Human Milk Microbiota Profile Affected by Prematurity in Argentinian Lactating Women
Abstract
1. Introduction
2. Materials and Methods
2.1. Subjects, Design, and Sample Collection
2.2. DNA Extraction and 16S rRNA Amplicon Sequencing
2.3. Data Analysis
2.4. Statistical Analysis
3. Results
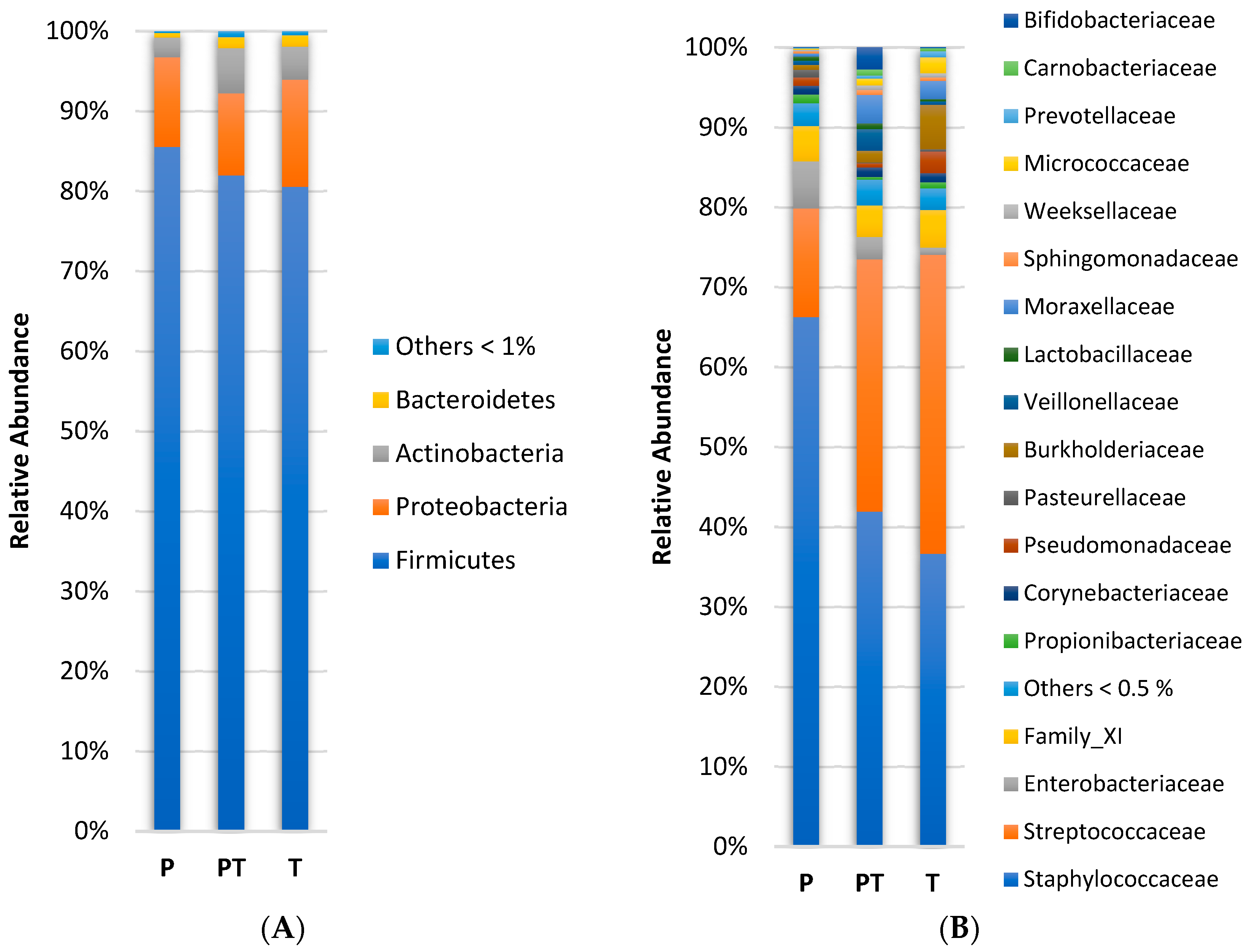
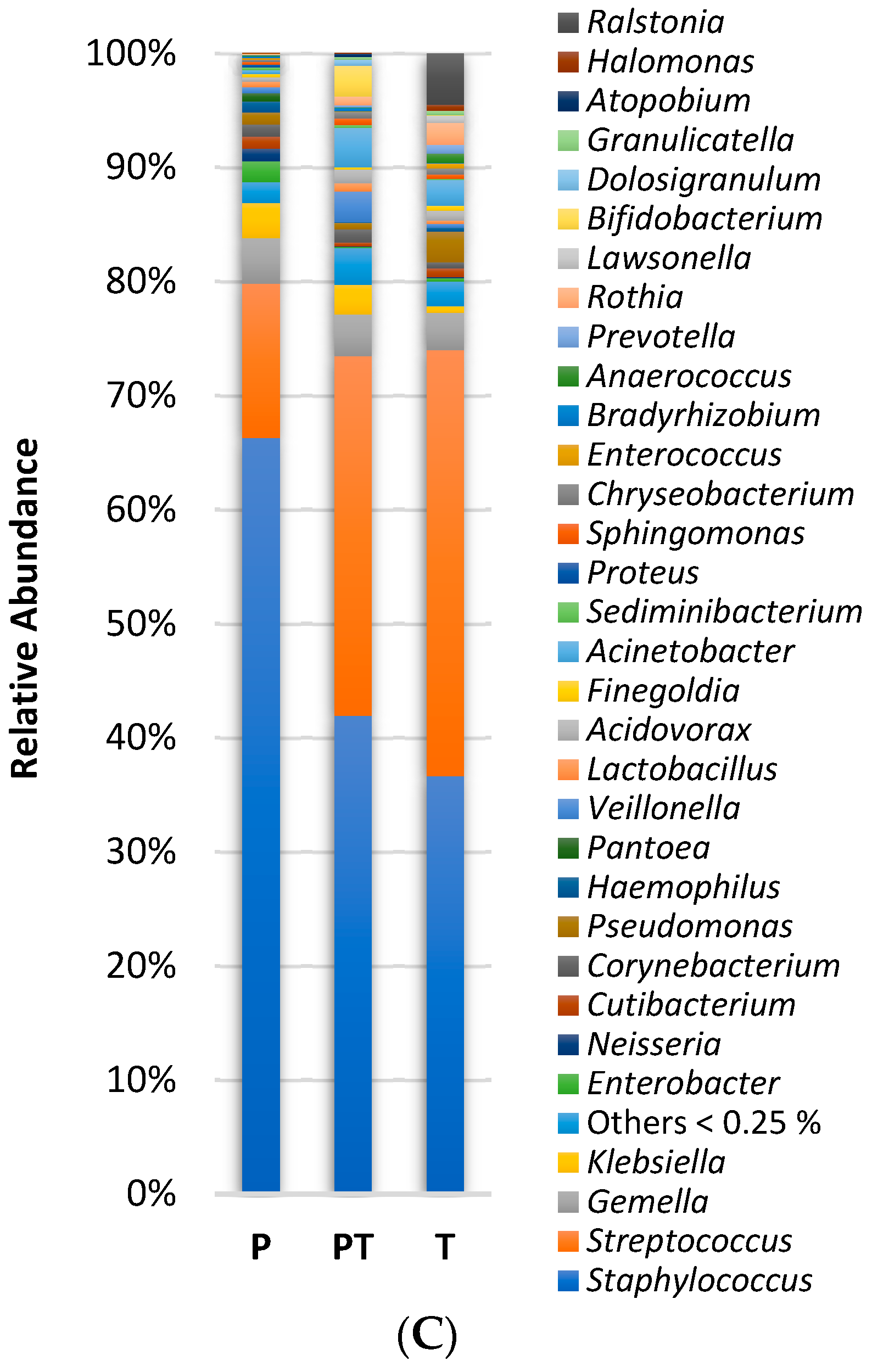
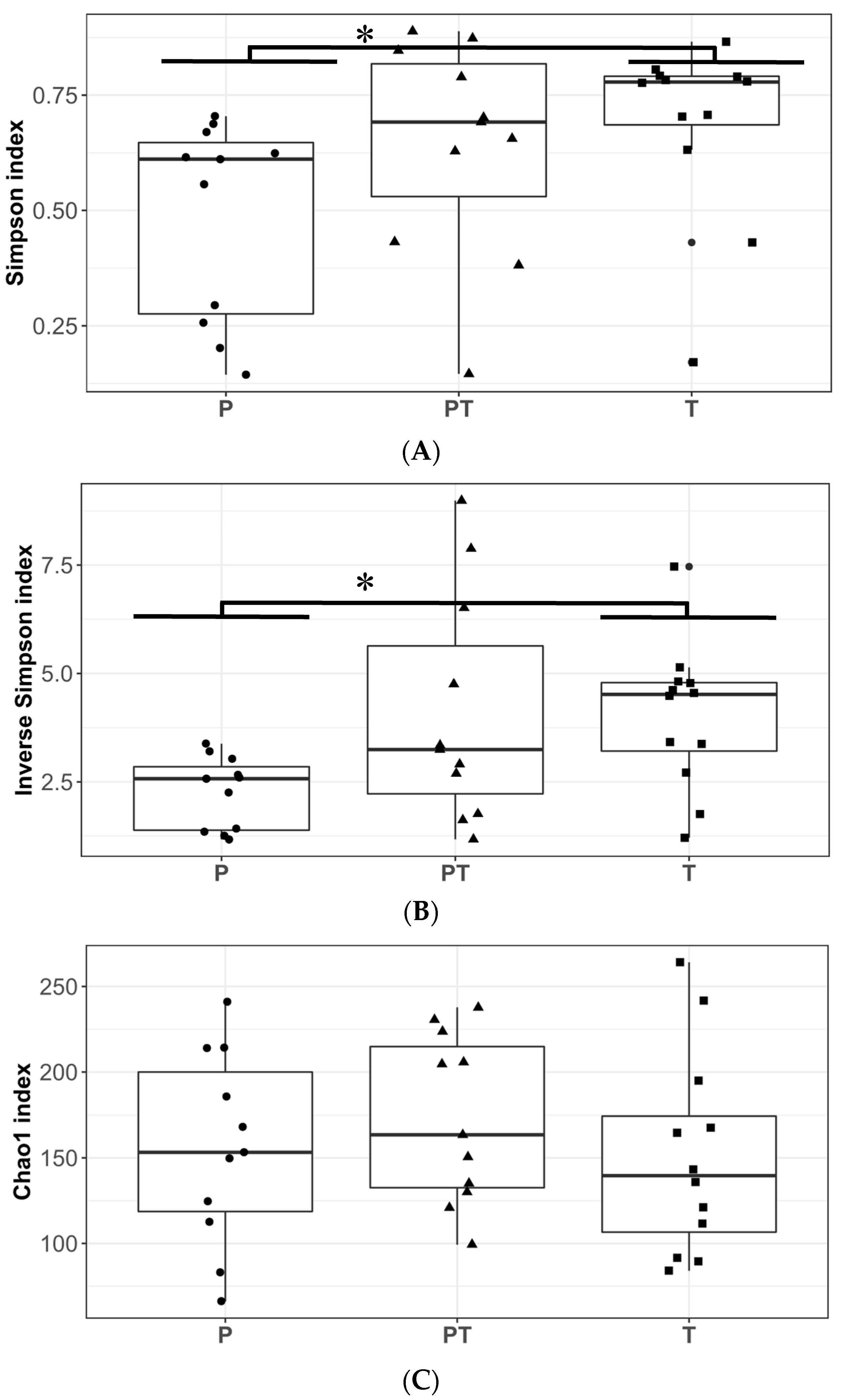
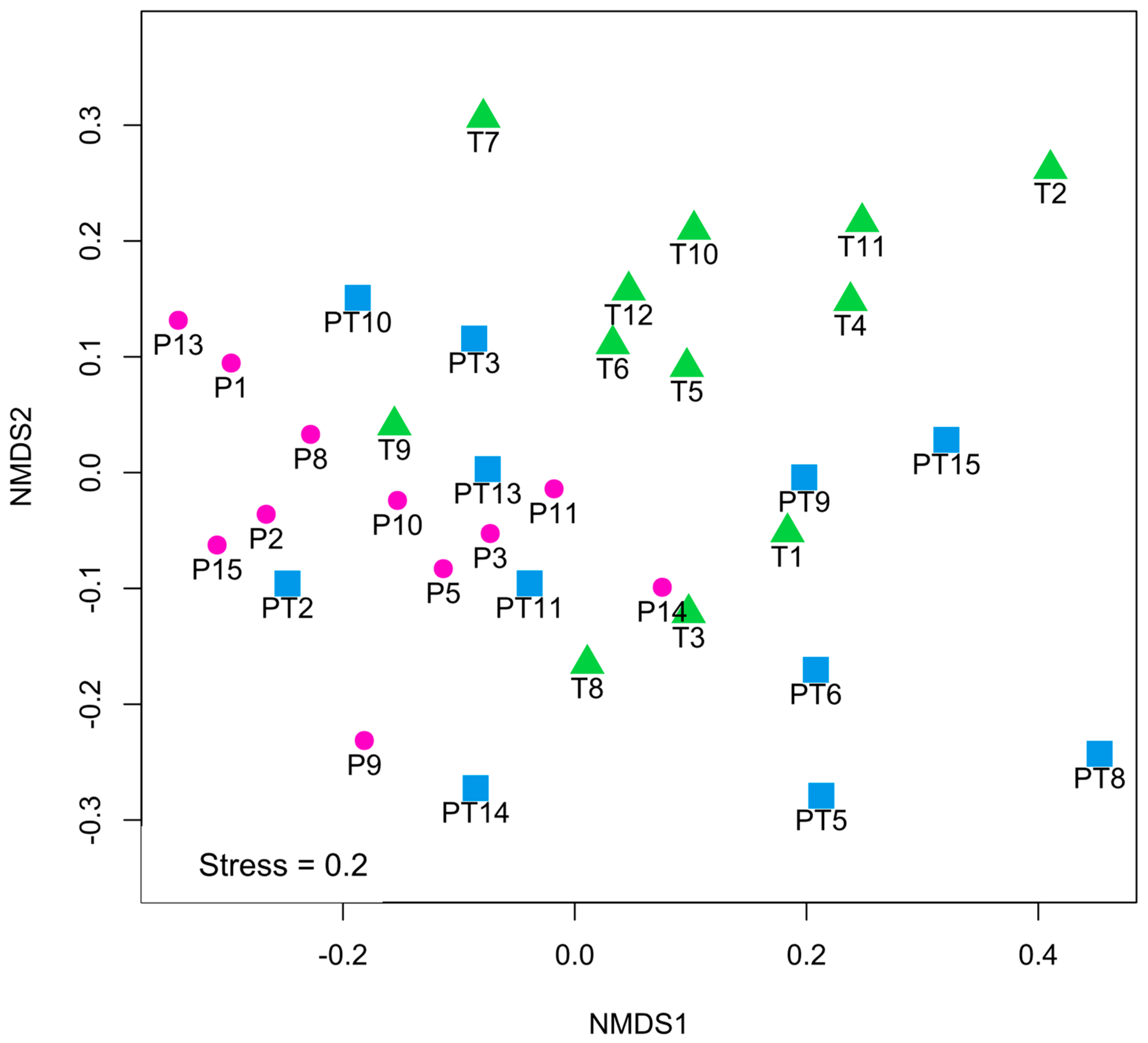
3.1. Clinical Data Profile and Human Milk Microbiota Profile
3.2. Changes over Chronological Age of Pre-Term Human Milk Microbiota (P vs. PT)
3.3. Differences in Human Milk Microbiota Profiles in Mothers with Full-Term and Pre-Term Infants at Similar Corrected Ages (T vs. PT)
3.4. Differences in Human Milk Microbiota Profiles in Mothers with Full-Term and Pre-Term Infants at the Same Chronological Ages (P vs. T)
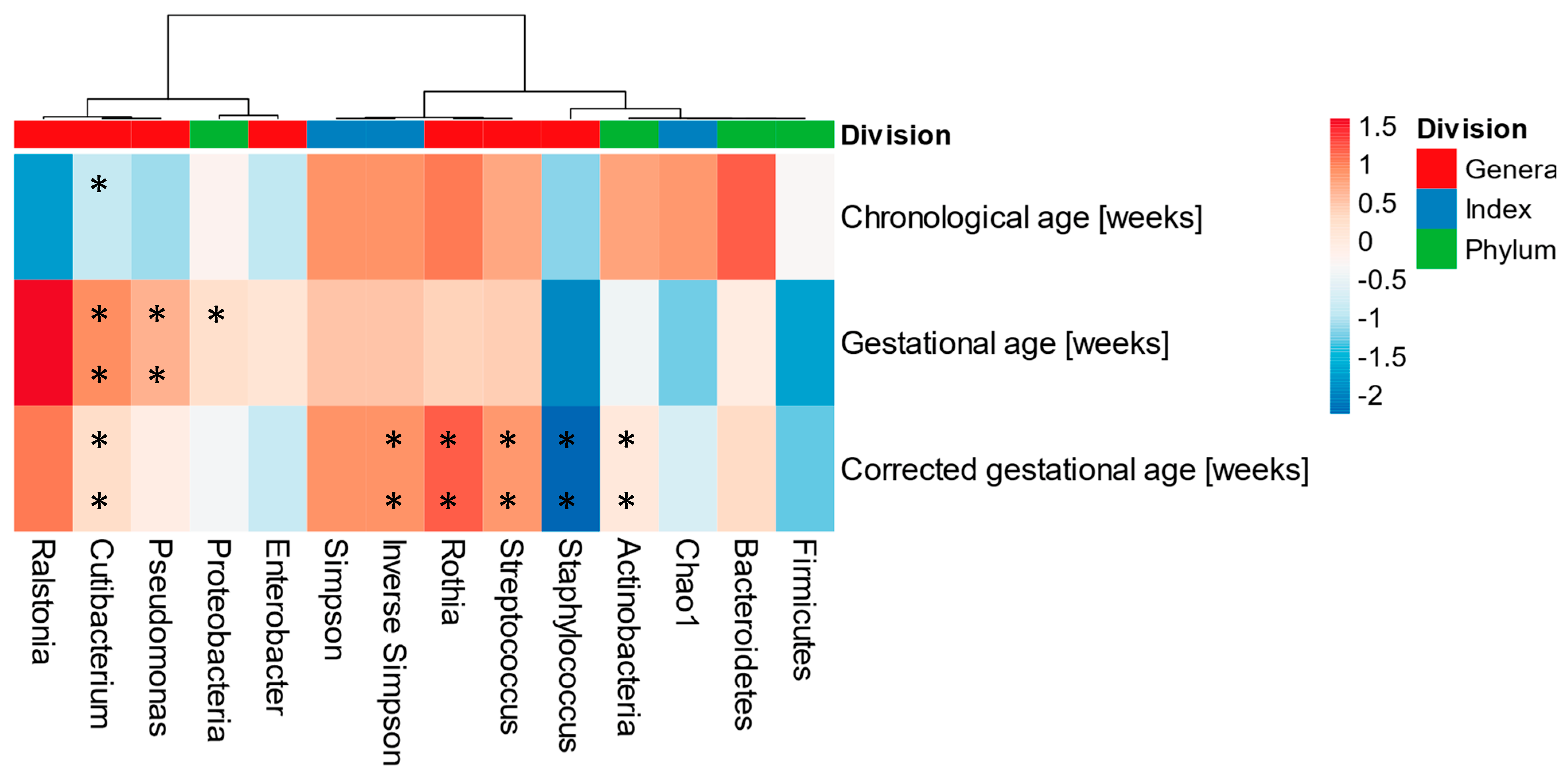
3.5. Human Milk Microbiota and Its Association with Gestational, Chronological, and Corrected Ages
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lyons, K.E.; Ryan, C.A.; Dempsey, E.M.; Ross, R.P.; Stanton, C. Breast Milk, a Source of Beneficial Microbes and Associated Benefits for Infant Health. Nutrients 2020, 12, 1039. [Google Scholar] [CrossRef] [PubMed]
- Le Doare, K.; Holder, B.; Bassett, A.; Pannaraj, P.S. Mother’s Milk: A Purposeful Contribution to the Development of the Infant Microbiota and Immunity. Front. Immunol. 2018, 9, 361. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhang, L.; Tang, J.; Shi, J.; Qu, Y.; Xiong, T.; Mu, D. Human Milk as a Protective Factor for Bronchopulmonary Dysplasia: A Systematic Review and Meta-Analysis. Arch. Dis. Child.-Fetal Neonatal Ed. 2019, 104, F128–F136. [Google Scholar] [CrossRef] [PubMed]
