Isolation, Characterization, and Genomic Analysis of Three Novel E. coli Bacteriophages That Effectively Infect E. coli O18
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Growth Conditions
2.2. Antimicrobial Susceptibility Testing
2.3. 16S Ribotyping
2.4. Phage Selection, Isolation, Purification, and Amplification
2.5. Host Range Determination
2.6. The Influence of Temperature and pH on Bacteriophages Stability
2.7. Time Killing Curve of the Bacteriophages
2.8. Examination of Phage Morphology by Transmission Electron Microscopy (TEM)
2.9. Phage DNA Sequencing and Bioinformatic Analysis
2.10. Accession Numbers
Statistical Analysis
3. Results
3.1. Antimicrobial Susceptibility Testing
3.2. Isolation of Bacteriophages
3.3. Host Range Determination
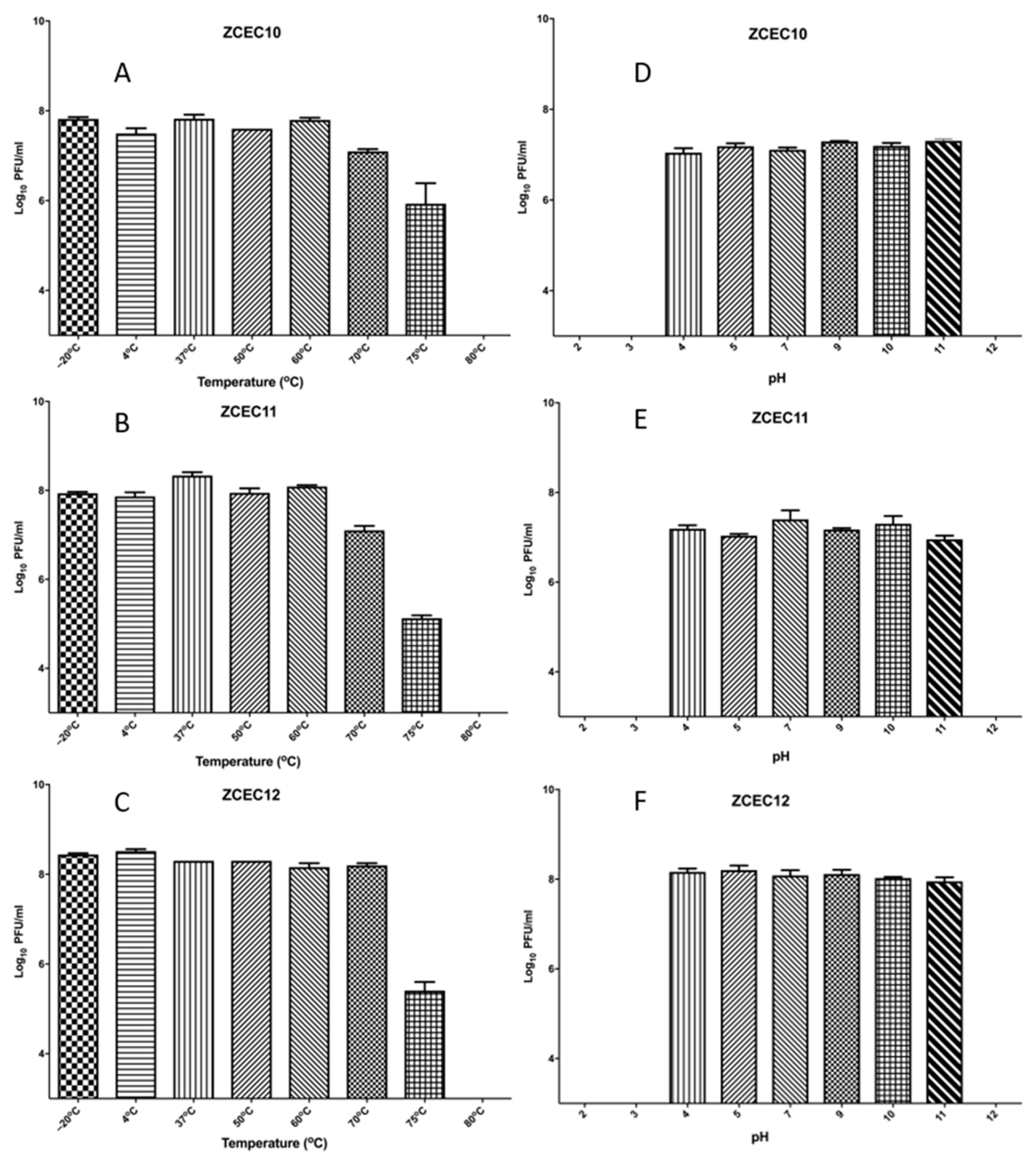
3.4. Sensitivity of Virions to pH and Temperature
3.5. Time Killing Curves
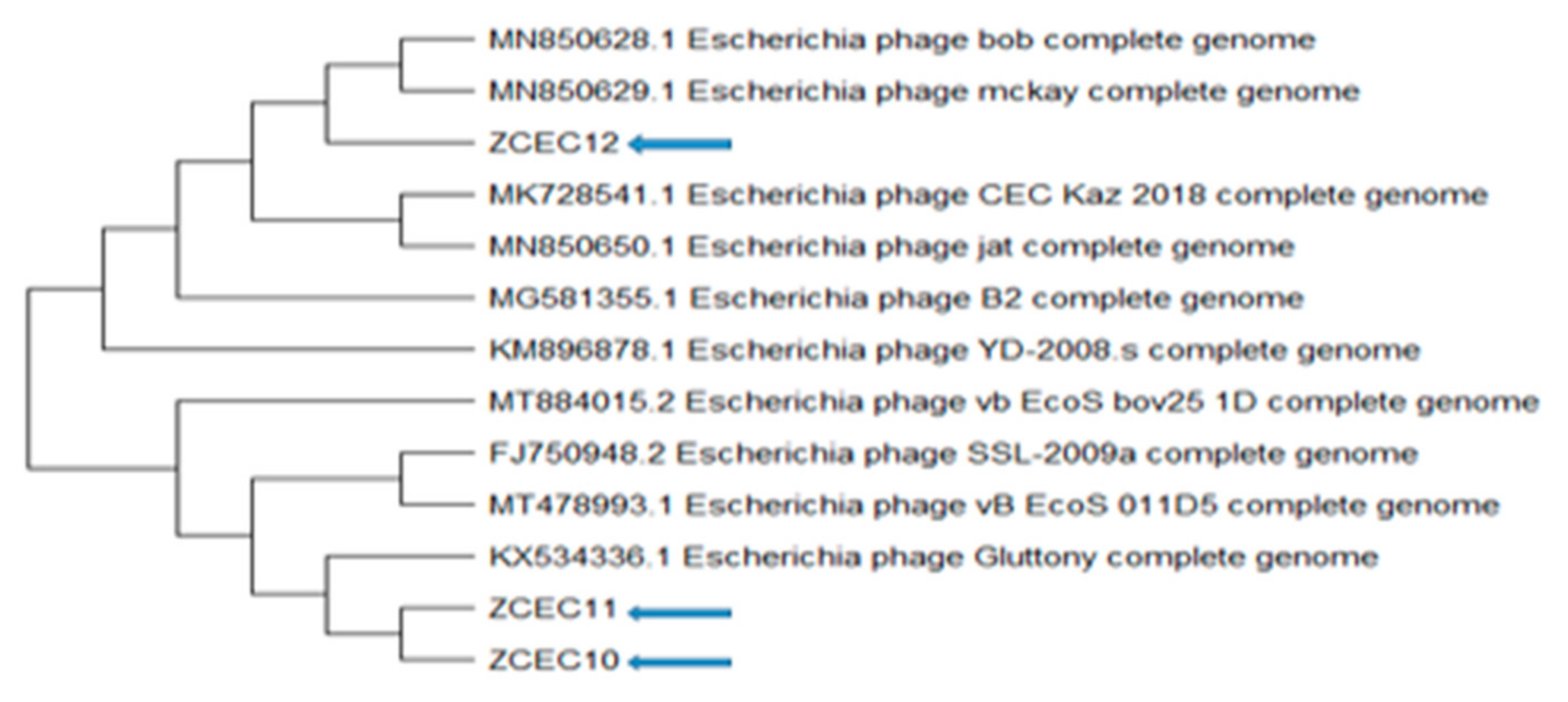
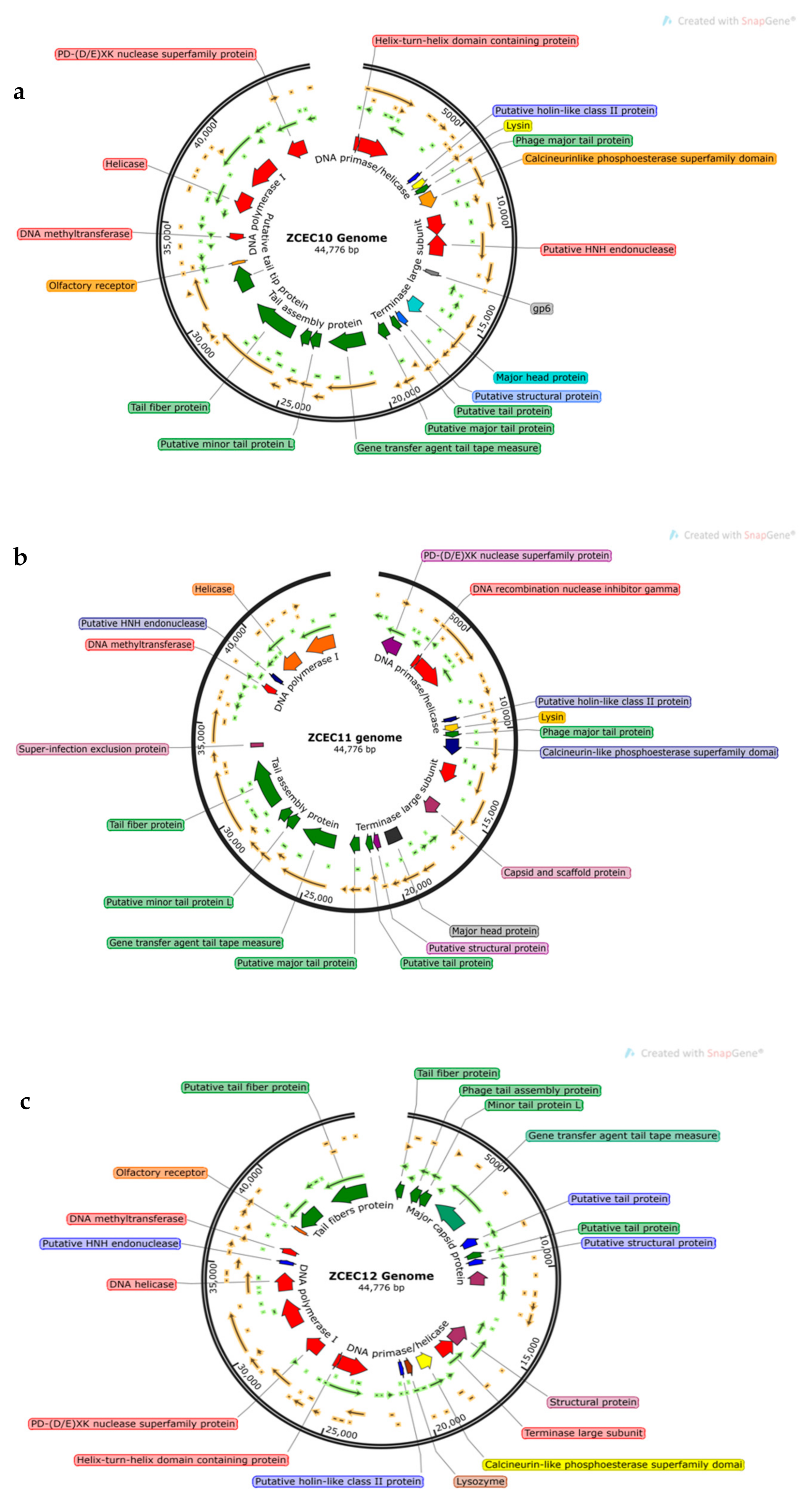
3.6. Bioinformatics Analysis and Characterization of Bacteriophages Genome
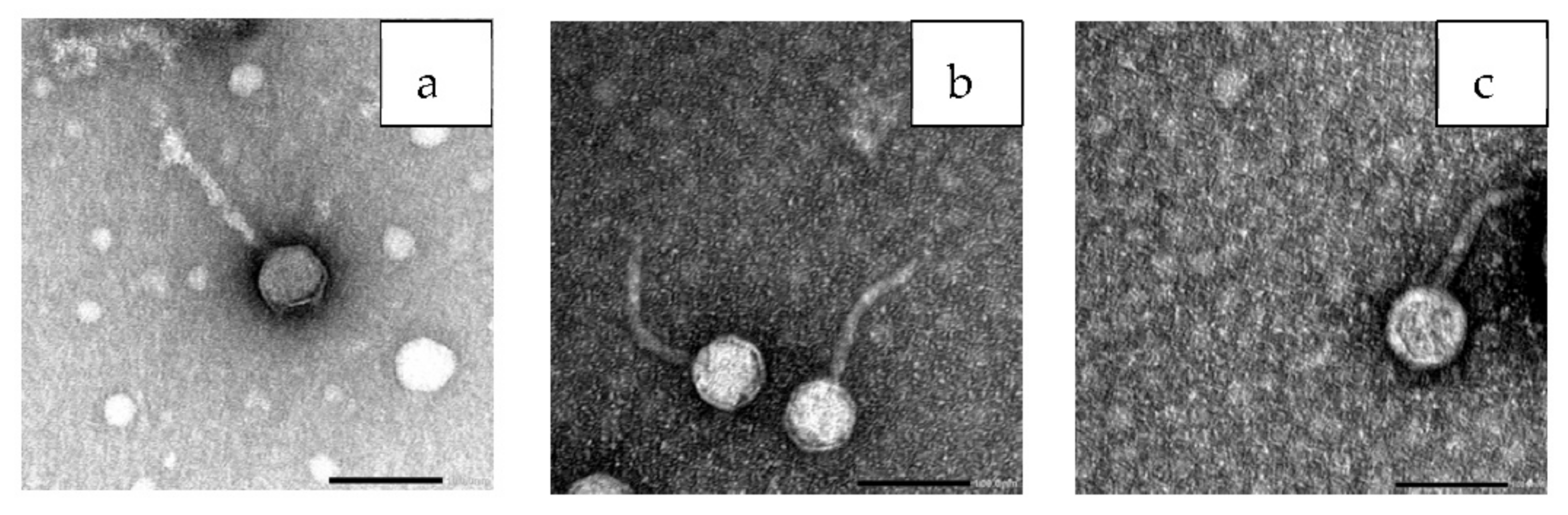
3.7. Bacteriophage Morphology
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ballou, A.L.; Ali, R.A.; Mendoza, M.A.; Ellis, J.C.; Hassan, H.M.; Croom, W.J.; Koci, M.D. Development of the Chick Microbiome: How Early Exposure Influences Future Microbial Diversity. Front. Vet. Sci. 2016, 3, 2. