Interkingdom Detection of Bacterial Quorum-Sensing Molecules by Mammalian Taste Receptors
Abstract
1. Introduction
2. Bacterial Quorum Sensing
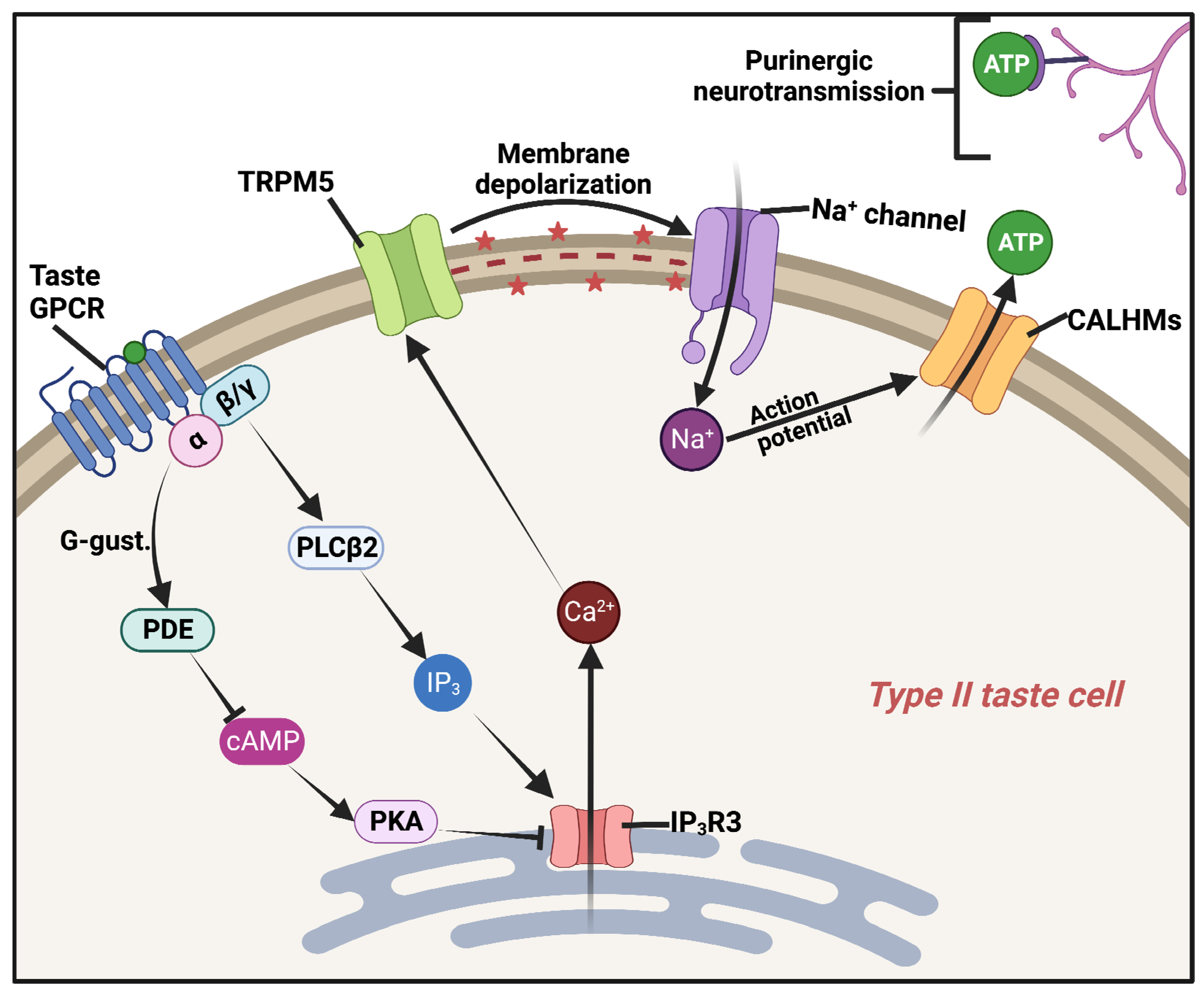
3. GPCR Taste Receptors
4. “Extraoral” Taste Receptors as Immune Detectors for Quorum-Sensing Molecules
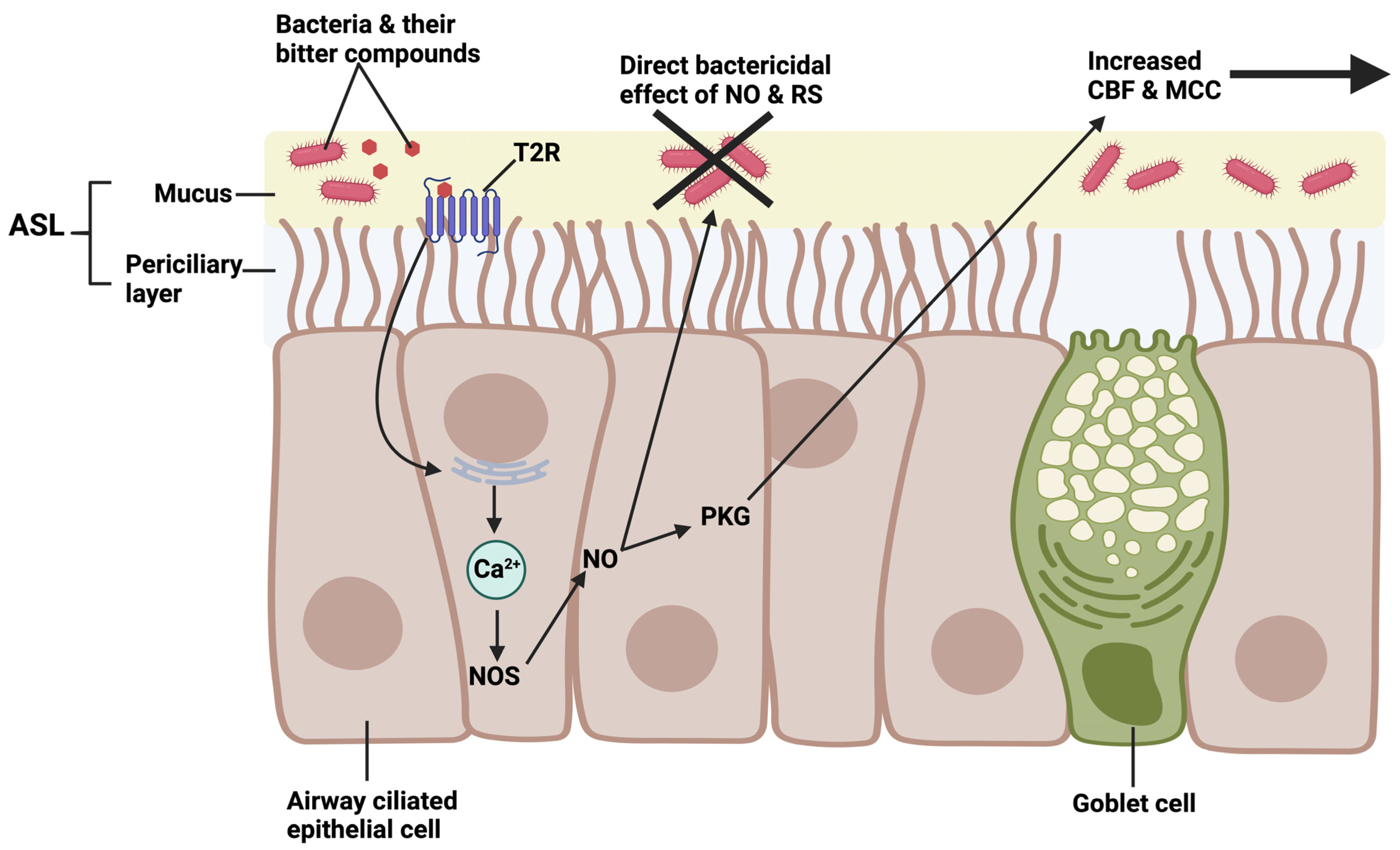
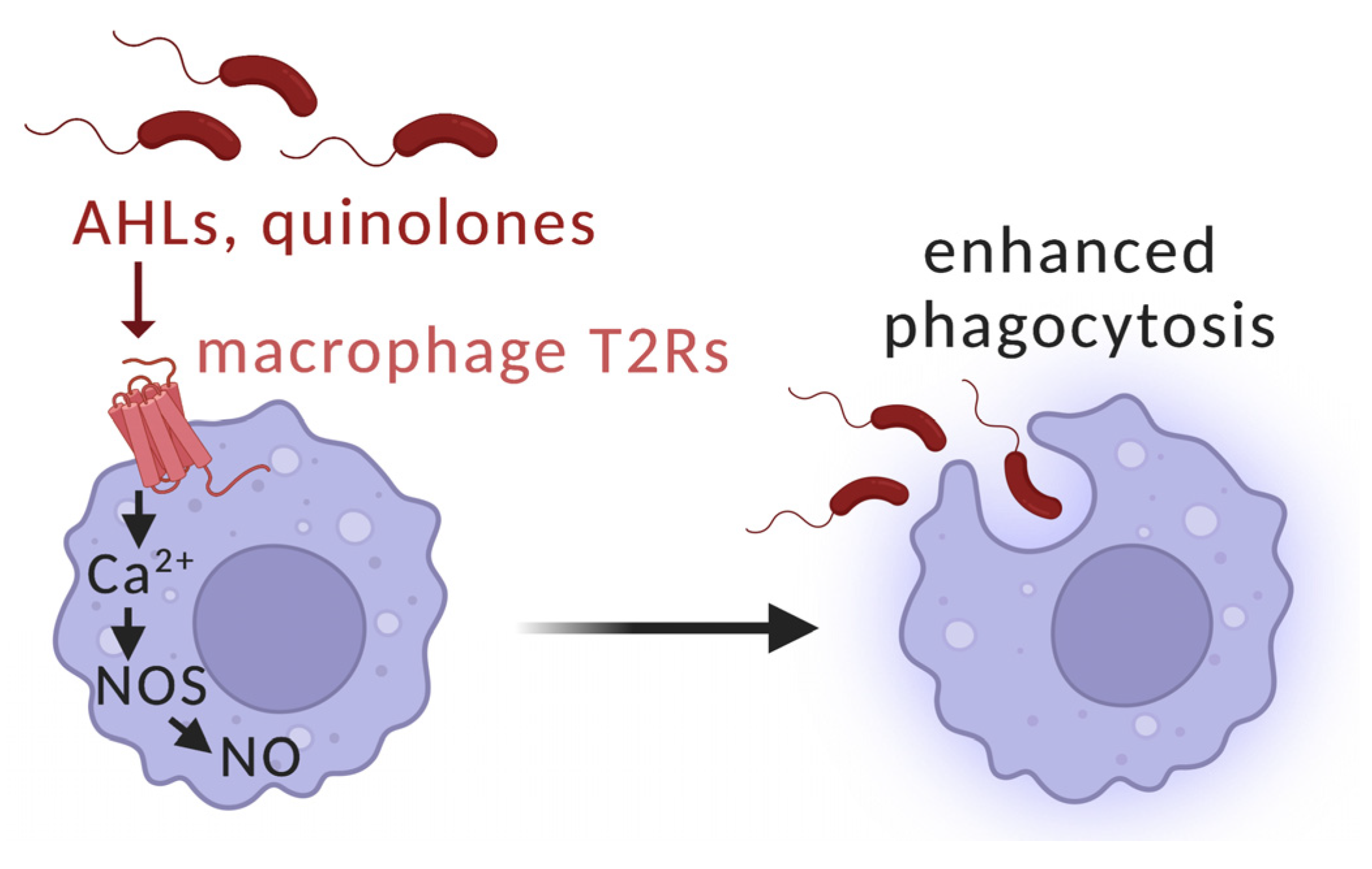
5. Interactions of T2Rs with Pseudomonas aeruginosa Acyl-Homoserine Lactone (AHL) and Quinolone Quorum-Sensing Molecules in Airway Ciliated Cells
6. Interactions of Bitter Taste Receptors in Gingival Epithelial Cells with Streptococcus mutans Competence Stimulating Peptides
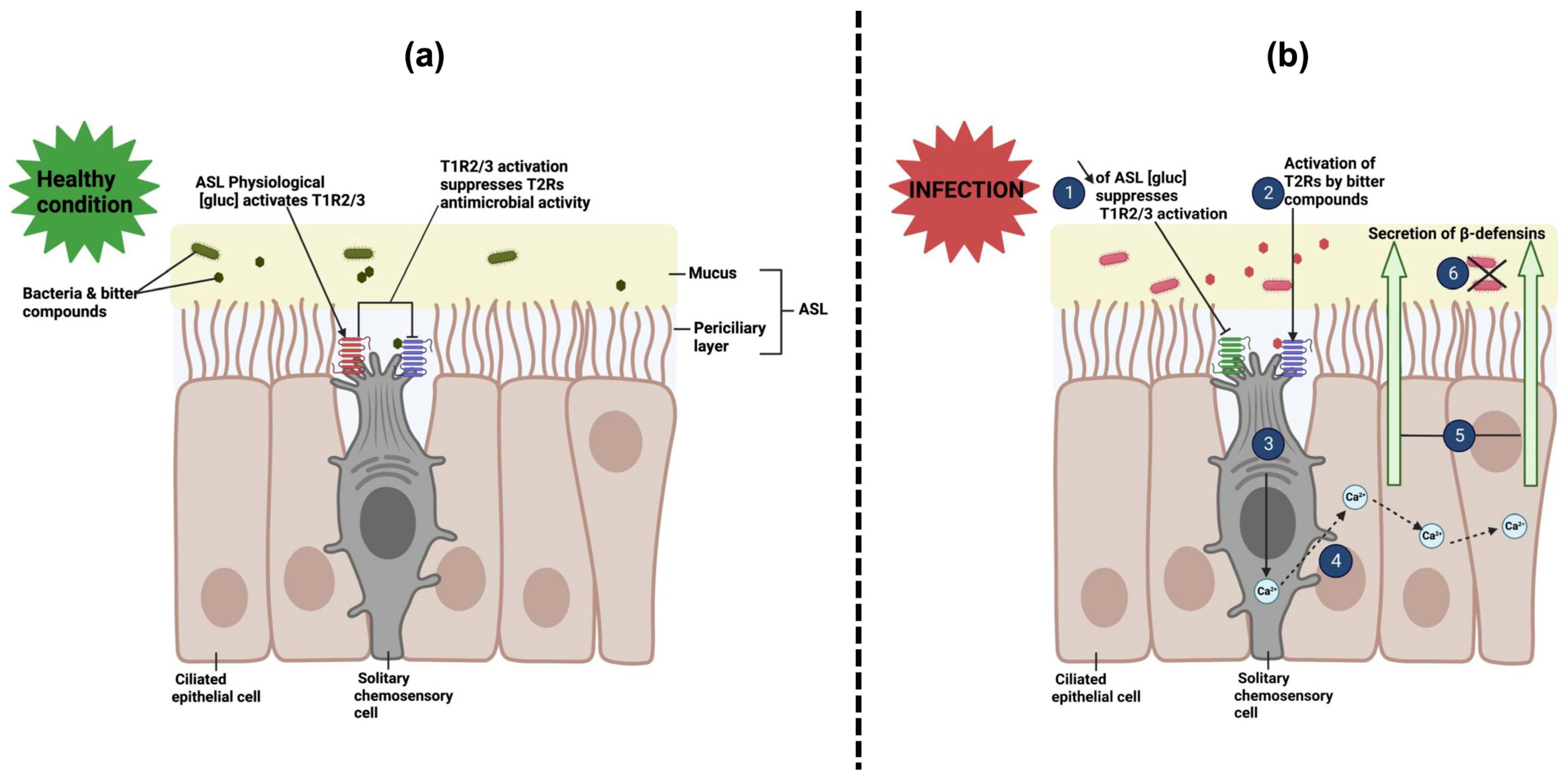
7. Interactions of Sweet Taste Receptors with Bacterial D-Amino Acids in Airway Solitary Chemosensory Cells
8. Conclusions and Remaining Questions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Ferencik, M.; Stvrtinova, V. Is the immune system our sixth sense? Relation between the immune and neuroendocrine systems. Bratisl. Lek. Listy 1997, 98, 187–198. [Google Scholar]
- Blalock, J.E. The immune system as the sixth sense. J. Intern. Med. 2005, 257, 126–138. [Google Scholar] [CrossRef]
- Patel, A.; Peralta-Yahya, P. Olfactory Receptors as an Emerging Chemical Sensing Scaffold. Biochemistry 2023, 62, 187–195. [Google Scholar] [CrossRef]
- Tong, T.; Wang, Y.; Kang, S.-G.; Huang, K. Ectopic Odorant Receptor Responding to Flavor Compounds: Versatile Roles in Health and Disease. Pharmaceutics 2021, 13, 1314. [Google Scholar] [CrossRef]
- Raka, R.N.; Wu, H.; Xiao, J.; Hossen, I.; Cao, Y.; Huang, M.; Jin, J. Human ectopic olfactory receptors and their food originated ligands: A review. Crit. Rev. Food Sci. Nutr. 2022, 62, 5424–5443. [Google Scholar] [CrossRef]
- Weidinger, D.; Jameel, K.J.; Alisch, D.; Jacobsen, J.; Bürger, P.; Ruhe, M.; Yusuf, F.; Rohde, S.; Störtkuhl, K.; Kaufmann, P.; et al. OR2AT4 and OR1A2 counterregulate molecular pathophysiological processes of steroid-resistant inflammatory lung diseases in human alveolar macrophages. Mol. Med. 2022, 28, 150. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Andreasson, K.I. Odorant receptors in macrophages: Potential targets for atherosclerosis. Trends Immunol. 2022, 43, 262–264. [Google Scholar] [CrossRef] [PubMed]
- Vadevoo, S.M.P.; Gunassekaran, G.R.; Lee, C.; Lee, N.; Lee, J.; Chae, S.; Park, J.-Y.; Koo, J.; Lee, B. The macrophage odorant receptor Olfr78 mediates the lactate-induced M2 phenotype of tumor-associated macrophages. Proc. Natl. Acad. Sci. USA 2021, 118, e2102434118. [Google Scholar] [CrossRef] [PubMed]
- Orecchioni, M.; Matsunami, H.; Ley, K. Olfactory receptors in macrophages and inflammation. Front. Immunol. 2022, 13, 1029244. [Google Scholar] [CrossRef] [PubMed]
- Orecchioni, M.; Kobiyama, K.; Winkels, H.; Ghosheh, Y.; McArdle, S.; Mikulski, Z.; Kiosses, W.B.; Fan, Z.; Wen, L.; Jung, Y.; et al. Olfactory receptor 2 in vascular macrophages drives atherosclerosis by NLRP3-dependent IL-1 production. Science 2022, 375, 214–221. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Wang, D.W. Olfactory receptor 2 activation in macrophages: Novel mediator of atherosclerosis progression. Signal Transduct. Target. Ther. 2022, 7, 247. [Google Scholar] [CrossRef]
- Chéret, J.; Bertolini, M.; Ponce, L.; Lehmann, J.; Tsai, T.; Alam, M.; Hatt, H.; Paus, R. Olfactory receptor OR2AT4 regulates human hair growth. Nat. Commun. 2018, 9, 3624. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.-J.; Depoortere, I.; Hatt, H. Therapeutic potential of ectopic olfactory and taste receptors. Nat. Rev. Drug Discov. 2019, 18, 116–138. [Google Scholar] [CrossRef]
- Kinnamon, S.C.; Finger, T.E. Recent advances in taste transduction and signaling. F1000Research 2019, 8, 2117. [Google Scholar] [CrossRef] [PubMed]
- Carey, R.M.; Lee, R.J. Taste Receptors in Upper Airway Innate Immunity. Nutrients 2019, 11, 2017. [Google Scholar] [CrossRef]
- Freund, J.R.; Lee, R.J. Taste receptors in the upper airway. World J. Otorhinolaryngol.-Head Neck Surg. 2018, 4, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Jeruzal-Świątecka, J.; Fendler, W.; Pietruszewska, W. Clinical Role of Extraoral Bitter Taste Receptors. Int. J. Mol. Sci. 2020, 21, 5156. [Google Scholar] [CrossRef] [PubMed]
- Behrens, M.; Lang, T. Extra-Oral Taste Receptors—Function, Disease, and Perspectives. Front. Nutr. 2022, 9, 881177. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Nair, S.K. Quorum sensing: How bacteria can coordinate activity and synchronize their response to external signals? Protein Sci. 2012, 10, 1403–1417. [Google Scholar] [CrossRef] [PubMed]
- Sionov, R.V.; Steinberg, D. Targeting the Holy Triangle of Quorum Sensing, Biofilm Formation, and Antibiotic Resistance in Pathogenic Bacteria. Microorganisms 2022, 10, 1239. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Bian, Z.; Wang, Y. Biofilm formation and inhibition mediated by bacterial quorum sensing. Appl. Microbiol. Biotechnol. 2022, 106, 6365–6381. [Google Scholar] [CrossRef]
- Otto, M. Critical Assessment of the Prospects of Quorum-Quenching Therapy for Staphylococcus aureus Infection. Int. J. Mol. Sci. 2023, 24, 4025. [Google Scholar] [CrossRef]
- Shi, X.; Zarkan, A. Bacterial survivors: Evaluating the mechanisms of antibiotic persistence. Microbiology 2022, 168, 001266. [Google Scholar] [CrossRef] [PubMed]
- Rather, M.A.; Saha, D.; Bhuyan, S.; Jha, A.N.; Mandal, M. Quorum Quenching: A Drug Discovery Approach Against Pseudomonas aeruginosa. Microbiol. Res. 2022, 264, 127173. [Google Scholar] [CrossRef] [PubMed]
- Liszt, K.I.; Wang, Q.; Farhadipour, M.; Segers, A.; Thijs, T.; Nys, L.; Deleus, E.; Van der Schueren, B.; Gerner, C.; Neuditschko, B.; et al. Human intestinal bitter taste receptors regulate innate immune responses and metabolic regulators in obesity. J. Clin. Investig. 2022, 132, e144828. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.; Soni, M.; Soyantar, B.; Shivangi, S.; Sutariya, S.; Saraf, M.; Goswami, D. A clash of quorum sensing vs quorum sensing inhibitors: An overview and risk of resistance. Arch. Microbiol. 2023, 205, 107. [Google Scholar] [CrossRef] [PubMed]
- Widmayer, P.; Partsch, V.; Pospiech, J.; Kusumakshi, S.; Boehm, U.; Breer, H. Distinct Cell Types With the Bitter Receptor Tas2r126 in Different Compartments of the Stomach. Front. Physiol. 2020, 11, 32. [Google Scholar] [CrossRef] [PubMed]
- Suntharalingam, P.; Cvitkovitch, D.G. Quorum sensing in streptococcal biofilm formation. Trends Microbiol. 2005, 13, 3–6. [Google Scholar] [CrossRef]
- Bernabè, G.; Pauletto, A.; Zamuner, A.; Cassari, L.; Castagliuolo, I.; Brun, P.; Dettin, M. Exploiting Conserved Quorum Sensing Signals in Streptococcus mutans and Streptococcus pneumoniae. Microorganisms 2022, 10, 2386. [Google Scholar] [CrossRef]
- Medapati, M.R.; Singh, N.; Bhagirath, A.Y.; Duan, K.; Triggs-Raine, B.; Batista, E.L.; Chelikani, P. Bitter taste receptor T2R14 detects quorum sensing molecules from cariogenic Streptococcus mutans and mediates innate immune responses in gingival epithelial cells. FASEB J. 2021, 35, e21375. [Google Scholar] [CrossRef] [PubMed]
- Medapati, M.R.; Bhagirath, A.Y.; Singh, N.; Schroth, R.J.; Bhullar, R.P.; Duan, K.; Chelikani, P. Bitter Taste Receptor T2R14 Modulates Gram-Positive Bacterial Internalization and Survival in Gingival Epithelial Cells. Int. J. Mol. Sci. 2021, 22, 9920. [Google Scholar] [CrossRef] [PubMed]
- Turner, H.N.; Liman, E.R. The Cellular and Molecular Basis of Sour Taste. Annu. Rev. Physiol. 2022, 84, 41–58. [Google Scholar] [CrossRef] [PubMed]
- Taruno, A.; Gordon, M.D. Molecular and Cellular Mechanisms of Salt Taste. Annu. Rev. Physiol. 2023, 85, 25–45. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Jin, H.; Zhang, W.; Ding, C.; O’keeffe, S.; Ye, M.; Zuker, C.S. Sour Sensing from the Tongue to the Brain. Cell 2019, 179, 392–402.e15. [Google Scholar] [CrossRef] [PubMed]
- Taruno, A.; Nomura, K.; Kusakizako, T.; Ma, Z.; Nureki, O.; Foskett, J.K. Taste transduction and channel synapses in taste buds. Pflügers Arch.-Eur. J. Physiol. 2021, 473, 3–13. [Google Scholar] [CrossRef]
- Teng, B.; Wilson, C.E.; Tu, Y.-H.; Joshi, N.R.; Kinnamon, S.C.; Liman, E.R. Cellular and Neural Responses to Sour Stimuli Require the Proton Channel Otop1. Curr. Biol. 2019, 29, 3647–3656.e5. [Google Scholar] [CrossRef] [PubMed]
- Nomura, K.; Nakanishi, M.; Ishidate, F.; Iwata, K.; Taruno, A. All-Electrical Ca2+-Independent Signal Transduction Mediates Attractive Sodium Taste in Taste Buds. Neuron 2020, 106, 816–829.e6. [Google Scholar] [CrossRef] [PubMed]
- Roper, S.D. Encoding Taste: From Receptors to Perception. Handb. Exp. Pharmacol. 2022, 275, 53–90. [Google Scholar] [CrossRef]
- Ahmad, R.; Dalziel, J.E. G Protein-Coupled Receptors in Taste Physiology and Pharmacology. Front. Pharmacol. 2020, 11, 587664. [Google Scholar] [CrossRef]
- Nieto Gutierrez, A.; McDonald, P.H. GPCRs: Emerging anti-cancer drug targets. Cell. Signal. 2018, 41, 65–74. [Google Scholar] [CrossRef]
- Kim, K.; Chung, K.Y. Many faces of the GPCR-arrestin interaction. Arch. Pharm. Res. 2020, 43, 890–899. [Google Scholar] [CrossRef] [PubMed]
- Talmon, M.; Pollastro, F.; Fresu, L.G. The Complex Journey of the Calcium Regulation Downstream of TAS2R Activation. Cells 2022, 11, 3638. [Google Scholar] [CrossRef]
