Monophasic Variant of Salmonella Typhimurium 4,[5],12:i:- (ACSSuGmTmpSxt Type) Outbreak in Central Italy Linked to the Consumption of a Roasted Pork Product (Porchetta)
Abstract
:1. Introduction
2. Materials and Methods
2.1. The Collection of Human, Food, and Environmental Strains and an Epidemiological Investigation
2.2. Microbiological Analysis and the Serotyping of Salmonella Strains
2.3. Antimicrobial Susceptibility Testing
2.4. Multiple Locus Variable Number Tandem Repeats Analysis (MLVA)
2.5. Whole Genome Sequencing (WGS)
2.5.1. In Silico Multilocus Sequence Typing (MLST)
2.5.2. Core Genome MLST
3. Results
3.1. Clinical Strains
3.1.1. Serotyping and PCR Analysis
3.1.2. Antimicrobial Susceptibility Testing
3.1.3. Multiple-Locus Variable-Number Tandem Repeats Analysis (MLVA)
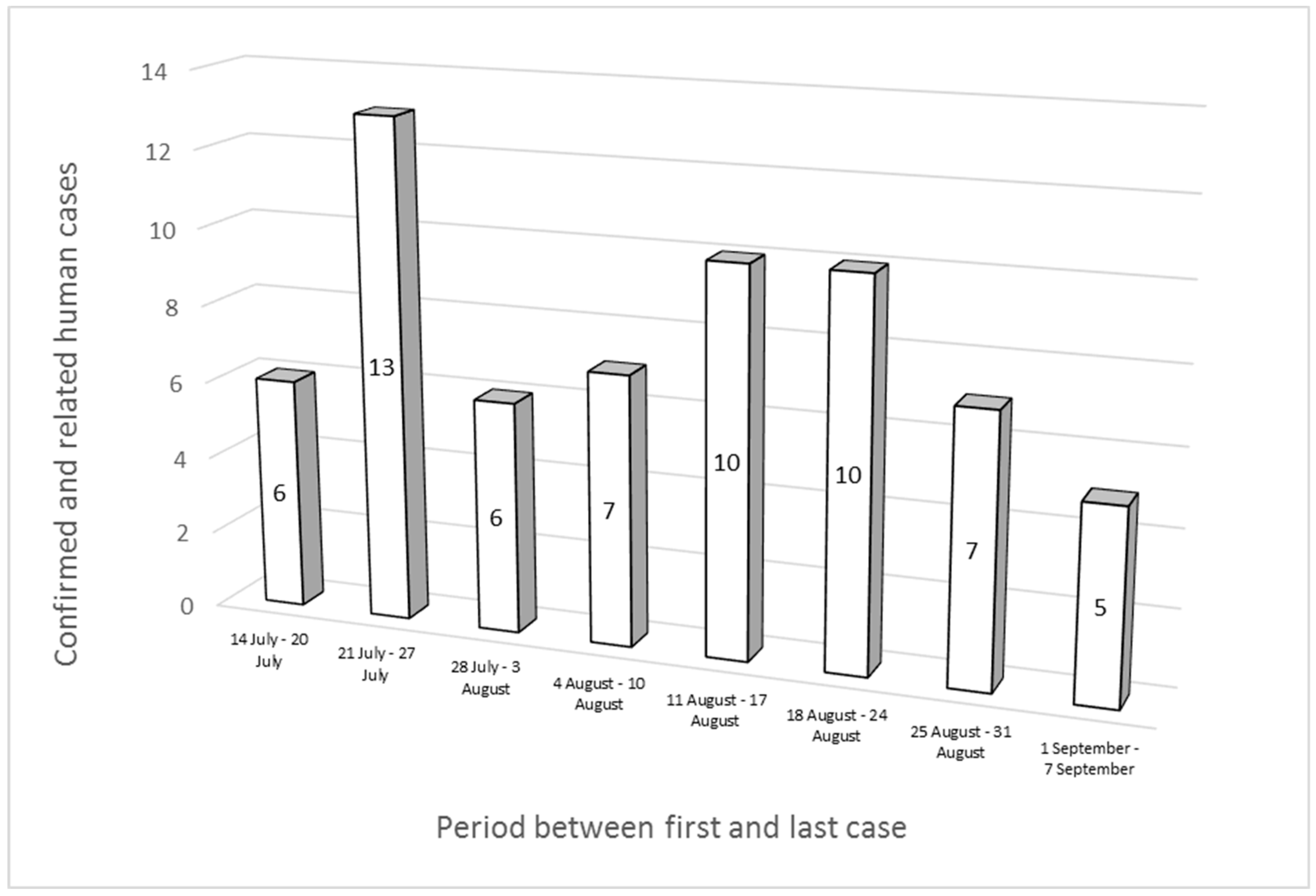
3.1.4. Epidemiological Investigations and Inquires in Neighboring Regions
3.2. Food-Related and Environmental Strains
3.2.1. Microbiological, Serotyping, and PCR Analyses
3.2.2. Antimicrobial Susceptibility and MLVA
3.3. Whole Genome Sequencing Analysis of the Clinical, Food-Related, and Environmental Strains
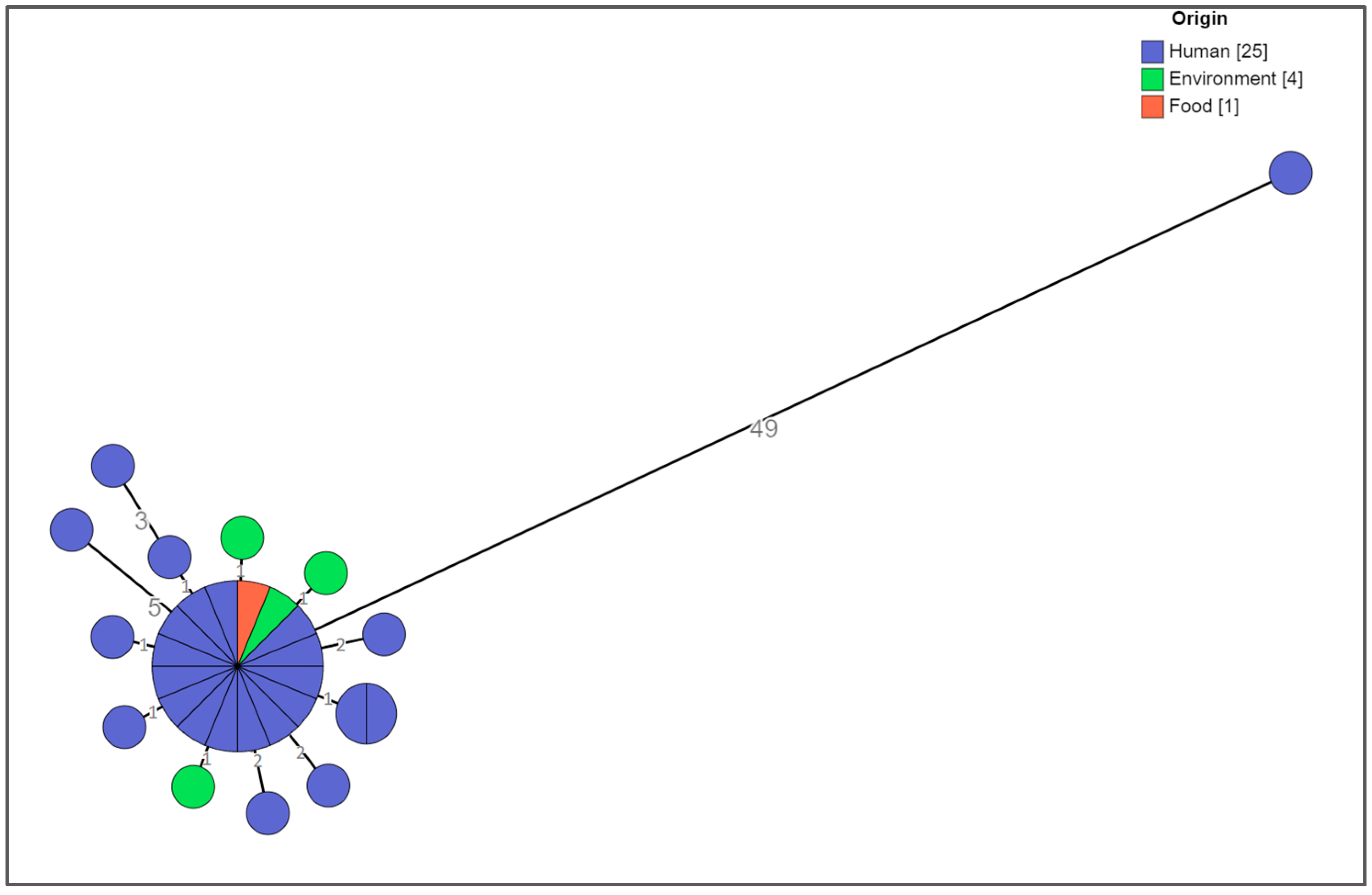
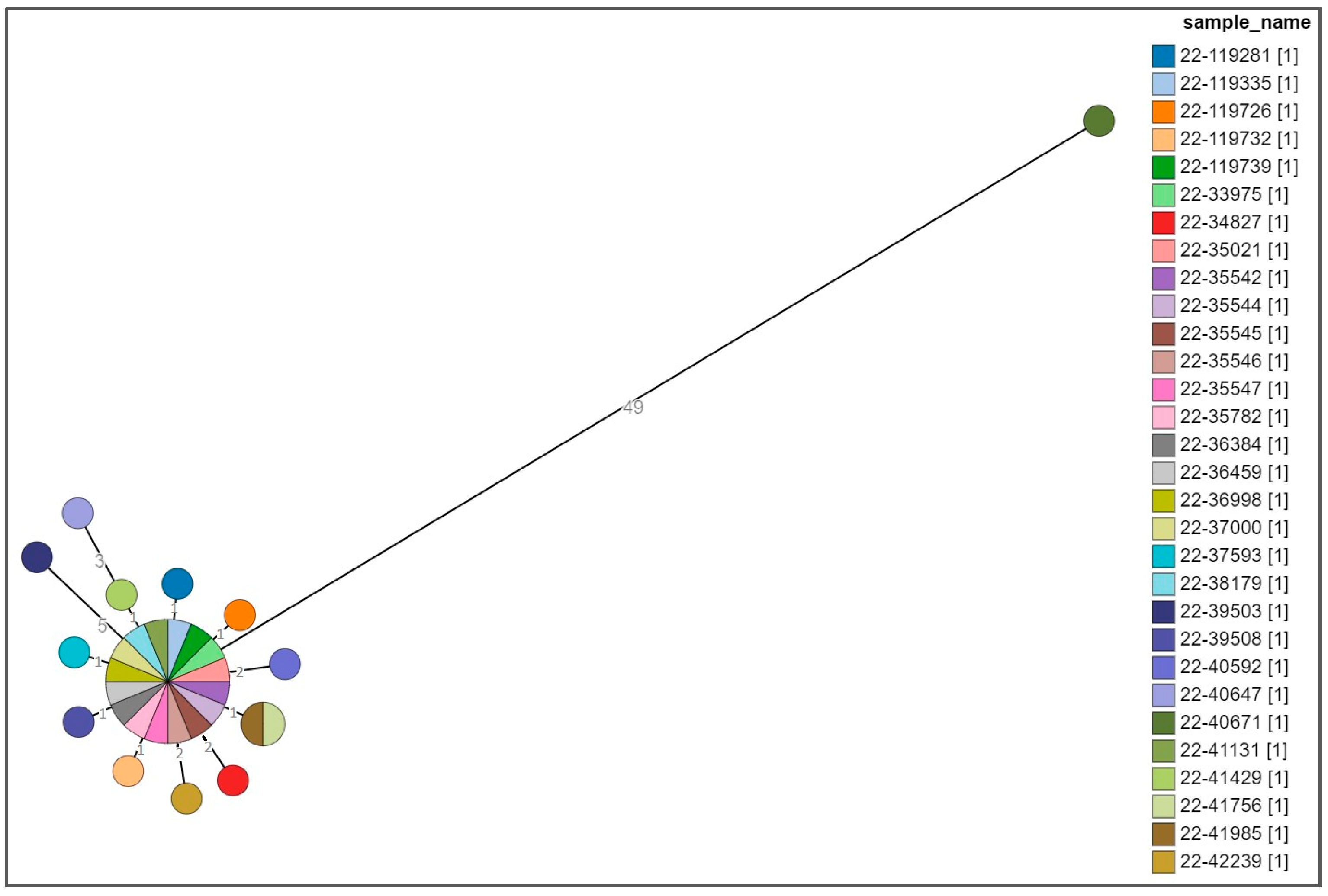
Cluster Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Crayford, G.; Coombes, J.L.; Humphrey, T.J.; Wigley, P. Monophasic Expression of FliC by Salmonella 4,[5],12:I:- DT193 Does Not Alter Its Pathogenicity during Infection of Porcine Intestinal Epithelial Cells. Microbiology 2014, 160, 2507–2516. [Google Scholar] [CrossRef]
- Machado, J.; Bernardo, F. Prevalence of Salmonella in Chicken Carcasses in Portugal. J. Appl. Bacteriol. 1990, 69, 477–480. [Google Scholar] [CrossRef]
- Sun, H.; Wan, Y.; Du, P.; Bai, L. The Epidemiology of Monophasic Salmonella Typhimurium. Foodborne Pathog. Dis. 2020, 17, 87–97. [Google Scholar] [CrossRef]
- Survellaince Atlas of Infectious Diseases. Available online: https://atlas.ecdc.europa.eu/public/index.aspx (accessed on 20 April 2023).
