Plant-Associated Representatives of the Bacillus cereus Group Are a Rich Source of Antimicrobial Compounds
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strain Isolation and Cultivation
2.2. Reconstruction of the Complete Genomes
2.3. Screening of the Virulence Genes
| Accession | Size | G + C | Genes | Sample | GTDB | Collection | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| bp | % | Coding | RNA | Name | Genomospecies | Plant | Organ | Site | Date | |
| CP085501.1 | 5,268,018 | 35.7 | 5533 | 149 | A24 | B. cereus ssp. A | black pepper | root | Vietnam | 24 May 2018 |
| CP085506.1 | 5,183,312 | 34.9 | 5450 | 154 | HD2.4B | B. cereus ssp. A | tomato plants | rhizosphere | Vietnam, Hoai Duc, Ha Noi | 23 April 2019 |
| CP085510.1 | 5,183,870 | 34.9 | 5450 | 157 | HD1.4B | B. cereus ssp. A | tomato plant | rhizosphere | Vietnam, Hoai Duc, Ha Noi | 23 April 2019 |
| JABSVB000000000.1 | 5,796,358 | 34.8 | 5664 | 80 | HB3.1 | B. cereus ssp. A | orange tree | rhizosphere | Vietnam, Cao Phong, Hoa Binh | 17 April 2019 |
| VDDR00000000.1 | 5,071,716 | 35.4 | 5619 | 63 | A8 | B. cereus ssp. A | coffee tree | root | Vietnam | 9 May 2018 |
| VEPT00000000.1 | 5,319,678 | 35.3 | 5332 | 63 | A31 | B. cereus ssp. A | black pepper | root | Vietnam | 22 May 2018 |
| VEPS00000000.2 | 6,195,299 | 34.7 | 6254 | 78 | TK1 | B. cereus ssp. B | black pepper | rhizosphere | Vietnam | 9 May 2018 |