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sarowska, J.; Futoma-Koloch, B.; Jama-Kmiecik, A.; Frej-Madrzak, M.; Ksiazczyk, M.; Bugla-PLoSkonska, G.; Choroszy-Krol, I. Virulence factors, prevalence and potential transmission of extraintestinal pathogenic Escherichia coli isolated from different sources: Recent reports. Gut Pathog. 2019, 11, 10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kabir, S.M.L. Avian Colibacillosis and Salmonellosis: A Closer Look at Epidemiology, Pathogenesis, Diagnosis, Control and Public Health Concerns. Int. J. Environ. Res. Public. Health 2010, 7, 89–114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dho-Moulin, M.; Fairbrother, J.M. Avian pathogenic Escherichia coli (APEC). Vet. Res. 1999, 30, 299–316. [Google Scholar]
- Mellata, M. Human and Avian Extraintestinal Pathogenic Escherichia coli: Infections, Zoonotic Risks, and Antibiotic Resistance Trends. Foodborne Pathog. Dis. 2013, 10, 916–932. [Google Scholar] [CrossRef] [Green Version]
- Johnson, J.R.; Sannes, M.R.; Croy, C.; Johnston, B.; Clabots, C.; Kuskowski, M.A.; Bender, J.; Smith, K.E.; Winokur, P.L.; Belongia, E.A. Antimicrobial drug-resistant Escherichia coli from humans and poultry products, Minnesota and Wisconsin, 2002–2004. Emerg. Infect. Dis. 2007, 13, 838–846. [Google Scholar] [CrossRef]
