Listeria monocytogenes Pathogenesis: The Role of Stress Adaptation
Abstract
1. Introduction
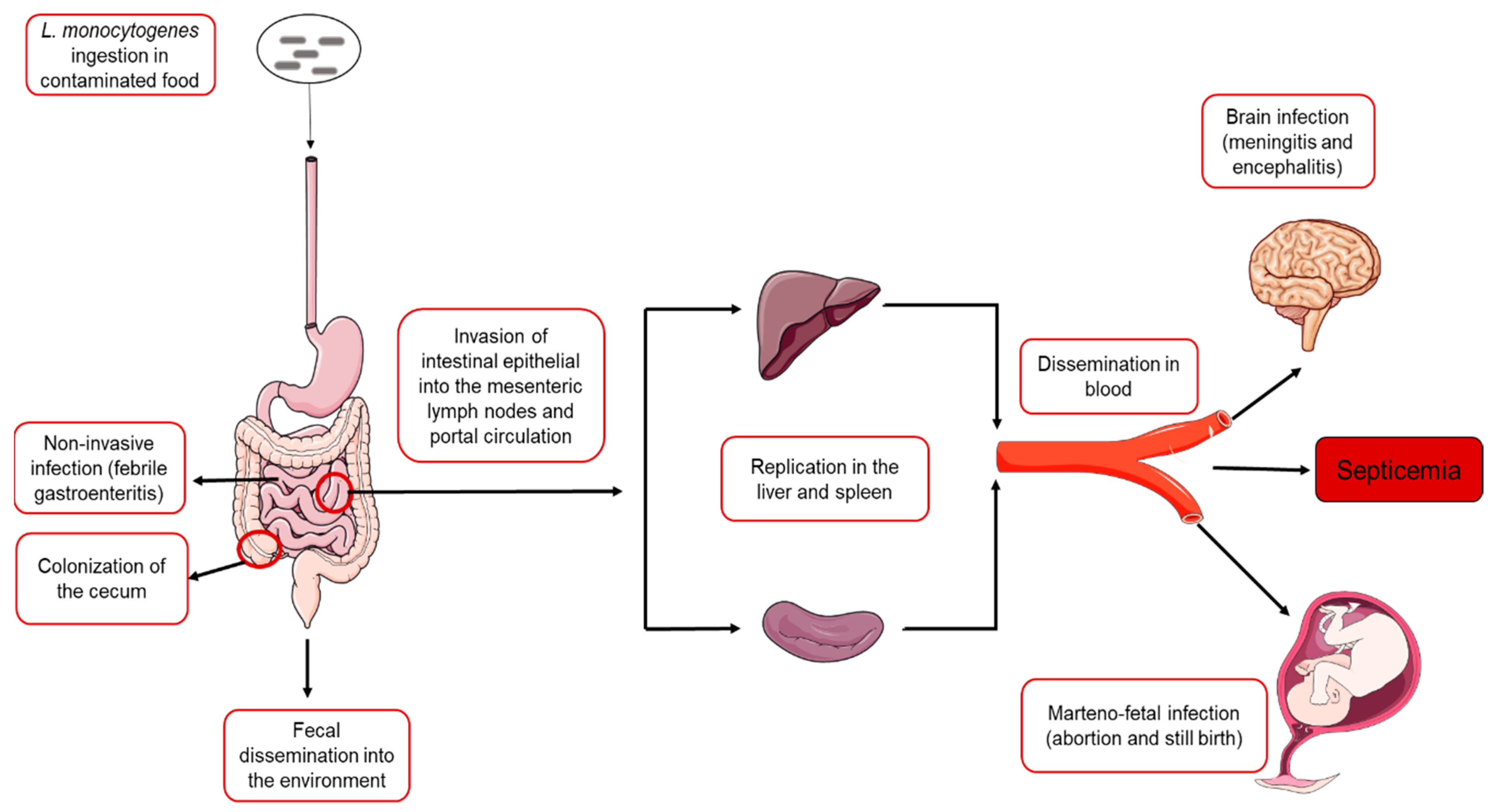
2. Overview of L. monocytogenes Infection Cycle
3. Pathogenesis of Invasive L. monocytogenes Infections
3.1. L. monocytogenes Virulence Factors
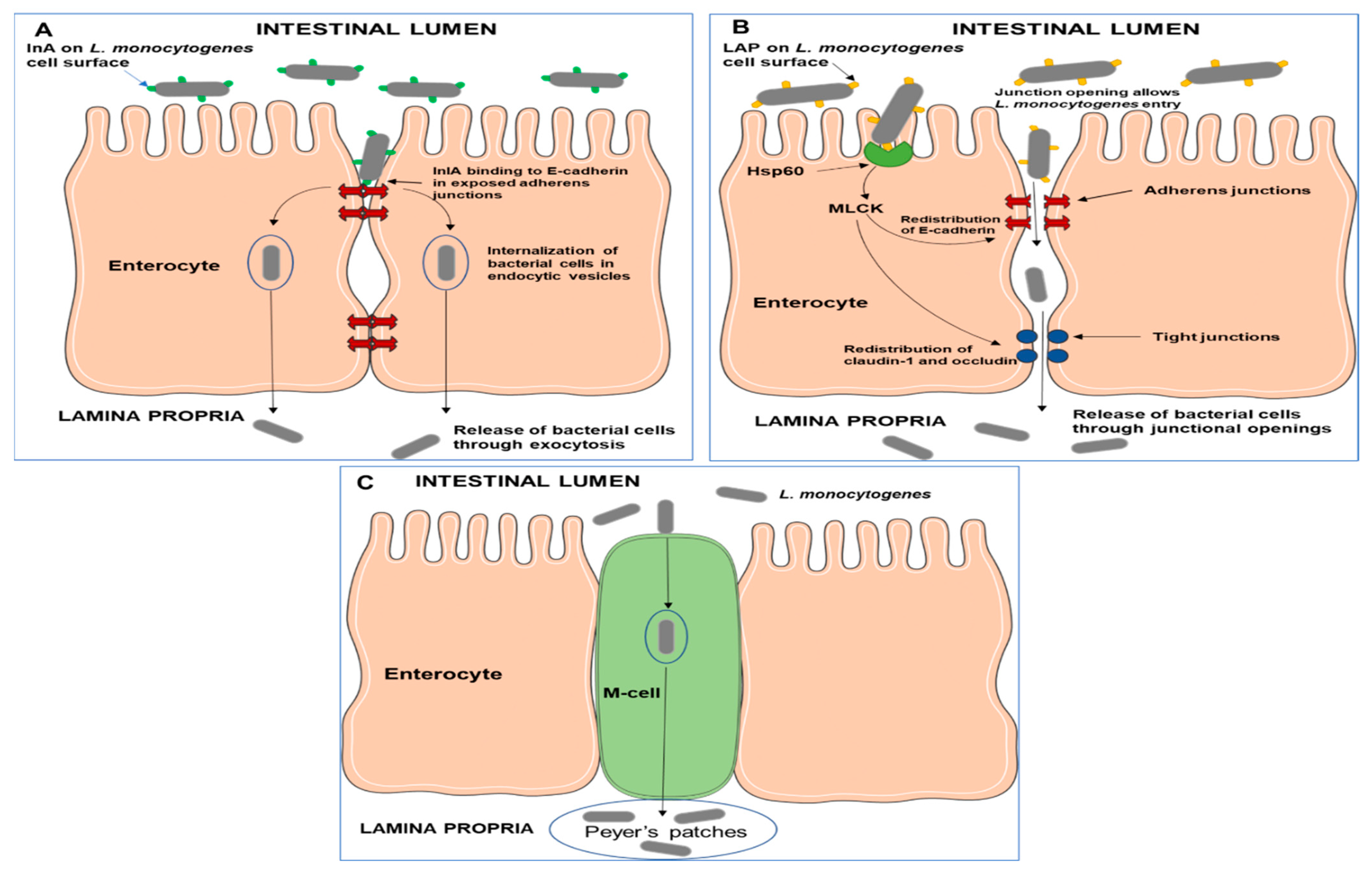
3.2. Gastrointestinal Tract Colonization and Invasion of Host Cells
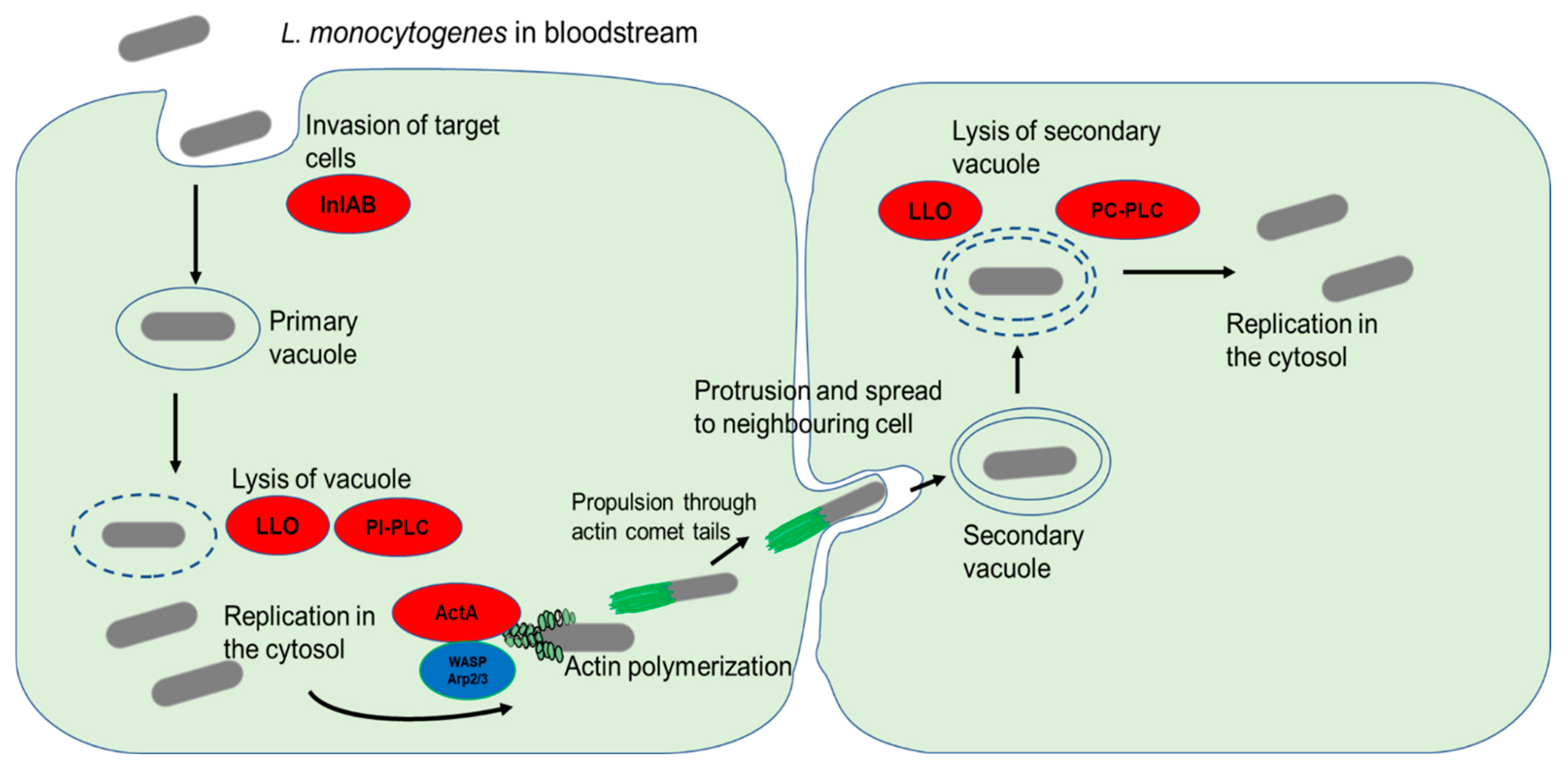
3.3. Intracellular Survival and Dissemination
3.4. Clinical Outcomes of Invasive L. monocytogenes Infections
4. L. monocytogenes Stress Responses and Adaptation
4.1. Adaptation to Stress in Foods and Food Processing Environments
4.1.1. Osmotic Stress Adaptation
4.1.2. Acid Stress Adaptation
4.1.3. Heat Stress Adaptation
4.1.4. Cold Stress Adaptation
4.1.5. Oxidative Stress Adaptation
4.2. Adaptation to Stress in the GIT
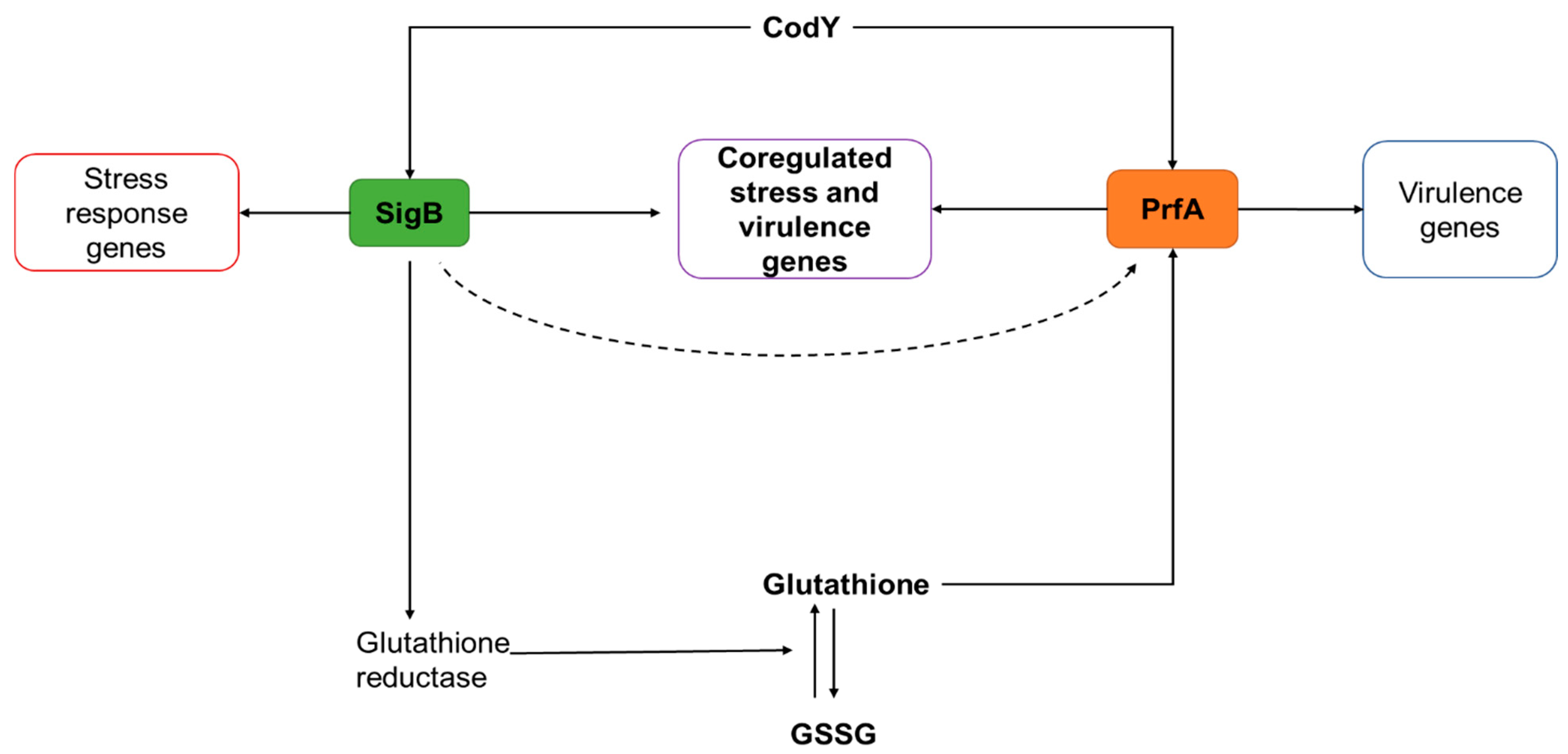
5. Crosslink between Stress Responses and Virulence
5.1. Regulation of L. monocytogenes Stress Response
Molecular Mechanisms of SigB-Dependent Regulation
5.2. Regulation of L. monocytogenes Virulence
5.3. Regulatory Intersection between Stress Response and Virulence
6. Strain and Lineage Variability in Stress Response and Virulence
7. Conclusions and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Linke, K.; Rückerl, I.; Brugger, K.; Karpiskova, R.; Walland, J.; Muri-Klinger, S.; Tichy, A.; Wagner, M.; Stessl, B. Reservoirs of Listeria species in three environmental ecosystems. Appl. Environ. Microbiol. 2014, 80, 5583–5592. [Google Scholar] [CrossRef]
- Halbedel, S.; Wilking, H.; Holzer, A.; Kleta, S.; Fischer, M.A.; Lüth, S.; Pietzka, A.; Huhulescu, S.; Lachmann, R.; Krings, A.; et al. Large nationawide outbreak of invasive listeriosis associated with blood sausage, Germany, 2018–2019. Emerg. Infect. Dis. 2020, 26, 1456–1464. [Google Scholar] [CrossRef]
- Thomas, J.; Govender, N.; McCarthy, K.M.; Erasmus, L.K.; Doyle, T.J.; Allam, M.; Ismail, A.; Ramalwa, N.; Sekwadi, P.; Ntshoe, G.; et al. Outbreak of listeriosis in South Africa associated with processed meat. N. Engl. J. Med. 2020, 382, 632–643. [Google Scholar] [CrossRef]
- Self, J.L.; Conrad, A.; Stroika, S.; Jackson, A.; Whitlock, L.; Jackson, K.A.; Al, J.L.S.E.; Wellman, A.; Fatica, M.K.; Bidol, S.; et al. Multistate outbreak of listeriosis associated with packaged leafy green salads, United States and Canada, 2015–2016. Emerg. Infect. Dis. 2019, 25, 1461–1468. [Google Scholar] [CrossRef] [PubMed]
- Warriner, K.; Namvar, A. What is the hysteria with Listeria? Trends Food Sci. Technol. 2009, 20, 245–254. [Google Scholar] [CrossRef]
- Quereda, J.J.; Morón-García, A.; Palacios-Gorba, C.; Dessaux, C.; Portillo, F.G.-D.; Pucciarelli, M.G.; Ortega, A.D. Pathogenicity and virulence of Listeria monocytogenes: A trip from environmental to medical microbiology. Virulence 2021, 12, 2509–2545. [Google Scholar] [CrossRef] [PubMed]
- Disson, O.; Lecuit, M. Targeting of the central nervous system by Listeria monocytogenes. Virulence 2012, 3, 213–221. [Google Scholar] [CrossRef]
- Pizarro-Cerdá, J.; Cossart, P. Invasion of host cells by Listeria monocytogenes. In Listeria monocytogenes: Pathogenesis and Host Response; Goldfine, H., Shen, H., Eds.; Springer Science Business Media, LLC: Berlin, Germany, 2007; pp. 159–176. [Google Scholar]
- Cossart, P.; Lebreton, A. A trip in the “new Microbiology” with the bacterial pathogen Listeria monocytogenes. FEBS Lett. 2014, 588, 2437–2445. [Google Scholar] [CrossRef]
- Wiktorczyk-Kapischke, N.; Skowron, K.; Grudlewska-Buda, K.; Wałecka-Zacharska, E.; Korkus, J.; Gospodarek-Komkowska, E. Adaptive response of Listeria monocytogenes to the stress factors in the food processing environment. Front. Microbiol. 2021, 12, 710085. [Google Scholar] [CrossRef] [PubMed]
- Vorob’eva, L.I. Stressors, stress reactions, and survival of bacteria: A review. Appl. Biochem. Microbiol. 2004, 40, 217–224. [Google Scholar] [CrossRef]
- Guerreiro, D.N.; Arcari, T.; O’Byrne, C.P. The σB-mediated general stress response of Listeria monocytogenes: Life and death decision making in a pathogen. Front. Microbiol. 2020, 11, 1505. [Google Scholar] [CrossRef]
- Bucur, F.I.; Grigore-Gurgu, L.; Crauwels, P.; Riedel, C.U.; Nicolau, A.I. Resistance of Listeria monocytogenes to stress conditions encountered in food and food processing environments. Front. Microbiol. 2018, 9, 2700. [Google Scholar] [CrossRef]
- Fang, F.C.; Frawley, E.R.; Tapscott, T.; Vázquez-Torres, A. Bacterial stress responses during host infection. Cell Host Microbe. 2016, 20, 133–143. [Google Scholar] [CrossRef] [PubMed]
- Begley, M.; Hill, C. Stress adaptation in foodborne pathogens. Annu. Rev. Food Sci. Technol. 2015, 6, 191–210. [Google Scholar] [CrossRef]
- Gahan, C.G.M.; Hill, C. Listeria monocytogenes: Survival and adaptation in the gastrointestinal tract. Front. Cell Infect. Microbiol. 2014, 4, 9. [Google Scholar] [CrossRef] [PubMed]
- Kazmierczak, M.J.; Mithoe, S.C.; Boor, K.J.; Wiedmann, M. Listeria monocytogenes σB regulates stress response and virulence functions. J. Bacteriol. 2003, 185, 5722–5734. [Google Scholar] [CrossRef] [PubMed]
- Kazmierczak, M.J.; Wiedmann, M.; Boor, K.J. Contributions of Listeria monocytogenes σB and PrfA to expression of virulence and stress reponse genes during extra- and intracellular growth. Microbiology 2006, 152, 1827–1838. [Google Scholar] [CrossRef] [PubMed]
- Gaballa, A.; Guariglia-Oropeza, V.; Wiedmann, M.; Boor, K.J. Cross talk between SigB and PrfA in Listeria monocytogenes facilitates transitions between extra- and intracellular environments. Microbiol. Mol. Biol. Rev. 2019, 83, e00034-19. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Orsi, R.H.; Gaballa, A.; Wiedmann, M.; Boor, K.J.; Guariglia-Oropeza, V. Systematic review of the Listeria monocytogenes σB regulon supports a role in stress response, virulence and metabolism. Future Microbiol. 2019, 14, 801–828. [Google Scholar] [CrossRef]
- Tiensuu, T.; Guerreiro, D.N.; Oliveira, A.H.; O’Byrne, C.; Johansson, J. Flick of a switch: Regulatory mechanisms allowing Listeria monocytogenes to transition from a saprophyte to a killer. Microbiology 2019, 165, 819–833. [Google Scholar] [CrossRef] [PubMed]
- Drolia, R.; Bhunia, A.K. Crossing the intestinal barrier via Listeria adhesion protein and Internalin A. Trends Microbiol. 2019, 27, 408–425. [Google Scholar] [CrossRef]
- McMullen, P.D.; Freitag, N.E. Listeria monocytogenes. In Molecular Medical Microbiology, 2nd ed.; Tang, Y.-W., Sussman, M., Liu, D., Poxton, I., Schwartzman, J., Eds.; Academic Press, Elsevier Ltd.: New York, NY, USA, 2014; pp. 1345–1361. [Google Scholar]
- Lecuit, M. Listeria monocytogenes, a model in infection biology. Cell. Microbiol. 2020, 22, e13186. [Google Scholar] [CrossRef]
- Liu, D.; Yin, X.; Olyha, S.J.; Nascimento, M.S.L.; Chen, P.; White, T.; Gowthaman, U.; Zhang, T.; Gertie, J.A.; Zhang, B.; et al. IL-10-dependent crosstalk between murine marginal zone B cells, macrophages, and CD8α+ dendritic cells promotes Listeria monocytogenes infection. Immunity 2019, 51, 64–76. [Google Scholar] [CrossRef]
- Travier, L.; Guadagnini, S.; Gouin, E.; Dufour, A.; Chenal-Francisque, V.; Cossart, P.; Olivo-Marin, J.C.; Ghigo, J.M.; Disson, O.; Lecuit, M. ActA promotes Listeria monocytogenes aggregation, intestinal colonization and carriage. PLoS Pathog. 2013, 9, e1003131. [Google Scholar] [CrossRef]