| CP085498.1 | 5,310,791 | 35.2 | 5640 | 143 | A22 | B. cereus ssp. B | coffee tree | root | Vietnam | 9 May 2018 |
| JABSVF000000000.1 | 5,869,336 | 34.8 | 5640 | 80 | M2.1B | B. cereus ssp. B | maize | rhizosphere | Vietnam, Phu An, Thanh Da | 6 December 2019 |
| VEPQ00000000.1 | 5,604,011 | 35.6 | 5466 | 63 | A42 | B. cereus ssp. B | black pepper | root | Vietnam | 12 May 2018 |
| VEPR00000000.1 | 5,594,617 | 35.6 | 5480 | 67 | SN4-3 | B. cereus ssp. B | maize | dead insect | Vietnam | 28 May 2018 |
| VEPV00000000.1 | 5,335,513 | 35.2 | 5339 | 58 | SN1 | B. tropicus ssp. B | Ostrinia nubilalis | Vietnam | 28 May 2018 | |
| VEPW00000000.1 | 5,958,606 | 35.8 | 5843 | 93 | CD3-2 | B. tropicus ssp. | brown mustard | rhizosphere | Vietnam | 28 May 2018 |
| VEPU00000000.1 | 5,443,801 | 35.2 | 5389 | 72 | SN4.1 | B. pacificus ssp. B | Ostrinia nubilalis | Vietnam | 28 May 2018 | |
| JABSVD000000000.1 | 5,695,940 | 35.1 | 5534 | 87 | HD1.3 | Bacillus sp. | tomato plant | rhizosphere | Vietnam, Hoai Duc, Ha Noi | 23 April 2019 |
| VEPX00000000.1 | 5,695,940 | 35.3 | 5136 | 65 | CD3-5 | Bacillus sp. | brown mustard | rhizosphere | Vietnam | 28 May 2018 |
| VEPY00000000.1 | 5,150,560 | 35.2 | 5175 | 61 | CD3-1a | Bacillus sp. | brown mustard | rhizosphere | Vietnam | 28 May 2018 |
2.4. Genotypic and Phenotypic Characterization of the Isolate B. cereus CD3-1a
2.5. Taxonomical Phylogeny Assessment
2.6. Genome Mining
