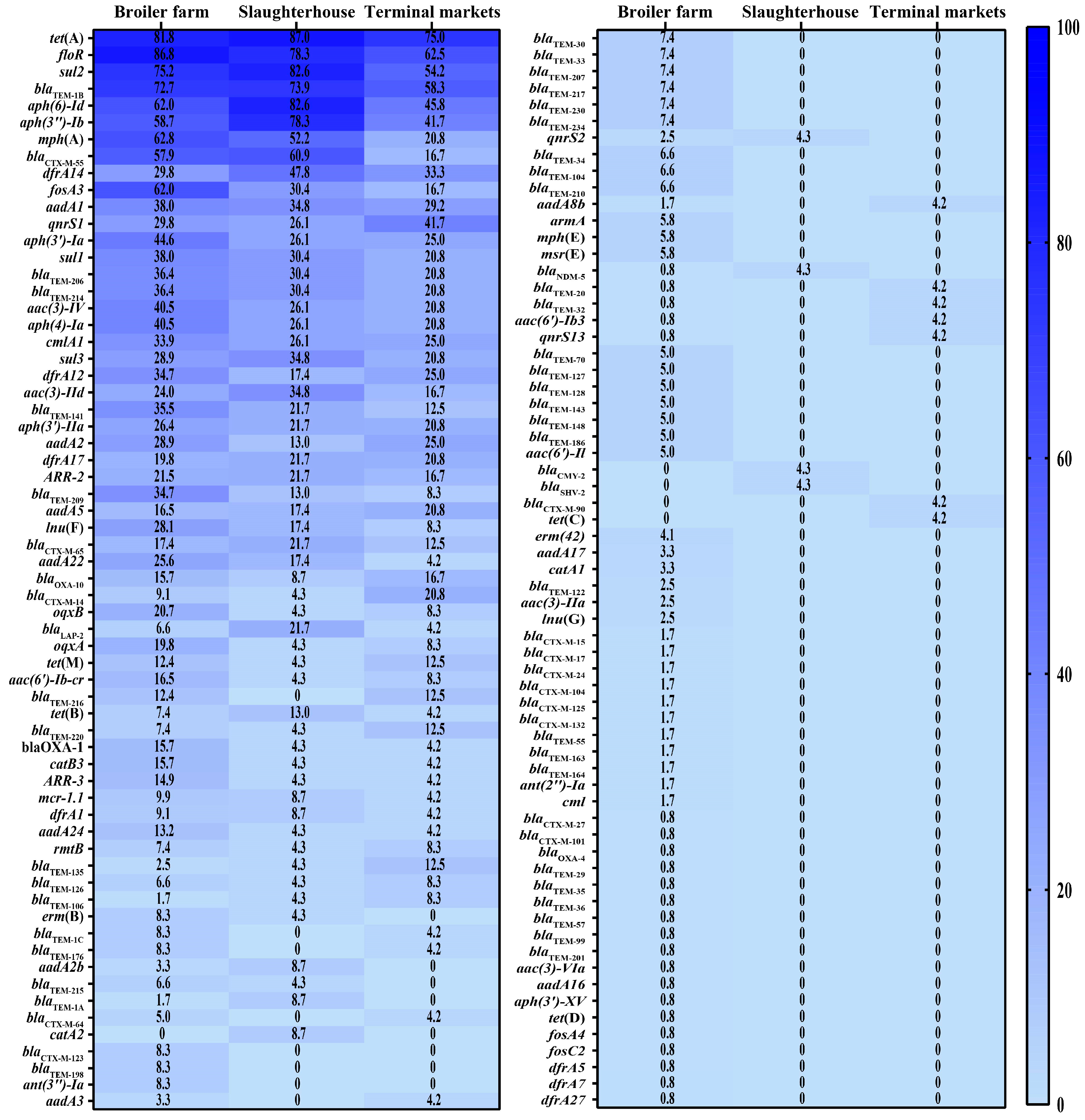
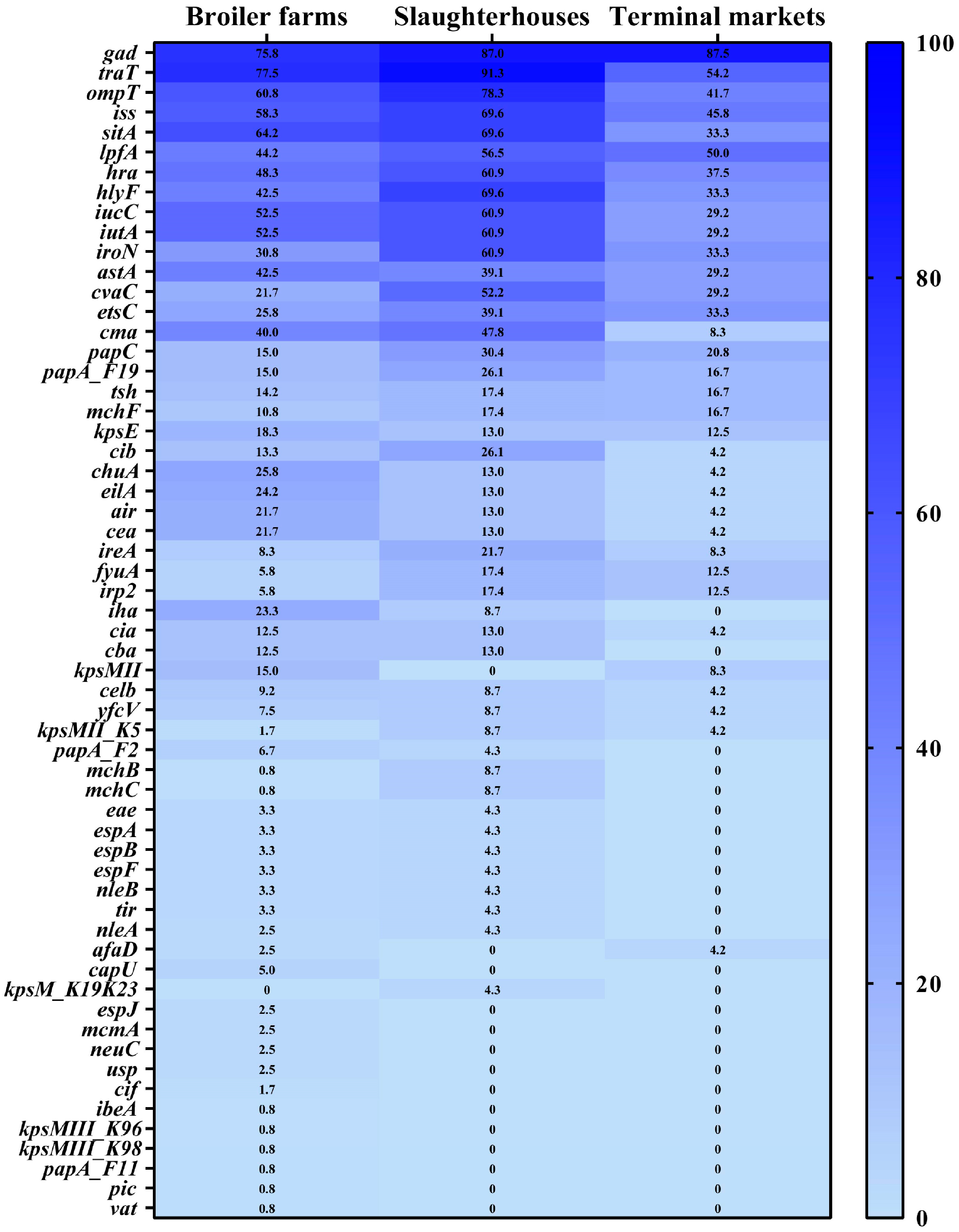
3.4. Prevalence of Antimicrobial Resistance Genes and Virulence Factor Genes
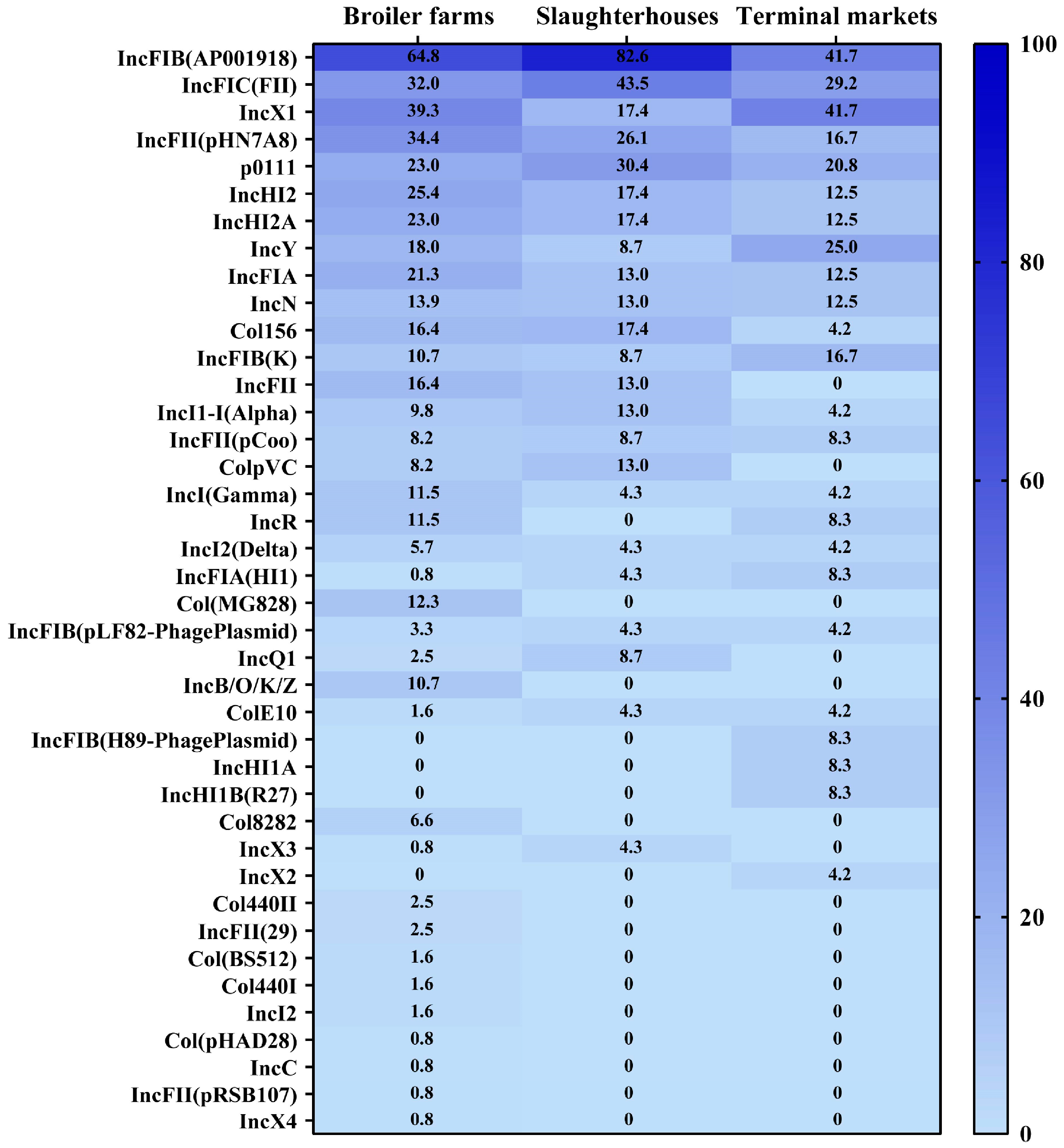
3.5. Prevalence and Distribution of Plasmid Profiles
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Van Boeckel, T.P.; Brower, C.; Gilbert, M.; Grenfell, B.T.; Levin, S.A.; Robinson, T.P.; Teillant, A.; Laxminarayan, R. Global trends in antimicrobial use in food animals. Proc. Natl. Acad. Sci. USA 2015, 112, 5649–5654. [Google Scholar] [CrossRef] [PubMed]
- CDC. Antibiotic Resistance Threats in the United States, 2019; Department of Health and Human Services, CDC: Atlanta, GA, USA, 2019. [Google Scholar]
- Kaper, J.B.; Nataro, J.P.; Mobley, H.L.T. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2004, 2, 123–140. [Google Scholar] [CrossRef] [PubMed]
- Gomes, T.A.T.; Elias, W.P.; Scaletsky, I.C.A.; Guth, B.E.C.; Rodrigues, J.F.; Piazza, R.M.F.; Ferreira, L.C.S.; Martinez, M.B. Diarrheagenic Escherichia coli. Braz. J. Microbiol. 2016, 47, 3–30. [Google Scholar] [CrossRef] [PubMed]