- Disson, O.; Moura, A.; Lecuit, M. Making sense of the biodiversity and virulence of Listeria monocytogenes. Trends Microbiol. 2021, 29, 811–822. [Google Scholar] [CrossRef] [PubMed]
- de las Heras, A.; Cain, R.J.; Bielecka, M.K.; Vázquez-Boland, J.A. Regulation of Listeria virulence: PrfA master and commander. Curr. Opin. Microbiol. 2011, 14, 118–127. [Google Scholar] [CrossRef] [PubMed]
- Jagadeesan, B.; Littlejohn, A.E.F.; Amalaradjou, M.A.R.; Singh, A.K.; Mishra, K.K.; La, D.; Kihara, D.; Bhunia, A.K. N-Terminal Gly224-Gly411 domain in Listeria adhesion protein interacts with host receptor HsP60. PLoS ONE 2011, 6, e20694. [Google Scholar] [CrossRef]
- Pandiripally, V.K.; Westbrook, D.G.; Sunki, G.R.; Bhunia, A.K. Surface protein p104 is involved in adhesion of Listeria monocytogenes to human intestinal cell line, Caco-2. J. Med. Microbiol. 1999, 48, 117–124. [Google Scholar] [CrossRef]
- Burkholder, K.M.; Bhunia, A.K. Listeria monocytogenes uses Listeria adhesion protein (LAP) to promote bacterial transepithelial translocation and induces expression of LAP receptor Hsp60. Infect. Immun. 2010, 78, 5062–5073. [Google Scholar] [CrossRef]
- Jagadeesan, B.; Koo, O.K.; Kim, K.P.; Burkholder, K.M.; Mishra, K.K.; Aroonnual, A.; Bhunia, A.K. LAP, an alcohol acetaldehyde dehydrogenase enzyme in Listeria, promotes bacterial adhesion to enterocyte-like Caco-2 cells only in pathogenic species. Microbiology 2010, 156, 2782–2795. [Google Scholar] [CrossRef] [PubMed]
- Burkholder, K.M.; Kim, K.-P.; Mishra, K.K.; Medina, S.; Hahm, B.-K.; Kim, H.; Bhunia, A.K. Expression of LAP, a SecA2-dependent secretory protein, is induced under anaerobic environment. Microbes Infect. 2009, 11, 859–867. [Google Scholar] [CrossRef]
- Drolia, R.; Tenguria, S.; Durkes, A.C.; Turner, J.R.; Bhunia, A.K. Listeria adhesion protein induces intestinal epithelial barrier dysfunction for bacterial translocation. Cell Host Microbe 2018, 23, 470–484.e7. [Google Scholar] [CrossRef] [PubMed]
- Hymes, J.P.; Klaenhammer, T.R. Stuck in the middle: Fibronectin-binding proteins in Gram-positive bacteria. Front. Microbiol. 2016, 7, 1504. [Google Scholar] [CrossRef] [PubMed]
- Henderson, B.; Nair, S.; Pallas, J.; Williams, M.A. Fibronectin: A multidomain host adhesin targeted by bacterial fibronectin-binding proteins. FEMS Microbiol. Rev. 2011, 35, 147–200. [Google Scholar] [CrossRef]
- Dramsi, S.; Bourdichon, F.; Cabanes, D.; Lecuit, M.; Fsihi, H.; Cossart, P. FbpA, a novel multifunctional Listeria monocytogenes virulence factor. Mol. Microbiol. 2004, 53, 639–649. [Google Scholar] [CrossRef]
- Gaillard, J.-L.; Berche, P.; Frehel, C.; Gouln, E.; Cossart, P. Entry of L. monocytogenes into cells is mediated by internalin, a repeat protein reminiscent of surface antigens from Gram-positive cocci. Cell 1991, 65, 1127–1141. [Google Scholar] [CrossRef]
- Ireton, K.; Mortuza, R.; Gyanwali, G.C.; Gianfelice, A.; Hussain, M. Role of internalin proteins in the pathogenesis of Listeria monocytogenes. Mol. Microbiol. 2021, 116, 1407–1419. [Google Scholar] [CrossRef]
- Schubert, W.-D.; Urbanke, C.; Ziehm, T.; Beier, V.; Machner, M.P.; Domann, E.; Wehland, J.; Chakraborty, T.; Heinz, D.W. Structure of internalin, a major invasion protein of Listeria monocytogenes, in complex with its human receptor E-cadherin. Cell 2002, 111, 825–836. [Google Scholar] [CrossRef]
- Dellafiora, L.; Filipello, V.; Dall’Asta, C.; Finazzi, G.; Galaverna, G.; Losio, M.N. A structural study on the Listeria monocytogenes internalin A—Human E-cadherin interaction: A molecular tool to investigate the effects of missense mutations. Toxins 2020, 12, 60. [Google Scholar] [CrossRef] [PubMed]
- Braun, L.; Dramsi, S.; Dehoux, P.; Bierne, H.; Lindahl, G.; Cossart, P. InIB: An invasion protein of Listeria monocytogenes with a novel type of surface association. Mol. Microbiol. 1997, 25, 285–294. [Google Scholar] [CrossRef] [PubMed]
- Bierne, H.; Sabet, C.; Personnic, N.; Cossart, P. Internalins: A complex family of leucine-rich repeat-containing proteins in Listeria monocytogenes. Microbes Infect. 2007, 9, 1156–1166. [Google Scholar] [CrossRef] [PubMed]
- Pizarro-Cerdá, J.; Kühbacher, A.; Cossart, P. Entry of Listeria monocytogenes in mammalian epithelial cells: An updated view. Cold Spring Harb. Perspect. Med. 2012, 2, a010009. [Google Scholar] [CrossRef]
- Milohanic, E.; Glaser, P.; Coppée, J.-Y.; Frangeul, L.; Vega, Y.; Vázquez-Boland, J.A.; Kunst, F.; Cossart, P.; Buchrieser, C. Transcriptome analysis of Listeria monocytogenes identifies three groups of genes differently regulated by PrfA. Mol. Microbiol. 2003, 47, 1613–1625. [Google Scholar] [CrossRef]
- Hamon, M.; Ribet, D.; Stavru, F.; Cossart, P. Listeriolysin O: The Swiss army knife of Listeria. Trends Microbiol. 2012, 20, 360–368. [Google Scholar] [CrossRef]
- Hof, H. Virulence of different strains of Listeria monocytogenes serovar 1/2a. Med. Microbiol. Immunol. 1984, 173, 207–218. [Google Scholar] [CrossRef]
- Geoffroy, C.; Gaillard, J.-L.; Alouf, J.E.; Berche, P. Purification, characterization, and toxicity of thesulfhydry-activated hemolysin Listerion O from Listeria monocytogenes. Infect. Immun. 1987, 55, 1641–1646. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, M.; Kathariou, S.; Goebel, W. Hemolysin supports survival but not entry of the intracellular bacterium Listeria monocytogenes. Infect. Immun. 1988, 56, 79–82. [Google Scholar] [CrossRef] [PubMed]
- Phelps, C.C.; Vadia, S.; Arnett, E.; Tan, Y.; Zhang, X.; Pathak-Sharma, S.; Gavrilin, M.A.; Seveau, S. Relative roles of Listeriolysin O, InlA, and InlB in Listeria monocytogenes uptake by host cells. Infect. Immun. 2018, 86, e00555-18. [Google Scholar] [CrossRef] [PubMed]
- Camilli, A.; Goldfine, H.; Portnoy, D.A. Listeria monocytogenes mutants lacking phosphatidylinositol-specific phospholipase C are avirulent. J. Exp. Med. 1991, 173, 751–754. [Google Scholar] [CrossRef]
- Smith, G.A.; Marquis, H.; Jones, S.; Johnston, N.C.; Portnoy, D.A.; Goldfine, H. The two distinct phospholipases C of Listeria monocytogenes have overlapping roles in escape from a vacuole and cell-to-cell spread. Infect. Immun. 1995, 63, 4231–4237. [Google Scholar] [CrossRef]
- Poussin, M.A.; Goldfine, H. Involvement of Listeria monocytogenes phosphatidylinositol-specific phospholipase C and host protein kinase C in permeabilization of the macrophage phagosome. Infect. Immun. 2005, 73, 4410–4413. [Google Scholar] [CrossRef]
- Gründling, A.; Gonzalez, M.D.; Higgins, D.E. Requirement of the Listeria monocytogenes broad-Range phospholipase PC-PLC during infection of human epithelial cells. J. Bacteriol. 2003, 185, 6295–6307. [Google Scholar]
- Coffey, A.; Burg, B.V.D.; Veltman, R.; Abee, T. Characteristics of the biologically active 35-kDa metalloprotease virulence factor from Listeria monocytogenes. J. Appl. Microbiol. 2000, 88, 132–141. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Suárez, M.; González-Zorn, B.; Vega, Y.; Chico-Calero, I.; Vázquez-Boland, J.A. A role for ActA in epithelial cell invasion by Listeria monocytogenes. Cell. Microbiol. 2001, 3, 853–864. [Google Scholar] [CrossRef] [PubMed]