2.7. Sample Preparation and Mass-Spectrometric Detection of the Bioactive Peptides
2.8. Antifungal, Nematocidal, and Plant-Growth-Promoting Activity Assays
2.9. Data Analysis
2.10. Gene Bank Accession Numbers of the Complete Genome Sequences
3. Results and Discussion
3.1. Comparative Genome Analysis of the Isolates from Vietnamese Crop Plants Representing the Bacillus cereus s.l. Complex
3.1.1. Genome-Based Species and Subspecies Delineation of the Plant-Associated Isolates Belonging to the B. cereus Group
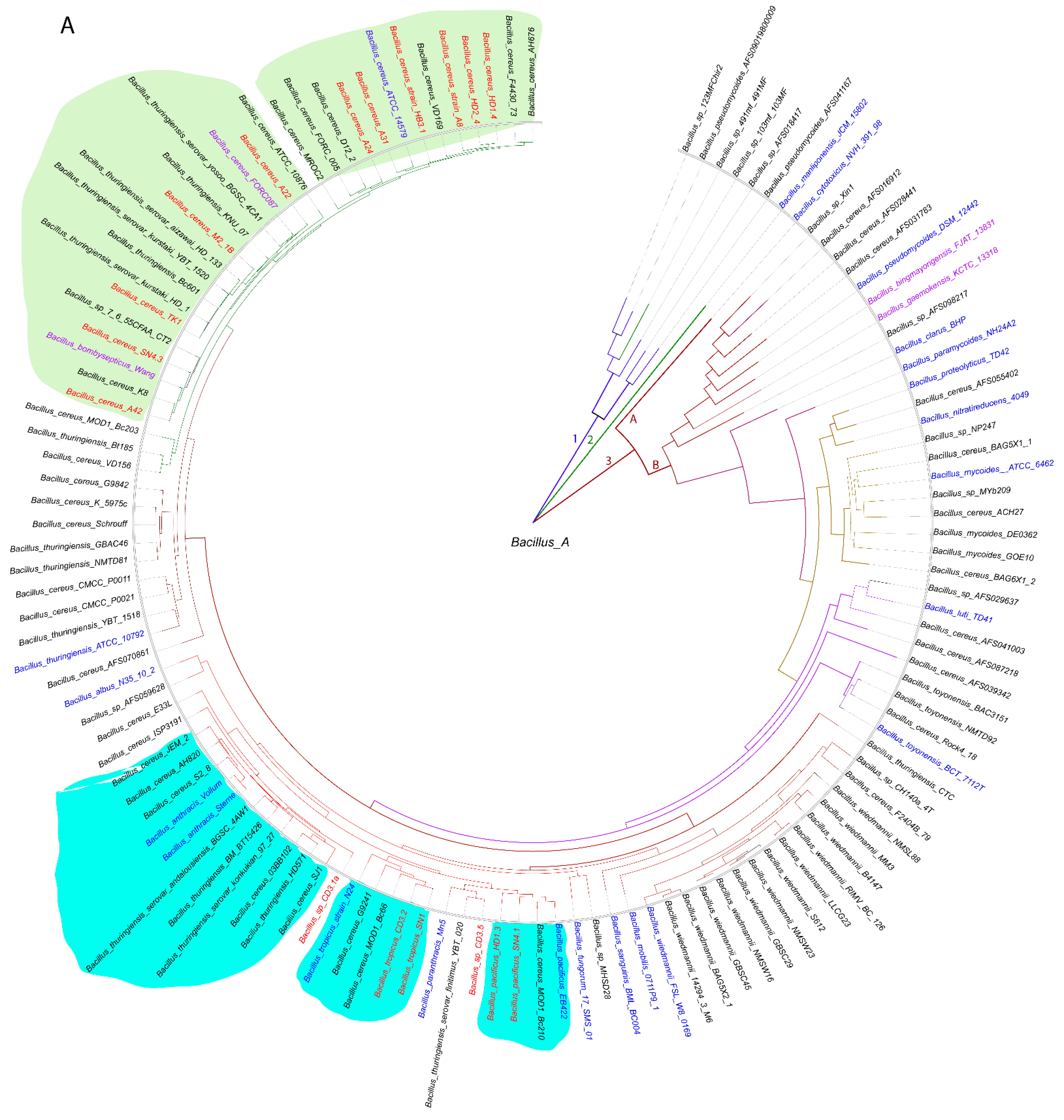
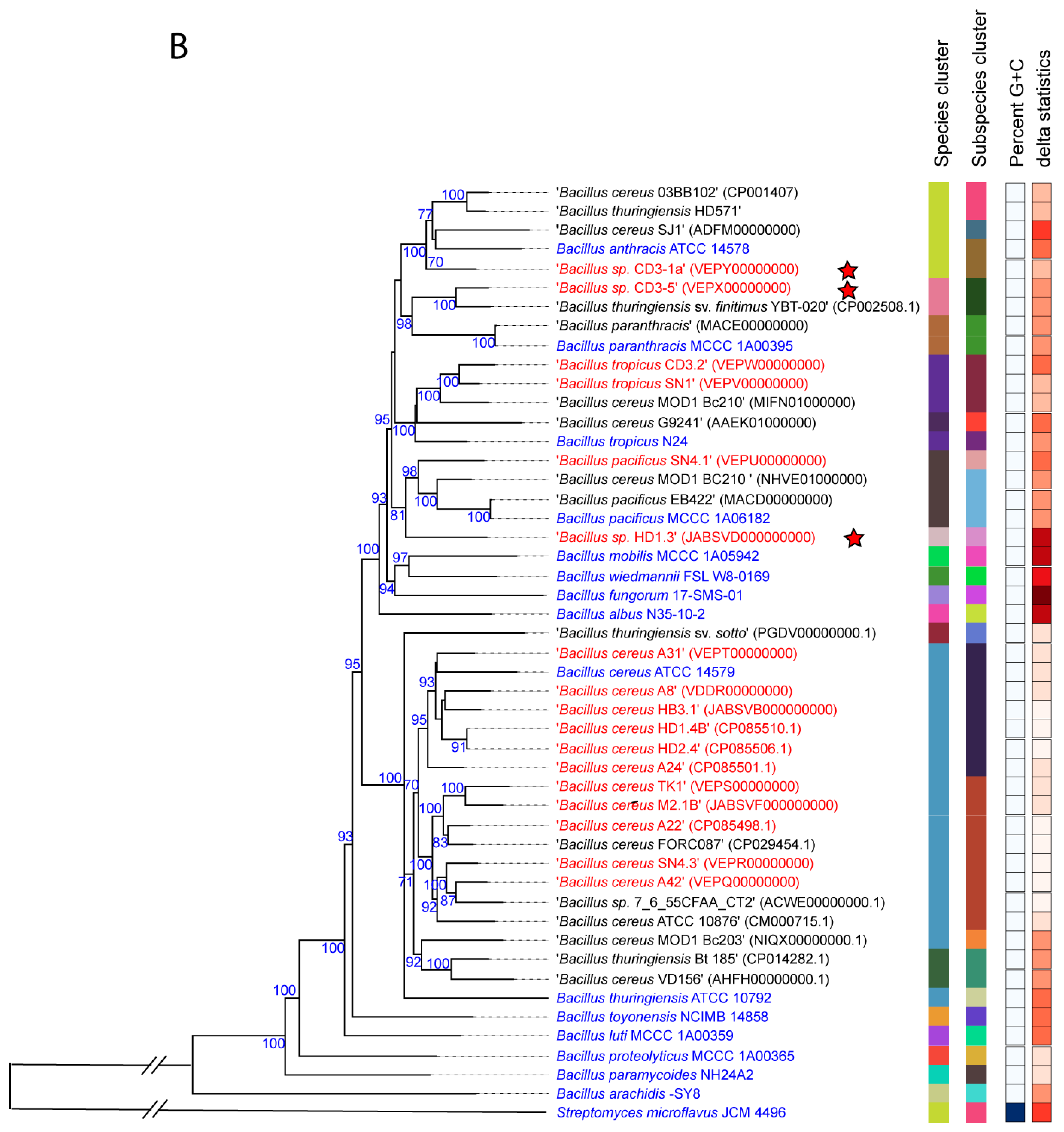
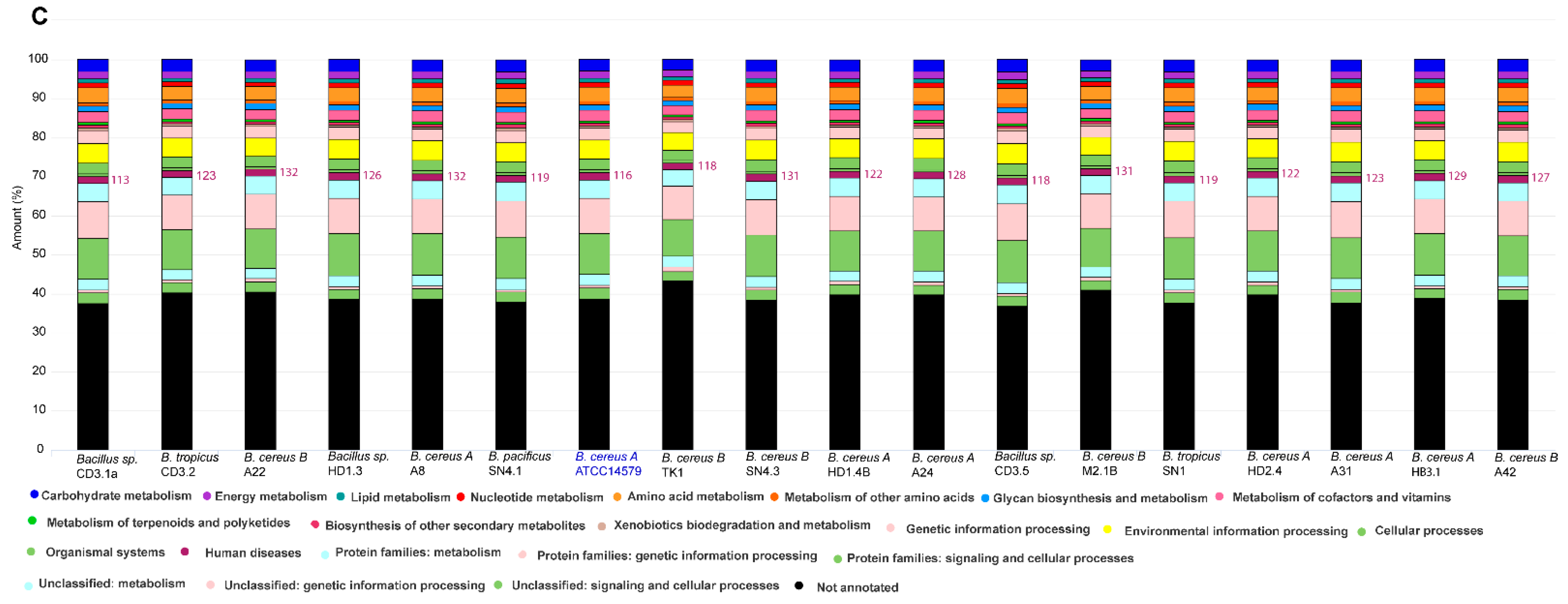
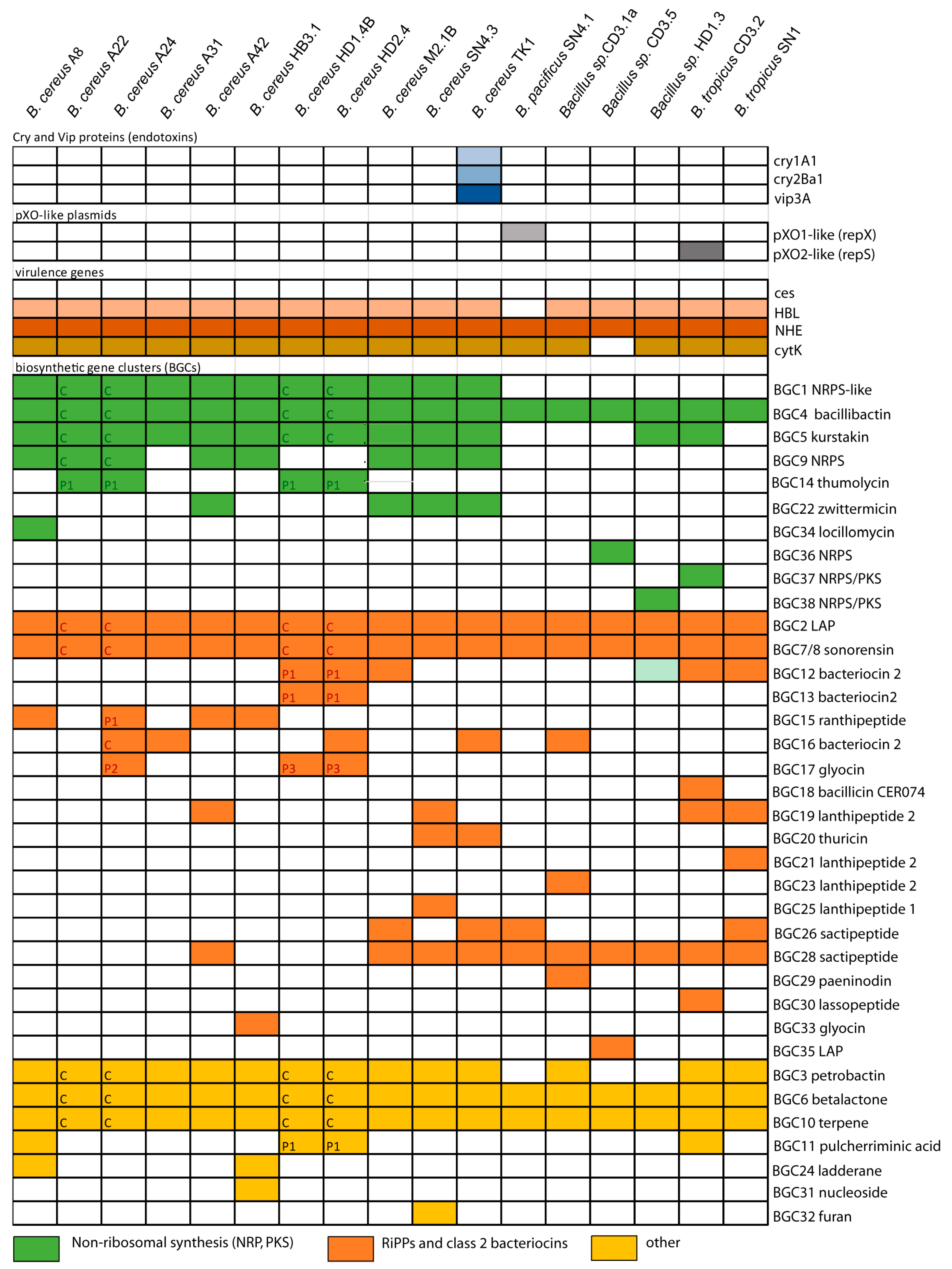
3.1.2. Occurrence of Virulence Genes Might Restrict the Application of B. cereus s.l. Isolates
3.1.3. Genes Encoding Insecticidal Proteins in B. cereus subsp. Bombysepticus TK1
3.1.4. Plasmid-Encoded Virulence Genes and Biosynthetic Gene Clusters in B. cereus Isolates A22, A24, HD1.4B, and HD2.4
3.2. Genome Mining for Biosynthetic Gene Clusters (BGCs) Encoding Secondary Metabolites
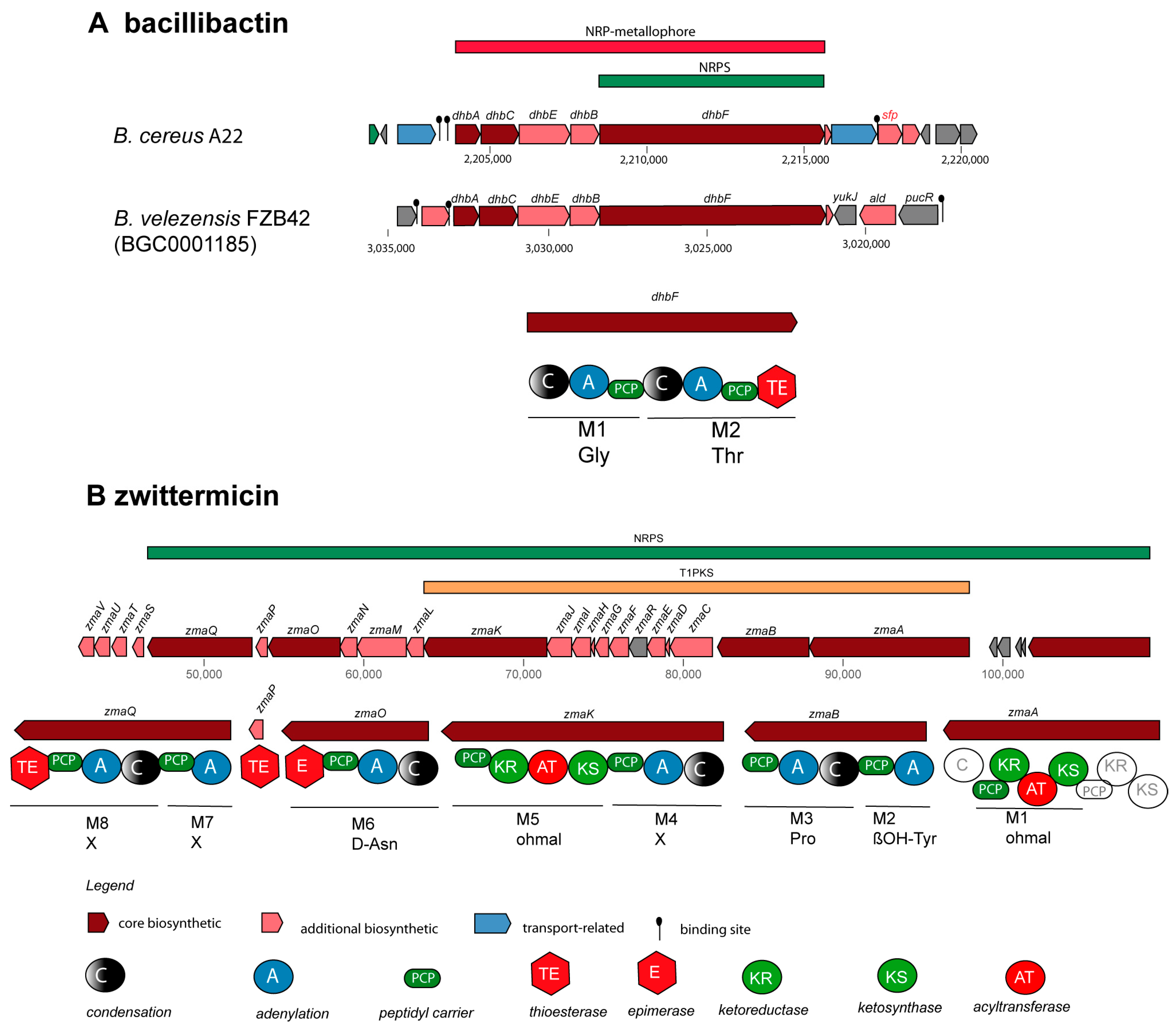
3.2.1. Non-Ribosomally Synthesized Antimicrobial Peptides (NRPs) and Polyketides (PKs)
3.2.2. Gene Clusters Representing RiPPs and Bacteriocins
- Class I: post-translationally modified peptides smaller than 10 kDa.
- Class II: small (<10 Da), unmodified peptides with or without a leader sequence.
- Class III: peptides larger than 10 kDa.