- Biran, D.; Ron, E.Z. Extraintestinal pathogenic Escherichia coli. Curr. Top. Microbiol. Immunol. 2018, 416, 149–161. [Google Scholar] [CrossRef] [PubMed]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile genetic elements associated with antimicrobial resistance. Clin. Microbiol. Rev. 2018, 31, e00088-17. [Google Scholar] [CrossRef] [PubMed]
- Brisola, M.C.; Crecencio, R.B.; Bitner, D.S.; Frigo, A.; Rampazzo, L.; Stefani, L.M.; Faria, G.A. Escherichia coli used as a biomarker of antimicrobial resistance in pig farms of Southern Brazil. Sci. Total Environ. 2019, 647, 362–368. [Google Scholar] [CrossRef] [PubMed]
- Poirel, L.; Madec, J.Y.; Lupo, A.; Schink, A.K.; Kieffer, N.; Nordmann, P.; Schwarz, S. Antimicrobial resistance in Escherichia coli. Microbiol. Spectr. 2018, 6, 10-1128. [Google Scholar] [CrossRef] [PubMed]
- Jenkins, C.; Dallman, T.J.; Grant, K.A. Impact of whole genome sequencing on the investigation of food-borne outbreaks of Shiga toxin-producing Escherichia coli serogroup O157:H7, England, 2013 to 2017. Eurosurveillance 2019, 24, 84–90. [Google Scholar] [CrossRef]
- Peng, Z.; Hu, Z.Z.; Li, Z.G.; Zhang, X.X.; Jia, C.Y.; Li, T.Z.; Dai, M.H.; Tan, C.; Xu, Z.F.; Wu, B.; et al. Antimicrobial resistance and population genomics of multidrug-resistant Escherichia coli in pig farms in mainland China. Nat. Commun. 2022, 13, 1116. [Google Scholar] [CrossRef]