- Moubareck, C.A. Human Milk Microbiota and Oligosaccharides: A Glimpse into Benefits, Diversity, and Correlations. Nutrients 2021, 13, 1123. [Google Scholar] [CrossRef] [PubMed]
- Kumar, H.; du Toit, E.; Kulkarni, A.; Aakko, J.; Linderborg, K.M.; Zhang, Y.; Nicol, M.P.; Isolauri, E.; Yang, B.; Collado, M.C.; et al. Distinct Patterns in Human Milk Microbiota and Fatty Acid Profiles across Specific Geographic Locations. Front. Microbiol. 2016, 7, 1619. [Google Scholar] [CrossRef] [PubMed]
- Ayoub Moubareck, C.; Lootah, M.; Tahlak, M.; Venema, K. Profiles of Human Milk Oligosaccharides and Their Relations to the Milk Microbiota of Breastfeeding Mothers in Dubai. Nutrients 2020, 12, 1727. [Google Scholar] [CrossRef]
- Bauer, J.; Gerss, J. Longitudinal Analysis of Macronutrients and Minerals in Human Milk Produced by Mothers of Preterm Infants. Clin. Nutr. 2011, 30, 215–220. [Google Scholar] [CrossRef]
- Chrustek, A.; Dombrowska-Pali, A.; Olszewska-Słonina, D. Analysis of the Composition and Antioxidant Status of Breast Milk in Women Giving Birth Prematurely and on Time. PLoS ONE 2021, 16, e0255252. [Google Scholar] [CrossRef]
- Gustafsson, A.; Johansson, E.; Henckel, E.; Lange, S.; Bohlin, K. Changes in Antisecretory Factor in Human Milk during the Postpartum and Length of Gestation. J. Hum. Lact. 2021, 38, 131–140. [Google Scholar] [CrossRef]
- Mank, E.; van Toledo, L.; Heijboer, A.C.; van den Akker, C.H.P.; van Goudoever, J.B. Insulin Concentration in Human Milk in the First Ten Days Postpartum. J. Pediatr. Gastroenterol. Nutr. 2021, 73, e115–e119. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Gálvez, A.; Calvo, M.V.; Megino-Tello, J.; Aguayo-Maldonado, J.; Jiménez-Flores, R.; Fontecha, J. Effect of Gestational Age (Preterm or Full Term) on Lipid Composition of the Milk Fat Globule and Its Membrane in Human Colostrum. J. Dairy Sci. 2020, 103, 7742–7751. [Google Scholar] [CrossRef] [PubMed]