- Jakobsen, L.; Spangholm, D.J.; Pedersen, K.; Jensen, L.B.; Emborg, H.D.; Agersø, Y.; Aarestrup, F.M.; Hammerum, A.M.; Frimodt-Møller, N. Broiler chickens, broiler chicken meat, pigs and pork as sources of ExPEC related virulence genes and resistance in Escherichia coli isolates from community-dwelling humans and UTI patients. Int. J. Food Microbiol. 2010, 142, 264–272. [Google Scholar] [CrossRef]
- Halfaoui, Z.; Menoueri, N.M.; Bendali, L.M. Serogrouping and antibiotic resistance of Escherichia coli isolated from broiler chicken with colibacillosis in center of Algeria. Vet. World 2017, 10, 830–835. [Google Scholar] [CrossRef] [Green Version]
- Alvi, I.A.; Asif, M.; Tabassum, R.; Aslam, R.; Abbas, Z.; Rehman, S.U. RLP, a bacteriophage of the family Podoviridae, rescues mice from bacteremia caused by multi-drug-resistant Pseudomonas aeruginosa. Arch. Virol. 2020, 165, 1289–1297. [Google Scholar] [CrossRef]
- Batinovic, S.; Wassef, F.; Knowler, S.A.; Rice, D.T.F.; Stanton, C.R.; Rose, J.; Tucci, J.; Nittami, T.; Vinh, A.; Drummond, G.R.; et al. Bacteriophages in natural and artificial environments. Pathogens 2019, 8, 100. [Google Scholar] [CrossRef] [Green Version]
- El-Shibiny, A.; Dawoud, A. Bacteriophage Applications for Food Safety. In Biocommunication of Phages; Witzany, G., Ed.; Springer: Cham, Switzerland, 2020; pp. 463–484. ISBN 978-3-030-45884-3. [Google Scholar] [CrossRef]