- Romito, O.; Gueguinou, M.; Raoul, W.; Champion, O.; Robert, A.; Trebak, M.; Goupille, C.; Potier-Cartereau, M. Calcium signaling: A therapeutic target to overcome resistance to therapies in cancer. Cell Calcium 2022, 108, 102673. [Google Scholar] [CrossRef] [PubMed]
- Wong, G.T.; Gannon, K.S.; Margolskee, R.F. Transduction of bitter and sweet taste by gustducin. Nature 1996, 381, 796–800. [Google Scholar] [CrossRef]
- Giovannucci, D.R.; Groblewski, G.E.; Sneyd, J.; Yule, D.I. Targeted phosphorylation of inositol 1,4,5-trisphosphate receptors selectively inhibits localized Ca2+ release and shapes oscillatory Ca2+ signals. J. Biol. Chem. 2000, 275, 33704–33711. [Google Scholar] [CrossRef]
- Yule, D.I.; Straub, S.V.; Bruce, J.I. Modulation of Ca2+ oscillations by phosphorylation of Ins(1,4,5)P3 receptors. Biochem. Soc. Trans. 2003, 31, 954–957. [Google Scholar] [CrossRef] [PubMed]
- Clapp, T.R.; Stone, L.M.; Margolskee, R.F.; Kinnamon, S.C. Immunocytochemical evidence for co-expression of Type III IP3 receptor with signaling components of bitter taste transduction. BMC Neurosci. 2001, 2, 6. [Google Scholar] [CrossRef] [PubMed]
- Hisatsune, C.; Yasumatsu, K.; Takahashi-Iwanaga, H.; Ogawa, N.; Kuroda, Y.; Yoshida, R.; Ninomiya, Y.; Mikoshiba, K. Abnormal taste perception in mice lacking the type 3 inositol 1,4,5-trisphosphate receptor. J. Biol. Chem. 2007, 282, 37225–37231. [Google Scholar] [CrossRef] [PubMed]
- Miyoshi, M.A.; Abe, K.; Emori, Y. IP(3) receptor type 3 and PLCbeta2 are co-expressed with taste receptors T1R and T2R in rat taste bud cells. Chem. Senses 2001, 26, 259–265. [Google Scholar] [CrossRef]
- Bruce, J.I.; Shuttleworth, T.J.; Giovannucci, D.R.; Yule, D.I. Phosphorylation of inositol 1,4,5-trisphosphate receptors in parotid acinar cells. A mechanism for the synergistic effects of cAMP on Ca2+ signaling. J. Biol. Chem. 2002, 277, 1340–1348. [Google Scholar] [CrossRef]
- Bruce, J.I.; Straub, S.V.; Yule, D.I. Crosstalk between cAMP and Ca2+ signaling in non-excitable cells. Cell Calcium 2003, 34, 431–444. [Google Scholar] [CrossRef]
- Brown, D.A.; Bruce, J.I.; Straub, S.V.; Yule, D.I. cAMP potentiates ATP-evoked calcium signaling in human parotid acinar cells. J. Biol. Chem. 2004, 279, 39485–39494. [Google Scholar] [CrossRef] [PubMed]
- Straub, S.V.; Giovannucci, D.R.; Bruce, J.I.; Yule, D.I. A role for phosphorylation of inositol 1,4,5-trisphosphate receptors in defining calcium signals induced by Peptide agonists in pancreatic acinar cells. J. Biol. Chem. 2002, 277, 31949–31956. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhao, Z.; Margolskee, R.; Liman, E. The transduction channel TRPM5 is gated by intracellular calcium in taste cells. J. Neurosci. 2007, 27, 5777–5786. [Google Scholar] [CrossRef] [PubMed]
- Perez, C.A.; Huang, L.; Rong, M.; Kozak, J.A.; Preuss, A.K.; Zhang, H.; Max, M.; Margolskee, R.F. A transient receptor potential channel expressed in taste receptor cells. Nat. Neurosci. 2002, 5, 1169–1176. [Google Scholar] [CrossRef]
- Gao, N.; Lu, M.; Echeverri, F.; Laita, B.; Kalabat, D.; Williams, M.E.; Hevezi, P.; Zlotnik, A.; Moyer, B.D. Voltage-gated sodium channels in taste bud cells. BMC Neurosci. 2009, 10, 20. [Google Scholar] [CrossRef]
- Kinnamon, S.C. Taste receptor signalling—From tongues to lungs. Acta Physiol. 2012, 204, 158–168. [Google Scholar] [CrossRef]
- Taruno, A.; Matsumoto, I.; Ma, Z.; Marambaud, P.; Foskett, J.K. How do taste cells lacking synapses mediate neurotransmission? CALHM1, a voltage-gated ATP channel. Bioessays 2013, 35, 1111–1118. [Google Scholar] [CrossRef]
- Taruno, A.; Vingtdeux, V.; Ohmoto, M.; Ma, Z.; Dvoryanchikov, G.; Li, A.; Adrien, L.; Zhao, H.; Leung, S.; Abernethy, M.; et al. CALHM1 ion channel mediates purinergic neurotransmission of sweet, bitter and umami tastes. Nature 2013, 495, 223–226. [Google Scholar] [CrossRef]
- Ma, Z.; Taruno, A.; Ohmoto, M.; Jyotaki, M.; Lim, J.C.; Miyazaki, H.; Niisato, N.; Marunaka, Y.; Lee, R.J.; Hoff, H.; et al. CALHM3 Is Essential for Rapid Ion Channel-Mediated Purinergic Neurotransmission of GPCR-Mediated Tastes. Neuron 2018, 98, 547–561.e510. [Google Scholar] [CrossRef] [PubMed]
- Romanov, R.A.; Lasher, R.S.; High, B.; Savidge, L.E.; Lawson, A.; Rogachevskaja, O.A.; Zhao, H.; Rogachevsky, V.V.; Bystrova, M.F.; Churbanov, G.D.; et al. Chemical synapses without synaptic vesicles: Purinergic neurotransmission through a CALHM1 channel-mitochondrial signaling complex. Sci. Signal. 2018, 11, eaao1815. [Google Scholar] [CrossRef] [PubMed]
- Kinnamon, S.; Finger, T. The Role of ATP and Purinergic Receptors in Taste Signaling. Handb. Exp. Pharmacol. 2022, 275, 91–107. [Google Scholar] [CrossRef]
- Finger, T.; Kinnamon, S. Purinergic neurotransmission in the gustatory system. Auton. Neurosci. 2021, 236, 102874. [Google Scholar] [CrossRef]
- Zhang, Y.; Hoon, M.A.; Chandrashekar, J.; Mueller, K.L.; Cook, B.; Wu, D.; Zuker, C.S.; Ryba, N.J. Coding of sweet, bitter, and umami tastes: Different receptor cells sharing similar signaling pathways. Cell 2003, 112, 293–301. [Google Scholar] [CrossRef]
- Zhao, G.Q.; Zhang, Y.; Hoon, M.A.; Chandrashekar, J.; Erlenbach, I.; Ryba, N.J.; Zuker, C.S. The receptors for mammalian sweet and umami taste. Cell 2003, 115, 255–266. [Google Scholar] [CrossRef] [PubMed]
- Nelson, G.; Hoon, M.A.; Chandrashekar, J.; Zhang, Y.; Ryba, N.J.; Zuker, C.S. Mammalian sweet taste receptors. Cell 2001, 106, 381–390. [Google Scholar] [CrossRef]
- Sukumaran, S.K.; Palayyan, S.R. Sweet Taste Signaling: The Core Pathways and Regulatory Mechanisms. Int. J. Mol. Sci. 2022, 23, 8225. [Google Scholar] [CrossRef] [PubMed]
- Cui, M.; Jiang, P.; Maillet, E.; Max, M.; Margolskee, R.F.; Osman, R. The heterodimeric sweet taste receptor has multiple potential ligand binding sites. Curr. Pharm. Des. 2006, 12, 4591–4600. [Google Scholar] [CrossRef]
- Dubovski, N.; Ben-Shoshan Galezcki, Y.; Malach, E.; Niv, M.Y. Sensitivity of human sweet taste receptor subunits T1R2 and T1R3 to activation by glucose enantiomers. Chem. Senses 2023, 48, bjad005. [Google Scholar] [CrossRef]
- Dubovski, N.; Ben Shoshan-Galeczki, Y.; Malach, E.; Niv, M.Y. Taste and chirality: L-glucose sweetness is mediated by TAS1R2/TAS2R3 receptor. Food Chem. 2022, 373, 131393. [Google Scholar] [CrossRef] [PubMed]
- Di Pizio, A.; Ben Shoshan-Galeczki, Y.; Hayes, J.E.; Niv, M.Y. Bitter and sweet tasting molecules: It’s complicated. Neurosci. Lett. 2019, 700, 56–63. [Google Scholar] [CrossRef]
- Shallenberger, R.S.; Acree, T.E.; Lee, C.Y. Sweet taste of D and L-sugars and amino-acids and the steric nature of their chemo-receptor site. Nature 1969, 221, 555–556. [Google Scholar] [CrossRef] [PubMed]
- Maillet, E.L.; Cui, M.; Jiang, P.; Mezei, M.; Hecht, E.; Quijada, J.; Margolskee, R.F.; Osman, R.; Max, M. Characterization of the Binding Site of Aspartame in the Human Sweet Taste Receptor. Chem. Senses 2015, 40, 577–586. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.J.; Hariri, B.M.; McMahon, D.B.; Chen, B.; Doghramji, L.; Adappa, N.D.; Palmer, J.N.; Kennedy, D.W.; Jiang, P.; Margolskee, R.F.; et al. Bacterial d-amino acids suppress sinonasal innate immunity through sweet taste receptors in solitary chemosensory cells. Sci. Signal. 2017, 10, eaam7703. [Google Scholar] [CrossRef]
- Diepeveen, J.; Moerdijk-Poortvliet, T.C.W.; van der Leij, F.R. Molecular insights into human taste perception and umami tastants: A review. J. Food Sci. 2022, 87, 1449–1465. [Google Scholar] [CrossRef]
- Scott, K. The sweet and the bitter of mammalian taste. Curr. Opin. Neurobiol. 2004, 14, 423–427. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, Y.; Ohtsu, Y.; Nagasawa, M.; Shibata, H.; Kojima, I. Glucose promotes its own metabolism by acting on the cell-surface glucose-sensing receptor T1R3. Endocr. J. 2014, 61, 119–131. [Google Scholar] [CrossRef]
- Medina, A.; Nakagawa, Y.; Ma, J.; Li, L.; Hamano, K.; Akimoto, T.; Ninomiya, Y.; Kojima, I. Expression of the glucose-sensing receptor T1R3 in pancreatic islet: Changes in the expression levels in various nutritional and metabolic states. Endocr. J. 2014, 61, 797–805. [Google Scholar] [CrossRef]
- Masubuchi, Y.; Nakagawa, Y.; Ma, J.; Sasaki, T.; Kitamura, T.; Yamamoto, Y.; Kurose, H.; Kojima, I.; Shibata, H. A novel regulatory function of sweet taste-sensing receptor in adipogenic differentiation of 3T3-L1 cells. PLoS ONE 2013, 8, e54500. [Google Scholar] [CrossRef]
- Tordoff, M.G.; Shao, H.; Alarcon, L.K.; Margolskee, R.F.; Mosinger, B.; Bachmanov, A.A.; Reed, D.R.; McCaughey, S. Involvement of T1R3 in calcium-magnesium taste. Physiol. Genom. 2008, 34, 338–348. [Google Scholar] [CrossRef]
- Atsumi, N.; Yasumatsu, K.; Takashina, Y.; Ito, C.; Yasui, N.; Margolskee, R.F.; Yamashita, A. Chloride ions evoke taste sensations by binding to the extracellular ligand-binding domain of sweet/umami taste receptors. Elife 2023, 12, e84291. [Google Scholar] [CrossRef]
- Bachmanov, A.A.; Tordoff, M.G.; Beauchamp, G.K. Sweetener preference of C57BL/6ByJ and 129P3/J mice. Chem. Senses 2001, 26, 905–913. [Google Scholar] [CrossRef]
- Jiang, P.; Cui, M.; Zhao, B.; Liu, Z.; Snyder, L.A.; Benard, L.M.; Osman, R.; Margolskee, R.F.; Max, M. Lactisole interacts with the transmembrane domains of human T1R3 to inhibit sweet taste. J. Biol. Chem. 2005, 280, 15238–15246. [Google Scholar] [CrossRef]