- Leati, M.; Cibin, V.; Mancin, M.; Pestelli, P.; Barco, L. Enter-Vet Report Dati 2019, Centro Di Referenza Nazionale per Le Salmonellosi; Istituto Zooprofiliattico Sperimentale Delle Venezi: Legnaro, Italy, 2019. [Google Scholar]
- European Food Safety Authority; European Centre for Disease Prevention and Control. The European Union One Health 2021 Zoonoses Report. EFSA J. 2022, 20, e07666. [Google Scholar] [CrossRef]
- D’Incau, M.; Salogni, C.; Giovannini, S.; Ruggeri, J.; Scali, F.; Tonni, M.; Formenti, N.; Guarneri, F.; Pasquali, P.; Alborali, G.L. Occurrence of Salmonella Typhimurium and Its Monophasic Variant (4,[5],12:I:-) In Healthy and Clinically Ill Pigs in Northern Italy. Porc. Health Manag. 2021, 7, 34. [Google Scholar] [CrossRef] [PubMed]
- Kapetanovic, R.; Bokil, N.J.; Achard, M.E.S.; Ong, C.Y.; Peters, K.M.; Stocks, C.J.; Phan, M.; Monteleone, M.; Schroder, K.; Irvine, K.M.; et al. Salmonella Employs Multiple Mechanisms to Subvert the TLR-inducible Zinc-mediated Antimicrobial Response of Human Macrophages. FASEB J. 2016, 30, 1901–1912. [Google Scholar] [CrossRef] [PubMed]
- Neyrolles, O.; Wolschendorf, F.; Mitra, A.; Niederweis, M. Mycobacteria, Metals, and the Macrophage. Immunol. Rev. 2015, 264, 249–263. [Google Scholar] [CrossRef] [PubMed]
- Mastrorilli, E.; Pietrucci, D.; Barco, L.; Ammendola, S.; Petrin, S.; Longo, A.; Mantovani, C.; Battistoni, A.; Ricci, A.; Desideri, A.; et al. A Comparative Genomic Analysis Provides Novel Insights Into the Ecological Success of the Monophasic Salmonella Serovar 4,[5],12:I:-. Front. Microbiol. 2018, 9, 715. [Google Scholar] [CrossRef]
- Medardus, J.J.; Molla, B.Z.; Nicol, M.; Morrow, W.M.; Rajala-Schultz, P.J.; Kazwala, R.; Gebreyes, W.A. In-Feed Use of Heavy Metal Micronutrients in U.S. Swine Production Systems and Its Role in Persistence of Multidrug-Resistant Salmonellae. Appl. Environ. Microbiol. 2014, 80, 2317–2325. [Google Scholar] [CrossRef]
- Mourão, J.; Machado, J.; Novais, C.; Antunes, P.; Peixe, L. Characterization of the Emerging Clinically-Relevant Multidrug-Resistant Salmonella Enterica Serotype 4,[5],12:I:-(Monophasic Variant of S. Typhimurium) Clones. Eur. J. Clin. Microbiol. Infect. Dis. 2014, 33, 2249–2257. [Google Scholar] [CrossRef]
- European Medicines Agency. Sales of Veterinary Antimicrobial Agents in 31 European Countries in 2021: Trends from 2010 to 2021: Twelfth ESVAC Report; Publications Office: Luxembourg, 2022. [Google Scholar]
- Plasmid-Mediated. Antimicrobial Resistance in Salmonella Enterica. Curr. Issues Mol. Biol. 2003, 5, 113–122. [Google Scholar] [CrossRef]
- Barco, L.; Ramon, E.; Cortini, E.; Longo, A.; Dalla Pozza, M.C.; Lettini, A.A.; Dionisi, A.M.; Olsen, J.E.; Ricci, A. Molecular Characterization of Salmonella enterica Serovar 4,[5],12:I:-DT193 ASSuT Strains from Two Outbreaks in Italy. Foodborne Pathog. Dis. 2014, 11, 138–144. [Google Scholar] [CrossRef]
- Andreoli, G.; Merla, C.; Valle, C.D.; Corpus, F.; Morganti, M.; D’incau, M.; Colmegna, S.; Marone, P.; Fabbi, M.; Barco, L.; et al. Foodborne Salmonellosis in Italy: Characterization of Salmonella enterica Serovar Typhimurium and Monophasic Variant 4,[5],12:I−Isolated from Salami and Human Patients. J. Food Prot. 2017, 80, 632–639. [Google Scholar] [CrossRef] [PubMed]
- Gossner, C.M.; van Cauteren, D.; Le Hello, S.; Weill, F.X.; Terrien, E.; Tessier, S.; Janin, C.; Brisabois, A.; Dusch, V.; Vaillant, V.; et al. Nationwide Outbreak of Salmonella enterica Serotype 4,[5],12:I:-Infection Associated with Consumption of Dried Pork Sausage, France, November to December 2011. Eurosurveillance 2012, 17, 20071. [Google Scholar] [CrossRef] [PubMed]