- Mu, A.; McDonald, D.; Jarmusch, A.K.; Martino, C.; Brennan, C.; Bryant, M.; Humphrey, G.C.; Toronczak, J.; Schwartz, T.; Nguyen, D.; et al. Assessment of the microbiome during bacteriophage therapy in combination with systemic antibiotics to treat a case of staphylococcal device infection. Microbiome 2021, 9, 92. [Google Scholar] [CrossRef]
- Abdelsattar, A.S.; Abdelrahman, F.; Dawoud, A.; Connerton, I.F.; El-Shibiny, A. Encapsulation of E. coli phage ZCEC5 in chitosan–alginate beads as a delivery system in phage therapy. AMB Express 2019, 9, 87. [Google Scholar] [CrossRef] [Green Version]
- Sulakvelidze, A.; Alavidze, Z.; Morris, J.G. Bacteriophage Therapy. Antimicrob. Agents Chemother. 2001, 45, 649–659. [Google Scholar] [CrossRef] [Green Version]
- Smith, H.W.; Huggins, M.B. Effectiveness of Phages in Treating Experimental Escherichia coli Diarrhoea in Calves, Piglets and Lambs. Microbiology 1983, 129, 2659–2675. [Google Scholar] [CrossRef] [Green Version]
- Huff, W.; Huff, G.; Rath, N.; Balog, J.; Donoghue, A. Prevention of Escherichia coli infection in broiler chickens with a bacteriophage aerosol spray. Poult. Sci. 2002, 81, 1486–1491. [Google Scholar] [CrossRef]
- Huff, W.E.; Huff, G.R.; Rath, N.C.; Donoghue, A.M. Evaluation of the Influence of Bacteriophage Titer on the Treatment of Colibacillosis in Broiler Chickens. Poult. Sci. 2006, 85, 1373–1377. [Google Scholar] [CrossRef]
- Higgins, J.P.; Higgins, S.E.; Guenther, K.L.; Huff, W.; Donoghue, A.M.; Donoghue, D.J.; Hargis, B.M. Use of a specific bacteriophage treatment to reduce Salmonella in poultry products. Poult. Sci. 2005, 84, 1141–1145. [Google Scholar] [CrossRef]