- Ghunaim, H.; Abu-Madi, M.A.; Kariyawasam, S. Advances in vaccination against avian pathogenic Escherichia coli respiratory disease: Potentials and limitations. Vet. Microbiol. 2014, 172, 13–22. [Google Scholar] [CrossRef]
- Liu, Z.; Wang, K.; Zhang, Y.; Xia, L.; Zhao, L.; Guo, C.; Liu, X.; Qin, L.; Hao, Z. High prevalence and diversity characteristics of blaNDM, mcr, and blaESBLs harboring multidrug-resistant Escherichia coli from chicken, pig, and cattle in China. Front. Cell. Infect. Microbiol. 2022, 11, 755545. [Google Scholar] [CrossRef]
- Hussain, H.I.; Iqbal, Z.; Iqbal, M.; Kuang, X.H.; Wang, Y.L.; Yang, L.Q.; Ihsan, A.; Aqib, A.I.; Kaleem, Q.M.; Gu, Y.F.; et al. Coexistence of virulence and β-lactamase genes in avian pathogenic Escherichia coli. Microb. Pathog. 2022, 163, 105389. [Google Scholar] [CrossRef]
- Rui, Y.P.; Qiu, G. Analysis of gut microbial communities and resistance genes in pigs and chickens in Central China. Animals 2022, 12, 3404. [Google Scholar] [CrossRef]
- Hu, Z.Z.; Peng, Z.; Zhang, X.X.; Li, Z.G.; Jia, C.Y.; Li, X.S.; Lv, Y.J.; Tan, C.; Chen, H.C.; Wang, X.R. Prevalence and molecular characterization of antimicrobial-resistant Escherichia coli in pig farms, slaughterhouses, and terminal markets in Henan province of China. Foodborne Pathog. Dis. 2021, 18, 733–743. [Google Scholar] [CrossRef]
- Sabat, G.; Rose, P.; Hickey, W.J.; Harkin, J.M. Selective and sensitive method for PCR amplification of Escherichia coli 16S rRNA genes in soil. Appl. Environ. Microbiol. 2000, 66, 844–849. [Google Scholar] [CrossRef]
- Wirth, T.; Falush, D.; Lan, R.T.; Colles, F.; Mensa, P.; Wieler, L.H.; Karch, H.; Reeves, P.R.; Maiden, M.C.J.; Ochman, H.; et al. Sex and virulence in Escherichia coli: An evolutionary perspective. Mol. Microbiol. 2006, 60, 1136–1151. [Google Scholar] [CrossRef]
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 28th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2018. [Google Scholar]
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 30th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2020. [Google Scholar]
- CLSI. Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated from Animals, 5th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2020. [Google Scholar]
- EUCAST. Breakpoint Tables for Interpretation of MICs and Zone Diameters. Version 10.0. Available online: https://www.eucast.org/clinical_breakpoints (accessed on 1 January 2020).