- Jiang, P.; Cui, M.; Zhao, B.; Snyder, L.A.; Benard, L.M.; Osman, R.; Max, M.; Margolskee, R.F. Identification of the cyclamate interaction site within the transmembrane domain of the human sweet taste receptor subunit T1R3. J. Biol. Chem. 2005, 280, 34296–34305. [Google Scholar] [CrossRef]
- Sanematsu, K.; Kusakabe, Y.; Shigemura, N.; Hirokawa, T.; Nakamura, S.; Imoto, T.; Ninomiya, Y. Molecular Mechanisms for Sweet-suppressing Effect of Gymnemic Acids. J. Biol. Chem. 2014, 289, 25711–25720. [Google Scholar] [CrossRef] [PubMed]
- Lang, T.; Pizio, A.D.; Risso, D.; Drayna, D.; Behrens, M. Activation Profile of Tas2r2, The 26th Human Bitter Taste Receptor. Mol. Nutr. Food Res. 2023, e2200775. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, C.; Bufe, B.; Batram, C.; Meyerhof, W. Oligomerization of TAS2R bitter taste receptors. Chem. Senses 2010, 35, 395–406. [Google Scholar] [CrossRef] [PubMed]
- Dagan-Wiener, A.; Di Pizio, A.; Nissim, I.; Bahia, M.S.; Dubovski, N.; Margulis, E.; Niv, M.Y. BitterDB: Taste ligands and receptors database in 2019. Nucleic Acids Res. 2019, 47, D1179–D1185. [Google Scholar] [CrossRef] [PubMed]
- Meyerhof, W.; Batram, C.; Kuhn, C.; Brockhoff, A.; Chudoba, E.; Bufe, B.; Appendino, G.; Behrens, M. The molecular receptive ranges of human TAS2R bitter taste receptors. Chem. Senses 2010, 35, 157–170. [Google Scholar] [CrossRef]
- Jiang, P.; Josue, J.; Li, X.; Glaser, D.; Li, W.; Brand, J.G.; Margolskee, R.F.; Reed, D.R.; Beauchamp, G.K. Major taste loss in carnivorous mammals. Proc. Natl. Acad. Sci. USA 2012, 109, 4956–4961. [Google Scholar] [CrossRef]
- Lossow, K.; Hubner, S.; Roudnitzky, N.; Slack, J.P.; Pollastro, F.; Behrens, M.; Meyerhof, W. Comprehensive Analysis of Mouse Bitter Taste Receptors Reveals Different Molecular Receptive Ranges for Orthologous Receptors in Mice and Humans. J. Biol. Chem. 2016, 291, 15358–15377. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Zhang, J. Diet shapes the evolution of the vertebrate bitter taste receptor gene repertoire. Mol. Biol. Evol. 2014, 31, 303–309. [Google Scholar] [CrossRef]
- Bufe, B.; Breslin, P.A.; Kuhn, C.; Reed, D.R.; Tharp, C.D.; Slack, J.P.; Kim, U.K.; Drayna, D.; Meyerhof, W. The molecular basis of individual differences in phenylthiocarbamide and propylthiouracil bitterness perception. Curr. Biol. 2005, 15, 322–327. [Google Scholar] [CrossRef]
- Risso, D.S.; Mezzavilla, M.; Pagani, L.; Robino, A.; Morini, G.; Tofanelli, S.; Carrai, M.; Campa, D.; Barale, R.; Caradonna, F.; et al. Global diversity in the TAS2R38 bitter taste receptor: Revisiting a classic evolutionary PROPosal. Sci. Rep. 2016, 6, 25506. [Google Scholar] [CrossRef]
- Tan, J.; Abrol, R.; Trzaskowski, B.; Goddard, W.A., 3rd. 3D Structure Prediction of TAS2R38 Bitter Receptors Bound to Agonists Phenylthiocarbamide (PTC) and 6-n-Propylthiouracil (PROP). J. Chem. Inf. Model. 2012, 52, 1875–1885. [Google Scholar] [CrossRef]
- Biarnes, X.; Marchiori, A.; Giorgetti, A.; Lanzara, C.; Gasparini, P.; Carloni, P.; Born, S.; Brockhoff, A.; Behrens, M.; Meyerhof, W. Insights into the binding of Phenyltiocarbamide (PTC) agonist to its target human TAS2R38 bitter receptor. PLoS ONE 2010, 5, e12394. [Google Scholar] [CrossRef]
- Floriano, W.B.; Hall, S.; Vaidehi, N.; Kim, U.; Drayna, D.; Goddard, W.A., 3rd. Modeling the human PTC bitter-taste receptor interactions with bitter tastants. J. Mol. Model. 2006, 12, 931–941. [Google Scholar] [CrossRef]
- Lipchock, S.V.; Mennella, J.A.; Spielman, A.I.; Reed, D.R. Human bitter perception correlates with bitter receptor messenger RNA expression in taste cells. Am. J. Clin. Nutr. 2013, 98, 1136–1143. [Google Scholar] [CrossRef]
- Bachmanov, A.A.; Bosak, N.P.; Lin, C.; Matsumoto, I.; Ohmoto, M.; Reed, D.R.; Nelson, T.M. Genetics of taste receptors. Curr. Pharm. Des. 2014, 20, 2669–2683. [Google Scholar] [CrossRef]
- Mennella, J.A.; Pepino, M.Y.; Reed, D.R. Genetic and environmental determinants of bitter perception and sweet preferences. Pediatrics 2005, 115, e216–e222. [Google Scholar] [CrossRef]
- Chamoun, E.; Mutch, D.M.; Allen-Vercoe, E.; Buchholz, A.C.; Duncan, A.M.; Spriet, L.L.; Haines, J.; Ma, D.W.L.; Guelph Family Health, S. A review of the associations between single nucleotide polymorphisms in taste receptors, eating behaviors, and health. Crit. Rev. Food Sci. Nutr. 2018, 58, 194–207. [Google Scholar] [CrossRef]
- Ramos-Lopez, O.; Panduro, A.; Martinez-Lopez, E.; Roman, S. Sweet Taste Receptor TAS1R2 Polymorphism (Val191Val) Is Associated with a Higher Carbohydrate Intake and Hypertriglyceridemia among the Population of West Mexico. Nutrients 2016, 8, 101. [Google Scholar] [CrossRef]
- Lee, R.J.; Cohen, N.A. Bitter taste bodyguards. Sci. Am. 2016, 314, 38–43. [Google Scholar] [CrossRef]
- Lee, R.J.; Cohen, N.A. Taste receptors in innate immunity. Cell. Mol. Life Sci. 2015, 72, 217–236. [Google Scholar] [CrossRef]
- Chalmers, J.A.; Jang, J.J.; Belsham, D.D. Glucose sensing mechanisms in hypothalamic cell models: Glucose inhibition of AgRP synthesis and secretion. Mol. Cell. Endocrinol. 2014, 382, 262–270. [Google Scholar] [CrossRef]
- Dehkordi, O.; Rose, J.E.; Fatemi, M.; Allard, J.S.; Balan, K.V.; Young, J.K.; Fatima, S.; Millis, R.M.; Jayam-Trouth, A. Neuronal expression of bitter taste receptors and downstream signaling molecules in the rat brainstem. Brain Res. 2012, 1475, 1–10. [Google Scholar] [CrossRef]
- Lemon, C.H.; Margolskee, R.F. Contribution of the T1r3 taste receptor to the response properties of central gustatory neurons. J. Neurophysiol. 2009, 101, 2459–2471. [Google Scholar] [CrossRef]
- Ren, X.; Zhou, L.; Terwilliger, R.; Newton, S.S.; de Araujo, I.E. Sweet taste signaling functions as a hypothalamic glucose sensor. Front. Integr. Neurosci. 2009, 3, 12. [Google Scholar] [CrossRef]
- Martin, B.; Wang, R.; Cong, W.N.; Daimon, C.M.; Wu, W.W.; Ni, B.; Becker, K.G.; Lehrmann, E.; Wood, W.H., 3rd; Zhang, Y.; et al. Altered learning, memory, and social behavior in type 1 taste receptor subunit 3 knock-out mice are associated with neuronal dysfunction. J. Biol. Chem. 2017, 292, 11508–11530. [Google Scholar] [CrossRef]
- Kojima, I.; Nakagawa, Y.; Ohtsu, Y.; Medina, A.; Nagasawa, M. Sweet Taste-Sensing Receptors Expressed in Pancreatic beta-Cells: Sweet Molecules Act as Biased Agonists. Endocrinol. Metab. 2014, 29, 12–19. [Google Scholar] [CrossRef]
- Kyriazis, G.A.; Soundarapandian, M.M.; Tyrberg, B. Sweet taste receptor signaling in beta cells mediates fructose-induced potentiation of glucose-stimulated insulin secretion. Proc. Natl. Acad. Sci. USA 2012, 109, E524–E532. [Google Scholar] [CrossRef]
- Nakagawa, Y.; Nagasawa, M.; Yamada, S.; Hara, A.; Mogami, H.; Nikolaev, V.O.; Lohse, M.J.; Shigemura, N.; Ninomiya, Y.; Kojima, I. Sweet taste receptor expressed in pancreatic beta-cells activates the calcium and cyclic AMP signaling systems and stimulates insulin secretion. PLoS ONE 2009, 4, e5106. [Google Scholar] [CrossRef]
- Malaisse, W.J.; Vanonderbergen, A.; Louchami, K.; Jijakli, H.; Malaisse-Lagae, F. Effects of artificial sweeteners on insulin release and cationic fluxes in rat pancreatic islets. Cell. Signal. 1998, 10, 727–733. [Google Scholar] [CrossRef]
- Ozdener, M.H.; Subramaniam, S.; Sundaresan, S.; Sery, O.; Hashimoto, T.; Asakawa, Y.; Besnard, P.; Abumrad, N.A.; Khan, N.A. CD36- and GPR120-mediated Ca2+ signaling in human taste bud cells mediates differential responses to fatty acids and is altered in obese mice. Gastroenterology 2014, 146, 995–1005. [Google Scholar] [CrossRef]
- Cartoni, C.; Yasumatsu, K.; Ohkuri, T.; Shigemura, N.; Yoshida, R.; Godinot, N.; le Coutre, J.; Ninomiya, Y.; Damak, S. Taste preference for fatty acids is mediated by GPR40 and GPR120. J. Neurosci. 2010, 30, 8376–8382. [Google Scholar] [CrossRef]
- Khan, N.A.; Besnard, P. Oro-sensory perception of dietary lipids: New insights into the fat taste transduction. Biochim. Biophys. Acta 2009, 1791, 149–155. [Google Scholar] [CrossRef]
- Sclafani, A.; Zukerman, S.; Glendinning, J.I.; Margolskee, R.F. Fat and carbohydrate preferences in mice: The contribution of alpha-gustducin and Trpm5 taste-signaling proteins. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2007, 293, R1504–R1513. [Google Scholar] [CrossRef]
- Laugerette, F.; Passilly-Degrace, P.; Patris, B.; Niot, I.; Febbraio, M.; Montmayeur, J.P.; Besnard, P. CD36 involvement in orosensory detection of dietary lipids, spontaneous fat preference, and digestive secretions. J. Clin. Investig. 2005, 115, 3177–3184. [Google Scholar] [CrossRef]
- Pirkwieser, P.; Behrens, M.; Somoza, V. Metallic Sensation-Just an Off-Flavor or a Biologically Relevant Sensing Pathway? J. Agric. Food Chem. 2021, 69, 1775–1780. [Google Scholar] [CrossRef] [PubMed]
- Ecarma, M.J.Y.; Nolden, A.A. A review of the flavor profile of metal salts: Understanding the complexity of metallic sensation. Chem. Senses 2021, 46, bjab043. [Google Scholar] [CrossRef]
- Ohsu, T.; Amino, Y.; Nagasaki, H.; Yamanaka, T.; Takeshita, S.; Hatanaka, T.; Maruyama, Y.; Miyamura, N.; Eto, Y. Involvement of the calcium-sensing receptor in human taste perception. J. Biol. Chem. 2010, 285, 1016–1022. [Google Scholar] [CrossRef]
- Maruyama, Y.; Yasuda, R.; Kuroda, M.; Eto, Y. Kokumi substances, enhancers of basic tastes, induce responses in calcium-sensing receptor expressing taste cells. PLoS ONE 2012, 7, e34489. [Google Scholar] [CrossRef]