- Kocks, C.; Hellio, R.; Gounon, P.; Ohayon, H.; Cossart, P. Polarized distribution of Listeria monocytogenes surface protein ActA at the site of directional actin assembly. J. Cell Sci. 1993, 105, 699–710. [Google Scholar] [CrossRef] [PubMed]
- Travier, L.; Lecuit, M. Listeria monocytogenes ActA: A new function for a “classic” virulence factor. Curr. Opin. Microbiol. 2014, 17, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Skoble, J.; Auerbuch, V.; Goley, E.D.; Welch, M.D.; Portnoy, D.A. Pivotal role of VASP in Arp2/3 complex-mediated actin nucleation, actin branch-formation, and Listeria monocytogenes motility. J. Cell Biol. 2001, 155, 89–100. [Google Scholar] [CrossRef] [PubMed]
- Kühn, S.; Enninga, J. The actin comet guides the way: How Listeria actin subversion has impacted cell biology, infection biology and structural biology. Cell. Microbiol. 2020, 22, e13190. [Google Scholar] [CrossRef] [PubMed]
- Maurella, C.; Gallina, S.; Ru, G.; Adriano, D.; Bellio, A.; Bianchi, D.M.; Chiavacci, L.; Crescio, M.I.; Croce, M.; D'Errico, V.; et al. Outbreak of febrile gastroenteritis caused by Listeria monocytogenes 1/2a in sliced cold beef ham, Italy, May 2016. Eurosurveillance 2018, 23, 17–00155. [Google Scholar] [CrossRef]
- Ooi, S.T.; Lorber, B. Gastroenteritis due to Listeria monocytogenes. Clin. Infect. Dis. 2005, 40, 1327–1332. [Google Scholar] [CrossRef] [PubMed]
- Jacks, A.; Pihlajasaari, A.; Vahe, M.; Myntti, A.; Kaukoranta, S.-S.; Elomaa, N.; Salmenlinna, S.; Rantala, L.; Lahti, K.; Huusko, S.; et al. Outbreak of hospital-acquired gastroenteritis and invasive infection caused by Listeria monocytogenes, Finland, 2012. Epidemiol. Infect. 2016, 144, 2732–2742. [Google Scholar] [CrossRef]
- Sim, J.; Hood, D.; Finnie, L.; Wilson, M.; Graham, C.; Brett, M.; Hudson, J. Series of incidents of Listeria monocytogenes non-invasive febrile gastroenteritis involving ready-to-eat meats. Lett. Appl. Microbiol. 2002, 35, 409–413. [Google Scholar] [CrossRef]
- Halbedel, S.; Prager, R.; Banerji, S.; Kleta, S.; Trost, E.; Nishanth, G.; Alles, G.; Hölzel, C.; Schlesiger, F.; Pietzka, A.; et al. A Listeria monocytogenes ST2 clone lacking chitinase ChiB from an outbreak of non-invasive gastroenteritis. Emerg. Microbes Infect. 2019, 8, 17–28. [Google Scholar] [CrossRef]
- Radoshevich, L.; Cossart, P. Listeria monocytogenes: Towards a complete picture of its physiology and pathogenesis. Nat. Rev. Microbiol. 2018, 16, 32–46. [Google Scholar] [CrossRef]
- Nikitas, G.; Deschamps, C.; Disson, O.; Niault, T.; Cossart, P.; Lecuit, M. Transcytosis of Listeria monocytogenes across the intestinal barrier upon specific targeting of goblet cell accessible E-cadherin. J. Exp. Med. 2011, 208, 2263–2277. [Google Scholar] [CrossRef] [PubMed]
- Pentecost, M.; Otto, G.; Theriot, J.; Amieva, M.R. Listeria monocytogenes invades the epithelial junctions at sites of cell extrusion. PLoS Pathog. 2006, 2, 0029–0040. [Google Scholar] [CrossRef]
- Bonazzi, M.; Lecuit, M.; Cossart, P. Listeria monocytogenes internalin and E-cadherin: From structure to pathogenesis. Cell. Microbiol. 2009, 11, 693–702. [Google Scholar] [CrossRef] [PubMed]
- Saila, S.; Gyanwali, G.C.; Hussain, M.; Gianfelice, A.; Ireton, K. The host GTPase Arf1 and its effectors AP1 and PICK1 stimulate actin polymerization and exocytosis to promote entry of Listeria monocytogenes. Infect. Immun. 2020, 88, e00578-19. [Google Scholar] [CrossRef] [PubMed]
- Holch, A.; Ingmer, H.; Licht, T.R.; Gram, L. Listeria monocytogenes strains encoding premature stop codons in inlA invade mice and guinea pig fetuses in orally dosed dams. J. Med. Microbiol. 2013, 62, 1799–1806. [Google Scholar] [CrossRef]
- Gelbíčová, T.; Koláčková, I.; Pantůček, R.; Karpíšková, R. A novel mutation leading to a premature stop codon in inlA of Listeria monocytogenes isolated from neonatal listeriosis. New Microbiol. 2015, 38, 293–296. [Google Scholar]
- Hase, K.; Kawano, K.; Nochi, T.; Pontes, G.S.; Fukuda, S.; Ebisawa, M.; Kadokura, K.; Tobe, T.; Fujimura, Y.; Kawano, S.; et al. Uptake through glycoprotein 2 of FimH+ bacteria by M cells initiates mucosal immune response. Nature 2009, 462, 226–230. [Google Scholar] [CrossRef] [PubMed]
- Rey, C.; Chang, Y.-Y.; Latour-Lambert, P.; Varet, H.; Proux, C.; Legendre, R.; Coppée, J.-Y.; Enninga, J. Transcytosis subversion by M cell-to-enterocyte spread promotes Shigella flexneri and Listeria monocytogenes intracellular bacterial dissemination. PLoS Pathog. 2020, 16, e1008446. [Google Scholar] [CrossRef] [PubMed]
- Marco, A.J.; Altimira, J.; Prats, N.; López, S.; Dominguez, L.; Domingo, M.; Briones, V. Penetration of Listeria monocytogenes in mice infected by the oral route. Microb. Pathog. 1997, 23, 255–263. [Google Scholar] [CrossRef] [PubMed]
- Corr, S.; Hill, C.; Gahan, C.G. An in vitro cell-culture model demonstrates internalin- and hemolysin-independent translocation of Listeria monocytogenes across M cells. Microb. Pathog. 2006, 41, 241–250. [Google Scholar] [CrossRef] [PubMed]
- Charlier, C.; Disson, O.; Lecuit, M. Maternal-neonatal listeriosis. Virulence 2020, 11, 391–397. [Google Scholar] [CrossRef]
- Disson, O.; Grayo, S.; Huillet, E.; Nikitas, G.; Langa-Vives, F.; Dussurget, O.; Ragon, M.; Le Monnier, A.; Babinet, C.; Cossart, P.; et al. Conjugated action of two species-specific invasion proteins for fetoplacental listeriosis. Nat. Lett. 2008, 455, 1114–1118. [Google Scholar] [CrossRef] [PubMed]
- Kortebi, M.; Milohanic, E.; Mitchell, G.; Péchoux, C.; Prevost, M.-C.; Cossart, P.; Bierne, H. Listeria monocytogenes switches from dissemination to persistence by adopting a vacuolar lifestyle in epithelial cells. PLoS Pathog. 2017, 13, e1006734. [Google Scholar] [CrossRef] [PubMed]
- Schnupf, P.; Portnoy, D.A. Listeriolysin O: A phagosome-specific lysin. Microbes Infect. 2007, 9, 1176–1187. [Google Scholar] [CrossRef] [PubMed]
- Charlier, C.; Perrodeau, É.; Leclercq, A.; Cazenave, B.; Pilmis, B.; Henry, B.; Lopes, A.; Maury, M.M.; Moura, A.; Goffinet, F.; et al. Clinical features and prognostic factors of listeriosis: The MONALISA national prospective cohort study. Lancet Infect. Dis. 2017, 17, 510–519. [Google Scholar] [CrossRef]
- de Noordhout, C.M.; Devleesschauwer, B.; Angulo, F.J.; Verbeke, G.; Haagsma, J.; Kirk, M.; Havelaar, A.; Speybroeck, N. The global burden of listeriosis: A systematic review and meta-analysis. Lancet Infect. Dis. 2014, 14, 1073–1082. [Google Scholar] [CrossRef]
- Scobie, A.; Kanagarajah, S.; Harris, R.J.; Byrne, L.; Amar, C.; Grant, K.; Godbole, G. Mortality risk factors for listeriosis—A 10 year review of non-pregnancy associated cases in England 2006–2015. J. Infect. 2019, 78, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Wadhwa Desai, R.; Smith, M.A. Pregnancy-related listeriosis. Birth Defects Res. 2017, 109, 324–335. [Google Scholar] [CrossRef]
- Lepe, J.A. Current aspects of listeriosis. Med. Clínica 2020, 154, 453–458. [Google Scholar] [CrossRef] [PubMed]
- Shoai-Tehrani, M.; Pilmis, B.; Maury, M.M.; Robineau, O.; Disson, O.; Jouvion, G.; Coulpier, G.; Thouvenot, P.; Bracq-Dieye, H.; Valès, G.; et al. Listeria monocytogenes-associated endovascular infections: A study of 71 consecutive cases. J. Infect. 2019, 79, 322–331. [Google Scholar] [CrossRef]