- EUCAST. Routine and Extended Internal Quality Control for MIC Determination and Disk Diffusion as Recommended by EUCAST. Version 10.0. Available online: https://www.eucast.org/ast_of_bacteria/quality_control (accessed on 1 January 2020).
- Larsen, M.V.; Cosentino, S.; Rasmussen, S.; Friis, C.; Hasman, H.; Marvig, R.L.; Jelsbak, L.; Sicheritz-Pontén, T.; Ussery, D.W.; Aarestrup, F.M.; et al. Multilocus sequence typing of total-genome-sequenced bacteria. J. Clin. Microbiol. 2012, 50, 1355–1361. [Google Scholar] [CrossRef]
- Joensen, K.G.; Tetzschner, A.M.M.; Iguchi, A.; Aarestrup, F.M.; Scheutz, F. Rapid and easy In Silico serotyping of Escherichia coli isolates by use of whole-genome sequencing data. J. Clin. Microbiol. 2015, 53, 2410–2426. [Google Scholar] [CrossRef] [PubMed]
- Beghain, J.; Bridier-Nahmias, A.; Le Nagard, H.; Denamur, E.; Clermont, O. ClermonTyping: An easy-to-use and accurate in silico method for Escherichia genus strain phylotyping. Microb. Genom. 2018, 4, e000192. [Google Scholar] [CrossRef] [PubMed]
- Bortolaia, V.; Kaas, R.S.; Ruppe, E.; Roberts, M.C.; Schwarz, S.; Cattoir, V.; Philippon, A.; Allesoe, R.L.; Rebelo, A.R.; Florensa, A.F.; et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 2020, 75, 3491–3500. [Google Scholar] [CrossRef] [PubMed]
- Joensen, K.G.; Scheutz, F.; Lund, O.; Hasman, H.; Kaas, R.S.; Nielsen, E.M.; Aarestrup, F.M. Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J. Clin. Microbiol. 2014, 52, 1501–1510. [Google Scholar] [CrossRef]
- Malberg Tetzschner, A.M.; Johnson, J.R.; Johnston, B.D.; Lund, O.; Scheutz, F.; Dekker, J.P. In Silico genotyping of Escherichia coli isolates for extraintestinal virulence genes by use of whole-genome sequencing data. J. Clin. Microbiol. 2020, 58, e01269-20. [Google Scholar] [CrossRef] [PubMed]
- Carattoli, A.; Zankari, E.; Garcia-Fernandez, A.; Larsen, M.V.; Lund, O.; Villa, L.; Aarestrup, F.M.; Hasman, H. Detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 2014, 58, 3895–3903. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Jiu, Y.G.; Meng, X.R.; Hong, X.; Huang, Q.; Wang, C.F.; Chen, Z.; Zhao, L.J.; Liu, X.Y.; Lu, Y.C.; Li, S.W. Prevalence and characterization of Salmonella in three typical commercial pig abattoirs in Wuhan, China. Foodborne Pathog. Dis. 2020, 17, 620–627. [Google Scholar] [CrossRef] [PubMed]
- Agerso, Y.; Aarestrup, F.M.; Pedersen, K.; Seyfarth, A.M.; Struve, T.; Hasman, H. Prevalence of extended-spectrum cephalosporinase (ESC)-producing Escherichia coli in Danish slaughter pigs and retail meat identified by selective enrichment and association with cephalosporin usage. J. Antimicrob. Chemother. 2012, 67, 582–588. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Xi, M.L.; Lv, X.Y.; Xu, Y.F.; Feng, Y.Q.; Li, Q.; Yang, Q.N.; Xia, X.D. Presence and antimicrobial susceptibility of Escherichia coli recovered from retail chicken in China. J. Food Prot. 2014, 77, 1773–1777. [Google Scholar] [CrossRef]