- Raguenaud, M.E.; Le Hello, S.; Salah, S.; Weill, F.X.; Brisabois, A.; Delmas, G.; Germonneau, P. Epidemiological and Microbiological Investigation of a Large Outbreak of Monophasic Salmonella Typhimurium 4,5,12:I:-In Schools Associated with Imported Beef in Poitiers, France, October 2010. EuroSurveill 2012, 17, 20289. [Google Scholar] [CrossRef]
- Arnedo-Pena, A.; Sabater-Vidal, S.; Herrera-León, S.; Bellido-Blasco, J.B.; Silvestre-Silvestre, E.; Meseguer-Ferrer, N.; Yague-Muñoz, A.; Gil-Fortuño, M.; Romeu-García, A.; Moreno-Muñoz, R. An Outbreak of Monophasic and Biphasic Salmonella Typhimurium, and Salmonella Derby Associated with the Consumption of Dried Pork Sausage in Castellon (Spain). Enfermedades Infecc. Y Microbiol. Clínica 2016, 34, 544–550. [Google Scholar] [CrossRef]
- Pardos De La Gandara, M.; Fournet, N.; Bonifait, L.; Lefèvre, S.; Chemaly, M.; Grastilleur, C.; Cadel-Six, S.; Fach, P.; Pignault, A.; Brisabois, A.; et al. Countrywide Multi-Serotype Outbreak of Salmonella Bovismorbificans ST142 and Monophasic Salmonella Typhimurium ST34 Associated with Dried Pork Sausages in France, September 2020* to January 2021. Eurosurveillance 2023, 28, 2200123. [Google Scholar] [CrossRef]
- de Frutos, M.; López-Urrutia, L.; Berbel, C.; Allue, M.; Herrera, S.; Azcona, J.M.; Beristaín, X.; Aznar, E.; Albert, M.; Ruiz, C.; et al. [Monophasic Salmonella Typhimurium outbreak due to the consumption of roast pork meat]. Rev. Esp. Quim. 2018, 31, 156–159. [Google Scholar]
- Kawakami, V.M.; Bottichio, L.; Angelo, K.; Linton, N.; Kissler, B.; Basler, C.; Lloyd, J.; Inouye, W.; Gonzales, E.; Rietberg, K.; et al. Notes from the Field: Outbreak of Multidrug-Resistant Salmonella Infections Linked to Pork—Washington, 2015. MMWR Morb. Mortal. Wkly. Rep. 2016, 65, 379–381. [Google Scholar] [CrossRef]
- Biswas, S.; Li, Y.; Elbediwi, M.; Yue, M. Emergence and Dissemination of Mcr-Carrying Clinically Relevant Salmonella Typhimurium Monophasic Clone ST34. Microorganisms 2019, 7, 298. [Google Scholar] [CrossRef]
- Russini, V.; Corradini, C.; Rasile, E.; Terracciano, G.; Senese, M.; Bellagamba, F.; Amoruso, R.; Bottoni, F.; De Santis, P.; Bilei, S.; et al. A Familiar Outbreak of Monophasic Salmonella Serovar Typhimurium (ST34) Involving Three Dogs and Their Owner’s Children. Pathogens 2022, 11, 1500. [Google Scholar] [CrossRef]
- Lund, S.; Tahir, M.; Vohra, L.I.; Hamdana, A.H.; Ahmad, S. Outbreak of Monophasic Salmonella Typhimurium Sequence Type 34 Linked to Chocolate Products. Ann. Med. Surg. 2022, 82, 104597. [Google Scholar] [CrossRef] [PubMed]
- Larkin, L.; Pardos De La Gandara, M.; Hoban, A.; Pulford, C.; Jourdan-Da Silva, N.; De Valk, H.; Browning, L.; Falkenhorst, G.; Simon, S.; Lachmann, R.; et al. Investigation of an International Outbreak of Multidrug-Resistant Monophasic Salmonella Typhimurium Associated with Chocolate Products, EU/EEA and United Kingdom, February to April 2022. Eurosurveillance 2022, 27, 2200314. [Google Scholar] [CrossRef] [PubMed]
- Colombe, S.; Jernberg, C.; Löf, E.; Angervall, A.L.; Mellström-Dahlgren, H.; Dotevall, L.; Bengnér, M.; Hall, I.; Sundqvist, L.; Kühlmann-Berenzon, S.; et al. Outbreak of Unusual H2S-Negative Monophasic Salmonella Typhimurium Strain Likely Associated with Small Tomatoes, Sweden, August to October 2019. Eurosurveillance 2019, 24, 1900643. [Google Scholar] [CrossRef]
- ISO 6579-1:2017/Amd 1:2020; Microbiology of the Food Chain—Horizontal Method for the Detection, Enumeration and Serotyping of Salmonella—Part 1: Detection of Salmonella spp.—Amendment 1: Broader Range of Incubation Temperatures, Amendment to the Status of Annex D, and Correction of the Composition of MSRV and SC. ISO: Geneva, Switzerland, 2020.