- Yamamoto, T.; Inui-Yamamoto, C. The flavor-enhancing action of glutamate and its mechanism involving the notion of kokumi. NPJ Sci. Food 2023, 7, 3. [Google Scholar] [CrossRef] [PubMed]
- Laffitte, A.; Gibbs, M.; Hernangomez de Alvaro, C.; Addison, J.; Lonsdale, Z.N.; Giribaldi, M.G.; Rossignoli, A.; Vennegeerts, T.; Winnig, M.; Klebansky, B.; et al. Kokumi taste perception is functional in a model carnivore, the domestic cat (Felis catus). Sci. Rep. 2021, 11, 10527. [Google Scholar] [CrossRef]
- Oka, Y.; Butnaru, M.; von Buchholtz, L.; Ryba, N.J.; Zuker, C.S. High salt recruits aversive taste pathways. Nature 2013, 494, 472–475. [Google Scholar] [CrossRef] [PubMed]
- Gaillard, D.; Kinnamon, S.C. New evidence for fat as a primary taste quality. Acta Physiol. 2019, 226, e13246. [Google Scholar] [CrossRef] [PubMed]
- Simons, C.T.; Klein, A.H.; Carstens, E. Chemogenic Subqualities of Mouthfeel. Chem. Senses 2019, 44, 281–288. [Google Scholar] [CrossRef]
- Sclafani, A.; Zukerman, S.; Ackroff, K. GPR40 and GPR120 fatty acid sensors are critical for postoral but not oral mediation of fat preferences in the mouse. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2013, 305, R1490–R1497. [Google Scholar] [CrossRef]
- Abdoul-Azize, S.; Selvakumar, S.; Sadou, H.; Besnard, P.; Khan, N.A. Ca2+ signaling in taste bud cells and spontaneous preference for fat: Unresolved roles of CD36 and GPR120. Biochimie 2014, 96, 8–13. [Google Scholar] [CrossRef]
- Gerbe, F.; Sidot, E.; Smyth, D.J.; Ohmoto, M.; Matsumoto, I.; Dardalhon, V.; Cesses, P.; Garnier, L.; Pouzolles, M.; Brulin, B.; et al. Intestinal epithelial tuft cells initiate type 2 mucosal immunity to helminth parasites. Nature 2016, 529, 226–230. [Google Scholar] [CrossRef]
- Howitt, M.R.; Lavoie, S.; Michaud, M.; Blum, A.M.; Tran, S.V.; Weinstock, J.V.; Gallini, C.A.; Redding, K.; Margolskee, R.F.; Osborne, L.C.; et al. Tuft cells, taste-chemosensory cells, orchestrate parasite type 2 immunity in the gut. Science 2016, 351, 1329–1333. [Google Scholar] [CrossRef]
- Barham, H.P.; Cooper, S.E.; Anderson, C.B.; Tizzano, M.; Kingdom, T.T.; Finger, T.E.; Kinnamon, S.C.; Ramakrishnan, V.R. Solitary chemosensory cells and bitter taste receptor signaling in human sinonasal mucosa. Int. Forum Allergy Rhinol. 2013, 3, 450–457. [Google Scholar] [CrossRef]
- Tizzano, M.; Cristofoletti, M.; Sbarbati, A.; Finger, T.E. Expression of taste receptors in solitary chemosensory cells of rodent airways. BMC Pulm. Med. 2011, 11, 3. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.J.; Kofonow, J.M.; Rosen, P.L.; Siebert, A.P.; Chen, B.; Doghramji, L.; Xiong, G.; Adappa, N.D.; Palmer, J.N.; Kennedy, D.W.; et al. Bitter and sweet taste receptors regulate human upper respiratory innate immunity. J. Clin. Investig. 2014, 124, 1393–1405. [Google Scholar] [CrossRef] [PubMed]
- Clark, A.A.; Liggett, S.B.; Munger, S.D. Extraoral bitter taste receptors as mediators of off-target drug effects. FASEB J. 2012, 26, 4827–4831. [Google Scholar] [CrossRef]
- Jaggupilli, A.; Howard, R.; Upadhyaya, J.D.; Bhullar, R.P.; Chelikani, P. Bitter taste receptors: Novel insights into the biochemistry and pharmacology. Int. J. Biochem. Cell Biol. 2016, 77, 184–196. [Google Scholar] [CrossRef]
- Levit, A.; Nowak, S.; Peters, M.; Wiener, A.; Meyerhof, W.; Behrens, M.; Niv, M.Y. The bitter pill: Clinical drugs that activate the human bitter taste receptor TAS2R14. FASEB J. 2014, 28, 1181–1197. [Google Scholar] [CrossRef] [PubMed]
- Mennella, J.A.; Spector, A.C.; Reed, D.R.; Coldwell, S.E. The bad taste of medicines: Overview of basic research on bitter taste. Clin. Ther. 2013, 35, 1225–1246. [Google Scholar] [CrossRef]
- Bhatia, V.; de Jesus, V.C.; Shaik, F.A.; Jaggupilli, A.; Singh, N.; Chelikani, P.; Atukorallaya, D. Extraoral expression and characterization of bitter taste receptors in Astyanax mexicanus (Mexican tetra fish). FASEB Bioadv. 2022, 4, 574–584. [Google Scholar] [CrossRef]
- Wooding, S.P.; Ramirez, V.A.; Behrens, M. Bitter taste receptors: Genes, evolution and health. Evol. Med. Public Health 2021, 9, 431–447. [Google Scholar] [CrossRef]
- Nakagawa, Y.; Nagasawa, M.; Mogami, H.; Lohse, M.; Ninomiya, Y.; Kojima, I. Multimodal function of the sweet taste receptor expressed in pancreatic beta-cells: Generation of diverse patterns of intracellular signals by sweet agonists. Endocr. J. 2013, 60, 1191–1206. [Google Scholar] [CrossRef]
- Meyer-Gerspach, A.C.; Wolnerhanssen, B.; Beglinger, C. Gut sweet taste receptors and their role in metabolism. Front. Horm. Res. 2014, 42, 123–133. [Google Scholar] [CrossRef] [PubMed]
- Kokrashvili, Z.; Mosinger, B.; Margolskee, R.F. Taste signaling elements expressed in gut enteroendocrine cells regulate nutrient-responsive secretion of gut hormones. Am. J. Clin. Nutr. 2009, 90, 822S–825S. [Google Scholar] [CrossRef] [PubMed]
- Dyer, J.; Salmon, K.S.; Zibrik, L.; Shirazi-Beechey, S.P. Expression of sweet taste receptors of the T1R family in the intestinal tract and enteroendocrine cells. Biochem. Soc. Trans. 2005, 33, 302–305. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Cao, J.; Iguchi, N.; Riethmacher, D.; Huang, L. Functional characterization of bitter-taste receptors expressed in mammalian testis. Mo.l Hum. Reprod. 2013, 19, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Mosinger, B.; Redding, K.M.; Parker, M.R.; Yevshayeva, V.; Yee, K.K.; Dyomina, K.; Li, Y.; Margolskee, R.F. Genetic loss or pharmacological blockade of testes-expressed taste genes causes male sterility. Proc. Natl. Acad. Sci. USA 2013, 110, 12319–12324. [Google Scholar] [CrossRef]
- Li, F.; Zhou, M. Depletion of bitter taste transduction leads to massive spermatid loss in transgenic mice. Mol. Hum. Reprod. 2012, 18, 289–297. [Google Scholar] [CrossRef]
- Luo, X.C.; Chen, Z.H.; Xue, J.B.; Zhao, D.X.; Lu, C.; Li, Y.H.; Li, S.M.; Du, Y.W.; Liu, Q.; Wang, P.; et al. Infection by the parasitic helminth Trichinella spiralis activates a Tas2r-mediated signaling pathway in intestinal tuft cells. Proc. Natl. Acad. Sci. USA 2019, 116, 5564–5569. [Google Scholar] [CrossRef]
- Sun, S.; Yang, Y.; Xiong, R.; Ni, Y.; Ma, X.; Hou, M.; Chen, L.; Xu, Z.; Chen, L.; Ji, M. Oral berberine ameliorates high-fat diet-induced obesity by activating TAS2Rs in tuft and endocrine cells in the gut. Life Sci. 2022, 311, 121141. [Google Scholar] [CrossRef]
- Howitt, M.R.; Cao, Y.G.; Gologorsky, M.B.; Li, J.A.; Haber, A.L.; Biton, M.; Lang, J.; Michaud, M.; Regev, A.; Garrett, W.S. The Taste Receptor TAS1R3 Regulates Small Intestinal Tuft Cell Homeostasis. Immunohorizons 2020, 4, 23–32. [Google Scholar] [CrossRef]
- Bezencon, C.; le Coutre, J.; Damak, S. Taste-signaling proteins are coexpressed in solitary intestinal epithelial cells. Chem. Senses 2007, 32, 41–49. [Google Scholar] [CrossRef]
- Strine, M.S.; Wilen, C.B. Tuft cells are key mediators of interkingdom interactions at mucosal barrier surfaces. PLoS Pathog. 2022, 18, e1010318. [Google Scholar] [CrossRef]
- Lei, W.; Ren, W.; Ohmoto, M.; Urban, J.F., Jr.; Matsumoto, I.; Margolskee, R.F.; Jiang, P. Activation of intestinal tuft cell-expressed Sucnr1 triggers type 2 immunity in the mouse small intestine. Proc. Natl. Acad. Sci. USA 2018, 115, 5552–5557. [Google Scholar] [CrossRef] [PubMed]
- Nadjsombati, M.S.; McGinty, J.W.; Lyons-Cohen, M.R.; Jaffe, J.B.; DiPeso, L.; Schneider, C.; Miller, C.N.; Pollack, J.L.; Nagana Gowda, G.A.; Fontana, M.F.; et al. Detection of Succinate by Intestinal Tuft Cells Triggers a Type 2 Innate Immune Circuit. Immunity 2018, 49, 33–41.e7. [Google Scholar] [CrossRef] [PubMed]
- Schneider, C.; O’Leary, C.E.; von Moltke, J.; Liang, H.E.; Ang, Q.Y.; Turnbaugh, P.J.; Radhakrishnan, S.; Pellizzon, M.; Ma, A.; Locksley, R.M. A Metabolite-Triggered Tuft Cell-ILC2 Circuit Drives Small Intestinal Remodeling. Cell 2018, 174, 271–284.e214. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A.; Herring, C.A.; Chen, B.; Kim, H.; Simmons, A.J.; Southard-Smith, A.N.; Allaman, M.M.; White, J.R.; Macedonia, M.C.; McKinley, E.T.; et al. Succinate Produced by Intestinal Microbes Promotes Specification of Tuft Cells to Suppress Ileal Inflammation. Gastroenterology 2020, 159, 2101–2115.e2105. [Google Scholar] [CrossRef]
- Schneider, C.; O’Leary, C.E.; Locksley, R.M. Regulation of immune responses by tuft cells. Nat. Rev. Immunol. 2019, 19, 584–593. [Google Scholar] [CrossRef]
- Qin, Y.; Palayyan, S.R.; Zheng, X.; Tian, S.; Margolskee, R.F.; Sukumaran, S.K. Type II taste cells participate in mucosal immune surveillance. PLoS Biol. 2023, 21, e3001647. [Google Scholar] [CrossRef]
- O’Toole, G.A.; Kolter, R. Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol. Microbiol. 1998, 30, 295–304. [Google Scholar] [CrossRef]
- Lee, R.J.; Xiong, G.; Kofonow, J.M.; Chen, B.; Lysenko, A.; Jiang, P.; Abraham, V.; Doghramji, L.; Adappa, N.D.; Palmer, J.N.; et al. T2R38 taste receptor polymorphisms underlie susceptibility to upper respiratory infection. J. Clin. Investig. 2012, 122, 4145–4159. [Google Scholar] [CrossRef]
- Freund, J.R.; Mansfield, C.J.; Doghramji, L.J.; Adappa, N.D.; Palmer, J.N.; Kennedy, D.W.; Reed, D.R.; Jiang, P.; Lee, R.J. Activation of airway epithelial bitter taste receptors by Pseudomonas aeruginosa quinolones modulates calcium, cyclic-AMP, and nitric oxide signaling. J. Biol. Chem. 2018, 293, 9824–9840. [Google Scholar] [CrossRef] [PubMed]