- Morgand, M.; Leclercq, A.; Maury, M.; Bracq-Dieye, H.; Thouvenot, P.; Vales, G.; Lecuit, M.; Charlier, C. Listeria monocytogenes-associated respiratory infections: A study of 38 consecutive cases. Clin. Microbiol. Infect. 2018, 24, 1339-e1. [Google Scholar] [CrossRef]
- Charlier, C.; Fevre, C.; Travier, L.; Cazenave, B.; Bracq-Dieye, H.; Podevin, J.; Assomany, D.; Guilbert, L.; Bossard, C.; Carpentier, F.; et al. Listeria monocytogenes-associated biliary tract infections: A study of 12 consecutive cases and review. Medicine 2014, 93, e105. [Google Scholar] [CrossRef] [PubMed]
- Charlier, C.; Leclercq, A.; Cazenave, B.; Desplaces, N.; Travier, L.; Cantinelli, T.; Lortholary, O.; Goulet, V.; Le Monnier, A.; Lecuit, M.; et al. Listeria monocytogenes-associated joint and bone infections: A study of 43 consecutive cases. Clin. Infect. Dis. 2012, 54, 240–248. [Google Scholar] [CrossRef] [PubMed]
- Wesche, A.M.; Gurtler, J.B.; Marks, B.P.; Ryser, E.T. Stress, sublethal injury, resuscitation, and virulence of bacterial foodborne pathogens. J. Food Prot. 2009, 72, 1121–1138. [Google Scholar] [CrossRef]
- Singh, S.; Shalini, R. Effect of Hurdle Technology in Food Preservation: A Review. Crit. Rev. Food Sci. Nutr. 2016, 56, 641–649. [Google Scholar] [CrossRef] [PubMed]
- Chorianopoulos, N.; Giaouris, E.; Grigoraki, I.; Skandamis, P.; Nychas, G.-J. Effect of acid tolerance response (ATR) on attachment of Listeria monocytogenes Scott A to stainless steel under extended exposure to acid or/and salt stress and resistance of sessile cells to subsequent strong acid challenge. Int. J. Food Microbiol. 2011, 145, 400–406. [Google Scholar] [CrossRef]
- Davis, M.J.; Coote, P.J.; O’Byrne, C.P. Acid tolerance in Listeria monocytogenes: The adaptive acid tolerance response (ATR) and growth-phase-dependent acid resistance. Microbiology 1996, 142, 2975–2982. [Google Scholar] [CrossRef] [PubMed]
- O’Driscoll, B.; Gahan, C.G.M.; Hill, C. Adaptive acid tolerance response in Listeria monocytogenes: Isolation of an acid-tolerant mutant which demonstrates increased virulence. Appl. Environ. Microbiol. 1996, 62, 1693–1698. [Google Scholar] [CrossRef]
- Abeysundara, P.D.A.; Nannapaneni, R.; Soni, K.A.; Sharma, C.S.; Mahmoud, B. Induction and stability of oxidative stress adaptation in Listeria monocytogenes EGD (Bug600) and F1057 in sublethal concentrations of H2O2 and NaOH. Int. J. Food Microbiol. 2016, 238, 288–294. [Google Scholar] [CrossRef] [PubMed]
- Sibanda, T.; Buys, E.M. Modelling the survival of Listeria monocytogenes strains in soft lactic cheese following acid and salt stress exposures. Lett. Appl. Microbiol. 2019, 69, 230–236. [Google Scholar] [CrossRef] [PubMed]
- Skandamis, P.N.; Yoon, Y.; Stopforth, J.D.; Kendall, P.A.; Sofos, J.N. Heat and acid tolerance of Listeria monocytogenes after exposure to single and multiple sublethal stresses. Food Microbiol. 2008, 25, 294–303. [Google Scholar] [CrossRef] [PubMed]
- Faleiro, M.L.; Andrew, P.W.; Power, D. Stress response of Listeria monocytogenes isolated from cheese and other foods. Int. J. Food Microbiol. 2003, 84, 207–216. [Google Scholar] [CrossRef]
- Alvarez-Ordóñez, A.; Broussolle, V.; Colin, P.; Nguyen-The, C.; Prieto, M. The adaptive response of bacterial food-borne pathogens in the environment, host and food: Implications for food safety. Int. J. Food Microbiol. 2015, 213, 99–109. [Google Scholar] [CrossRef] [PubMed]
- Phadtare, S. Recent developments in bacterial cold-shock response. Curr. Issues Mol. Biol. 2004, 6, 125–136. [Google Scholar] [PubMed]
- Roncarati, D.; Scarlato, V. Regulation of heat-shock genes in bacteria: From signal sensing to gene expression output. FEMS Microbiol. Rev. 2017, 41, 549–574. [Google Scholar] [CrossRef]
- Ryan, S.; Begley, M.; Gahan, C.G.M.; Hill, C. Molecular characterization of the arginine deiminase system in Listeria monocytogenes: Regulation and role in acid tolerance. Environ. Microbiol. 2009, 11, 432–445. [Google Scholar] [CrossRef] [PubMed]
- Tasara, T.; Stephan, R. Cold stress tolerance of Listeria monocytogenes: A review of molecular adaptive mechanisms and food safety implications. J. Food Prot. 2006, 69, 1473–1484. [Google Scholar] [CrossRef]
- Cotter, P.D.; O’Reilly, K.; Hill, C. Role of the glutamate decarboxylase acid resistance system in the survival of Listeria monocytogenes LO28 in low pH foods. J. Food Prot. 2001, 64, 1362–1368. [Google Scholar] [CrossRef] [PubMed]
- Bremer, E.; Krämer, R. Responses of microorganisms to osmotic stress. Annu. Rev. Microbiol. 2019, 73, 313–334. [Google Scholar] [CrossRef] [PubMed]
- Yan, N.; Marschner, P.; Cao, W.; Zuo, C.; Qin, W. Influence of salinity and water content on soil microorganisms. Int. Soil Water Conserv. Res. 2015, 3, 316–323. [Google Scholar] [CrossRef]
- Manzanera, M. Dealing with water stress and microbial preservation. Environ. Microbiol. 2021, 23, 3351–3359. [Google Scholar] [CrossRef]
- Cole, M.B.; Jones, M.V.; Holyoak, C. The effect of pH, salt concentration and temperature on the survival and growth of Listeria monocytogenes. J. Appl. Microbiol. 1990, 69, 63–72. [Google Scholar]
- Angelidis, A.S.; Smith, G.M. Three transporters mediate uptake of glycine betaine and carnitine by Listeria monocytogenes in response to hyperosmotic stress. Appl. Environ. Microbiol. 2003, 69, 1013–1022. [Google Scholar] [CrossRef] [PubMed]
- Bayles, D.O.; Wilkinson, B.J. Osmoprotectants and cryoprotectants for Listeria monocytogenes. Lett. Appl. Microbiol. 2000, 30, 23–27. [Google Scholar] [CrossRef] [PubMed]
- Ko, R.; Smith, L.T. Identification of an ATP-driven, osmoregulated glycine betaine transport system in Listeria monocytogenes. Appl. Environ. Microbiol. 1999, 65, 4040–4048. [Google Scholar] [CrossRef] [PubMed]
- Wood, J.M. Osmosensing by bacteria: Signals and membrane-based sensors. Microbiol. Mol. Biol. Rev. 1999, 63, 230–262. [Google Scholar] [CrossRef] [PubMed]
- Gibhardt, J.; Heidemann, J.L.; Bremenkamp, R.; Rosenberg, J.; Seifert, R.; Kaever, V.; Ficner, R.; Commichau, F.M. An extracytoplasmic protein and a moonlighting enzyme modulate synthesis of c-di-AMP in Listeria monocytogenes. Environ. Microbiol. 2020, 22, 2771–2791. [Google Scholar] [CrossRef]
- Gerhardt, P.N.; Tombras Smith, L.; Smith, G.M. Osmotic and chill activation of glycine betaine porter II in Listeria monocytogenes membrane vesicles. J. Bacteriol. 2000, 182, 2544–2550. [Google Scholar] [CrossRef] [PubMed]
- Sleator, R.D.; Gahan, C.G.; Abee, T.; Hill, C. Identification and disruption of betL, a secondary glycine betaine transport system linked to the salt tolerance of Listeria monocytogenes LO28. Appl. Environ. Microbiol. 1999, 65, 2078–2083. [Google Scholar] [CrossRef] [PubMed]
- Sleator, R.D.; Wouters, J.; Gahan, C.G.; Abee, T.; Hill, C. Analysis of the role of OpuC, an osmolyte transport system, in salt tolerance and virulence potential of Listeria monocytogenes. Appl. Environ. Microbiol. 2002, 67, 2692–2698. [Google Scholar] [CrossRef]
- Roe, A.J.; McLaggan, D.; Davidson, I.; O’Byrne, C.; Booth, I.R. Perturbation of anion balance during inhibition of growth of Escherichia coli by weak acids. J. Bacteriol. 1998, 180, 767–772. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.A.; Rhee, M.S. Marked synergistic bactericidal effects and mode of action of medium-chain fatty acids in combination with organic acids against Escherichia coli O157:H7. Appl. Environ. Microbiol. 2013, 79, 6552–6560. [Google Scholar] [CrossRef] [PubMed]