3.2.3. Other Antimicrobial Secondary Metabolites
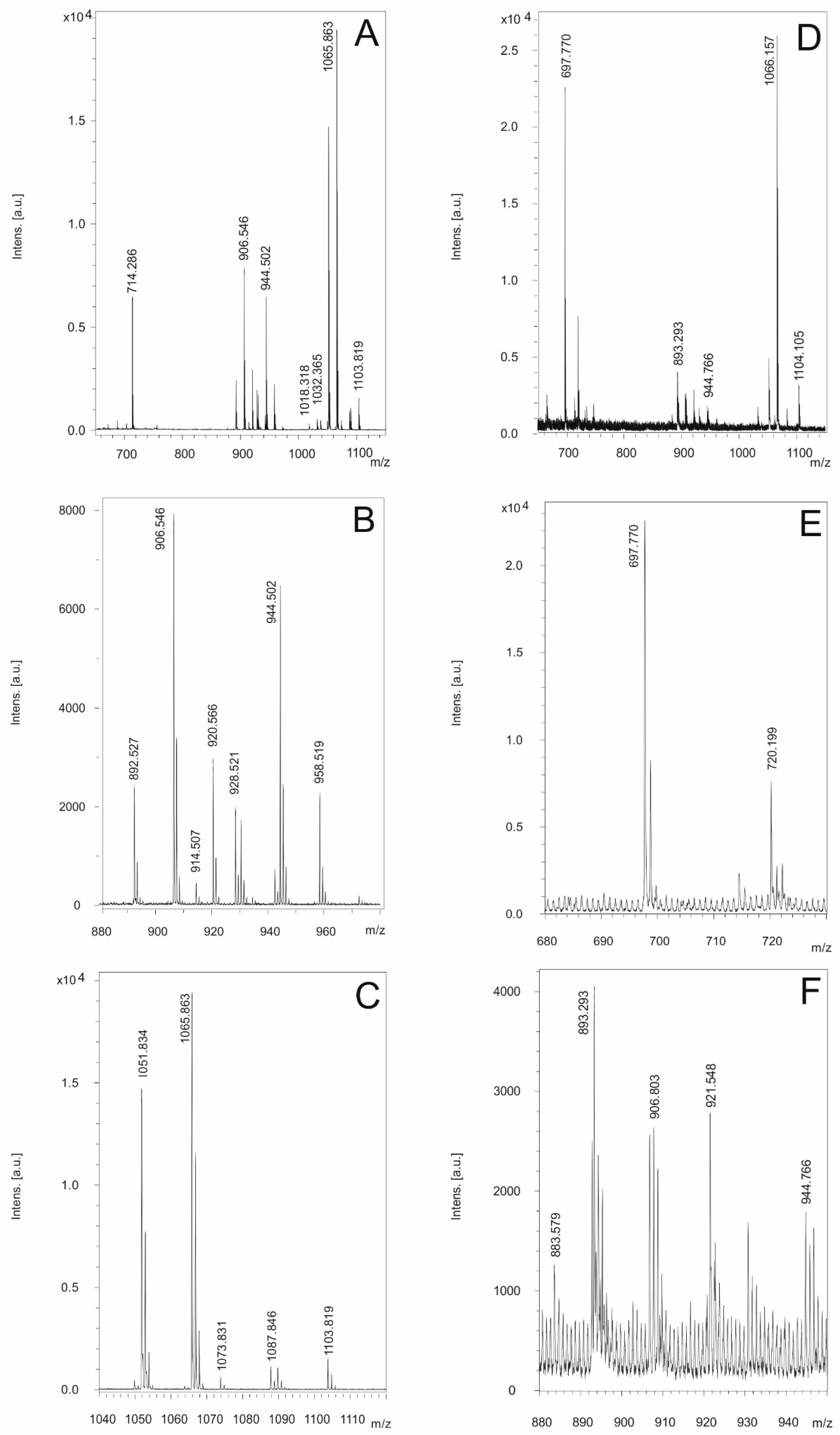
3.3. Detection of Bioactive Peptides by MALDI-TOF Mass Spectrometry
| bn₋H2O (found) | - | 280.123 | 337.149 | 408.107 | 495.159 | 632.284 | 760.445 | - |
| bn (found) | 197.030 | 298.133 | 355.112 | - | - | - | 778.446 | 906.504 |
| bn (calc.) | 197.190 | 298.238 | 355.259 | 426.296 | 513.328 | 650.387 | 778.446 | 906.504 |
| C13-FA | Thr (1) | Gly (2) | Ala (3) | Ser (4) | His (5) | Gln (6) | Gln (7) | |
| yn (calc.) | 906.502 | 710.322 | 609.275 | 552.253 | 481.216 | 394.184 | 257.125 | 129.066 |
| yn (found) | 906.504 | 710.266 | 609.163 | 552.133 | 481.106 | 394.084 | 257.050 | 129.066 |
| yn₋H2O (found) | - | 692.185 | - | - | 463.103 | - | - | - |
| (A) Dipeptide fragments | m/z | m/z |
| Calc. | found | |
| C13-FA-Thr | 298.238 | 298.141/280.129 |
| Thr-Gly | 159.077 | 159.007/141.022 |
| Gly-Ala | 129.066 | 129.016 |
| Ala-Ser | 159.077 | 159.007/141.022 |
| Ser-His | 225.099 | 225.034/207.019 |
| His-Gln | 266.125 | 266.060 |
| Gln-Gln | 257.125 | 257.050 |
| (B) Tripeptide fragments | ||
| C13-FA-Thr-Gly | 341.244 | 341.170/323.167 |
| Thr-Gly-Ala | 230.114 | 230.026/212.025 |
| Gly-Ala-Ser | 216.098 | 216.024/198.022 |
| Ala-Ser-His | 296.136 | -/278.050 |
| Ser-His-Gln | 353.157 | 353.084/335.070 |
| His-Gln-Gln | 394.184 | 394.101/335.070 |
| (C) Tetrapeptide fragments | ||
| C13-FA-Thr-Gly-Ala | 426.296 | -/408.121 |
| Thr-Gly-Ala-Ser | 317.146 | 317.057/299.070 |
| Gly-Ala-Ser-His | 353.157 | 353.084/335.070 |
| Ala-Ser-His-Gln | 424.194 | - |
| Ser-His-Gln-Gln | 481.216 | 481.120/463.120 |
3.4. B. cereus s.l. Strains Suppressed Plant Pathogens and Promoted Plant Growth
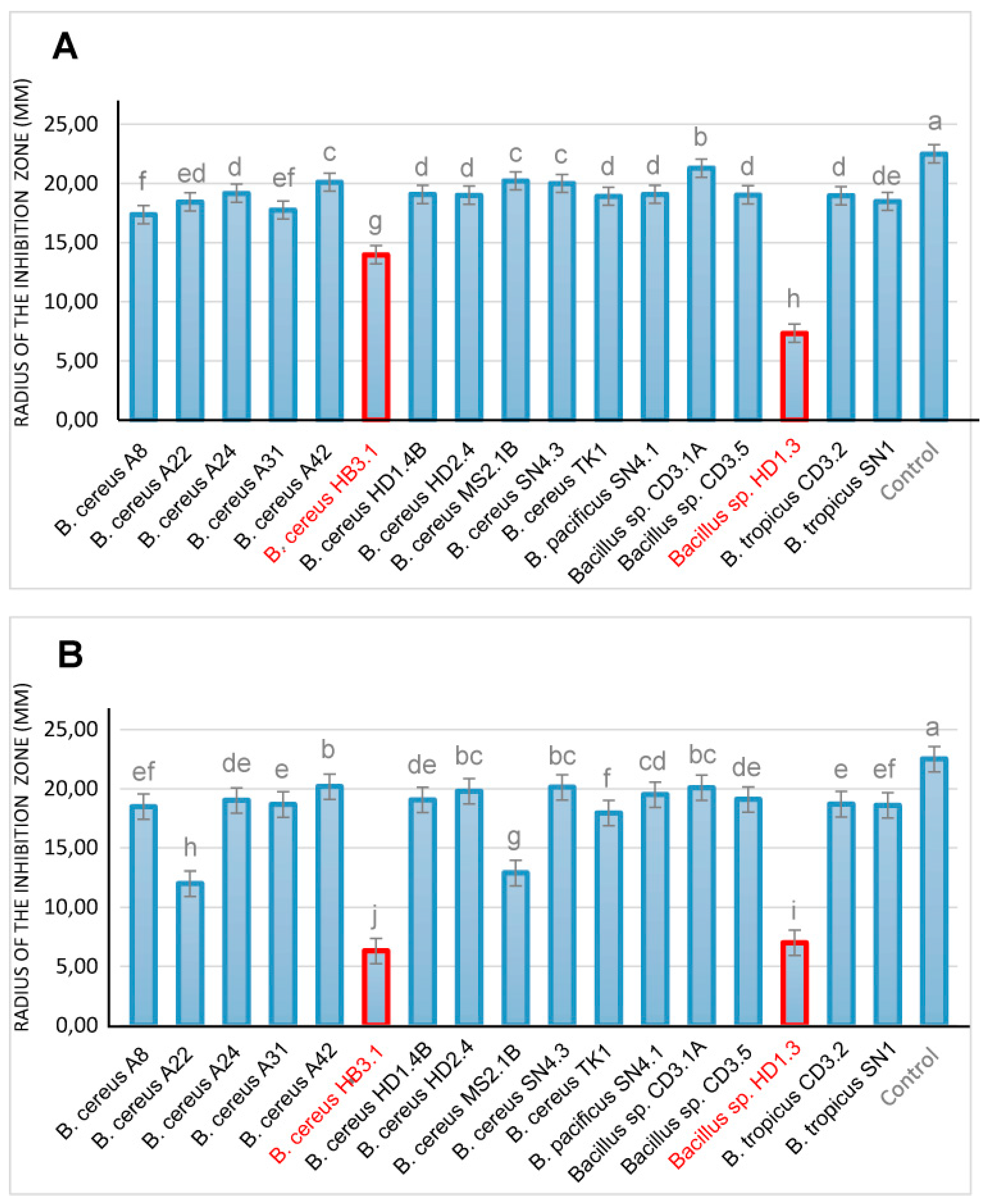
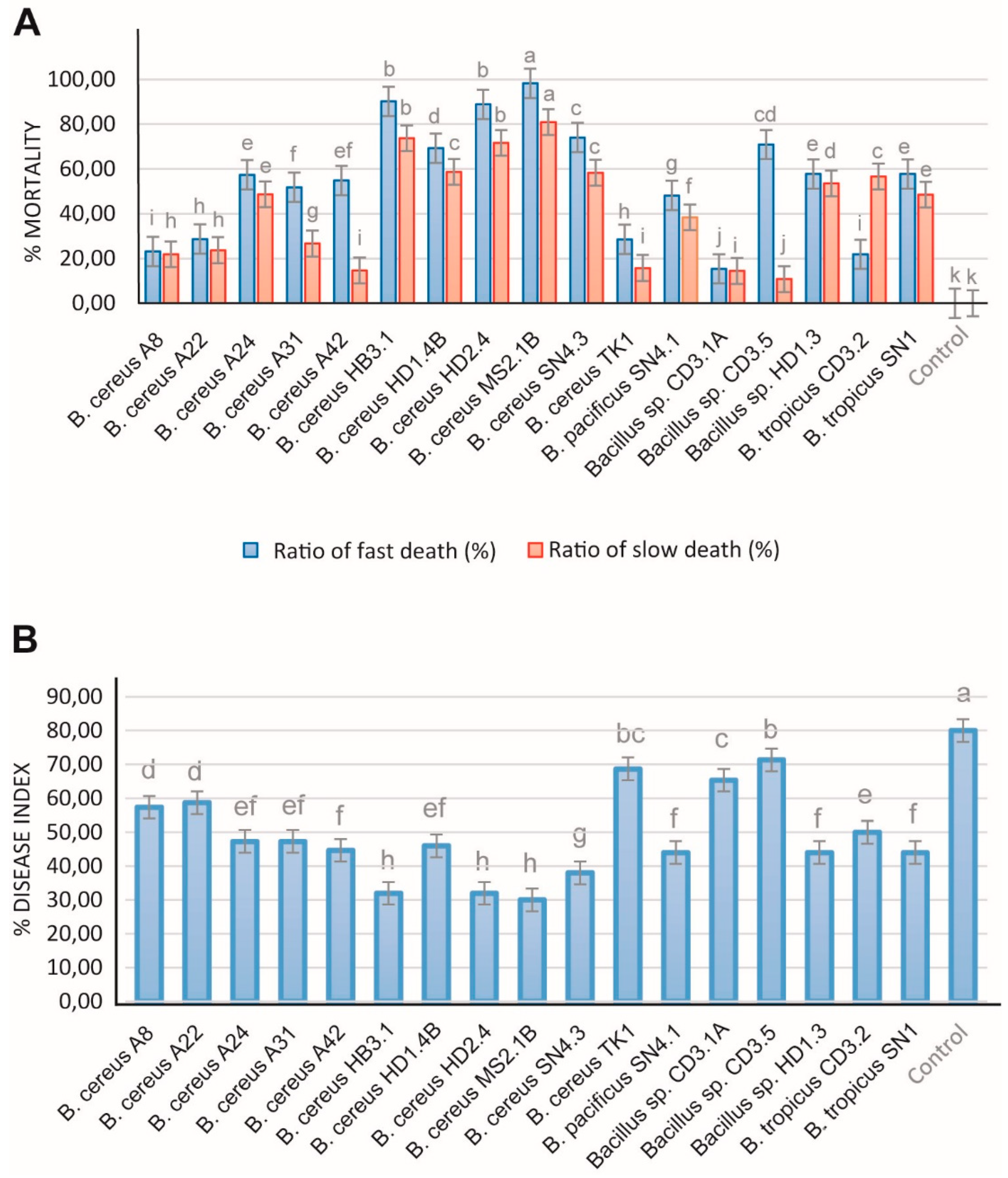
3.4.1. Antifungal and Nematocidal Activity
3.4.2. Plant Growth Promotion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Borriss, R. Use of plant-associated Bacillus strains as biofertilizers and biocontrol agents. In Bacteria in Agrobiology: Plant Growth Responses; Maheshwari, D.K., Ed.; Springer: Berlin/Heidelberg, Germany, 2011; pp. 41–76. [Google Scholar] [CrossRef]
- Tam, L.T.T.; Jähne, J.; Luong, P.T.; Thao, L.T.P.; Chung, L.T.K.; Schneider, A.; Blumenscheit, C.; Lasch, P.; Schweder, T.; Borriss, R. Draft genome sequences of 59 endospore-forming Gram-positive bacteria associated with crop plants grown in Vietnam. Microbiol. Resour. Announc. 2020, 9, e01154-20. [Google Scholar] [CrossRef]
- Jähne, J.; Le Thi, T.T.; Blumenscheit, C.; Schneider, A.; Pham, T.L.; Le Thi, P.T.; Blom, J.; Vater, J.; Schweder, T.; Lasch, P.; et al. Novel Plant-Associated Brevibacillus and Lysinibacillus Genomospecies Harbor a Rich Biosynthetic Potential of Antimicrobial Compounds. Microorganisms 2023, 11, 168. [Google Scholar] [CrossRef]
- Thanh Tam, L.T.; Jähne, J.; Luong, P.T.; Phuong Thao, L.T.; Nhat, L.M.; Blumenscheit, C.; Schneider, A.; Blom, J.; Kim Chung, L.T.; Anh Minh, P.L.; et al. Two plant-associated Bacillus velezensis strains selected after genome analysis, metabolite profiling, and with proved biocontrol potential, were enhancing harvest yield of coffee and black pepper in large field trials. Front. Plant. Sci. 2023, 14, 1194887. [Google Scholar] [CrossRef] [PubMed]
- Carroll, L.M.; Cheng, R.A.; Wiedmann, M.; Kovac, J. Keeping up with the Bacillus cereus group: Taxonomy through the genomics era and beyond. Crit. Rev. Food Sci. Nutr. 2022, 62, 7677–7702. [Google Scholar] [CrossRef]
- Ehling-Schulz, M.; Lereclus, D.; Koehler, T.M. The Bacillus cereus Group: Bacillus Species with Pathogenic Potential. Microbiol. Spectr. 2019, 7. [Google Scholar] [CrossRef]
- Koch, R. Die Ätiologie der Milzbrand-Krankheit, begründet auf die Entwicklungsgeschichte des Bacillus Anthracis. Cohns beiträge ur Biol. Der Pflanz. 1876, 2, 277. [Google Scholar]
- Moayeri, M.; Leppla, S.H.; Vrentas, C.; Pomerantsev, A.P.; Liu, S. Anthrax Pathogenesis. Annu. Rev. Microbiol. 2015, 69, 185–208. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Wang, Y.; Liu, Y.; Jia, K.; Zhang, Z.; Dong, Q. Cereulide and Emetic Bacillus cereus: Characterizations, Impacts and Public Precautions. Foods 2023, 12, 833. [Google Scholar] [CrossRef]
- Berliner, E. Über die Schlafsucht der Mehlmottenraupe (Ephestia kühniella Zell) und ihren Erreger Bacillus thuringiensis n. sp. Z. fAngew. Entomol. 1915, 2, 29–56. [Google Scholar] [CrossRef]
- Palma, L.; Muñoz, D.; Berry, C.; Murillo, J.; Caballero, P. Bacillus thuringiensis toxins: An overview of their biocidal activity. Toxins 2014, 6, 3296–3325. [Google Scholar] [CrossRef]
- Baldwin, V.M. You Can’t B. cereus—A Review of Bacillus cereus Strains That Cause Anthrax-Like Disease. Front. Microbiol. 2020, 11, 1731. [Google Scholar] [CrossRef]
- Kolstø, A.B.; Tourasse, N.J.; Økstad, O.A. What sets Bacillus anthracis apart from other Bacillus species? Annu. Rev. Microbiol. 2009, 63, 451–476. [Google Scholar] [CrossRef] [PubMed]