- Harro, J.M.; Daugherty, S.; Bruno, V.M.; Jabra-Rizk, M.A.; Rasko, D.A.; Shirtliff, M.E. Draft Genome Sequence of the Methicillin-Resistant Staphylococcus aureus Isolate MRSA-M2. Genome Announc. 2013, 1, e00037-12. [Google Scholar] [CrossRef] [PubMed]
- Carey, R.M.; Adappa, N.D.; Palmer, J.N.; Cohen, N.A.; Lee, R.J. Taste receptor T1R3 in nasal cilia detects <em>Staphylococcus aureus</em> D-amino acids to enhance apical glucose uptake. bioRxiv 2022. [Google Scholar] [CrossRef]
- Carey, R.M.; Workman, A.D.; Chen, B.; Adappa, N.D.; Palmer, J.N.; Kennedy, D.W.; Lee, R.J.; Cohen, N.A. Staphylococcus aureus triggers nitric oxide production in human upper airway epithelium. Int. Forum Allergy Rhinol. 2015, 5, 808–813. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.C.; Huang, S.D.; Tu, J.H.; Yu, J.S.; Nurlatifah, A.O.; Chiu, W.C.; Su, Y.H.; Chang, H.L.; Putri, D.A.; Cheng, H.L. Exopolysaccharides of Bacillus amyloliquefaciens modulate glycemic level in mice and promote glucose uptake of cells through the activation of Akt. Int. J. Biol. Macromol. 2020, 146, 202–211. [Google Scholar] [CrossRef]
- Sung, W.W.; Tu, J.H.; Yu, J.S.; Ulfa, M.Z.; Chang, J.H.; Cheng, H.L. Bacillus amyloliquefaciens exopolysaccharide preparation induces glucagon-like peptide 1 secretion through the activation of bitter taste receptors. Int. J. Biol. Macromol. 2021, 185, 562–571. [Google Scholar] [CrossRef]
- Vaid, S.; Vaid, N. Sinonasal Anatomy. Neuroimaging Clin. N. Am. 2022, 32, 713–734. [Google Scholar] [CrossRef]
- Stevens, W.W.; Lee, R.J.; Schleimer, R.P.; Cohen, N.A. Chronic rhinosinusitis pathogenesis. J. Allergy Clin. Immunol. 2015, 136, 1442–1453. [Google Scholar] [CrossRef]
- Piatti, G.; Ambrosetti, U.; Robino, A.; Girotto, G.; Gasparini, P. Primary Ciliary Dyskinesia: The Impact of Taste Receptor (TAS2R38) Gene Polymorphisms on Disease Outcome and Severity. Int. Arch. Allergy Immunol. 2020, 181, 727–731. [Google Scholar] [CrossRef]
- Myer, H.; Chupita, S.; Jnah, A. Cystic Fibrosis: Back to the Basics. Neonatal Netw. 2023, 42, 23–30. [Google Scholar] [CrossRef]
- Hariri, B.M.; Cohen, N.A. New insights into upper airway innate immunity. Am. J. Rhinol. Allergy 2016, 30, 319–323. [Google Scholar] [CrossRef] [PubMed]
- Laidlaw, T.M.; Buchheit, K.M. Biologics in chronic rhinosinusitis with nasal polyposis. Ann. Allergy Asthma Immunol. 2020, 124, 326–332. [Google Scholar] [CrossRef]
- Naclerio, R.; Baroody, F.; Bachert, C.; Bleier, B.; Borish, L.; Brittain, E.; Chupp, G.; Fisher, A.; Fokkens, W.; Gevaert, P.; et al. Clinical Research Needs for the Management of Chronic Rhinosinusitis with Nasal Polyps in the New Era of Biologics: A National Institute of Allergy and Infectious Diseases Workshop. J. Allergy Clin. Immunol. Pract. 2020, 8, 1532–1549.e1531. [Google Scholar] [CrossRef] [PubMed]
- Staudacher, A.G.; Peters, A.T.; Kato, A.; Stevens, W.W. Use of endotypes, phenotypes, and inflammatory markers to guide treatment decisions in chronic rhinosinusitis. Ann. Allergy Asthma Immunol. 2020, 124, 318–325. [Google Scholar] [CrossRef]
- Matera, M.G.; Rinaldi, B.; de Novellis, V.; Rogliani, P.; Cazzola, M. Current and emerging treatment modalities for bacterial rhinosinusitis in adults: A comprehensive review. Expert Opin. Pharmacother. 2022, 23, 2013–2022. [Google Scholar] [CrossRef]
- Rudmik, L. Economics of Chronic Rhinosinusitis. Curr. Allergy Asthma Rep. 2017, 17, 20. [Google Scholar] [CrossRef] [PubMed]
- Chapurin, N.; Khan, S.; Gutierrez, J.; Soler, Z.M. Economics of Medical and Surgical Management of Chronic Rhinosinusitis with Nasal Polyps: A Contemporary Review. Am. J. Rhinol. Allergy 2023, 37, 227–231. [Google Scholar] [CrossRef]
- Lee, R.J.; Chen, B.; Redding, K.M.; Margolskee, R.F.; Cohen, N.A. Mouse nasal epithelial innate immune responses to Pseudomonas aeruginosa quorum-sensing molecules require taste signaling components. Innate Immun. 2014, 20, 606–617. [Google Scholar] [CrossRef]
- Hariri, B.M.; McMahon, D.B.; Chen, B.; Freund, J.R.; Mansfield, C.J.; Doghramji, L.J.; Adappa, N.D.; Palmer, J.N.; Kennedy, D.W.; Reed, D.R.; et al. Flavones modulate respiratory epithelial innate immunity: Anti-inflammatory effects and activation of the T2R14 receptor. J. Biol. Chem. 2017, 292, 8484–8497. [Google Scholar] [CrossRef]
- Carey, R.M.; Adappa, N.D.; Palmer, J.N.; Lee, R.J. Neuropeptide Y Reduces Nasal Epithelial T2R Bitter Taste Receptor-Stimulated Nitric Oxide Production. Nutrients 2021, 13, 3392. [Google Scholar] [CrossRef]
- Carey, R.M.; Hariri, B.M.; Adappa, N.D.; Palmer, J.N.; Lee, R.J. HSP90 Modulates T2R Bitter Taste Receptor Nitric Oxide Production and Innate Immune Responses in Human Airway Epithelial Cells and Macrophages. Cells 2022, 11, 1478. [Google Scholar] [CrossRef]
- McMahon, D.B.; Kuek, L.E.; Johnson, M.E.; Johnson, P.O.; Horn, R.L.J.; Carey, R.M.; Adappa, N.D.; Palmer, J.N.; Lee, R.J. The bitter end: T2R bitter receptor agonists elevate nuclear calcium and induce apoptosis in non-ciliated airway epithelial cells. Cell Calcium 2022, 101, 102499. [Google Scholar] [CrossRef]
- Carey, R.M.; Palmer, J.N.; Adappa, N.D.; Lee, R.J. Loss of CFTR function is associated with reduced bitter taste receptor-stimulated nitric oxide innate immune responses in nasal epithelial cells and macrophages. Front. Immunol. 2023, 14, 1096242. [Google Scholar] [CrossRef]
- Shah, A.S.; Ben-Shahar, Y.; Moninger, T.O.; Kline, J.N.; Welsh, M.J. Motile cilia of human airway epithelia are chemosensory. Science 2009, 325, 1131–1134. [Google Scholar] [CrossRef]
- Kuek, L.E.; Lee, R.J. First contact: The role of respiratory cilia in host-pathogen interactions in the airways. Am. J. Physiol.-Lung Cell. Mol. Physiol. 2020, 319, L603–L619. [Google Scholar] [CrossRef] [PubMed]
- Tizzano, M.; Gulbransen, B.D.; Vandenbeuch, A.; Clapp, T.R.; Herman, J.P.; Sibhatu, H.M.; Churchill, M.E.; Silver, W.L.; Kinnamon, S.C.; Finger, T.E. Nasal chemosensory cells use bitter taste signaling to detect irritants and bacterial signals. Proc. Natl. Acad. Sci. USA 2010, 107, 3210–3215. [Google Scholar] [CrossRef] [PubMed]
- Jones-Carson, J.; Laughlin, J.R.; Stewart, A.L.; Voskuil, M.I.; Vazquez-Torres, A. Nitric oxide-dependent killing of aerobic, anaerobic and persistent Burkholderia pseudomallei. Nitric Oxide 2012, 27, 25–31. [Google Scholar] [CrossRef]
- Bogdan, C. Nitric oxide synthase in innate and adaptive immunity: An update. Trends Immunol. 2015, 36, 161–178. [Google Scholar] [CrossRef]
- Akaberi, D.; Krambrich, J.; Ling, J.; Luni, C.; Hedenstierna, G.; Jarhult, J.D.; Lennerstrand, J.; Lundkvist, A. Mitigation of the replication of SARS-CoV-2 by nitric oxide in vitro. Redox Biol. 2020, 37, 101734. [Google Scholar] [CrossRef] [PubMed]
- Wiegand, S.B.; Traeger, L.; Nguyen, H.K.; Rouillard, K.R.; Fischbach, A.; Zadek, F.; Ichinose, F.; Schoenfisch, M.H.; Carroll, R.W.; Bloch, D.B.; et al. Antimicrobial effects of nitric oxide in murine models of Klebsiella pneumonia. Redox Biol. 2021, 39, 101826. [Google Scholar] [CrossRef]
- Adappa, N.D.; Howland, T.J.; Palmer, J.N.; Kennedy, D.W.; Doghramji, L.; Lysenko, A.; Reed, D.R.; Lee, R.J.; Cohen, N.A. Genetics of the taste receptor T2R38 correlates with chronic rhinosinusitis necessitating surgical intervention. Int. Forum Allergy Rhinol. 2013, 3, 184–187. [Google Scholar] [CrossRef] [PubMed]
- Adappa, N.D.; Zhang, Z.; Palmer, J.N.; Kennedy, D.W.; Doghramji, L.; Lysenko, A.; Reed, D.R.; Scott, T.; Zhao, N.W.; Owens, D.; et al. The bitter taste receptor T2R38 is an independent risk factor for chronic rhinosinusitis requiring sinus surgery. Int. Forum Allergy Rhinol. 2014, 4, 3–7. [Google Scholar] [CrossRef] [PubMed]
- Adappa, N.D.; Truesdale, C.M.; Workman, A.D.; Doghramji, L.; Mansfield, C.; Kennedy, D.W.; Palmer, J.N.; Cowart, B.J.; Cohen, N.A. Correlation of T2R38 taste phenotype and in vitro biofilm formation from nonpolypoid chronic rhinosinusitis patients. Int. Forum Allergy Rhinol. 2016, 6, 783–791. [Google Scholar] [CrossRef]
- Gallo, S.; Grossi, S.; Montrasio, G.; Binelli, G.; Cinquetti, R.; Simmen, D.; Castelnuovo, P.; Campomenosi, P. TAS2R38 taste receptor gene and chronic rhinosinusitis: New data from an Italian population. BMC Med. Genet. 2016, 17, 54. [Google Scholar] [CrossRef]
- Dzaman, K.; Zagor, M.; Sarnowska, E.; Krzeski, A.; Kantor, I. The correlation of TAS2R38 gene variants with higher risk for chronic rhinosinusitis in Polish patients. Otolaryngol. Pol. 2016, 70, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Mfuna Endam, L.; Filali-Mouhim, A.; Boisvert, P.; Boulet, L.P.; Bosse, Y.; Desrosiers, M. Genetic variations in taste receptors are associated with chronic rhinosinusitis: A replication study. Int. Forum Allergy Rhinol. 2014, 4, 200–206. [Google Scholar] [CrossRef]
- Purnell, P.R.; Addicks, B.L.; Zalzal, H.G.; Shapiro, S.; Wen, S.; Ramadan, H.H.; Setola, V.; Siderovski, D.P. Single Nucleotide Polymorphisms in Chemosensory Pathway Genes GNB3, TAS2R19, and TAS2R38 Are Associated with Chronic Rhinosinusitis. Int. Arch. Allergy Immunol. 2019, 180, 72–78. [Google Scholar] [CrossRef]
- Rom, D.I.; Christensen, J.M.; Alvarado, R.; Sacks, R.; Harvey, R.J. The impact of bitter taste receptor genetics on culturable bacteria in chronic rhinosinusitis. Rhinology 2017, 55, 90–94. [Google Scholar] [CrossRef]