- Ning, Y.; Yan, A.; Yang, K.; Wang, Z.; Li, X.; Jia, Y. Antibacterial activity of phenyllactic acid against Listeria monocytogenes and Escherichia coli by dual mechanisms. Food Chem. 2017, 228, 533–540. [Google Scholar] [CrossRef] [PubMed]
- Arcari, T.; Feger, M.-L.; Guerreiro, D.N.; Wu, J.; O’Byrne, C.P. Comparative review of the responses of Listeria monocytogenes and Escherichia coli to Low pH stress. Genes 2020, 11, 1330. [Google Scholar] [CrossRef]
- Cotter, P.D.; Ryan, S.; Gahan, C.G.M.; Hill, C. Presence of GadD1 glutamate decarboxylase in selected Listeria monocytogenes strains is associated with an ability to grow at low pH. Appl. Environ. Microbiol. 2005, 71, 2832–2839. [Google Scholar] [CrossRef] [PubMed]
- Spano, G.; Chieppa, G.; Beneduce, L.; Massa, S. Expression analysis of putative arcA, arcB and arcC genes partially cloned from Lactobacillus plantarum isolated from wine. J. Appl. Microbiol. 2004, 96, 185–193. [Google Scholar] [CrossRef]
- Ryan, S.; Hill, C.; Gahan, C.G.M. Acid stress responses in Listeria monocytogenes. In Advances in Applied Microbiology; Elsevier Masson SAS: Amsterdam, The Netherlands, 2008; pp. 67–91. [Google Scholar]
- Shen, Q.; Jangam, P.M.; Soni, K.A.; Nannapaneni, R.; Schilling, W.; Silva, J.L. Low, medium, and high heat tolerant strains of Listeria monocytogenes and increased heat stress resistance after exposure to sublethal heat. J. Food Prot. 2014, 77, 1298–1307. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Haykir, O.; Mohácsi-Farkas, C.; Engelhardt, T. Enhanced heat resistance of Listeria innocua as a surrogate of Listeria monocytogenes after sublethal heat treatment. Acta Aliment. 2022, 51, 241–248. [Google Scholar] [CrossRef]
- Lou, Y.; Yousef, A.E. Resistance of Listeria monocytogenes to heat after adaptation to environmental stresses. J. Food Prot. 1996, 59, 465–471. [Google Scholar] [CrossRef] [PubMed]
- Taormina, P.J.; Beuchat, L.R. Survival and heat resistance of Listeria monocytogenes after exposure to alkali and chlorine. Appl. Environ. Microbiol. 2001, 67, 2555–2563. [Google Scholar] [CrossRef] [PubMed]
- Van Der Veen, S.; Hain, T.; Wouters, J.A.; Hossain, H.; De Vos, W.M.; Abee, T.; Chakraborty, T.; Wells-Bennik, M.H.J. The heat-shock response of Listeria monocytogenes comprises genes involved in heat shock, cell division, cell wall synthesis, and the SOS response. Microbiology 2007, 153, 3593–3607. [Google Scholar] [CrossRef] [PubMed]
- Hanawa, T.; Kai, M.; Kamiya, S.; Yamamoto, T. Cloning, sequencing, and transcriptional analysis of the dnaK heat shock operon of Listeria monocytogenes. Cell Stress Chaperones 2000, 5, 21–29. [Google Scholar] [CrossRef]
- Gahan, C.G.M.; O’Mahony, J.; Hill, C. Characterization of the groESL operon in Listeria monocytogenes: Utilization of two reporter systems (gfp and hly) for evaluating in vivo expression. Infect. Immun. 2001, 69, 3924–3932. [Google Scholar] [CrossRef]
- Nair, S.; Derré, I.; Msadek, T.; Gaillot, O.; Berche, P. CtsR controls class III heat shock gene expression in the human pathogen Listeria monocytogenes. Mol. Microbiol. 2000, 35, 800–811. [Google Scholar] [CrossRef] [PubMed]
- Cabrita, P.; Trigo, M.J.; Ferreira, R.B.; Brito, L. Differences in the expression of cold stress-related genes and in the swarming motility among persistent and sporadic strains of Listeria monocytogenes. Foodborne Pathog. Dis. 2015, 12, 576–584. [Google Scholar] [CrossRef]
- Bayles, D.O.; Annous, B.A.; Wilkinson, B.J. Cold stress proteins induced in Listeria monocytogenes in response to temperature downshock and growth at low temperatures. Appl. Environ. Microbiol. 1996, 62, 1116–1119. [Google Scholar] [CrossRef]
- Russell, N.J. Bacterial membranes: The effects of chill storage and food processing. An overview. Int. J. Food. Microbiol. 2002, 79, 27–34. [Google Scholar] [CrossRef]
- Yoon, Y.; Lee, H.; Lee, S.; Kim, S.; Choi, K.-H. Membrane fluidity-related adaptive response mechanisms of foodborne bacterial pathogens under environmental stresses. Food Res. Int. 2015, 72, 25–36. [Google Scholar] [CrossRef]
- Annous, B.A.; Becker, L.A.; Bayles, D.O.; Labeda, D.P.; Wilkinson, B.J. Critical role of anteiso-C15:0 fatty acid in the growth of Listeria monocytogenes at low temperatures. Appl. Environ. Microbiol. 1997, 63, 3887–3894. [Google Scholar] [CrossRef]
- Choi, K.H.; Heath, R.J.; Rock, C.O. β-ketoacyl-acyl carrier protein synthase III (FabH) is a determining factor in branched-chain fatty acid biosynthesis. J. Bacteriol. 2000, 182, 365–370. [Google Scholar] [CrossRef]
- Nelson, K.E.; Fouts, D.E.; Mongodin, E.F.; Ravel, J.; DeBoy, R.T.; Kolonay, J.F.; Rasko, D.A.; Angiuoli, S.V.; Gill, S.R.; Paulsen, I.T.; et al. Whole genome comparisons of serotype 4b and 1/2a strains of the food-borne pathogen Listeria monocytogenes reveal new insights into the core genome components of this species. Nucleic Acids Res. 2004, 32, 2386–2395. [Google Scholar] [CrossRef] [PubMed]
- Schmid, B.; Klumpp, J.; Raimann, E.; Loessner, M.J.; Stephan, R.; Tasara, T. Role of cold shock proteins in growth of Listeria monocytogenes under cold and osmotic stress conditions. Appl. Environ. Microbiol. 2009, 75, 1621–1627. [Google Scholar] [CrossRef] [PubMed]
- Wemekamp-Kamphuis, H.H.; Karatzas, A.K.; Wouters, J.A.; Abee, T. Enhanced levels of cold shock proteins in Listeria monocytogenes LO28 upon exposure to low temperature and high hydrostatic pressure. Appl. Environ. Microbiol. 2002, 68, 456–463. [Google Scholar] [CrossRef] [PubMed]
- Netterling, S.; Vaitkevicius, K.; Nord, S.; Johansson, J. A Listeria monocytogenes RNA helicase essential for growth and ribosomal maturation at low temperatures uses its C terminus for appropriate interaction with the ribosome. J. Bacteriol. 2012, 194, 4377–4385. [Google Scholar] [CrossRef]
- Markkula, A.; Lindstrom, M.; Johansson, P.; Bjorkroth, J.; Korkeala, H. Roles of four putative DEAD-box RNA helicase genes in growth of Listeria monocytogenes EGD-e under heat, pH, osmotic, ethanol, and oxidative stress conditions. Appl. Environ. Microbiol. 2012, 78, 6875–6882. [Google Scholar] [CrossRef] [PubMed]
- Bäreclev, C.; Vaitkevicius, K.; Netterling, S.; Johansson, J. DExD-box RNA-helicases in Listeria monocytogenes are important for growth, ribosomal maturation, rRNA processing and virulence factor expression. RNA Biol. 2014, 11, 1457–1466. [Google Scholar] [CrossRef] [PubMed]
- Mendum, M.L.; Smith, L.T. Characterization of glycine betaine porter I from Listeria monocytogenes and its roles in salt and chill tolerance. Appl. Environ. Microbiol. 2002, 68, 813–819. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Graham, J.E.; Bigelow, L.; Morse, P.D.; Wilkinson, B.J. Identification of Listeria monocytogenes genes expressed in response to growth at low temperature. Appl. Environ. Microbiol. 2002, 68, 1697–1705. [Google Scholar] [CrossRef]
- Chan, Y.C.; Wiedmann, M. Physiology and genetics of Listeria monocytogenes survival and growth at cold temperatures. Crit. Rev. Food Sci. Nutr. 2008, 49, 237–253. [Google Scholar] [CrossRef]