- ISO/TR 6579-3:2014; Microbiology of the Food Chain—Horizontal Method for the Detection, Enumeration and Serotyping of Salmonella—Part 3: Guidelines for Serotyping of Salmonella spp. ISO: Geneva, Switzerland, 2014.
- Lim, Y.-H.; Hirose, K.; Izumiya, H.; Arakawa, E.; Takahashi, H.; Terajima, J.; Itoh, K.; Tamura, K.; Kim, S.-I.; Watanabe, H. Multiplex Polymerase Chain Reaction Assay for Selective Detection of Salmonella enterica Serovar Typhimurium. Jpn. J. Infect. Dis. 2003, 56, 151–155. [Google Scholar]
- Rahn, K.; De Grandis, S.A.; Clarke, R.C.; McEwen, S.A.; Galán, J.E.; Ginocchio, C.; Curtiss, R.; Gyles, C.L. Amplification of an invA Gene Sequence of Salmonella Typhimurium by Polymerase Chain Reaction as a Specific Method of Detection of Salmonella. Mol. Cell. Probes 1992, 6, 271–279. [Google Scholar] [CrossRef]
- Weinstein, M.P.; Clinical and Laboratory Standards Institute CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 33rd ed.; CLSI Supplement M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2023. [Google Scholar]
- ECDC; Statens Serum Institut. Laboratory Standard Operating Procedure for MLVA of Salmonella Enterica Serotype Typhimurium; Publications Office: Luxembourg, 2011. [Google Scholar]
- Chen, S. Ultrafast One-pass FASTQ Data Preprocessing, Quality Control, and Deduplication Using Fastp. iMeta 2023, 2, e107. [Google Scholar] [CrossRef]
- Silva, M.; Machado, M.P.; Silva, D.N.; Rossi, M.; Moran-Gilad, J.; Santos, S.; Ramirez, M.; Carriço, J.A. chewBBACA: A Complete Suite for Gene-by-Gene Schema Creation and Strain Identification. Microb. Genom. 2018, 4, e000166. [Google Scholar] [CrossRef]
- Zhou, Z.; Alikhan, N.-F.; Sergeant, M.J.; Luhmann, N.; Vaz, C.; Francisco, A.P.; Carriço, J.A.; Achtman, M. GrapeTree: Visualization of Core Genomic Relationships among 100,000 Bacterial Pathogens. Genome Res. 2018, 28, 1395–1404. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA); Costa, G.; Di Piazza, G.; Koevoets, P.; Iacono, G.; Liebana, E.; Pasinato, L.; Rizzi, V.; Rossi, M. Guidelines for Reporting Whole Genome Sequencing-based Typing Data through the EFSA One Health WGS System. EFS3 2022, 19, 7413E. [Google Scholar] [CrossRef]
- De la Torre, E.; Zapata, D.; Tello, M.; Mejía, W.; Frías, N.; García Peña, F.J.; Mateu, E.M.; Torre, E. Several Salmonella Enterica Subsp. Enterica Serotype 4,5,12:I:−Phage Types Isolated from Swine Samples Originate from Serotype Typhimurium DT U302. J. Clin. Microbiol. 2003, 41, 2395–2400. [Google Scholar] [CrossRef] [PubMed]
- Lettini, A.A.; Saccardin, C.; Ramon, E.; Longo, A.; Cortini, E.; Dalla Pozza, M.C.; Barco, L.; Guerra, B.; Luzzi, I.; Ricci, A. Characterization of an Unusual Salmonella Phage Type DT7a and Report of a Foodborne Outbreak of Salmonellosis. Int. J. Food Microbiol. 2014, 189, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Borch, E.; Nesbakken, T.; Christensen, H. Hazard Identification in Swine Slaughter with Respect to Foodborne Bacteria. Int. J. Food Microbiol. 1996, 30, 9–25. [Google Scholar] [CrossRef]
- Lauteri, C.; Festino, A.R.; Conter, M.; Vergara, A. Prevalence and Antimicrobial Resistance Profile in Salmonella Spp. Isolates from Swine Food Chain. Ital. J. Food Saf. 2022, 11, 9980. [Google Scholar] [CrossRef]
- Lucarelli, C.; Dionisi, A.M.; Torpdahl, M.; Villa, L.; Graziani, C.; Hopkins, K.; Threlfall, J.; Caprioli, A.; Luzzi, I. Evidence for a Second Genomic Island Conferring Multidrug Resistance in a Clonal Group of Strains of Salmonella Enterica Serovar Typhimurium and Its Monophasic Variant Circulating in Italy, Denmark, and the United Kingdom. J. Clin. Microbiol. 2010, 48, 2103–2109. [Google Scholar] [CrossRef]