- Cantone, E.; Negri, R.; Roscetto, E.; Grassia, R.; Catania, M.R.; Capasso, P.; Maffei, M.; Soriano, A.A.; Leone, C.A.; Iengo, M.; et al. In Vivo Biofilm Formation, Gram-Negative Infections and TAS2R38 Polymorphisms in CRSw NP Patients. Laryngoscope 2018, 128, E339–E345. [Google Scholar] [CrossRef]
- Piatti, G.; Ambrosetti, U.; Alde, M.; Girotto, G.; Concas, M.P.; Torretta, S. Chronic Rhinosinusitis: T2r38 Genotyping and Nasal Cytology in Primary Ciliary Dyskinesia. Laryngoscope 2023, 133, 248–254. [Google Scholar] [CrossRef]
- Takemoto, K.; Lomude, L.S.; Takeno, S.; Kawasumi, T.; Okamoto, Y.; Hamamoto, T.; Ishino, T.; Ando, Y.; Ishikawa, C.; Ueda, T. Functional Alteration and Differential Expression of the Bitter Taste Receptor T2R38 in Human Paranasal Sinus in Patients with Chronic Rhinosinusitis. Int. J. Mol. Sci. 2023, 24, 4499. [Google Scholar] [CrossRef] [PubMed]
- Jaggupilli, A.; Singh, N.; Jesus, V.C.; Duan, K.; Chelikani, P. Characterization of the Binding Sites for Bacterial Acyl Homoserine Lactones (AHLs) on Human Bitter Taste Receptors (T2Rs). ACS Infect. Dis. 2018, 4, 1146–1156. [Google Scholar] [CrossRef] [PubMed]
- Tran, H.T.T.; Herz, C.; Ruf, P.; Stetter, R.; Lamy, E. Human T2R38 Bitter Taste Receptor Expression in Resting and Activated Lymphocytes. Front. Immunol. 2018, 9, 2949. [Google Scholar] [CrossRef] [PubMed]
- Gaida, M.M.; Dapunt, U.; Hansch, G.M. Sensing developing biofilms: The bitter receptor T2R38 on myeloid cells. Pathog. Dis. 2016, 74, ftw004. [Google Scholar] [CrossRef]
- Maurer, S.; Wabnitz, G.H.; Kahle, N.A.; Stegmaier, S.; Prior, B.; Giese, T.; Gaida, M.M.; Samstag, Y.; Hansch, G.M. Tasting Pseudomonas aeruginosa Biofilms: Human Neutrophils Express the Bitter Receptor T2R38 as Sensor for the Quorum Sensing Molecule N-(3-Oxododecanoyl)-l-Homoserine Lactone. Front. Immunol. 2015, 6, 369. [Google Scholar] [CrossRef]
- Malki, A.; Fiedler, J.; Fricke, K.; Ballweg, I.; Pfaffl, M.W.; Krautwurst, D. Class I odorant receptors, TAS1R and TAS2R taste receptors, are markers for subpopulations of circulating leukocytes. J. Leukoc. Biol. 2015, 97, 533–545. [Google Scholar] [CrossRef]
- Grassin-Delyle, S.; Salvator, H.; Mantov, N.; Abrial, C.; Brollo, M.; Faisy, C.; Naline, E.; Couderc, L.J.; Devillier, P. Bitter Taste Receptors (TAS2Rs) in Human Lung Macrophages: Receptor Expression and Inhibitory Effects of TAS2R Agonists. Front. Physiol. 2019, 10, 1267. [Google Scholar] [CrossRef]
- Kobayashi, D.; Watarai, T.; Ozawa, M.; Kanda, Y.; Saika, F.; Kiguchi, N.; Takeuchi, A.; Ikawa, M.; Matsuzaki, S.; Katakai, T. Tas2R signaling enhances mouse neutrophil migration via a ROCK-dependent pathway. Front. Immunol. 2022, 13, 973880. [Google Scholar] [CrossRef]
- Gopallawa, I.; Freund, J.R.; Lee, R.J. Bitter taste receptors stimulate phagocytosis in human macrophages through calcium, nitric oxide, and cyclic-GMP signaling. Cell. Mol. Life Sci. 2021, 78, 271–286. [Google Scholar] [CrossRef]
- Ribeiro, C.M.P.; Higgs, M.G.; Muhlebach, M.S.; Wolfgang, M.C.; Borgatti, M.; Lampronti, I.; Cabrini, G. Revisiting Host-Pathogen Interactions in Cystic Fibrosis Lungs in the Era of CFTR Modulators. Int. J. Mol. Sci. 2023, 24, 5010. [Google Scholar] [CrossRef]
- Adappa, N.D.; Workman, A.D.; Hadjiliadis, D.; Dorgan, D.J.; Frame, D.; Brooks, S.; Doghramji, L.; Palmer, J.N.; Mansfield, C.; Reed, D.R.; et al. T2R38 genotype is correlated with sinonasal quality of life in homozygous DeltaF508 cystic fibrosis patients. Int. Forum Allergy Rhinol. 2016, 6, 356–361. [Google Scholar] [CrossRef]
- Turnbull, A.R.; Murphy, R.; Behrends, V.; Lund-Palau, H.; Simbo, A.; Mariveles, M.; Alton, E.; Bush, A.; Shoemark, A.; Davies, J.C. Impact of T2R38 Receptor Polymorphisms on Pseudomonas aeruginosa Infection in Cystic Fibrosis. Am. J. Respir. Crit. Care Med. 2018, 197, 1635–1638. [Google Scholar] [CrossRef]
- Castaldo, A.; Cernera, G.; Iacotucci, P.; Cimbalo, C.; Gelzo, M.; Comegna, M.; Di Lullo, A.M.; Tosco, A.; Carnovale, V.; Raia, V.; et al. TAS2R38 is a novel modifier gene in patients with cystic fibrosis. Sci. Rep. 2020, 10, 5806. [Google Scholar] [CrossRef]
- Totani, L.; Plebani, R.; Piccoli, A.; Di Silvestre, S.; Lanuti, P.; Recchiuti, A.; Cianci, E.; Dell’Elba, G.; Sacchetti, S.; Patruno, S.; et al. Mechanisms of endothelial cell dysfunction in cystic fibrosis. Biochim. Biophys. Acta 2017, 1863, 3243–3253. [Google Scholar] [CrossRef]
- Wilson, A.; Altman, K.; Schindler, T.; Schwarzenberg, S.J. Updates in Nutrition Management of Cystic Fibrosis in the Highly Effective Modulator Era. Clin. Chest Med. 2022, 43, 727–742. [Google Scholar] [CrossRef] [PubMed]
- Pezzulo, A.A.; Tudas, R.A.; Stewart, C.G.; Buonfiglio, L.G.V.; Lindsay, B.D.; Taft, P.J.; Gansemer, N.D.; Zabner, J. HSP90 inhibitor geldanamycin reverts IL-13- and IL-17-induced airway goblet cell metaplasia. J. Clin. Investig. 2019, 129, 744–758. [Google Scholar] [CrossRef]
- Lemos, J.A.; Palmer, S.R.; Zeng, L.; Wen, Z.T.; Kajfasz, J.K.; Freires, I.A.; Abranches, J.; Brady, L.J. The Biology of Streptococcus mutans. Microbiol. Spectr. 2019, 7, 7. [Google Scholar] [CrossRef]
- Arguedas, A.; Trzcinski, K.; O’Brien, K.L.; Ferreira, D.M.; Wyllie, A.L.; Weinberger, D.; Danon, L.; Pelton, S.I.; Azzari, C.; Hammitt, L.L.; et al. Upper respiratory tract colonization with Streptococcus pneumoniae in adults. Expert Rev. Vaccines 2020, 19, 353–366. [Google Scholar] [CrossRef] [PubMed]
- Grousd, J.A.; Rich, H.E.; Alcorn, J.F. Host-Pathogen Interactions in Gram-Positive Bacterial Pneumonia. Clin. Microbiol. Rev. 2019, 32, e00107-18. [Google Scholar] [CrossRef] [PubMed]
- Orlova, E.; Dudding, T.; Chernus, J.M.; Alotaibi, R.N.; Haworth, S.; Crout, R.J.; Lee, M.K.; Mukhopadhyay, N.; Feingold, E.; Levy, S.M.; et al. Association of Early Childhood Caries with Bitter Taste Receptors: A Meta-Analysis of Genome-Wide Association Studies and Transcriptome-Wide Association Study. Genes 2022, 14, 59. [Google Scholar] [CrossRef] [PubMed]
- de Jesus, V.C.; Mittermuller, B.A.; Hu, P.; Schroth, R.J.; Chelikani, P. Genetic variants in taste genes play a role in oral microbial composition and severe early childhood caries. iScience 2022, 25, 105489. [Google Scholar] [CrossRef] [PubMed]
- de Jesus, V.C.; Singh, M.; Schroth, R.J.; Chelikani, P.; Hitchon, C.A. Association of Bitter Taste Receptor T2R38 Polymorphisms, Oral Microbiota, and Rheumatoid Arthritis. Curr. Issues Mol. Biol. 2021, 43, 1460–1472. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Xi, R.; Liu, J.; Peng, X.; Zhao, L.; Zhou, X.; Li, J.; Zheng, X.; Xu, X. TAS2R16 Activation Suppresses LPS-Induced Cytokine Expression in Human Gingival Fibroblasts. Front. Immunol. 2021, 12, 726546. [Google Scholar] [CrossRef] [PubMed]
- Braun, T.; Mack, B.; Kramer, M.F. Solitary chemosensory cells in the respiratory and vomeronasal epithelium of the human nose: A pilot study. Rhinology 2011, 49, 507–512. [Google Scholar] [CrossRef]
- Finger, T.E.; Bottger, B.; Hansen, A.; Anderson, K.T.; Alimohammadi, H.; Silver, W.L. Solitary chemoreceptor cells in the nasal cavity serve as sentinels of respiration. Proc. Natl. Acad. Sci. USA 2003, 100, 8981–8986. [Google Scholar] [CrossRef]
- Gulbransen, B.; Silver, W.; Finger, T.E. Solitary chemoreceptor cell survival is independent of intact trigeminal innervation. J. Comp. Neurol. 2008, 508, 62–71. [Google Scholar] [CrossRef]
- Saunders, C.J.; Reynolds, S.D.; Finger, T.E. Chemosensory brush cells of the trachea. A stable population in a dynamic epithelium. Am. J. Respir Cell. Mol. Biol. 2013, 49, 190–196. [Google Scholar] [CrossRef]
- Sbarbati, A.; Osculati, F. Solitary chemosensory cells in mammals? Cells Tissues Organs 2003, 175, 51–55. [Google Scholar] [CrossRef]
- Tizzano, M.; Merigo, F.; Sbarbati, A. Evidence of solitary chemosensory cells in a large mammal: The diffuse chemosensory system in Bos taurus airways. J. Anat. 2006, 209, 333–337. [Google Scholar] [CrossRef]
- Saunders, C.J.; Christensen, M.; Finger, T.E.; Tizzano, M. Cholinergic neurotransmission links solitary chemosensory cells to nasal inflammation. Proc. Natl. Acad. Sci. USA 2014, 111, 6075–6080. [Google Scholar] [CrossRef]
- Reid, L.; Meyrick, B.; Antony, V.B.; Chang, L.Y.; Crapo, J.D.; Reynolds, H.Y. The mysterious pulmonary brush cell: A cell in search of a function. Am. J. Respir. Crit. Care Med. 2005, 172, 136–139. [Google Scholar] [CrossRef]
- Brody, A.R. The brush cell. Am. J. Respir. Crit. Care Med. 2005, 172, 1349. [Google Scholar] [CrossRef]
- Gomi, T.; Kimura, A.; Kikuchi, Y.; Higashi, K.; Tsuchiya, H.; Sasa, S.; Kishi, K. Electron-microscopic observations of the alveolar brush cell of the rat. Cells Tissues Organs 1991, 141, 294–301. [Google Scholar] [CrossRef] [PubMed]
- Meyrick, B.; Reid, L. The alveolar brush cell in rat lung--a third pneumonocyte. J. Ultrastruct. Res. 1968, 23, 71–80. [Google Scholar] [CrossRef]
- Chang, L.Y.; Mercer, R.R.; Crapo, J.D. Differential distribution of brush cells in the rat lung. Anat. Rec. 1986, 216, 49–54. [Google Scholar] [CrossRef] [PubMed]
- Hijiya, K. Electron microscope study of the alveolar brush cell. J. Electron Microsc. 1978, 27, 223–227. [Google Scholar]