- O’Flynn, G.; Coffey, A.; Fitzgerald, G.F.; Ross, R.P. The newly isolated lytic bacteriophages st104a and st104b are highly virulent against Salmonella enterica. J. Appl. Microbiol. 2006, 101, 251–259. [Google Scholar] [CrossRef]
- Atterbury, R.J.; Van Bergen, M.A.P.; Ortiz, F.; Lovell, M.A.; Harris, J.A.; De Boer, A.; Wagenaar, J.A.; Allen, V.M.; Barrow, P.A. Bacteriophage Therapy to Reduce Salmonella Colonization of Broiler Chickens. Appl. Environ. Microbiol. 2007, 73, 4543–4549. [Google Scholar] [CrossRef] [Green Version]
- Loc Carrillo, C.; Atterbury, R.J.; El-Shibiny, A.; Connerton, P.L.; Dillon, E.; Scott, A.; Connerton, I.F. Bacteriophage Therapy to Reduce Campylobacter jejuni Colonization of Broiler Chickens. Appl. Environ. Microbiol. 2005, 71, 6554–6563. [Google Scholar] [CrossRef] [Green Version]
- Wagenaar, J.A.; Van Bergen, M.A.P.; Mueller, M.A.; Wassenaar, T.M.; Carlton, R.M. Phage therapy reduces Campylobacter jejuni colonization in broilers. Vet. Microbiol. 2005, 109, 275–283. [Google Scholar] [CrossRef]
- Hudzicki, J. Kirby-Bauer Disk Diffusion Susceptibility Test Protocol Author Information. Am. Soc. Microbiol. 2009, 15, 55–63. [Google Scholar]
- CLSI M100-ED29; 2019 Performance Standards for Antimicrobial Susceptibility Testing, 29th ed.; CLSI: Wayne, PA, USA, 2019; ISBN 9781684400324.