- Elnekave, E.; Hong, S.; Mather, A.E.; Boxrud, D.; Taylor, A.J.; Lappi, V.; Johnson, T.J.; Vannucci, F.; Davies, P.; Hedberg, C.; et al. Salmonella enterica Serotype 4,[5],12:I:- In Swine in the United States Midwest: An Emerging Multidrug-Resistant Clade. Clin. Infect. Dis. 2018, 66, 877–885. [Google Scholar] [CrossRef] [PubMed]
- OECD; Food and Agriculture Organization of the United Nations. OECD-FAO Agricultural Outlook 2021–2030; OECD-FAO Agricultural Outlook; OECD: Paris, France, 2021; ISBN 978-92-64-43607-7. [Google Scholar]
- Harrison, O.L.; Rensing, S.; Jones, C.K.; Trinetta, V. Salmonella enterica 4,[5],12:I:-, an Emerging Threat for the Swine Feed and Pork Production Industry. J. Food Prot. 2022, 85, 660–663. [Google Scholar] [CrossRef] [PubMed]
| No. of Strains | Origin of Sample | Antibiotic Resistance Type * | MLVA Profile | Inclusiveness in the MVST Cluster |
|---|---|---|---|---|
| 56 | Feces (55) + Urine (1) | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 | Yes |
| 1 | Feces | ACSSu+Gm+Tmp+Sxt | 3-13-10-na-211 | Yes |
| 3 | Feces | ACSSu+Tmp+Sxt | 3-14-10-na-211 | closely related |
| 1 | Feces | ACSSu | 3-14-10-na-211 | closely related |
| 1 | Feces | ACSSu+Gm | 3-14-10-na-211 | closely related |
| 1 | Feces | ACSSu+Amc+Gm+Tmp+Sxt | 3-14-10-na-211 | closely related |
| 1 | Feces | CSSu+Gm+Tmp+Sxt | 3-14-10-na-211 | closely related |
| 3 | Feces | ASSuT | 3-13-10-na-211 | No |
| 1 | Feces | ASSuT | 2-11-10-na-211 | No |
| 1 | Feces | ASSuT | 3-11-10-na-211 | No |
| 1 | Feces | ASSuT | 3-11-11-na-211 | No |
| 1 | Feces | ASSuT | 3-12-11-na-211 | No |
| 1 | Feces | ASSuT | 3-12-17-na-211 | No |
| 1 | Feces | ASFox | 3-12-8-na-211 | No |
| 1 | Feces | ASSu | 3-12-14-na-211 | No |
| 1 | Feces | ASSu | 3-12-9-na-211 | No |
| 1 | Feces | ASSuFox | 3-12-16-na-211 | No |
| 1 | Feces | ASSuGm | 3-12-8-na-211 | No |
| 1 | Feces | ASSuTPef | 3-15-10-na-211 | No |
| No. of Strains | Place of Sampling | Sample Detail | Origin of Sample | Antibiotic Resistance Type ** | MLVA Profile |
|---|---|---|---|---|---|
| 1 | RS(A) | Sponge swab on an unsanitized surface | Teflon chopping board for supporting and cutting porchetta (FCS) * | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 1 | RS(B) | Ready-to-eat food | Porchetta | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 1 | RS(B) | Sponge swab on an unsanitized surface | Wooden chopping board for porchetta(FCS) * | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 1 | RS(B) | Sponge swab on an unsanitized surface | Porchetta knife (FCS) * | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 1 | FPP | Sponge swab on an unsanitized surface | Transporting board for cooked porchetta (FCS) * | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| ID Strain | Origin of Sample | Identification Detail | Consumption of Porchetta by Cases | Antibiotic Resistance Type | MLVA Profile |
|---|---|---|---|---|---|
| 22-35542 | Human | Feces | Confirmed | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-35544 | Human | Feces | Confirmed | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-35546 | Human | Feces | Confirmed | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-36384 | Human | Feces | Confirmed | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-36459 | Human | Feces | Confirmed | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-35782 | Human | Feces | Confirmed | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-36998 | Human | Feces | Confirmed | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-37000 | Human | Feces | Confirmed | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-38179 | Human | Feces | Confirmed | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-35021 | Human | Feces | Information not available | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-35545 | Human | Feces | Information not available | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-35547 | Human | Feces | Information not available | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-34827 | Human | Feces | Information not available | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-41131 | Human | Feces | Information not available | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-41429 | Human | Feces | Information not available | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-40647 | Human | Urine | Information not available | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-41985 | Human | Feces | Information not available | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-37593 | Human | Feces | Information not available | ACSSu+Gm+Tmp+Sxt | 3-13-10-na-211 |
| 22-40592 | Human | Feces | Information not available | ACSSu+Tmp+Sxt | 3-14-10-na-211 |
| 22-41756 | Human | Feces | Information not available | ACSSu+Tmp+Sxt | 3-14-10-na-211 |
| 22-42239 | Human | Feces | Information not available | ACSSu+Tmp+Sxt | 3-14-10-na-211 |
| 22-40671 | Human | Feces | Information not available | ACSSu | 3-14-10-na-211 |
| 22-39503 | Human | Feces | Information not available | ACSSu+Gm | 3-14-10-na-211 |
| 22-33975 | Human | Feces | Information not available | ACSSu+Amc+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-39508 | Human | Feces | Information not available | CSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-119335 | Food | Porchetta | // | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-119726 | Environment | Teflon chopping board for supporting and cutting porchetta | // | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-119281 | Environment | Wooden chopping board for porchetta | // | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-119732 | Environment | Porchetta knife | // | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
| 22-119739 | Environment | Transporting board for cooked porchetta | // | ACSSu+Gm+Tmp+Sxt | 3-14-10-na-211 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Napoleoni, M.; Villa, L.; Barco, L.; Lucarelli, C.; Tiengo, A.; Baggio, G.; Dionisi, A.M.; Angellotti, A.; Ferretti, E.; Ruggeri, S.; et al. Monophasic Variant of Salmonella Typhimurium 4,[5],12:i:- (ACSSuGmTmpSxt Type) Outbreak in Central Italy Linked to the Consumption of a Roasted Pork Product (Porchetta). Microorganisms 2023, 11, 2567. https://doi.org/10.3390/microorganisms11102567
Napoleoni M, Villa L, Barco L, Lucarelli C, Tiengo A, Baggio G, Dionisi AM, Angellotti A, Ferretti E, Ruggeri S, et al. Monophasic Variant of Salmonella Typhimurium 4,[5],12:i:- (ACSSuGmTmpSxt Type) Outbreak in Central Italy Linked to the Consumption of a Roasted Pork Product (Porchetta). Microorganisms. 2023; 11(10):2567. https://doi.org/10.3390/microorganisms11102567
Chicago/Turabian StyleNapoleoni, Maira, Laura Villa, Lisa Barco, Claudia Lucarelli, Alessia Tiengo, Giulia Baggio, Anna Maria Dionisi, Antonio Angellotti, Ezio Ferretti, Simonetta Ruggeri, and et al. 2023. "Monophasic Variant of Salmonella Typhimurium 4,[5],12:i:- (ACSSuGmTmpSxt Type) Outbreak in Central Italy Linked to the Consumption of a Roasted Pork Product (Porchetta)" Microorganisms 11, no. 10: 2567. https://doi.org/10.3390/microorganisms11102567
APA StyleNapoleoni, M., Villa, L., Barco, L., Lucarelli, C., Tiengo, A., Baggio, G., Dionisi, A. M., Angellotti, A., Ferretti, E., Ruggeri, S., Staffolani, M., Rocchegiani, E., Silenzi, V., Morandi, B., & Blasi, G. (2023). Monophasic Variant of Salmonella Typhimurium 4,[5],12:i:- (ACSSuGmTmpSxt Type) Outbreak in Central Italy Linked to the Consumption of a Roasted Pork Product (Porchetta). Microorganisms, 11(10), 2567. https://doi.org/10.3390/microorganisms11102567