- Collado, M.; Santaella, M.; Mira-Pascual, L.; Martínez-Arias, E.; Khodayar-Pardo, P.; Ros, G.; Martínez-Costa, C. Longitudinal Study of Cytokine Expression, Lipid Profile and Neuronal Growth Factors in Human Breast Milk from Term and Preterm Deliveries. Nutrients 2015, 7, 8577–8591. [Google Scholar] [CrossRef] [PubMed]
- Caldeo, V.; Downey, E.; O’Shea, C.-A.; Affolter, M.; Volger, S.; Courtet-Compondu, M.-C.; De Castros, C.A.; O’Mahony, J.A.; Ryan, C.A.; Kelly, A.L. Protein Levels and Protease Activity in Milk from Mothers of Pre-Term Infants: A Prospective Longitudinal Study of Human Milk Macronutrient Composition. Clin. Nutr. 2021, 40, 3567–3577. [Google Scholar] [CrossRef]
- Aylward, G.P. Is It Correct to Correct for Prematurity? Theoretic Analysis of the Bayley-4 Normative Data. J. Dev. Behav. Pediatr. 2020, 41, 128–133. [Google Scholar] [CrossRef] [PubMed]
- Age Terminology during the Perinatal Period. Pediatrics 2004, 114, 1362–1364. [CrossRef]
- Logares, R. Ramalok/Amplicon_processing: Workflow for Analysing MiSeq Amplicons Based on Uparse. Zenodo10 2017, 5281. [Google Scholar] [CrossRef]
- Edgar, R.C. Updating the 97% Identity Threshold for 16S Ribosomal RNA OTUs. Bioinformatics 2018, 34, 2371–2375. [Google Scholar] [CrossRef]
- Simpson, E.H. Measurement of Diversity. Nature 1949, 163, 688. [Google Scholar] [CrossRef]
- Hughes, J.B.; Hellmann, J.J.; Ricketts, T.H.; Bohannan, B.J.M. Counting the Uncountable: Statistical Approaches to Estimating Microbial Diversity. Appl. Environ. Microbiol. 2001, 67, 4399–4406. [Google Scholar] [CrossRef]
- R Team. A Language and Environment for Statistical Computing; R Team: Vienna, Austria, 2017. [Google Scholar]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontol. Electron. 2001, 4, 9. [Google Scholar]