- DiMaio, M.F.; Dische, R.; Gordon, R.E.; Kattan, M. Alveolar brush cells in an infant with desquamative interstitial pneumonitis. Pediatr. Pulmonol. 1988, 4, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Montoro, D.T.; Haber, A.L.; Biton, M.; Vinarsky, V.; Lin, B.; Birket, S.E.; Yuan, F.; Chen, S.; Leung, H.M.; Villoria, J.; et al. A revised airway epithelial hierarchy includes CFTR-expressing ionocytes. Nature 2018, 560, 319–324. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.J.; Cohen, N.A. Role of the bitter taste receptor T2R38 in upper respiratory infection and chronic rhinosinusitis. Curr. Opin. Allergy Clin. Immunol. 2015, 15, 14–20. [Google Scholar] [CrossRef]
- Lee, R.J.; Cohen, N.A. Sinonasal solitary chemosensory cells "taste" the upper respiratory environment to regulate innate immunity. Am. J. Rhinol. Allergy 2014, 28, 366–373. [Google Scholar] [CrossRef]
- Baker, E.H.; Baines, D.L. Airway Glucose Homeostasis: A New Target in the Prevention and Treatment of Pulmonary Infection. Chest 2018, 153, 507–514. [Google Scholar] [CrossRef] [PubMed]
- Hatten, K.M.; Palmer, J.N.; Lee, R.J.; Adappa, N.D.; Kennedy, D.W.; Cohen, N.A. Corticosteroid use does not alter nasal mucus glucose in chronic rhinosinusitis. Otolaryngol.—Head Neck Surg. 2015, 152, 1140–1144. [Google Scholar] [CrossRef]
- Smith, N.J.; Grant, J.N.; Moon, J.I.; So, S.S.; Finch, A.M. Critically evaluating sweet taste receptor expression and signaling through a molecular pharmacology lens. FEBS J. 2021, 288, 2660–2672. [Google Scholar] [CrossRef] [PubMed]
- von Molitor, E.; Riedel, K.; Krohn, M.; Hafner, M.; Rudolf, R.; Cesetti, T. Sweet Taste Is Complex: Signaling Cascades and Circuits Involved in Sweet Sensation. Front. Hum. Neurosci. 2021, 15, 667709. [Google Scholar] [CrossRef]
- Wang, H.; Matsumoto, I.; Jiang, P. Immune Regulatory Roles of Cells Expressing Taste Signaling Elements in Nongustatory Tissues. Handb. Exp. Pharmacol. 2022, 275, 271–293. [Google Scholar] [CrossRef]
- Kim, D.; Woo, J.A.; Geffken, E.; An, S.S.; Liggett, S.B. Coupling of Airway Smooth Muscle Bitter Taste Receptors to Intracellular Signaling and Relaxation Is via Galphai1,2,3. Am. J. Respir. Cell Mol. Biol. 2017, 56, 762–771. [Google Scholar] [CrossRef]
- Gusach, A.; Garcia-Nafria, J.; Tate, C.G. New insights into GPCR coupling and dimerisation from cryo-EM structures. Curr. Opin. Struct. Biol. 2023, 80, 102574. [Google Scholar] [CrossRef] [PubMed]
- Lam, H.; Oh, D.C.; Cava, F.; Takacs, C.N.; Clardy, J.; de Pedro, M.A.; Waldor, M.K. D-amino acids govern stationary phase cell wall remodeling in bacteria. Science 2009, 325, 1552–1555. [Google Scholar] [CrossRef] [PubMed]
- Brandenburg, K.S.; Rodriguez, K.J.; McAnulty, J.F.; Murphy, C.J.; Abbott, N.L.; Schurr, M.J.; Czuprynski, C.J. Tryptophan inhibits biofilm formation by Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2013, 57, 1921–1925. [Google Scholar] [CrossRef]
- Yu, C.; Wu, J.; Contreras, A.E.; Li, Q. Control of nanofiltration membrane biofouling by Pseudomonas aeruginosa using d-tyrosine. J. Membr. Sci. 2012, 423, 487–494. [Google Scholar] [CrossRef]
- Harmata, A.J.; Ma, Y.; Sanchez, C.J.; Zienkiewicz, K.J.; Elefteriou, F.; Wenke, J.C.; Guelcher, S.A. D-amino acid inhibits biofilm but not new bone formation in an ovine model. Clin. Orthop. Relat. Res. 2015, 473, 3951–3961. [Google Scholar] [CrossRef] [PubMed]
- Hochbaum, A.I.; Kolodkin-Gal, I.; Foulston, L.; Kolter, R.; Aizenberg, J.; Losick, R. Inhibitory effects of D-amino acids on Staphylococcus aureus biofilm development. J. Bacteriol. 2011, 193, 5616–5622. [Google Scholar] [CrossRef] [PubMed]
- Ramon-Perez, M.L.; Diaz-Cedillo, F.; Ibarra, J.A.; Torales-Cardena, A.; Rodriguez-Martinez, S.; Jan-Roblero, J.; Cancino-Diaz, M.E.; Cancino-Diaz, J.C. D-Amino acids inhibit biofilm formation in Staphylococcus epidermidis strains from ocular infections. J. Med. Microbiol. 2014, 63, 1369–1376. [Google Scholar] [CrossRef]
- Kolodkin-Gal, I.; Romero, D.; Cao, S.; Clardy, J.; Kolter, R.; Losick, R. D-amino acids trigger biofilm disassembly. Science 2010, 328, 627–629. [Google Scholar] [CrossRef]
- Sanchez, C.J., Jr.; Akers, K.S.; Romano, D.R.; Woodbury, R.L.; Hardy, S.K.; Murray, C.K.; Wenke, J.C. D-amino acids enhance the activity of antimicrobials against biofilms of clinical wound isolates of Staphylococcus aureus and Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2014, 58, 4353–4361. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Liu, Y. d-Amino acid mitigated membrane biofouling and promoted biofilm detachment. J. Membr. Sci. 2011, 376, 266–275. [Google Scholar] [CrossRef]
- Xua, H.; Liu, Y. Reduced microbial attachment by d-amino acid-inhibited AI-2 and EPS production. Water Res. 2011, 45, 5796–5804. [Google Scholar] [CrossRef]
- Leiman, S.A.; May, J.M.; Lebar, M.D.; Kahne, D.; Kolter, R.; Losick, R. D-amino acids indirectly inhibit biofilm formation in Bacillus subtilis by interfering with protein synthesis. J. Bacteriol. 2013, 195, 5391–5395. [Google Scholar] [CrossRef]
- Sarkar, S.; Pires, M.M. d-Amino acids do not inhibit biofilm formation in Staphylococcus aureus. PLoS ONE 2015, 10, e0117613. [Google Scholar] [CrossRef]
- Yu, C.; Li, X.; Zhang, N.; Wen, D.; Liu, C.; Li, Q. Inhibition of biofilm formation by D-tyrosine: Effect of bacterial type and D-tyrosine concentration. Water Res. 2016, 92, 173–179. [Google Scholar] [CrossRef]
- Li, Y.; Jia, R.; Al-Mahamedh, H.H.; Xu, D.; Gu, T. Enhanced Biocide Mitigation of Field Biofilm Consortia by a Mixture of D-Amino Acids. Front. Microbiol. 2016, 7, 896. [Google Scholar] [CrossRef] [PubMed]
- She, P.; Chen, L.; Liu, H.; Zou, Y.; Luo, Z.; Koronfel, A.; Wu, Y. The effects of D-Tyrosine combined with amikacin on the biofilms of Pseudomonas aeruginosa. Microb. Pathog. 2015, 86, 38–44. [Google Scholar] [CrossRef] [PubMed]
- Cava, F.; Lam, H.; de Pedro, M.A.; Waldor, M.K. Emerging knowledge of regulatory roles of D-amino acids in bacteria. Cell. Mol. Life Sci. 2011, 68, 817–831. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, M.; Kunisawa, A.; Hattori, T.; Kawana, S.; Kitada, Y.; Tamada, H.; Kawano, S.; Hayakawa, Y.; Iida, J.; Fukusaki, E. Free D-amino acids produced by commensal bacteria in the colonic lumen. Sci. Rep. 2018, 8, 17915. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, J. d-Amino Acids and Lactic Acid Bacteria. Microorganisms 2019, 7, 690. [Google Scholar] [CrossRef]
- Marcone, G.L.; Rosini, E.; Crespi, E.; Pollegioni, L. D-amino acids in foods. Appl. Microbiol. Biotechnol. 2020, 104, 555–574. [Google Scholar] [CrossRef]
- Kapil, S.; Bhattu, M.; Sharma, V.; Kumar, T. Racemization rate and biomolecular characterization of D-serine synthesizing bacteria Bacillus tequilensis A1C1. Lett. Appl. Microbiol. 2023, 76, ovac017. [Google Scholar] [CrossRef]
- Bassoli, A.; Borgonovo, G.; Caremoli, F.; Mancuso, G. The taste of D- and L-amino acids: In vitro binding assays with cloned human bitter (TAS2Rs) and sweet (TAS1R2/TAS1R3) receptors. Food Chem. 2014, 150, 27–33. [Google Scholar] [CrossRef]
- Radkov, A.D.; Moe, L.A. Bacterial synthesis of D-amino acids. Appl. Microbiol. Biotechnol. 2014, 98, 5363–5374. [Google Scholar] [CrossRef]
- Suzuki, M.; Sujino, T.; Chiba, S.; Harada, Y.; Goto, M.; Takahashi, R.; Mita, M.; Hamase, K.; Kanai, T.; Ito, M.; et al. Host-microbe cross-talk governs amino acid chirality to regulate survival and differentiation of B cells. Sci. Adv. 2021, 7, eabd6480. [Google Scholar] [CrossRef]
- Kohanski, M.A.; Workman, A.D.; Patel, N.N.; Hung, L.Y.; Shtraks, J.P.; Chen, B.; Blasetti, M.; Doghramji, L.; Kennedy, D.W.; Adappa, N.D.; et al. Solitary chemosensory cells are a primary epithelial source of IL-25 in patients with chronic rhinosinusitis with nasal polyps. J. Allergy Clin. Immunol. 2018, 142, 460–469. [Google Scholar] [CrossRef]
- Patel, N.N.; Triantafillou, V.; Maina, I.W.; Workman, A.D.; Tong, C.C.L.; Kuan, E.C.; Papagiannopoulos, P.; Bosso, J.V.; Adappa, N.D.; Palmer, J.N.; et al. Fungal extracts stimulate solitary chemosensory cell expansion in noninvasive fungal rhinosinusitis. Int. Forum Allergy Rhinol. 2019, 9, 730–737. [Google Scholar] [CrossRef]
- Tizzano, M.; Finger, T.E. Chemosensors in the nose: Guardians of the airways. Physiology 2013, 28, 51–60. [Google Scholar] [CrossRef]
- Krasteva, G.; Canning, B.J.; Hartmann, P.; Veres, T.Z.; Papadakis, T.; Muhlfeld, C.; Schliecker, K.; Tallini, Y.N.; Braun, A.; Hackstein, H.; et al. Cholinergic chemosensory cells in the trachea regulate breathing. Proc. Natl. Acad. Sci. USA 2011, 108, 9478–9483. [Google Scholar] [CrossRef]
- Tian, X.; Ding, H.; Ke, W.; Wang, L. Quorum Sensing in Fungal Species. Annu. Rev. Microbiol. 2021, 75, 449–469. [Google Scholar] [CrossRef] [PubMed]
- Mehmood, A.; Liu, G.; Wang, X.; Meng, G.; Wang, C.; Liu, Y. Fungal Quorum-Sensing Molecules and Inhibitors with Potential Antifungal Activity: A Review. Molecules 2019, 24, 1950. [Google Scholar] [CrossRef]
- Wiener, A.; Shudler, M.; Levit, A.; Niv, M.Y. BitterDB: A database of bitter compounds. Nucleic Acids Res. 2012, 40, D413–D419. [Google Scholar] [CrossRef] [PubMed]
- Kuek, L.E.; McMahon, D.B.; Ma, R.Z.; Miller, Z.A.; Jolivert, J.F.; Adappa, N.D.; Palmer, J.N.; Lee, R.J. Cilia Stimulatory and Antibacterial Activities of T2R Bitter Taste Receptor Agonist Diphenhydramine: Insights into Repurposing Bitter Drugs for Nasal Infections. Pharmaceuticals 2022, 15, 452. [Google Scholar] [CrossRef]