- Awad, A.M.; El-Shall, N.A.; Khalil, D.S.; El-Hack, M.E.A.; Swelum, A.A.; Mahmoud, A.H.; Ebaid, H.; Komany, A.; Sammour, R.H.; Sedeik, M.E. Incidence, Pathotyping, and Antibiotic Susceptibility of Avian Pathogenic Escherichia coli among Diseased Broiler Chicks. Pathogens 2020, 9, 114. [Google Scholar] [CrossRef] [Green Version]
- Kropinski, A.M.; Mazzocco, A.; Waddell, T.E.; Lingohr, E.; Johnson, R.P. Enumeration of Bacteriophages by Double Agar Overlay Plaque Assay. Methods Mol. Biol. 2009, 501, 69–76. [Google Scholar] [CrossRef]
- Mazzocco, A.; Waddell, T.E.; Lingohr, E.; Johnson, R.P. Enumeration of bacteriophages using the small drop plaque assay system. Methods Mol. Biol. 2009, 501, 81–85. [Google Scholar] [CrossRef]
- Cao, Z.; Zhang, J.; Niu, Y.D.; Cui, N.; Ma, Y.; Cao, F.; Jin, L.; Li, Z.; Xu, Y. Isolation and characterization of a “phiKMVlike” bacteriophage and its therapeutic effect on mink hemorrhagic pneumonia. PLoS ONE 2015, 10, e0116571. [Google Scholar] [CrossRef]
- Fayez, M.S.; Hakim, T.A.; Agwa, M.M.; Abdelmoteleb, M.; Aly, R.G.; Montaser, N.N.; Abdelsattar, A.S.; Rezk, N.; El-Shibiny, A. Topically Applied Bacteriophage to Control Multi-Drug Resistant Klebsiella pneumoniae Infected Wound in a Rat Model. Antibiotics 2021, 10, 1048. [Google Scholar] [CrossRef]
- Capra, M.L.; Quiberoni, A.; Reinheimer, J.A. Thermal and chemical resistance of Lactobacillus casei and Lactobacillus paracasei bacteriophages. Lett. Appl. Microbiol. 2004, 38, 499–504. [Google Scholar] [CrossRef]
- Hammerl, J.A.; Jäckel, C.; Alter, T.; Janzcyk, P.; Stingl, K.; Knüver, M.T.; Hertwig, S. Reduction of Campylobacter jejuni in Broiler Chicken by Successive Application of Group II and Group III Phages. PLoS ONE 2014, 9, e114785. [Google Scholar] [CrossRef]
- Kusradze, I.; Karumidze, N.; Rigvava, S.; Dvalidze, T.; Katsitadze, M.; Amiranashvili, I.; Goderdzishvili, M. Characterization and Testing the Efficiency of Acinetobacter baumannii Phage vB-GEC_Ab-M-G7 as an Antibacterial Agent. Front. Microbiol. 2016, 7, 1590. [Google Scholar] [CrossRef] [Green Version]
- Chen, L.; Yuan, S.; Liu, Q.; Mai, G.; Yang, J.; Deng, D. In Vitro Design and Evaluation of Phage Cocktails against Aeromonas salmonicida. Front. Microbiol. 2018, 9, 1476. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nale, J.Y.; Spencer, J.; Hargreaves, K.R.; Buckley, A.M.; Trzepiński, P.; Douce, G.R.; Clokie, M.R.J. Bacteriophage Combinations Significantly Reduce Clostridium difficile Growth In Vitro and Proliferation In Vivo. Antimicrob. Agents Chemother. 2016, 60, 968–981. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- LaMar, D. FastQC. 2015. Available online: https://qubeshub.org/resources/fastqc (accessed on 21 December 2021).