- Gloor, G.B.; Reid, G. Compositional Analysis: A Valid Approach to Analyze Microbiome High-Throughput Sequencing Data. Can. J. Microbiol. 2016, 62, 692–703. [Google Scholar] [CrossRef] [PubMed]
- Metsalu, T.; Vilo, J. ClustVis: A Web Tool for Visualizing Clustering of Multivariate Data Using Principal Component Analysis and Heatmap. Nucleic Acids Res. 2015, 43, W566–W570. [Google Scholar] [CrossRef] [PubMed]
- Jeurink, P.V.; van Bergenhenegouwen, J.; Jiménez, E.; Knippels, L.M.J.; Fernández, L.; Garssen, J.; Knol, J.; Rodríguez, J.M.; Martín, R. Human Milk: A Source of More Life than We Imagine. Benef. Microbes 2013, 4, 17–30. [Google Scholar] [CrossRef] [PubMed]
- Pannaraj, P.S.; Li, F.; Cerini, C.; Bender, J.M.; Yang, S.; Rollie, A.; Adisetiyo, H.; Zabih, S.; Lincez, P.J.; Bittinger, K.; et al. Association between Breast Milk Bacterial Communities and Establishment and Development of the Infant Gut Microbiome. JAMA Pediatr. 2017, 171, 647–654. [Google Scholar] [CrossRef] [PubMed]
- Milani, C.; Duranti, S.; Bottacini, F.; Casey, E.; Turroni, F.; Mahony, J.; Belzer, C.; Delgado Palacio, S.; Arboleya Montes, S.; Mancabelli, L.; et al. The First Microbial Colonizers of the Human Gut: Composition, Activities, and Health Implications of the Infant Gut Microbiota. Microbiol. Mol. Biol. Rev. 2017, 81, e00036-17. [Google Scholar] [CrossRef]
- Moossavi, S.; Sepehri, S.; Robertson, B.; Bode, L.; Goruk, S.; Field, C.J.; Lix, L.M.; de Souza, R.J.; Becker, A.B.; Mandhane, P.J.; et al. Composition and Variation of the Human Milk Microbiota Are Influenced by Maternal and Early-Life Factors. Cell Host Microbe 2019, 25, 324–335.e4. [Google Scholar] [CrossRef]