- Miller, Z.A.; Jolivert, J.F.; Ma, R.Z.; Muthuswami, S.; Mueller, A.; McMahon, D.B.; Carey, R.M.; Lee, R.J. Lidocaine Induces Apoptosis in Head and Neck Squamous Cell Carcinoma Cells Through Activation of Bitter Taste Receptor T2R14. bioRxiv 2022. [Google Scholar] [CrossRef]
- Hariri, B.M.; McMahon, D.B.; Chen, B.; Adappa, N.D.; Palmer, J.N.; Kennedy, D.W.; Lee, R.J. Plant flavones enhance antimicrobial activity of respiratory epithelial cell secretions against Pseudomonas aeruginosa. PLoS ONE 2017, 12, e0185203. [Google Scholar] [CrossRef] [PubMed]
- Tran, H.T.T.; Stetter, R.; Herz, C.; Spottel, J.; Krell, M.; Hanschen, F.S.; Schreiner, M.; Rohn, S.; Behrens, M.; Lamy, E. Allyl Isothiocyanate: A TAS2R38 Receptor-Dependent Immune Modulator at the Interface Between Personalized Medicine and Nutrition. Front. Immunol. 2021, 12, 669005. [Google Scholar] [CrossRef] [PubMed]
- Roland, W.S.; Vincken, J.P.; Gouka, R.J.; van Buren, L.; Gruppen, H.; Smit, G. Soy isoflavones and other isoflavonoids activate the human bitter taste receptors hTAS2R14 and hTAS2R39. J. Agric. Food Chem. 2011, 59, 11764–11771. [Google Scholar] [CrossRef] [PubMed]
- Behrens, M.; Gu, M.; Fan, S.; Huang, C.; Meyerhof, W. Bitter substances from plants used in traditional Chinese medicine exert biased activation of human bitter taste receptors. Chem. Biol. Drug Des. 2017, 91, 422–433. [Google Scholar] [CrossRef]
- Roland, W.S.; Gouka, R.J.; Gruppen, H.; Driesse, M.; van Buren, L.; Smit, G.; Vincken, J.P. 6-methoxyflavanones as bitter taste receptor blockers for hTAS2R39. PLoS ONE 2014, 9, e94451. [Google Scholar] [CrossRef]
- Fierro, F.; Peri, L.; Hübner, H.; Tabor-Schkade, A.; Waterloo, L.; Löber, S.; Pfeiffer, T.; Weikert, D.; Dingjan, T.; Margulis, E.; et al. Inhibiting a promiscuous GPCR: Iterative discovery of bitter taste receptor ligands. bioRxiv 2022. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Alashi, A.M.; Singh, N.; Liu, K.; Chelikani, P.; Aluko, R.E. Beef Protein-Derived Peptides as Bitter Taste Receptor T2R4 Blockers. J. Agric. Food Chem. 2018, 66, 4902–4912. [Google Scholar] [CrossRef]
- Pydi, S.P.; Sobotkiewicz, T.; Billakanti, R.; Bhullar, R.P.; Loewen, M.C.; Chelikani, P. Amino acid derivatives as bitter taste receptor (T2R) blockers. J. Biol. Chem. 2014, 289, 25054–25066. [Google Scholar] [CrossRef]
- Greene, T.A.; Alarcon, S.; Thomas, A.; Berdougo, E.; Doranz, B.J.; Breslin, P.A.; Rucker, J.B. Probenecid inhibits the human bitter taste receptor TAS2R16 and suppresses bitter perception of salicin. PLoS ONE 2011, 6, e20123. [Google Scholar] [CrossRef]
- Kim, M.J.; Son, H.J.; Kim, Y.; Misaka, T.; Rhyu, M.R. Umami-bitter interactions: The suppression of bitterness by umami peptides via human bitter taste receptor. Biochem. Biophys. Res. Commun. 2015, 456, 586–590. [Google Scholar] [CrossRef]
- McMahon, D.B.; Jolivert, J.F.; Kuek, L.E.; Adappa, N.D.; Palmer, J.N.; Lee, R.J. Savory Signaling: T1R Umami Receptor Modulates Endoplasmic Reticulum Calcium Store Content and Release Dynamics in Airway Epithelial Cells. Nutrients 2023, 15, 493. [Google Scholar] [CrossRef]
- McMahon, D.B.; Jolivert, J.F.; Kuek, L.E.; Adappa, N.D.; Palmer, J.N.; Lee, R.J. Utilizing the Off-Target Effects of T1R3 Antagonist Lactisole to Enhance Nitric Oxide Production in Basal Airway Epithelial Cells. Nutrients 2023, 15, 517. [Google Scholar] [CrossRef] [PubMed]
- Patel, N.N.; Kohanski, M.A.; Maina, I.W.; Triantafillou, V.; Workman, A.D.; Tong, C.C.L.; Kuan, E.C.; Bosso, J.V.; Adappa, N.D.; Palmer, J.N.; et al. Solitary chemosensory cells producing interleukin-25 and group-2 innate lymphoid cells are enriched in chronic rhinosinusitis with nasal polyps. Int. Forum Allergy Rhinol. 2018, 8, 900–906. [Google Scholar] [CrossRef] [PubMed]
- Carey, R.M.; Freund, J.R.; Hariri, B.M.; Adappa, N.D.; Palmer, J.N.; Lee, R.J. Polarization of protease-activated receptor 2 (PAR-2) signaling is altered during airway epithelial remodeling and deciliation. J. Biol. Chem. 2020, 295, 6721–6740. [Google Scholar] [CrossRef]
- McMahon, D.B.; Carey, R.M.; Kohanski, M.A.; Adappa, N.D.; Palmer, J.N.; Lee, R.J. PAR-2-activated secretion by airway gland serous cells: Role for CFTR and inhibition by Pseudomonas aeruginosa. Am. J. Physiol.-Lung Cell. Mol. Physiol. 2021, 320, L845–L879. [Google Scholar] [CrossRef]
- McMahon, D.B.; Workman, A.D.; Kohanski, M.A.; Carey, R.M.; Freund, J.R.; Hariri, B.M.; Chen, B.; Doghramji, L.J.; Adappa, N.D.; Palmer, J.N.; et al. Protease-activated receptor 2 activates airway apical membrane chloride permeability and increases ciliary beating. FASEB J. 2018, 32, 155–167. [Google Scholar] [CrossRef] [PubMed]
- Rane, C.K.; Jackson, S.R.; Pastore, C.F.; Zhao, G.; Weiner, A.I.; Patel, N.N.; Herbert, D.R.; Cohen, N.A.; Vaughan, A.E. Development of solitary chemosensory cells in the distal lung after severe influenza injury. Am. J. Physiol.-Lung Cell. Mol. Physiol. 2019, 316, L1141–L1149. [Google Scholar] [CrossRef] [PubMed]
- Richter, P.; Sebald, K.; Fischer, K.; Behrens, M.; Schnieke, A.; Somoza, V. Bitter Peptides YFYPEL, VAPFPEVF, and YQEPVLGPVRGPFPIIV, Released during Gastric Digestion of Casein, Stimulate Mechanisms of Gastric Acid Secretion via Bitter Taste Receptors TAS2R16 and TAS2R38. J. Agric. Food Chem. 2022, 70, 11591–11602. [Google Scholar] [CrossRef]
- Zhao, W.; Li, D.; Wang, Y.; Kan, R.; Ji, H.; Su, L.; Yu, Z.; Li, J. Identification and molecular docking of peptides from Mizuhopecten yessoensis myosin as human bitter taste receptor T2R14 blockers. Food Funct. 2021, 12, 11966–11973. [Google Scholar] [CrossRef]
- Xu, Q.; Singh, N.; Hong, H.; Yan, X.; Yu, W.; Jiang, X.; Chelikani, P.; Wu, J. Hen protein-derived peptides as the blockers of human bitter taste receptors T2R4, T2R7 and T2R14. Food Chem. 2019, 283, 621–627. [Google Scholar] [CrossRef]
- Carey, R.M.; McMahon, D.B.; Miller, Z.A.; Kim, T.; Rajasekaran, K.; Gopallawa, I.; Newman, J.G.; Basu, D.; Nead, K.T.; White, E.A.; et al. T2R bitter taste receptors regulate apoptosis and may be associated with survival in head and neck squamous cell carcinoma. Mol. Oncol. 2022, 16, 1474–1492. [Google Scholar] [CrossRef]
- Carey, R.M.; Kim, T.; Cohen, N.A.; Lee, R.J.; Nead, K.T. Impact of sweet, umami, and bitter taste receptor (TAS1R and TAS2R) genomic and expression alterations in solid tumors on survival. Sci. Rep. 2022, 12, 8937. [Google Scholar] [CrossRef]
- Costa, A.R.; Duarte, A.C.; Costa-Brito, A.R.; Goncalves, I.; Santos, C.R.A. Bitter taste signaling in cancer. Life Sci. 2023, 315, 121363. [Google Scholar] [CrossRef]
- Zehentner, S.; Reiner, A.T.; Grimm, C.; Somoza, V. The Role of Bitter Taste Receptors in Cancer: A Systematic Review. Cancers 2021, 13, 5891. [Google Scholar] [CrossRef] [PubMed]
- Sharma, T.; Gupta, A.; Chauhan, R.; Bhat, A.A.; Nisar, S.; Hashem, S.; Akhtar, S.; Ahmad, A.; Haris, M.; Singh, M.; et al. Cross-talk between the microbiome and chronic inflammation in esophageal cancer: Potential driver of oncogenesis. Cancer Metastasis Rev. 2022, 41, 281–299. [Google Scholar] [CrossRef] [PubMed]
- Xia, C.; Cai, Y.; Ren, S.; Xia, C. Role of microbes in colorectal cancer therapy: Cross-talk between the microbiome and tumor microenvironment. Front. Pharmacol. 2022, 13, 1051330. [Google Scholar] [CrossRef] [PubMed]


| Bacteria Species | Strain(s) | Molecule Detected by Taste Receptors | Effects |
|---|---|---|---|
| Pseudomonas aeruginosa | PAO1, Sad36 [159] | Acyl-homoserine lactones (AHLs) | T2R activation, increased nasal cell nitric oxide production, ciliary beating, bacterial killing [160] |
| PAO1 | Pseudomonas quinolone signal (PQS) | T2R activation, increased nasal cell nitric oxide production, ciliary beating [161] | |
| Staphylococcus aureus | M2 [162] | D-amino acids | Activation of T1R2/3, suppression of solitary chemosensory cells [74] |
| M2, clinical isolates | D-amino acids | Activation of T1R3, increased airway glucose transporter expression [163] | |
| M2 | unknown | T2R activation, increased nasal cell nitric oxide production [163,164] | |
| Streptococcus mutans | UA159 | competence stimulating peptides | T2R activation, cytoskeletal remodeling, increased bacterial internalization in gingival keratinocytes [31] |
| Bacillus cereus | ATCC 14579 | unknown | T2R activation, increased nasal cell nitric oxide production |
| Bacillus amyloliquefaciens | amy-1 [165] | exopolysaccharides | T2R activation, glucagon-like peptide 1 secretion [166] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kouakou, Y.I.; Lee, R.J. Interkingdom Detection of Bacterial Quorum-Sensing Molecules by Mammalian Taste Receptors. Microorganisms 2023, 11, 1295. https://doi.org/10.3390/microorganisms11051295
Kouakou YI, Lee RJ. Interkingdom Detection of Bacterial Quorum-Sensing Molecules by Mammalian Taste Receptors. Microorganisms. 2023; 11(5):1295. https://doi.org/10.3390/microorganisms11051295
Chicago/Turabian StyleKouakou, Yobouet Ines, and Robert J. Lee. 2023. "Interkingdom Detection of Bacterial Quorum-Sensing Molecules by Mammalian Taste Receptors" Microorganisms 11, no. 5: 1295. https://doi.org/10.3390/microorganisms11051295
APA StyleKouakou, Y. I., & Lee, R. J. (2023). Interkingdom Detection of Bacterial Quorum-Sensing Molecules by Mammalian Taste Receptors. Microorganisms, 11(5), 1295. https://doi.org/10.3390/microorganisms11051295