- Blumenscheit, C.; Jähne, J.; Schneider, A.; Blom, J.; Schweder, T.; Lasch, P.; Borriss, R. Genome sequence data of Bacillus velezensis BP1.2A and BT2.4. Data Brief. 2022, 41, 107978. [Google Scholar] [CrossRef] [PubMed]
- Olson, R.D.; Assaf, R.; Brettin, T.; Conrad, N.; Cucinell, C.; Davis, J.J.; Dempsey, D.M.; Dickerman, A.; Dietrich, E.M.; Kenyon, R.W.; et al. Introducing the Bacterial and Viral Bioinformatics Resource Center (BV-BRC): A resource combining PATRIC, IRD and ViPR. Nucleic Acids Res. 2023, 51, D678–D689. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Lai, Q.; Göker, M.; Meier-Kolthoff, J.P.; Wang, M.; Sun, Y.; Wang, L.; Shao, Z. Genomic insights into the taxonomic status of the Bacillus cereus group. Sci. Rep. 2015, 5, 14082. [Google Scholar] [CrossRef]
- Radnedge, L.; Agron, P.G.; Hill, K.K.; Jackson, P.J.; Ticknor, L.O.; Keim, P.; Andersen, G.L. Genome differences that distinguish Bacillus anthracis from Bacillus cereus and Bacillus thuringiensis. Appl. Environ. Microbiol. 2003, 69, 2755–2764. [Google Scholar] [CrossRef]
- Klee, S.R.; Ozel, M.; Appel, B.; Boesch, C.; Ellerbrok, H.; Jacob, D.; Holland, G.; Leendertz, F.H.; Pauli, G.; Grunow, R.; et al. Characterization of Bacillus anthracis-like bacteria isolated from wild great apes from Cote d’Ivoire and Cameroon. J. Bacteriol. 2006, 188, 5333–5344. [Google Scholar] [CrossRef]
- Meier-Kolthoff, J.P.; Sardà Carbasse, J.; Peinado-Olarte, R.L.; Göker, M. TYGS and LPSN: A database tandem for fast and reliable genome-based classification and nomenclature of prokaryotes. Nucleic Acid Res. 2022, 50, D801–D807. [Google Scholar] [CrossRef]
- Dieckmann, M.A.; Beyvers, S.; Nkouamedjo-Fankep, R.C.; Hanel, P.H.G.; Jelonek, L.; Blom, J.; Goesmann, A. EDGAR3.0, Comparative genomics and phylogenomics on a scalable infrastructure. Nucleic Acids Res. 2021, 49, W185–W192. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Kloosterman, A.M.; Charlop-Powers, Z.; van Wezel, G.P.; Medema, M.H.; Weber, T. antiSMASH 6.0, Improving cluster detection and comparison capabilities. Nucleic Acids Res. 2021, 49, W29–W35. [Google Scholar] [CrossRef]
- Bachmann, B.O.; Ravel, J. Chapter 8. Methods for in silico prediction of microbial polyketide and nonribosomal peptide biosynthetic pathways from DNA sequence data. Methods Enzymol. 2009, 458, 181–217. [Google Scholar] [CrossRef] [PubMed]
- van Heel, A.J.; de Jong, A.; Song, C.; Viel, J.H.; Kok, J.; Kuipers, O.P. BAGEL4, A user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res. 2018, 46, W278–W281. [Google Scholar] [CrossRef] [PubMed]
- Vater, J.; Herfort, S.; Doellinger, J.; Weydmann, M.; Borriss, R.; Lasch, P. Genome Mining of the Lipopeptide Biosynthesis of Paenibacillus polymyxa E681 in Combination with Mass Spectrometry: Discovery of the Lipoheptapeptide Paenilipoheptin. Chembiochem. 2018, 19, 744–753. [Google Scholar] [CrossRef]
- Mülner, P.; Schwarz, E.; Dietel, K.; Junge, H.; Herfort, S.; Weydmann, M.; Lasch, P.; Cernava, T.; Berg, G.; Vater, J. Profiling for Bioactive Peptides and Volatiles of Plant Growth Promoting Strains of the Bacillus subtilis Complex of Industrial Relevance. Front. Microbiol. 2020, 11, 1432. [Google Scholar] [CrossRef] [PubMed]
- Suckau, D.; Resemann, A.; Schuerenberg, M.; Hufnagel, P.; Franzen, J.; Holle, A. A novel MALDI LIFT-TOF/TOF mass spectrometer for proteomics. Anal. Bioanal. Chem. 2003, 376, 952–956. [Google Scholar] [CrossRef] [PubMed]
- Hooper, D.J.; Hallmann, J.; Subbotin, S.A. Methods for extraction, processing and detection of plant and soil nematodes. In Plant Parasitic Nematodes in Subtropical and Tropical Agriculture; Luc, M., Sikora, R., Bridge, J., Eds.; CAB International: Wallingford, UK, 2005; pp. 53–86. [Google Scholar]
- Bridge, J.; Page, S.L.J. Estimation of Root Knot Nematode Infestation Levels in Roots Using a Rating Chart. Trop. Pest Manag. 1980, 26, 296–298. [Google Scholar] [CrossRef]
- Budiharjo, A.; Chowdhury, S.P.; Dietel, K.; Beator, B.; Dolgova, O.; Fan, B.; Bleiss, W.; Ziegler, J.; Schmid, M.; Hartmann, A.; et al. Transposon mutagenesis of the plant-associated Bacillus amyloliquefaciens ssp. plantarum FZB42 revealed that the nfrA and RBAM17410 genes are involved in plant-microbe-interactions. PLoS ONE 2014, 9, e98267. [Google Scholar] [CrossRef]
- Lefort, V.; Desper, R.; Gascuel, O. FastME 2.0, A comprehensive, accurate, and fast distance-based phylogeny inference program. Mol. Biol. Evol. 2015, 32, 2798–2800. [Google Scholar] [CrossRef]
- Meier-Kolthoff, J.P.; Göker, M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat. Commun. 2019, 10, 2182. [Google Scholar] [CrossRef]
- Cheng, T.; Lin, P.; Jin, S.; Wu, Y.; Fu, B.; Long, R.; Liu, D.; Guo, Y.; Peng, L.; Xia, Q. Complete Genome Sequence of Bacillus bombysepticus, a Pathogen Leading to Bombyx mori Black Chest Septicemia. Genome Announc. 2014, 2, e00312-14. [Google Scholar] [CrossRef]