- Padilha, M.; Danneskiold-Samsøe, N.B.; Brejnrod, A.; Hoffmann, C.; Cabral, V.P.; de Melo Iaucci, J.; Sales, C.H.; Fisberg, R.M.; Cortez, R.V.; Brix, S.; et al. The Human Milk Microbiota Is Modulated by Maternal Diet. Microorganisms 2019, 7, 502. [Google Scholar] [CrossRef]
- Fitzstevens, J.L.; Smith, K.C.; Hagadorn, J.I.; Caimano, M.J.; Matson, A.P.; Brownell, E.A. Systematic Review of the Human Milk Microbiota. Nutr. Clin. Pract. 2017, 32, 354–364. [Google Scholar] [CrossRef]
- Becaria Coquet, J.; Caballero, V.R.; Camisasso, M.C.; González, M.F.; Niclis, C.; Román, M.D.; Muñoz, S.E.; Leone, C.M.; Procino, F.; Osella, A.R.; et al. Diet Quality, Obesity and Breast Cancer Risk: An Epidemiologic Study in Córdoba, Argentina. Nutr. Cancer 2020, 72, 1026–1035. [Google Scholar] [CrossRef]
- Instituto Nacional de Estadística y Censos. 4° Encuesta Nacional de Factores de Riesgo. Resultados Definitivos, 1st ed.; Todesca, J., Ed.; Instituto Nacional de Estadística y Censos (INDEC)—Secretaría de Gobierno de Salud de la Nación: Ciudad Autónoma de Buenos Aires, Argentina, 2019. [Google Scholar]
- Asbury, M.R.; Butcher, J.; Copeland, J.K.; Unger, S.; Bando, N.; Comelli, E.M.; Forte, V.; Kiss, A.; LeMay-Nedjelski, L.; Sherman, P.M.; et al. Mothers of Preterm Infants Have Individualized Breast Milk Microbiota That Changes Temporally Based on Maternal Characteristics. Cell Host Microbe 2020, 28, 669–682. [Google Scholar] [CrossRef]
- Zimmermann, P.; Curtis, N. Breast Milk Microbiota: A Review of the Factors That Influence Composition. J. Infect. 2020, 81, 17–47. [Google Scholar] [CrossRef] [PubMed]
- Urbaniak, C.; Angelini, M.; Gloor, G.B.; Reid, G. Human Milk Microbiota Profiles in Relation to Birthing Method, Gestation and Infant Gender. Microbiome 2016, 4, 1. [Google Scholar] [CrossRef] [PubMed]