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gurevich, A.; Saveliev, V.; Vyahhi, N.; Tesler, G. QUAST: Quality assessment tool for genome assemblies. Bioinformatics 2013, 29, 1072–1075. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [Green Version]
- Furuno, M.; Kasukawa, T.; Saito, R.; Adachi, J.; Suzuki, H.; Baldarelli, R.; Hayashizaki, Y.; Okazaki, Y. CDS Annotation in Full-Length cDNA Sequence. Genome Res. 2003, 13, 1478–1487. [Google Scholar] [CrossRef] [Green Version]
- Scarafile, G. Antibiotic resistance: Current issues and future strategies. Rev. Health Care 2016, 7, 3. [Google Scholar] [CrossRef] [Green Version]
- Rios, A.C.; Moutinho, C.G.; Pinto, F.C.; Del Fiol, F.S.; Jozala, A.; Chaud, M.V.; Vila, M.M.D.C.; Teixeira, J.A.; Balcão, V.M. Alternatives to overcoming bacterial resistances: State-of-the-art. Microbiol. Res. 2016, 191, 51–80. [Google Scholar] [CrossRef]
- Alsaadi, A.; Beamud, B.; Easwaran, M.; Abdelrahman, F.; El-Shibiny, A.; Alghoribi, M.F.; Domingo-Calap, P. Learning from Mistakes: The Role of Phages in Pandemics. Front. Microbiol. 2021, 12, 517. [Google Scholar] [CrossRef]
- El-Shibiny, A.; El-Sahhar, S.; Adel, M. Phage applications for improving food safety and infection control in Egypt. J. Appl. Microbiol. 2017, 123, 556–567. [Google Scholar] [CrossRef]
- Nishikawa, H.; Yasuda, M.; Uchiyama, J.; Rashel, M.; Maeda, Y.; Takemura, I.; Sugihara, S.; Ujihara, T.; Shimizu, Y.; Shuin, T.; et al. T-even-related bacteriophages as candidates for treatment of Escherichia coli urinary tract infections. Arch. Virol. 2008, 153, 507–515. [Google Scholar] [CrossRef]
- Bibi, Z.; Abbas, Z.; Rehman, S. A Phage, P. E1 Isolated from Hospital Sewage Reduces the Growth of Escherichia coli. Biocontrol Sci. Technol. 2015, 26, 181–188. [Google Scholar] [CrossRef]
- Jończyk, E.; Kłak, M.; Międzybrodzki, R.; Górski, A. The influence of external factors on bacteriophages—Review. Folia Microbiol. 2011, 56, 191–200. [Google Scholar] [CrossRef] [Green Version]
- Carey-Smith, G.V.; Billington, C.; Cornelius, A.J.; Hudson, J.A.; Heinemann, J.A. Isolation and characterization of bacteriophages infecting Salmonella spp. FEMS Microbiol. Lett. 2006, 258, 182–186. [Google Scholar] [CrossRef] [Green Version]
- Jamalludeen, N.; Johnson, R.P.; Friendship, R.; Kropinski, A.M.; Lingohr, E.J.; Gyles, C.L. Isolation and characterization of nine bacteriophages that lyse O149 enterotoxigenic Escherichia coli. Vet. Microbiol. 2007, 124, 47–57. [Google Scholar] [CrossRef]
- Yazdi, M.; Bouzari, M.; Ghaemi, E.A.; Shahin, K. Isolation, Characterization and Genomic Analysis of a Novel Bacteriophage VB_EcoS-Golestan Infecting Multidrug-Resistant Escherichia coli Isolated from Urinary Tract Infection. Sci. Rep. 2020, 10, 7690. [Google Scholar] [CrossRef]
- Yu, Y.-P.; Gong, T.; Jost, G.; Liu, W.-H.; Ye, D.-Z.; Luo, Z.-H. Isolation and characterization of five lytic bacteriophages infecting a Vibrio strain closely related to Vibrio owensii. FEMS Microbiol. Lett. 2013, 348, 112–119. [Google Scholar] [CrossRef] [Green Version]
- Feng, Y.Y.; Ong, S.L.; Hu, J.Y.; Tan, X.L.; Ng, W.J. Effects of pH and temperature on the survival of coliphages MS2 and Qβ. J. Ind. Microbiol. Biotechnol. 2003, 30, 549–552. [Google Scholar] [CrossRef]
- Ullah, A.; Qamash, T.; Khan, F.A.; Sultan, A.; Ahmad, S.; Abbas, M.; Khattak, M.a.K.; Begum, N.; Din, S.U.; Jamil, J.; et al. Characterization of a Coliphage AS1 isolated from sewage effluent in Pakistan. Braz. J. Biol. 2021, 82, e240943. [Google Scholar] [CrossRef]