- Vincent, C.; Boerlin, P.; Daignault, D.; Dozois, C.M.; Dutil, L.; Galanakis, C.; Reid-Smith, R.J.; Tellier, P.P.; Tellis, P.A.; Ziebell, K.; et al. Food reservoir for Escherichia coli causing urinary tract infections. Emerg. Infect. Dis. 2010, 16, 88–95. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, N.M.; Johnson, J.R.; Johnston, B.; Curtiss, R.; Mellataa, M. Zoonotic potential of Escherichia coli isolates from retail chicken meat products and eggs. Appl. Environ. Microbiol. 2015, 81, 1177–1187. [Google Scholar] [CrossRef]
- Jiu, Y.G.; Zhu, S.; Khan, S.B.; Sun, M.Z.; Zou, G.; Meng, X.R.; Wu, B.; Zhou, R.; Li, S.W. Phenotypic and genotypic resistance of Salmonella isolates from healthy and diseased pigs in China during 2008–2015. Microb. Drug Resist. 2017, 23, 651–659. [Google Scholar] [CrossRef]
- Peng, Z.; Li, X.S.; Hu, Z.Z.; Li, Z.G.; Lv, Y.J.; Lei, M.G.; Wu, B.; Chen, H.C.; Wang, X.R. Characteristics of carbapenem-resistant and colistin-resistant Escherichia coli co-producing NDM-1 and MCR-1 from pig farms in China. Microorganisms 2019, 7, 482. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.F.; Yang, L.; Lu, L.F.; Wang, K.Y.; Li, J.H.; Li, P.H.; Liu, Y.Q.; Liu, X.L.; Li, P.; Song, H.B. Genomic features of an Escherichia coli ST156 strain harboring chromosomelocated mcr-1 and plasmid-mediated blaNDM-5. Infect. Genet. Evol. 2020, 85, 104499. [Google Scholar] [CrossRef]
- Manges, A.R.; Geum, H.M.; Guo, A.; Edens, T.J.; Fibke, C.D.; Pitout, J.D.D. Global extraintestinal pathogenic Escherichia coli (ExPEC) lineages. Clin. Microbiol. Rev. 2019, 32, e00135-18. [Google Scholar] [CrossRef]
- Blaak, H.; van Hoek, A.H.A.M.; Hamidjaja, R.A.; van der Plaats, R.Q.J.; Kerkhof-de Heer, L.; Husman, A.M.D.; Schets, F.M. Distribution, numbers, and diversity of ESBL-producing E. coli in the poultry farm environment. PLoS ONE 2015, 10, e0135402. [Google Scholar] [CrossRef]
- Denamur, E.; Clermont, O.; Bonacorsi, S.; Gordon, D. The population genetics of pathogenic Escherichia coli. Nat. Rev. Microbiol. 2020, 19, 37–54. [Google Scholar] [CrossRef]
- Ethelberg, S.; Smith, B.; Torpdahl, M.; Lisby, M.; Boel, J.; Jensen, T.; Nielsen, E.M.; Molbak, K. Outbreak of non-O157 Shiga toxin–producing Escherichia coli infection from consumption of beef sausage. Clin. Infect. Dis. 2009, 48, E78–E81. [Google Scholar] [CrossRef] [PubMed]
- Beatty, M.E.; Bopp, C.A.; Wells, J.G.; Greene, K.D.; Puhr, N.D.; Mintz, E.D. Enterotoxin-producing Escherichia coli O169:H41, United States. Emerg. Infect. Dis. 2004, 10, 518–521. [Google Scholar] [CrossRef] [PubMed]
- Mellmann, A.; Bielaszewska, M.; Köck, R.; Friedrich, A.W.; Fruth, A.; Middendorf, B.; Harmsen, D.; Schmidt, M.A.; Karch, H. Analysis of collection of hemolytic uremic syndrome-associated enterohemorrhagic Escherichia coli Emerg. Infect. Dis. 2008, 14, 1287–1290. [Google Scholar] [CrossRef]
- Adams, N.L.; Byrne, L.; Smith, G.A.; Elson, R.; Harris, J.P.; Salmon, R.; Smith, R.; O’Brien, S.J.; Adak, G.K.; Jenkins, C. Shiga toxin–producing Escherichia coli O157, England and Wales, 1983–2012. Emerg. Infect. Dis. 2016, 22, 590–597. [Google Scholar] [CrossRef]
- Zbikowska, K.; Michalczuk, M.; Dolka, B. The use of bacteriophages in the poultry Industry. Animals 2020, 10, 872. [Google Scholar] [CrossRef]
- Chang, Y. Bacteriophage-derived endolysins applied as potent biocontrol agents to enhance food safety. Microorganisms 2020, 8, 724. [Google Scholar] [CrossRef] [PubMed]