- Zhao, X.; Drlica, K. Reactive oxygen species and the bacterial response to lethal stress. Curr. Opin. Microbiol. 2014, 21, 1–6. [Google Scholar] [CrossRef]
- Azizoglu, R.O.; Kathariou, S. Temperature-dependent requirement for catalase in aerobic growth of Listeria monocytogenes F2365. Appl. Environ. Microbiol. 2010, 76, 6998–7003. [Google Scholar] [CrossRef] [PubMed]
- Suo, Y.; Huang, Y.; Liu, Y.; Shi, C.; Shi, X. The expression of superoxide dismutase (SOD) and a putative ABC transporter permease is inversely correlated during biofilm formation in Listeria monocytogenes 4b G. PLoS ONE 2012, 7, e48467. [Google Scholar] [CrossRef] [PubMed]
- Mains, D.R.; Eallonardo, S.J.; Freitag, N.E. Identification of Listeria monocytogenes genes contributing to oxidative stress resistance under conditions relevant to host infection. Infect. Immun. 2021, 89, e00700-20. [Google Scholar] [CrossRef] [PubMed]
- Ruhland, B.R.; Reniere, M.L. Sense and sensor ability: Redox-responsive regulators in Listeria monocytogenes. Curr. Opin. Microbiol. 2019, 47, 20–25. [Google Scholar] [CrossRef] [PubMed]
- Rea, R.; Hill, C.; Gahan, C.G.M. Listeria monocytogenes PerR mutants display a small-colony phenotype, increased sensitivity to hydrogen peroxide, and significantly reduced murine virulence. Appl. Environ. Microbiol. 2005, 71, 8314–8322. [Google Scholar] [CrossRef] [PubMed]
- Sleator, R.D.; Watson, D.; Hill, C.; Gahan, C.G.M. The interaction between Listeria monocytogenes and the host gastrointestinal tract. Microbiology 2009, 155, 2463–2475. [Google Scholar] [CrossRef]
- Conte, M.P.; Petrone, G.; Di Biase, A.M.; Ammendolia, M.G.; Superti, F.; Seganti, L. Acid tolerance in Listeria monocytogenes influences invasiveness of enterocyte-like cells and macrophage-like cells. Microb. Pathog. 2000, 29, 137–144. [Google Scholar] [CrossRef]
- Saklani-Jusforgues, H.; Fontan, E.; Goossens, P.L. Effect of acid-adaptation on Listeria monocytogenes survival and translocation in a murine intragastric infection model. FEMS Microbiol. Lett. 2000, 193, 155–159. [Google Scholar] [CrossRef]
- O’Byrne, C.P.; Karatzas, K.A.G. The role of Sigma B (σB) in the stress adaptations of Listeria monocytogenes: Overlaps between stress adaptation and virulence. In Advances in Applied Microbiology; Elsevier Masson SAS: Amsterdam, The Netherlands, 2008; Volume 65, pp. 115–140. [Google Scholar]
- Begley, M.; Sleator, R.D.; Gahan, C.G.; Hill, C. Contribution of three bile-associated loci, bsh, pva, and btlB, to gastrointestinal persistence and bile tolerance of Listeria monocytogenes. Infect. Immun. 2005, 73, 894–904. [Google Scholar] [CrossRef]
- Sleator, R.D.; Wemekamp-Kamphuis, H.H.; Gahan, C.G.; Abee, T.; Hill, C. A PrfA-regulated bile exclusion system (BilE) is a novel virulence factor in Listeria monocytogenes. Mol. Microbiol. 2005, 55, 1183–1195. [Google Scholar] [CrossRef] [PubMed]
- Wiedmann, M.; Arvik, T.J.; Hurley, R.J.; Boor, K.J. General stress transcription factor σB and its role in acid tolerance and virulence of Listeria monocytogenes. J. Bacteriol. 1998, 180, 3650–3656. [Google Scholar] [CrossRef] [PubMed]
- Sue, D.; Fink, D.; Wiedmann, M.; Boor, K.J. σB-dependent gene induction and expression in Listeria monocytogenes during osmotic and acid stress conditions simulating the intestinal environment. Microbiology 2004, 150, 3843–3855. [Google Scholar] [CrossRef] [PubMed]
- Cetin, M.S.; Zhang, C.; Hutkins, R.W.; Benson, A.K. Regulation of transcription of compatible solute transporters by the general stress sigma factor, σB, in Listeria monocytogenes. J. Bacteriol. 2004, 186, 794–802. [Google Scholar] [CrossRef]
- Wemekamp-Kamphuis, H.H.; Wouters, J.A.; de Leeuw, P.P.; Hain, T.; Chakraborty, T.; Abee, T. Identification of sigma factor σB-controlled genes and their impact on acid stress, high hydrostatic pressure, and freeze survival in Listeria monocytogenes EGD-e. Appl. Environ. Microbiol. 2004, 70, 3457–3466. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Choi, C.W.; Lee, T.; Kim, S.I.; Lee, J.C.; Shin, J.H. Transcription factor σB plays an important role in the production of extracellular membrane-derived vesicles in Listeria monocytogenes. PLoS ONE 2013, 8, e73196. [Google Scholar] [CrossRef] [PubMed]
- Mujahid, S.; Orsi, R.H.; Vangay, P.; Boor, K.J.; Wiedmann, M. Refinement of the Listeria monocytogenes σB regulon through quantitative proteomic analysis. Microbiology 2013, 159, 1109–1119. [Google Scholar] [CrossRef][Green Version]
- Abram, F.; Starr, E.; Karatzas, K.A.G.; Matlawska-Wasowska, K.; Boyd, A.; Wiedmann, M.; Boor, K.J.; Connally, D.; O'Byrne, C.P. Identification of components of the sigma B regulon in Listeria monocytogenes that contribute to acid and salt tolerance. Appl. Environ. Microbiol. 2008, 74, 6848–6858. [Google Scholar] [CrossRef] [PubMed]
- Chaturongakul, S.; Boor, K.J. σB activation under environmental and energy stress conditions in Listeria monocytogenes. Appl. Environ. Microbiol. 2006, 72, 5197–5203. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, A.; O’Byrne, C.P.; Boor, K.J. Role of σB in heat, ethanol, acid, and oxidative stress resistance and during carbon starvation in Listeria monocytogenes. Appl. Environ. Microbiol. 2001, 67, 4454–4457. [Google Scholar] [CrossRef]
- Garner, M.R.; Njaa, B.L.; Wiedmann, M.; Boor, K.J. Sigma B contributes to Listeria monocytogenes gastrointestinal infection but not to systemic spread in the guinea pig infection model. Infect. Immun. 2006, 74, 876–886. [Google Scholar] [CrossRef]
- Guldimann, C.; Boor, K.J.; Wiedmann, M.; Guariglia-Oropeza, V. Resilience in the face of uncertainty: Sigma factor B fine-tunes gene expression to support homeostasis in gram-positive bacteria. Appl. Environ. Microbiol. 2016, 82, 4456–4469. [Google Scholar] [CrossRef] [PubMed]
- Dessaux, C.; Guerreiro, D.N.; Pucciarelli, M.G.; O’Byrne, C.P.; García-del Portillo, F. Impact of osmotic stress on the phosphorylation and subcellular location of Listeria monocytogenes stressosome proteins. Sci. Rep. 2020, 10, 20837. [Google Scholar] [CrossRef]
- Chaturongakul, S.; Boor, K.J. RsbT and RsbV contribute to σB-dependent survival under environmental, energy, and intracellular stress conditions in Listeria monocytogenes. Appl. Environ. Microbiol. 2004, 70, 5349–5356. [Google Scholar] [CrossRef]
- Freitag, N.E.; Port, G.C.; Miner, M.D. Listeria monocytogenes: From saprophyte to intracellular pathogen. Nat. Rev. Microbiol. 2009, 7, 623. [Google Scholar] [CrossRef]
- Freitag, N.E.; Rong, L.; Portnoy, D.A. Regulation of the prfA transcriptional activator of Listeria monocytogenes: Multiple promoter elements contribute to intracellular growth and cell-to-cell spread. Infect. Immun. 1993, 61, 2537–2544. [Google Scholar] [CrossRef] [PubMed]
- Gray, M.J.; Freitag, N.E.; Boor, K.J. How the bacterial pathogen Listeria monocytogenes mediates the switch from environmental Dr. Jekyll to pathogenic Mr. Hyde. Infect. Immun. 2006, 74, 2505–2512. [Google Scholar] [CrossRef]
- Miner, M.D.; Port, G.C.; Freitag, N.E. Regulation of Listeria monocytogenes virulence genes. In Listeria monocytogenes: Pathogenesis and Host Response; Goldfine, H., Shen, H., Eds.; Springer Science+Business Media, LLC: Berlin, Germany, 2007; pp. 139–158. [Google Scholar]