- Horváth, M.; Kovács, T.; Koderivalappil, S.; Ábrahám, H.; Rákhely, G.; Schneider, G. Identification of a newly isolated lytic bacteriophage against K24 capsular type, carbapenem resistant Klebsiella pneumoniae isolates. Sci. Rep. 2020, 10, 5891. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Class | Antibiotic Name | Abbreviation | Diameter (mm) | Susceptibility |
|---|---|---|---|---|
| Aminoglycoside | Gentamicin | CN 10 | 7 | Resistant |
| Beta-lactam antibiotics (monobactam) | Aztreonam | ATM 30 | 24 | Sensitive |
| Penicillin-like antibiotics | Amoxicillin/Clavulanic acid | AMC 30 | 16 | Intermediate |
| Beta-lactam antibiotics (First generation) | Ampicillin | AMP 10 | 11 | Resistant |
| Aminoglycoside | Amikacin | AK 30 | 16 | Intermediate |
| Cephalosporin (second generation) | Cefoxitin | FOX 30 | 18 | Sensitive |
| Pencillins/inhibitor combination | Ampicillin/sulbactam | SAM 20 | 0 | Resistant |
| Cephalosporin (third generation) | Cefotaxime | CTX 30 | 28 | Sensitive |
| Tetracycline | Tetracycline | TE 30 | 0 | Resistant |
| Cephalosporin (third generation) | Ceftriaxone | CRO 30 | 25 | Sensitive |
| Bacterial Strain | ZCEC10 | ZCEC11 | ZCEC12 |
|---|---|---|---|
| E. coli O157-H7 | + | + | + |
| E. coli O2 | + | + | + |
| E. coli O-127-H6 | + | + | + |
| E. coli- ATCC 8739 | + | + | + |
| Salmonella enterica Typhimurium- ATCC 14028 | − | − | − |
| Salmonella Kent | − | − | − |
| E. coli O25 | − | − | − |
| E. coli O88 | − | − | − |
| E. coli O78 | − | − | − |
| E. coli O18 | + | + | + |
| E. coli O1 | − | − | − |
| E. coli O12 | − | − | − |
| E. coli O6 | − | − | − |
| E. coli O186 | − | − | − |
| E. coli O-hemolytic | − | − | − |
| E. coli O125 | − | − | − |
| E. coli O55 | − | − | − |
| E. coli O125 | − | − | − |
| E. coli O115 | − | − | − |
| E. coli O27 | − | − | − |
| E. coli O168 | − | − | − |
| E. coli O164 | − | − | − |
| E. coli O114 | − | − | − |
| E. coli O151 | − | − | − |
| E. coli O169 | − | − | − |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abdelrahman, F.; Rezk, N.; Fayez, M.S.; Abdelmoteleb, M.; Atteya, R.; Elhadidy, M.; El-Shibiny, A. Isolation, Characterization, and Genomic Analysis of Three Novel E. coli Bacteriophages That Effectively Infect E. coli O18. Microorganisms 2022, 10, 589. https://doi.org/10.3390/microorganisms10030589
Abdelrahman F, Rezk N, Fayez MS, Abdelmoteleb M, Atteya R, Elhadidy M, El-Shibiny A. Isolation, Characterization, and Genomic Analysis of Three Novel E. coli Bacteriophages That Effectively Infect E. coli O18. Microorganisms. 2022; 10(3):589. https://doi.org/10.3390/microorganisms10030589
Chicago/Turabian StyleAbdelrahman, Fatma, Nouran Rezk, Mohamed S. Fayez, Mohamed Abdelmoteleb, Reham Atteya, Mohamed Elhadidy, and Ayman El-Shibiny. 2022. "Isolation, Characterization, and Genomic Analysis of Three Novel E. coli Bacteriophages That Effectively Infect E. coli O18" Microorganisms 10, no. 3: 589. https://doi.org/10.3390/microorganisms10030589
APA StyleAbdelrahman, F., Rezk, N., Fayez, M. S., Abdelmoteleb, M., Atteya, R., Elhadidy, M., & El-Shibiny, A. (2022). Isolation, Characterization, and Genomic Analysis of Three Novel E. coli Bacteriophages That Effectively Infect E. coli O18. Microorganisms, 10(3), 589. https://doi.org/10.3390/microorganisms10030589