- Sarowska, J.; Futoma-Koloch, B.; Jama-Kmiecik, A.; Frej-Madrzak, M.; Ksiazczyk, M.; Bugla-Ploskonska, G.; Choroszy-Krol, I. Virulence factors, prevalence and potential transmission of extraintestinal pathogenic Escherichia coli isolated from different sources: Recent reports. Gut Pathog. 2019, 11, 10. [Google Scholar] [CrossRef] [PubMed]
- Desvaux, M.; Dalmasso, G.; Beyrouthy, R.; Barnich, N.; Delmas, J.; Bonnet, R. Pathogenicity factors of genomic islands in intestinal and extraintestinal Escherichia coli. Front. Microbiol. 2020, 11, 2065. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Guo, X.Y.; Yan, Z.; Wang, W.; Chen, B.; Ge, F.; Ye, B.P. A comprehensive analysis on spread and distribution characteristic of antibiotic resistance genes in livestock farms of southeastern China. PLoS ONE 2016, 11, e0156889. [Google Scholar] [CrossRef]
- Azad, M.A.R.A.; Rahman, M.M.; Amin, R.; Begum, M.I.A.; Fries, R.; Husna, A.; Khairalla, A.S.; Badruzzaman, A.T.M.; El Zowalaty, M.E.; Lampang, K.N.; et al. Susceptibility and multidrug resistance patterns of Escherichia coli isolated from cloacal swabs of live broiler chickens in Bangladesh. Pathogens 2019, 8, 118. [Google Scholar] [CrossRef]
- Carattoli, A. Resistance plasmid families in Enterobacteriaceae. Antimicrob. Agents Chemother. 2009, 53, 2227–2238. [Google Scholar] [CrossRef]
- Rozwandowicz, M.; Brouwer, M.S.M.; Fischer, J.; Wagenaar, J.A.; Gonzalez-Zorn, B.; Guerra, B.; Mevius, D.J.; Hordijk, J. Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J. Antimicrob. Chemother. 2018, 73, 1121–1137. [Google Scholar] [CrossRef]
- Buckner, M.M.C.; Ciusa, M.L.; Piddock, L.J.V. Strategies to combat antimicrobial resistance: Anti-plasmid and plasmid curing. FEMS Microbiol. Rev. 2018, 42, 781–804. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Z.; Jia, C.; Hu, Z.; Jin, Y.; Li, T.; Zhang, X.; Peng, Z.; Yang, R.; Chen, H.; Wang, X. Antimicrobial Resistance and Genomic Characteristics of Escherichia coli Strains Isolated from the Poultry Industry in Henan Province, China. Microorganisms 2024, 12, 575. https://doi.org/10.3390/microorganisms12030575
Li Z, Jia C, Hu Z, Jin Y, Li T, Zhang X, Peng Z, Yang R, Chen H, Wang X. Antimicrobial Resistance and Genomic Characteristics of Escherichia coli Strains Isolated from the Poultry Industry in Henan Province, China. Microorganisms. 2024; 12(3):575. https://doi.org/10.3390/microorganisms12030575
Chicago/Turabian StyleLi, Zugang, Chaoying Jia, Zizhe Hu, Yancheng Jin, Tianzhi Li, Xiaoxue Zhang, Zhong Peng, Ruicheng Yang, Huanchun Chen, and Xiangru Wang. 2024. "Antimicrobial Resistance and Genomic Characteristics of Escherichia coli Strains Isolated from the Poultry Industry in Henan Province, China" Microorganisms 12, no. 3: 575. https://doi.org/10.3390/microorganisms12030575
APA StyleLi, Z., Jia, C., Hu, Z., Jin, Y., Li, T., Zhang, X., Peng, Z., Yang, R., Chen, H., & Wang, X. (2024). Antimicrobial Resistance and Genomic Characteristics of Escherichia coli Strains Isolated from the Poultry Industry in Henan Province, China. Microorganisms, 12(3), 575. https://doi.org/10.3390/microorganisms12030575