- Parks, D.H.; Chuvochina, M.; Rinke, C.; Mussig, A.J.; Chaumeil, P.A.; Hugenholtz, P. GTDB: An ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 2022, 50, D785–D794. [Google Scholar] [CrossRef] [PubMed]
- Parte, A.C.; Sardà Carbasse, J.; Meier-Kolthoff, J.P.; Reimer, L.C.; Göker, M. List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ. Int. J. Syst. Evol. Microbiol. 2020, 70, 5607–5612. [Google Scholar] [CrossRef]
- Agata, N.; Ohta, M.; Mori, M. Production of an emetic toxin, cereulide, is associated with a specific class of Bacillus cereus. Curr. Microbiol. 1996, 33, 67–69. [Google Scholar] [CrossRef] [PubMed]
- Baillie, L.; Read, T.D. Bacillus anthracis, a bug with attitude! Curr. Opin. Microbiol. 2001, 4, 78–81. [Google Scholar] [CrossRef] [PubMed]
- Bravo, A.; Gómez, I.; Porta, H.; García-Gómez, B.I.; Rodriguez-Almazan, C.; Pardo, L.; Soberón, M. Evolution of Bacillus thuringiensis Cry toxins insecticidal activity. Microb. Biotechnol. 2013, 6, 17–26. [Google Scholar] [CrossRef]
- Beecher, D.J.; Wong, A.C.L. Cooperative, synergistic and antagonistic haemolytic interactions between haemolysin BL, phosphatidylcholine phospholipase C and sphingomyelinase from Bacillus cereus. Microbiology 2000, 146 Pt 12, 3033–3039. [Google Scholar] [CrossRef]
- Dietrich, R.; Jessberger, N.; Ehling-Schulz, M.; Märtlbauer, E.; Granum, P.E. The Food Poisoning Toxins of Bacillus cereus. Toxins 2021, 13, 98. [Google Scholar] [CrossRef]
- Sanahuja, G.; Banakar, R.; Twyman, R.M.; Capell, T.; Christou, P. Bacillus thuringiensis: A century of research, development and commercial applications. Plant Biotechnol. J. 2011, 9, 283–300. [Google Scholar] [CrossRef]
- Gupta, M.; Kumar, H.; Kaur, S. Vegetative Insecticidal Protein (Vip): A Potential Contender From Bacillus thuringiensis for Efficient Management of Various Detrimental Agricultural Pests. Front. Microbiol. 2021, 12, 659736. [Google Scholar] [CrossRef]
- Zheng, D.; Zeng, Z.; Xue, B.; Deng, Y.; Sun, M.; Tang, Y.J.; Ruan, L. Bacillus thuringiensis produces the lipopeptide thumolycin to antagonize microbes and nematodes. Microbiol. Res. 2018, 215, 22–28. [Google Scholar] [CrossRef]
- Overbeek, R.; Olson, R.; Pusch, G.D.; Olsen, G.J.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Parrello, B.; Shukla, M.; et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2014, 42, D206-14. [Google Scholar] [CrossRef] [PubMed]
- Griffith, O.H.; Ryan, M. Bacterial phosphatidylinositol-specific phospholipase C: Structure, function, and interaction with lipids. Biochim. Biophys. Acta 1999, 1441, 237–254. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Cao, D.; Zhu, J.; Feng, H.; Luo, X.; Liu, S.; Yan, X.X.; Zhang, X.; Gao, P. Structural insights into assembly, operation and inhibition of a type I restriction-modification system. Nat. Microbiol. 2020, 5, 1107–1118. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; McPherson, S.A.; Wang, Y.; Li, M.; Wang, P.; Turnbough, C.L., Jr.; Pritchard, D.G. Characterization of the enzymes encoded by the anthrose biosynthetic operon of Bacillus anthracis. J. Bacteriol. 2010, 192, 5053–5062. [Google Scholar] [CrossRef]
- Dong, S.; McPherson, S.A.; Tan, L.; Chesnokova, O.N.; Turnbough, C.L., Jr.; Pritchard, D.G. Anthrose biosynthetic operon of Bacillus anthracis. J. Bacteriol. 2008, 190, 2350–2359. [Google Scholar] [CrossRef]
- Yoshida, K.; Yamaguchi, M.; Morinaga, T.; Kinehara, M.; Ikeuchi, M.; Ashida, H.; Fujita, Y. myo-Inositol catabolism in Bacillus subtilis. J. Biol. Chem. 2008, 283, 10415–10424. [Google Scholar] [CrossRef]
- Zhao, X.; Kuipers, O.P. Identification and classification of known and putative antimicrobial compounds produced by a wide variety of Bacillales species. BMC Genom. 2016, 17, 882. [Google Scholar] [CrossRef] [PubMed]
- Tracanna, V.; de Jong, A.; Medema, M.H.; Kuipers, O.P. Mining prokaryotes for antimicrobial compounds: From diversity to function. FEMS Microbiol. Rev. 2017, 41, 417–429. [Google Scholar] [CrossRef] [PubMed]
- Terlouw, B.R.; Blin, K.; Navarro-Muñoz, J.C.; Avalon, N.E.; Chevrette, M.G.; Egbert, S.; Lee, S.; Meijer, D.; Recchia, M.J.J.; Reitz, Z.L.; et al. MIBiG 3.0, a community-driven effort to annotate experimentally validated biosynthetic gene clusters. Nucleic Acids Res. 2023, 51, D603–D610. [Google Scholar] [CrossRef]
- Finking, R.; Marahiel, M.A. Biosynthesis of nonribosomal peptides. Annu. Rev. Microbiol. 2004, 58, 453–488. [Google Scholar] [CrossRef]
- Béchet, M.; Caradec, T.; Hussein, W.; Abderrahmani, A.; Chollet, M.; Leclère, V.; Dubois, T.; Lereclus, D.; Pupin, M.; Jacques, P. Structure, biosynthesis, and properties of kurstakins, nonribosomal lipopeptides from Bacillus spp. Appl. Microbiol. Biotechnol. 2012, 95, 593–600. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.Y.; Zhang, Y.Y.; Wang, T.; Huang, T.X.; Tang, S.Y.; Jin, Y.; Mi, D.D.; Zheng, Y.; Niu, D.D.; Guo, J.H.; et al. Kurstakin Triggers Multicellular Behaviors in Bacillus cereus AR156 and Enhances Disease Control Efficacy Against Rice Sheath Blight. Plant Dis. 2023, 107, 1463–1470. [Google Scholar] [CrossRef] [PubMed]