- Bruno, J.C.; Freitag, N.E. Constitutive activation of PrfA tilts the balance of Listeria monocytogenes fitness towards life within the host versus environmental survival. PLoS ONE 2010, 5, e15138. [Google Scholar] [CrossRef] [PubMed]
- Biswas, R.; Sonenshein, A.L.; Belitsky, B.R. Genome-wide identification of Listeria monocytogenes CodY-binding sites. Mol. Microbiol. 2020, 113, 841–858. [Google Scholar] [CrossRef]
- Lobel, L.; Sigal, N.; Borovok, I.; Belitsky, B.R.; Sonenshein, A.L.; Herskovits, A.A. The metabolic regulator CodY links Listeria monocytogenes metabolism to virulence by directly activating the virulence regulatory gene prfA. Mol. Microbiol. 2015, 95, 624–644. [Google Scholar] [CrossRef] [PubMed]
- Lobel, L.; Herskovits, A.A. Systems level analyses reveal multiple regulatory activities of CodY controlling metabolism, motility and virulence in Listeria monocytogenes. PLoS Genet. 2016, 12, e1005870. [Google Scholar] [CrossRef]
- Lobel, L.; Sigal, N.; Borovok, I.; Ruppin, E.; Herskovits, A. Integrative genomic analysis identifies isoleucine and CodY as regulators of Listeria monocytogenes virulence. PLoS Genet. 2012, 8, e1002887. [Google Scholar] [CrossRef]
- Chico-Calero, I.; Suárez, M.; González-Zorn, B.; Scortti, M.; Slaghuis, J.; Goebel, W.; Vázquez-Boland, J.A.; European Listeria Genome Consortium. Hpt, a bacterial homolog of the microsomal glucose-6-phosphate translocase, mediates rapid intracellular proliferation in Listeria. Proc. Natl. Acad. Sci. USA 2002, 99, 431–436. [Google Scholar] [CrossRef] [PubMed]
- Vega, Y.; Rauch, M.; Banfield, M.J.; Ermolaeva, S.; Scortti, M.; Goebel, W.; Vázquez-Boland, J.A. New Listeria monocytogenes prfA* mutants, transcriptional properties of PrfA* proteins and structure-function of the virulence regulator PrfA. Mol. Microbiol. 2004, 52, 1553–1565. [Google Scholar] [CrossRef] [PubMed]
- Xayarath, B.; Smart, J.I.; Mueller, K.J.; Freitag, N.E. A novel C-terminal mutation resulting in constitutive activation of the Listeria monocytogenes central virulence regulatory factor PrfA. Microbiology 2011, 157, 3138–3149. [Google Scholar] [CrossRef]
- Oropeza, V.G.; Orsi, R.H.; Guldimann, C.; Wiedmann, M.; Boor, K.J. The Listeria monocytogenes bile stimulon under acidic conditions is characterized by strain-specific patterns and the upregulation of motility, cell wall modification functions, and the PrfA regulon. Front. Microbiol. 2018, 9, 120. [Google Scholar] [CrossRef]
- Eshwar, A.K.; Guldimann, C.; Oevermann, A.; Tasara, T. Cold-shock domain family proteins (Csps) are involved in regulation of virulence, cellular aggregation, and flagella-based motility in Listeria monocytogenes. Front. Cell. Infect. Microbiol. 2017, 7, 453. [Google Scholar] [CrossRef]
- Wałecka, E.; Molenda, J.; Karpíšková, R.; Bania, J. Effect of osmotic stress and culture density on invasiveness of Listeria monocytogenes strains. Int. J. Food. Microbiol. 2011, 144, 440–445. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Marquis, H.; Boor, K.J. σB contributes to Listeria monocytogenes invasion by controlling expression of inlA and inlB. Microbiology 2005, 151, 3215–3222. [Google Scholar] [CrossRef] [PubMed]
- Dussurget, O.; Cabanes, D.; Dehoux, P.; Lecuit, M.; Buchrieser, C.; Glaser, P.; Cossart, P. European Listeria Genome Consortium Listeria monocytogenes bile salt hydrolase is a PrfA-regulated virulence factor involved in the intestinal and hepatic phases of listeriosis. Mol. Microbiol. 2002, 45, 1095–1106. [Google Scholar] [PubMed]
- Tiensuu, T.; Andersson, C.; Rydén, P.; Johansson, J. Cycles of light and dark co-ordinate reversible colony differentiation in Listeria monocytogenes. Mol. Microbiol. 2013, 87, 909–924. [Google Scholar] [CrossRef] [PubMed]
- Shetron-Rama, L.M.; Marquis, H.; Bouwer, H.G.A.; Freitag, N.E. Intracellular induction of Listeria monocytogenes actA expression. Infect. Immun. 2002, 70, 1087–1096. [Google Scholar] [CrossRef] [PubMed]
- Ollinger, J.; Bowen, B.; Wiedmann, M.; Boor, K.J.; Bergholz, T.M. Listeria monocytogenes σB modulates PrfA-mediated virulence factor expression. Infect. Immun. 2009, 77, 2113–2124. [Google Scholar] [CrossRef] [PubMed]
- Rauch, M.; Luo, Q.; Müller-Altrock, S.; Goebel, W. SigB-dependent in vitro transcription of prfA and some newly identified genes of Listeria monocytogenes whose expression is affected by PrfA in vivo. J. Bacteriol. 2005, 187, 800–804. [Google Scholar] [CrossRef]
- Lam, O.; Wheeler, J.; Tang, C.M. Thermal control of virulence factors in bacteria: A hot topic. Virulence 2014, 5, 852–862. [Google Scholar] [CrossRef] [PubMed]
- Loh, E.; Righetti, F.; Eichner, H.; Twittenhoff, C.; Narberhaus, F. RNA thermometers in bacterial pathogens. Microbiol. Spectr. 2018, 6. [Google Scholar] [CrossRef] [PubMed]
- Loh, E.; Memarpour, F.; Vaitkevicius, K.; Kallipolitis, B.H.; Johansson, J.; Sondén, B. An unstructured 5′-coding region of the prfA mRNA is required for efficient translation. Nucleic Acids Res. 2012, 40, 1818–1827. [Google Scholar] [CrossRef] [PubMed]
- Dorey, A.; Marinho, C.; Piveteau, P.; O’byrne, C. Role and regulation of the stress activated sigma factor sigma B (σB) in the saprophytic and host-associated life stages of Listeria monocytogenes. In Advances in Applied Microbiology, 1st ed.; Gadd, G.M., Sariaslani, S., Eds.; Elsevier Inc.: Amsterdam, The Netherlands, 2019; pp. 1–48. [Google Scholar]
- Bennett, H.J.; Pearce, D.M.; Glenn, S.; Taylor, C.M.; Kuhn, M.; Sonenshein, A.L.; Andrew, P.W.; Roberts, I.S. Characterization of relA and codY mutants of Listeria monocytogenes: Identification of the CodY regulon and its role in virulence. Mol. Microbiol. 2007, 63, 1453–1467. [Google Scholar] [CrossRef] [PubMed]
- Reniere, M.; Whiteley, A.; Hamilton, K.L.; John, S.M.; Lauer, P.; Brennan, R.; Portnoy, D.A. Glutathione activates virulence gene expression of an intracellular pathogen. Nature 2015, 517, 170–173. [Google Scholar] [CrossRef]
- Hall, M.; Grundström, C.; Begum, A.; Lindberg, M.J.; Sauer, U.H.; Almqvist, F.; Johansson, J.; Sauer-Eriksson, A.E. Structural basis for glutathione-mediated activation of the virulence regulatory protein PrfA in Listeria. Proc. Natl. Acad. Sci. USA 2016, 113, 14733–14738. [Google Scholar] [CrossRef] [PubMed]
- Gopal, S.; Borovok, I.; Ofer, A.; Yanku, M.; Cohen, G.; Goebel, W.; Kreft, J.; Aharonowitz, Y. A multidomain fusion protein in Listeria monocytogenes catalyzes the two primary activities for glutathione biosynthesis. J. Bacteriol. 2005, 187, 3839–3847. [Google Scholar] [CrossRef]
- Schärer, K.; Stephan, R.; Tasara, T. Cold shock proteins contribute to the regulation of listeriolysin O production in Listeria monocytogenes. Foodborne Pathog. Dis. 2013, 10, 1023–1029. [Google Scholar] [CrossRef]
- Cheng, Y.; Siletzky, R.M.; Kathariou, S. Genomic Divisions/Lineages, Epidemic Clones, and Population Structure. In Handbook of Listeria monocytogenes; Liu, D., Ed.; CRC Press: Boca Raton, FL, USA, 2008; pp. 337–358. [Google Scholar]
- Orsi, R.H.; den Bakker, H.C.; Wiedmann, M. Listeria monocytogenes lineages: Genomics, evolution, ecology, and phenotypic characteristics. Int. J. Med. Microbiol. 2011, 301, 79–96. [Google Scholar] [CrossRef] [PubMed]