- Soeorg, H.; Metsvaht, H.K.; Keränen, E.E.; Eelmäe, I.; Merila, M.; Ilmoja, M.-L.; Metsvaht, T.; Lutsar, I. Genetic Relatedness of Staphylococcus Haemolyticus in Gut and Skin of Preterm Neonates and Breast Milk of Their Mothers. Pediatr. Infect. Dis. J. 2019, 38, 308–313. [Google Scholar] [CrossRef] [PubMed]
- Soeorg, H.; Metsvaht, T.; Eelmäe, I.; Merila, M.; Treumuth, S.; Huik, K.; Jürna-Ellam, M.; Ilmoja, M.-L.; Lutsar, I. The Role of Breast Milk in the Colonization of Neonatal Gut and Skin with Coagulase-Negative Staphylococci. Pediatr. Res. 2017, 82, 759–767. [Google Scholar] [CrossRef] [PubMed]
- Patel, S.H.; Vaidya, Y.H.; Patel, R.J.; Pandit, R.J.; Joshi, C.G.; Kunjadiya, A.P. Culture Independent Assessment of Human Milk Microbial Community in Lactational Mastitis. Sci. Rep. 2017, 7, 7804. [Google Scholar] [CrossRef] [PubMed]
- Angelopoulou, A.; Field, D.; Ryan, C.A.; Stanton, C.; Hill, C.; Ross, R.P. The Microbiology and Treatment of Human Mastitis. Med. Microbiol. Immunol. 2018, 207, 83–94. [Google Scholar] [CrossRef]
- Wei, G.; Darwish, G.; Oppenheim, F.G.; Schuppan, D.; Helmerhorst, E.J. Commensal Bacterium Rothia aeria Degrades and Detoxifies Gluten via a Highly Effective Subtilisin Enzyme. Nutrients 2020, 12, 3724. [Google Scholar] [CrossRef]
- Arrieta, M.-C.; Stiemsma, L.T.; Dimitriu, P.A.; Thorson, L.; Russell, S.; Yurist-Doutsch, S.; Kuzeljevic, B.; Gold, M.J.; Britton, H.M.; Lefebvre, D.L.; et al. Early Infancy Microbial and Metabolic Alterations Affect Risk of Childhood Asthma. Sci. Transl. Med. 2015, 7, 307ra152. [Google Scholar] [CrossRef]
- Williams, J.E.; Carrothers, J.M.; Lackey, K.A.; Beatty, N.F.; York, M.A.; Brooker, S.L.; Shafii, B.; Price, W.J.; Settles, M.L.; McGuire, M.A.; et al. Human Milk Microbial Community Structure Is Relatively Stable and Related to Variations in Macronutrient and Micronutrient Intakes in Healthy Lactating Women. J. Nutr. 2017, 147, 1739–1748. [Google Scholar] [CrossRef]