- Luo, C.; Liu, X.; Zhou, X.; Guo, J.; Truong, J.; Wang, X.; Zhou, H.; Li, X.; Chen, Z. Unusual Biosynthesis and Structure of Locillomycins from Bacillus subtilis 916. Appl. Environ. Microbiol. 2015, 81, 6601–6609. [Google Scholar] [CrossRef]
- May, J.J.; Wendrich, T.M.; Marahiel, M.A. The dhb operon of Bacillus subtilis encodes the biosynthetic template for the catecholic siderophore 2,3-dihydroxybenzoate-glycine-threonine trimeric ester bacillibactin. J. Biol. Chem. 2001, 276, 7209–7217. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.H.; Koumoutsi, A.; Scholz, R.; Borriss, R. More than anticipated—Production of antibiotics and other secondary metabolites by Bacillus amyloliquefaciens FZB42. J. Mol. Microbiol. Biotechnol. 2009, 16, 14–24. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.H.; Koumoutsi, A.; Scholz, R.; Eisenreich, A.; Schneider, K.; Heinemeyer, I.; Morgenstern, B.; Voss, B.; Hess, W.R.; Reva, O.; et al. Comparative analysis of the complete genome sequence of the plant growth-promoting bacterium Bacillus amyloliquefaciens FZB42. Nat. Biotechnol. 2007, 25, 1007–1014. [Google Scholar] [CrossRef] [PubMed]
- Kevany, B.M.; Rasko, D.A.; Thomas, M.G. Characterization of the complete zwittermicin A biosynthesis gene cluster from Bacillus cereus. Appl. Environ. Microbiol. 2009, 75, 1144–1155. [Google Scholar] [CrossRef]
- Fellbrich, G.; Romanski, A.; Varet, A.; Blume, B.; Brunner, F.; Engelhardt, S.; Felix, G.; Kemmerling, B.; Krzymowska, M.; Nürnberger, T. NPP1, a Phytophthora-associated trigger of plant defense in parsley and Arabidopsis. Plant J. 2002, 32, 375–390. [Google Scholar] [CrossRef]
- Burkhart, B.J.; Hudson, G.A.; Dunbar, K.L.; Mitchell, D.A. A prevalent peptide-binding domain guides ribosomal natural product biosynthesis. Nat. Chem. Biol. 2015, 11, 564–570. [Google Scholar] [CrossRef]
- Kloosterman, A.M.; Shelton, K.E.; van Wezel, G.P.; Medema, M.H.; Mitchell, D.A. RRE-Finder: A Genome-Mining Tool for Class-Independent RiPP Discovery. mSystems 2020, 5, e00267. [Google Scholar] [CrossRef]
- Sahl, H.G.; Jack, R.W.; Bierbaum, G. Biosynthesis and biological activities of lantibiotics with unique post-translational modifications. Eur. J. Biochem. 1995, 230, 827–853. [Google Scholar] [CrossRef]
- Wang, J.; Ma, H.; Ge, X.; Zhang, J.; Teng, K.; Sun, Z.; Zhong, J. Bovicin HJ50-like lantibiotics, a novel subgroup of lantibiotics featured by an indispensable disulfide bridge. PLoS ONE 2014, 9, e97121. [Google Scholar] [CrossRef]
- Walker, M.C.; Eslami, S.M.; Hetrick, K.J.; Ackenhusen, S.E.; Mitchell, D.A.; van der Donk, W.A. Precursor peptide-targeted mining of more than one hundred thousand genomes expands the lanthipeptide natural product family. BMC Genom. 2020, 21, 387. [Google Scholar] [CrossRef]
- Arnison, P.G.; Bibb, M.J.; Bierbaum, G.; Bowers, A.A.; Bugni, T.S.; Bulaj, G.; Camarero, J.A.; Campopiano, D.J.; Challis, G.L.; Clardy, J.; et al. Ribosomally synthesized and post-translationally modified peptide natural products: Overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 2013, 30, 108–160. [Google Scholar] [CrossRef]
- Ren, H.; Biswas, S.; Ho, S.; van der Donk, W.A.; Zhao, H. Rapid Discovery of Glycocins through Pathway Refactoring in Escherichia coli. ACS Chem. Biol. 2018, 13, 2966–2972. [Google Scholar] [CrossRef] [PubMed]
- Oman, T.J.; Boettcher, J.M.; Wang, H.; Okalibe, X.N.; van der Donk, W.A. Sublancin is not a lantibiotic but an S-linked glycopeptide. Nat. Chem. Biol. 2011, 7, 78–80. [Google Scholar] [CrossRef] [PubMed]
- Hegemann, J.D.; Zimmermann, M.; Xie, X.; Marahiel, M.A. Lasso peptides: An intriguing class of bacterial natural products. Acc. Chem. Res. 2015, 48, 1909–1919. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Hegemann, J.D.; Fage, C.D.; Zimmermann, M.; Xie, X.; Linne, U.; Marahiel, M.A. Insights into the Unique Phosphorylation of the Lasso Peptide Paeninodin. J. Biol. Chem. 2016, 291, 13662–13678. [Google Scholar] [CrossRef] [PubMed]
- Flühe, L.; Knappe, T.A.; Gattner, M.J.; Schäfer, A.; Burghaus, O.; Linne, U.; Marahiel, M.A. The radical SAM enzyme AlbA catalyzes thioether bond formation in subtilosin A. Nat. Chem. Biol. 2012, 8, 350–357. [Google Scholar] [CrossRef]
- Hudson, G.A.; Burkhart, B.J.; DiCaprio, A.J.; Schwalen, C.J.; Kille, B.; Pogorelov, T.V.; Mitchell, D.A. Bioinformatic Mapping of Radical S-Adenosylmethionine-Dependent Ribosomally Synthesized and Post-Translationally Modified Peptides Identifies New Cα, Cβ, and Cγ-Linked Thioether-Containing Peptides. J. Am. Chem. Soc. 2019, 141, 8228–8238. [Google Scholar] [CrossRef]
- Chopra, L.; Singh, G.; Choudhary, V.; Sahoo, D.K. Sonorensin: An antimicrobial peptide, belonging to the heterocycloanthracin subfamily of bacteriocins, from a new marine isolate, Bacillus sonorensis MT93. Appl. Environ. Microbiol. 2014, 80, 2981–2990. [Google Scholar] [CrossRef] [PubMed]