- Moura, A.; Criscuolo, A.; Pouseele, H.; Maury, M.M.; Leclercq, A.; Tarr, C.; Björkman, J.T.; Dallman, T.; Reimer, A.; Enouf, V.; et al. Whole genome-based population biology and epidemiological surveillance of Listeria monocytogenes. Nat. Microbiol. 2016, 2, 16185. [Google Scholar] [CrossRef] [PubMed]
- Haase, J.K.; Didelot, X.; Lecuit, M.; Korkeala, H. L. monocytogenes MSLT Study Group; Achtman, M. The ubiquitous nature of Listeria monocytogenes clones: A large-scale Multilocus Sequence Typing study. Environ. Microbiol. 2014, 16, 405–416. [Google Scholar] [CrossRef]
- Doijad, S.; Weigel, M.; Barbuddhe, S.; Blom, J.; Goesmann, A.; Hain, T.; Chakraborty, T. Phylogenomic grouping of Listeria monocytogenes. Can. J. Microbiol. 2015, 61, 637–646. [Google Scholar] [CrossRef]
- Schiavano, G.F.; Ateba, C.N.; Petruzzelli, A.; Mele, V.; Amagliani, G.; Guidi, F.; De Santi, M.; Pomilio, F.; Blasi, G.; Gattuso, A.; et al. Whole-genome sequencing characterization of virulence profiles of Listeria monocytogenes food and human isolates and in vitro adhesion/invasion assessment. Microorganisms 2022, 10, 62. [Google Scholar] [CrossRef] [PubMed]
- Deng, X.; Phillippy, A.M.; Li, Z.; Salzberg, S.L.; Zhang, W. Probing the pan-genome of Listeria monocytogenes: New insights into intraspecific niche expansion and genomic diversification. BMC Genom. 2010, 11, 500. [Google Scholar] [CrossRef]
- Maury, M.M.; Tsai, Y.H.; Charlier, C.; Touchon, M.; Chenal-Francisque, V.; Leclercq, A.; Criscuolo, A.; Gaultier, C.; Roussel, S.; Brisabois, A.; et al. Uncovering Listeria monocytogenes hypervirulence by harnessing its biodiversity. Nat Genet. 2016, 48, 308. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Boland, J.A.; Wagner, M.; Scortti, M. Why are some Listeria monocytogenes genotypes more likely to cause invasive (brain, placental) infection? mBio 2020, 11, e03126-20. [Google Scholar] [CrossRef]
- Lee, S.; Chen, Y.; Gorski, L.; Ward, T.J.; Osborne, J.; Kathariou, S. Listeria monocytogenes source distribution analysis indicates regional heterogeneity and ecological niche preference among serotype 4b clones. mBio 2018, 9, e00396-18. [Google Scholar] [CrossRef]
- da Silva, M.F.; Ferreira, V.; Magalhães, R.; Almeida, G.; Alves, A.; Teixeira, P. Detection of premature stop codons leading to truncated internalin A among food and clinical strains of Listeria monocytogenes. Food Microbiol. 2017, 63, 6–11. [Google Scholar] [CrossRef]
- Nightingale, K.K.; Ivy, R.A.; Ho, A.J.; Fortes, E.D.; Njaa, B.L.; Peters, R.M.; Wiedmann, M. inlA premature stop codons are common among Listeria monocytogenes isolates from foods and yield virulence-attenuated strains that confer protection against fully virulent strains. Appl. Environ. Microbiol. 2008, 74, 6570–6583. [Google Scholar] [CrossRef] [PubMed]
- Van Stelten, A.; Simpson, J.M.; Ward, T.J.; Nightingale, K.K. Revelation by single-nucleotide polymorphism genotyping that mutations leading to a premature stop codon in inlA are common among Listeria monocytogenes isolates from ready-to-eat foods but not human listeriosis cases. Appl. Environ. Microbiol. 2010, 76, 2783–2790. [Google Scholar] [CrossRef] [PubMed]
- Cotter, P.D.; Draper, L.A.; Lawton, E.M.; Daly, K.M.; Groeger, D.S.; Casey, P.G.; Ross, R.P.; Hill, C. Listeriolysin S, a novel peptide haemolysin associated with a subset of lineage I Listeria monocytogenes. PLoS Pathog. 2008, 4, e1000144. [Google Scholar] [CrossRef]
- Quereda, J.J.; Meza-Torres, J.; Cossart, P.; Pizarro-Cerdá, J. Listeriolysin S: A bacteriocin from epidemic Listeria monocytogenes strains that targets the gut microbiota. Gut Microbes 2017, 8, 384–391. [Google Scholar] [CrossRef] [PubMed]
- Bergholz, T.M.; den Bakker, H.C.; Fortes, E.D.; Boor, K.J.; Wiedmann, M. Salt stress phenotypes in Listeria monocytogenes vary by genetic lineage and temperature. Foodborne Pathog. Dis. 2010, 7, 1537–1549. [Google Scholar] [CrossRef] [PubMed]
- Horlbog, J.A.; Kent, D.; Stephan, R.; Guldimann, C. Surviving host- and food relevant stresses: Phenotype of L. monocytogenes strains isolated from food and clinical sources. Nat. Sci. Rep. 2018, 8, 12931. [Google Scholar] [CrossRef]
- Pirone-Davies, C.; Chen, Y.; Pightling, A.; Ryan, G.; Wang, Y.; Yao, K.; Hoffmann, M.; Allard, M.W. Genes significantly associated with lineage II food isolates of Listeria monocytogenes. BMC Genom. 2018, 19, 708. [Google Scholar] [CrossRef]
- Naditz, A.L.; Dzieciol, M.; Wagner, M.; Schmitz-Esser, S. Plasmids contribute to food processing environment–associated stress survival in three Listeria monocytogenes ST121, ST8, and ST5 strains. Int. J. Food Microbiol. 2019, 299, 39–46. [Google Scholar] [CrossRef]
- Korsak, D.; Chmielowska, C.; Szuplewska, M.; Bartosik, D. Prevalence of plasmid-borne benzalkonium chloride resistance cassette bcrABC and cadmium resistance cadA genes in nonpathogenic Listeria spp. isolated from food and food-processing environments. Int. J. Food Microbiol. 2019, 290, 247–253. [Google Scholar] [CrossRef] [PubMed]
- Gelbicova, T.; Florianova, M.; Hluchanova, L.; Kalova, A.; Korena, K.; Strakova, N.; Karpiskova, R. Comparative analysis of genetic determinants encoding cadmium, arsenic, and benzalkonium chloride resistance in Listeria monocytogenes of human, food, and environmental origin. Front. Microbiol. 2021, 11, 599882. [Google Scholar] [CrossRef]
- Cerutti, F.; Mallet, L.; Painset, A.; Hoede, C.; Moisan, A.; Bécavin, C.; Duval, M.; Dussurget, O.; Cossart, P.; Gaspin, C.; et al. Unraveling the evolution and coevolution of small regulatory RNAs and coding genes in Listeria. BMC Genom. 2017, 18, 882. [Google Scholar] [CrossRef]

| Stage of Infection Cycle | Stress Response Proteins/Virulence Factor | Function | Transcriptional Regulator | References |
|---|---|---|---|---|
| Intestinal phase | GadD2T2 | Glutamate decarboxylase for survival of stomach acidity | SigB | [104] |
| OpuC | Carnitine transporter for osmotic stress survival | SigB | [116] | |
| Bsh | Bile salt hydrolase for bile stress survival | SigB and PrfA | [157,184] | |
| BilE | Bile exclusion system for bile stress survival | SigB and PrfA | [158] | |
| InlA | Adhesion and invasion of enterocytes | SigB and PrfA | [39,168] | |
| InlB | Adhesion and invasion of enterocytes | SigB and PrfA | [39] | |
| ActA | Bacterial aggregation and intestinal colonization | SigB and PrfA | [26,189] | |
| Intracellular phase | LLO | Primary vacuole lysis | PrfA | [46] |
| PI-PLC | Primary vacuole lysis | PrfA | [53] | |
| PC-PLC | Secondary vacuole lysis | PrfA | [54] | |
| ActA | Intracellular motility and cell-to-cell spread | PrfA | [56,190] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sibanda, T.; Buys, E.M. Listeria monocytogenes Pathogenesis: The Role of Stress Adaptation. Microorganisms 2022, 10, 1522. https://doi.org/10.3390/microorganisms10081522
Sibanda T, Buys EM. Listeria monocytogenes Pathogenesis: The Role of Stress Adaptation. Microorganisms. 2022; 10(8):1522. https://doi.org/10.3390/microorganisms10081522
Chicago/Turabian StyleSibanda, Thulani, and Elna M. Buys. 2022. "Listeria monocytogenes Pathogenesis: The Role of Stress Adaptation" Microorganisms 10, no. 8: 1522. https://doi.org/10.3390/microorganisms10081522
APA StyleSibanda, T., & Buys, E. M. (2022). Listeria monocytogenes Pathogenesis: The Role of Stress Adaptation. Microorganisms, 10(8), 1522. https://doi.org/10.3390/microorganisms10081522