- Kordy, K.; Gaufin, T.; Mwangi, M.; Li, F.; Cerini, C.; Lee, D.J.; Adisetiyo, H.; Woodward, C.; Pannaraj, P.S.; Tobin, N.H.; et al. Contributions to Human Breast Milk Microbiome and Enteromammary Transfer of Bifidobacterium Breve. PLoS ONE 2020, 15, e0219633. [Google Scholar] [CrossRef]
- Korpela, K.; Blakstad, E.W.; Moltu, S.J.; Strømmen, K.; Nakstad, B.; Rønnestad, A.E.; Brække, K.; Iversen, P.O.; Drevon, C.A.; de Vos, W. Intestinal Microbiota Development and Gestational Age in Preterm Neonates. Sci. Rep. 2018, 8, 2453. [Google Scholar] [CrossRef] [PubMed]
- Biesbroek, G.; Bosch, A.A.T.M.; Wang, X.; Keijser, B.J.F.; Veenhoven, R.H.; Sanders, E.A.M.; Bogaert, D. The Impact of Breastfeeding on Nasopharyngeal Microbial Communities in Infants. Am. J. Respir. Crit. Care Med. 2014, 190, 140612135546007. [Google Scholar] [CrossRef] [PubMed]
- Simpson, M.R.; Avershina, E.; Storrø, O.; Johnsen, R.; Rudi, K.; Øien, T. Breastfeeding-Associated Microbiota in Human Milk Following Supplementation with Lactobacillus rhamnosus GG, Lactobacillus acidophilus La-5, and Bifidobacterium animalis ssp. lactis Bb-12. J. Dairy Sci. 2018, 101, 889–899. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, J.M.; Murphy, K.; Stanton, C.; Ross, R.P.; Kober, O.I.; Juge, N.; Avershina, E.; Rudi, K.; Narbad, A.; Jenmalm, M.C.; et al. The Composition of the Gut Microbiota throughout Life, with an Emphasis on Early Life. Microb. Ecol. Health Dis. 2015, 26, 26050. [Google Scholar] [CrossRef]
- Nikolenko, S.I.; Korobeynikov, A.I.; Alekseyev, M.A. BayesHammer: Bayesian clustering for error correction in single-cell sequencing. BMC Genom. 2013, 14 (Suppl. S1), S7. [Google Scholar] [CrossRef]