- Franz, C.M.; van Belkum, M.J.; Holzapfel, W.H.; Abriouel, H.; Gálvez, A. Diversity of enterococcal bacteriocins and their grouping in a new classification scheme. FEMS Microbiol. Rev. 2007, 31, 293–310. [Google Scholar] [CrossRef]
- Aunpad, R.; Panbangred, W. Evidence for two putative holin-like peptides encoding genes of Bacillus pumilus strain WAPB4. Curr. Microbiol. 2012, 64, 343–348. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Janes, B.K.; Passalacqua, K.D.; Pfleger, B.F.; Bergman, N.H.; Liu, H.; Håkansson, K.; Somu, R.V.; Aldrich, C.C.; Cendrowski, S.; et al. Biosynthetic analysis of the petrobactin siderophore pathway from Bacillus anthracis. J. Bacteriol. 2007, 189, 1698–1710. [Google Scholar] [CrossRef] [PubMed]
- Koppisch, A.T.; Dhungana, S.; Hill, K.K.; Boukhalfa, H.; Heine, H.S.; Colip, L.A.; Romero, R.B.; Shou, Y.; Ticknor, L.O.; Marrone, B.L.; et al. Petrobactin is produced by both pathogenic and non-pathogenic isolates of the Bacillus cereus group of bacteria. Biometals. 2008, 21, 581–589. [Google Scholar] [CrossRef]
- Yuan, S.; Yong, X.; Zhao, T.; Li, Y.; Liu, J. Research Progress of the Biosynthesis of Natural Bio-Antibacterial Agent Pulcherriminic Acid in Bacillus. Molecules 2020, 25, 5611. [Google Scholar] [CrossRef]
- Corre, C.; Song, L.; O’Rourke, S.; Chater, K.F.; Challis, G.L. 2-Alkyl-4-hydroxymethylfuran-3-carboxylic acids, antibiotic production inducers discovered by Streptomyces coelicolor genome mining. Proc. Natl. Acad. Sci. USA 2008, 105, 17510–17515. [Google Scholar] [CrossRef]
- Arie, T. Fusarium diseases of cultivated plants, control, diagnosis, and molecular and genetic studies. J. Pestic. Sci. 2019, 44, 275–281. [Google Scholar] [CrossRef]
- Chi, N.M.; Thu, P.Q.; Nam, H.B.; Quang, D.Q.; Phong, L.V.; Van, N.D.; Trang, T.T.; Kien, T.T.; Tam, T.T.T.; Dell, B. Management of Phytophthora palmivora disease in Citrus reticulata with chemical fungicides. J. Gen. Plant Pathol. 2020, 86, 494–502. [Google Scholar] [CrossRef]
- Eisenback, J.D.; and Triantaphyllou, H.H. Root-knot nematodes: Meloidogyne species and races. In Manual of Agricultural Nematology; Nickle, W.R., Ed.; Marcell Dekker: New York, NY, USA, 1991; pp. 191–274. [Google Scholar]
- Hussey, P.S.; Barker, K.R. A comparison of methods of collecting inocula of Meloidogyne spp., including a new technique. Plant Dis. Report. 1973, 57, 1025–1028. [Google Scholar]
- Chowdhury, S.P.; Hartmann, A.; Gao, X.; Borriss, R. Biocontrol mechanism by root-associated Bacillus amyloliquefaciens FZB42—A review. Front. Microbiol. 2015, 6, 780. [Google Scholar] [CrossRef] [PubMed]
- Qiao, J.-Q.; Wu, H.-J.; Huo, R.; Gao, X.-W.; Borriss, R. Stimulation and biocontrol by Bacillus amyloliquefaciens subsp. plantarum FZB42 engineered for improved action. Chem. Biol. Technol. Agric. 2014, 1, 12. [Google Scholar] [CrossRef]
- Wu, L.; Wu, H.J.; Qiao, J.; Gao, X.; Borriss, R. Novel Routes for Improving Biocontrol Activity of Bacillus Based Bioinoculants. Front. Microbiol. 2015, 6, 1395. [Google Scholar] [CrossRef] [PubMed]




| b ions → | |||||
| bn (found) | - | 216.17 | 328.25 | 457.38 | (p)?? |
| bn (calc.) | 115.09 | 216.14 | 329.22 | 457.28 | (P)570.36 |
| Orn (1) | Thr (2) | Ile (3) | Gln (4) | Leu (5) | |
| yn (calc.) | (p)570.36 | 456.28 | 355.24 | 242.15 | 114.09 |
| yn (found) | (p)?? | 456.39 | 355.25 | 242.17 | - |
| <-- y ions |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vater, J.; Tam, L.T.T.; Jähne, J.; Herfort, S.; Blumenscheit, C.; Schneider, A.; Luong, P.T.; Thao, L.T.P.; Blom, J.; Klee, S.R.; et al. Plant-Associated Representatives of the Bacillus cereus Group Are a Rich Source of Antimicrobial Compounds. Microorganisms 2023, 11, 2677. https://doi.org/10.3390/microorganisms11112677
Vater J, Tam LTT, Jähne J, Herfort S, Blumenscheit C, Schneider A, Luong PT, Thao LTP, Blom J, Klee SR, et al. Plant-Associated Representatives of the Bacillus cereus Group Are a Rich Source of Antimicrobial Compounds. Microorganisms. 2023; 11(11):2677. https://doi.org/10.3390/microorganisms11112677
Chicago/Turabian StyleVater, Joachim, Le Thi Thanh Tam, Jennifer Jähne, Stefanie Herfort, Christian Blumenscheit, Andy Schneider, Pham Thi Luong, Le Thi Phuong Thao, Jochen Blom, Silke R. Klee, and et al. 2023. "Plant-Associated Representatives of the Bacillus cereus Group Are a Rich Source of Antimicrobial Compounds" Microorganisms 11, no. 11: 2677. https://doi.org/10.3390/microorganisms11112677
APA StyleVater, J., Tam, L. T. T., Jähne, J., Herfort, S., Blumenscheit, C., Schneider, A., Luong, P. T., Thao, L. T. P., Blom, J., Klee, S. R., Schweder, T., Lasch, P., & Borriss, R. (2023). Plant-Associated Representatives of the Bacillus cereus Group Are a Rich Source of Antimicrobial Compounds. Microorganisms, 11(11), 2677. https://doi.org/10.3390/microorganisms11112677