- Nurk, S.; Bankevich, A.; Antipov, D.; Gurevich, A.A.; Korobeynikov, A.; Lapidus, A.; Prjibelski, A.D.; Pyshkin, A.; Sirotkin, A.; Sirotkin, Y.; et al. Assembling Single-Cell Genomes and Mini-Metagenomes from Chimeric MDA Products. J. Comput. Biol. 2013, 20, 714–737. [Google Scholar] [CrossRef]
- Edgar, R.C.; Flyvbjerg, H. Error filtering, pair assembly and error correction for next-generation sequencing reads. Bioinformatics 2015, 31, 3476–3482. [Google Scholar] [CrossRef]
- Edgar, R.C. UNOISE2: Improved error-correction for Illumina 16S and ITS amplicon sequencing. bioRxiv 2016. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oddi, S.; Mantziari, A.; Huber, P.; Binetti, A.; Salminen, S.; Collado, M.C.; Vinderola, G. Human Milk Microbiota Profile Affected by Prematurity in Argentinian Lactating Women. Microorganisms 2023, 11, 1090. https://doi.org/10.3390/microorganisms11041090
Oddi S, Mantziari A, Huber P, Binetti A, Salminen S, Collado MC, Vinderola G. Human Milk Microbiota Profile Affected by Prematurity in Argentinian Lactating Women. Microorganisms. 2023; 11(4):1090. https://doi.org/10.3390/microorganisms11041090
Chicago/Turabian StyleOddi, Sofía, Anastasia Mantziari, Paula Huber, Ana Binetti, Seppo Salminen, Maria Carmen Collado, and Gabriel Vinderola. 2023. "Human Milk Microbiota Profile Affected by Prematurity in Argentinian Lactating Women" Microorganisms 11, no. 4: 1090. https://doi.org/10.3390/microorganisms11041090
APA StyleOddi, S., Mantziari, A., Huber, P., Binetti, A., Salminen, S., Collado, M. C., & Vinderola, G. (2023). Human Milk Microbiota Profile Affected by Prematurity in Argentinian Lactating Women. Microorganisms, 11(4), 1090. https://doi.org/10.3390/microorganisms11041090





