Reduction of Bacterial Enteric Pathogens and Hygiene Indicator Bacteria on Tomato Skin Surfaces by a Polymeric Nanoparticle-Loaded Plant-Derived Antimicrobial
Abstract
1. Introduction
2. Materials and Methods
2.1. Preparation of Microorganisms for Tomato Sample Inoculation
2.2. Preparation of Tomato Skin Samples for Inoculation by Pathogen Cocktail
2.3. Tomato Sample Inoculation by Cocktailed Pathogens
2.4. Tomato Inoculation and Sanitization Treatment Scenarios
- Scenario 1: The mixed E. coli O157:H7 and S. typhimurium organisms were inoculated prior to sanitization treatment, simulating contamination occurring immediately prior to or during tomato harvest;
- Scenario 2: Pathogens were inoculated twice onto tomato samples, once before sanitization treatment and then again after 3 days of refrigerated storage, simulating conditions of pathogen contamination both during harvest and again during post-harvest packing, and;
- Scenario 3: Pathogens were inoculated/contaminated onto already-treated tomato samples, wherein tomato samples were treated by one of four sanitization treatments (Section 2.5), covered, and then placed under refrigeration (5 °C) for 3 days. On day 3, samples were removed from refrigeration, inoculated with cocktailed pathogens, and then returned to refrigerated storage or immediately prepared for enumeration of surviving cells.
2.5. Preparation and Application of Sanitization Treatments to Tomato Sample Surfaces
2.6. Testing of Sanitization Treatments Efficacy against Hygiene Bacterial Organisms on Non-Inoculated Tomato Skin Samples
2.7. Microbiological Analysis of Inoculated or Uninoculated Tomato Samples
2.8. Experimental Design and Statistical Analysis
3. Results
3.1. Reduction in Bacterial Pathogen Survivors on Tomato Samples from Differing Contamination and Sanitization Scenarios
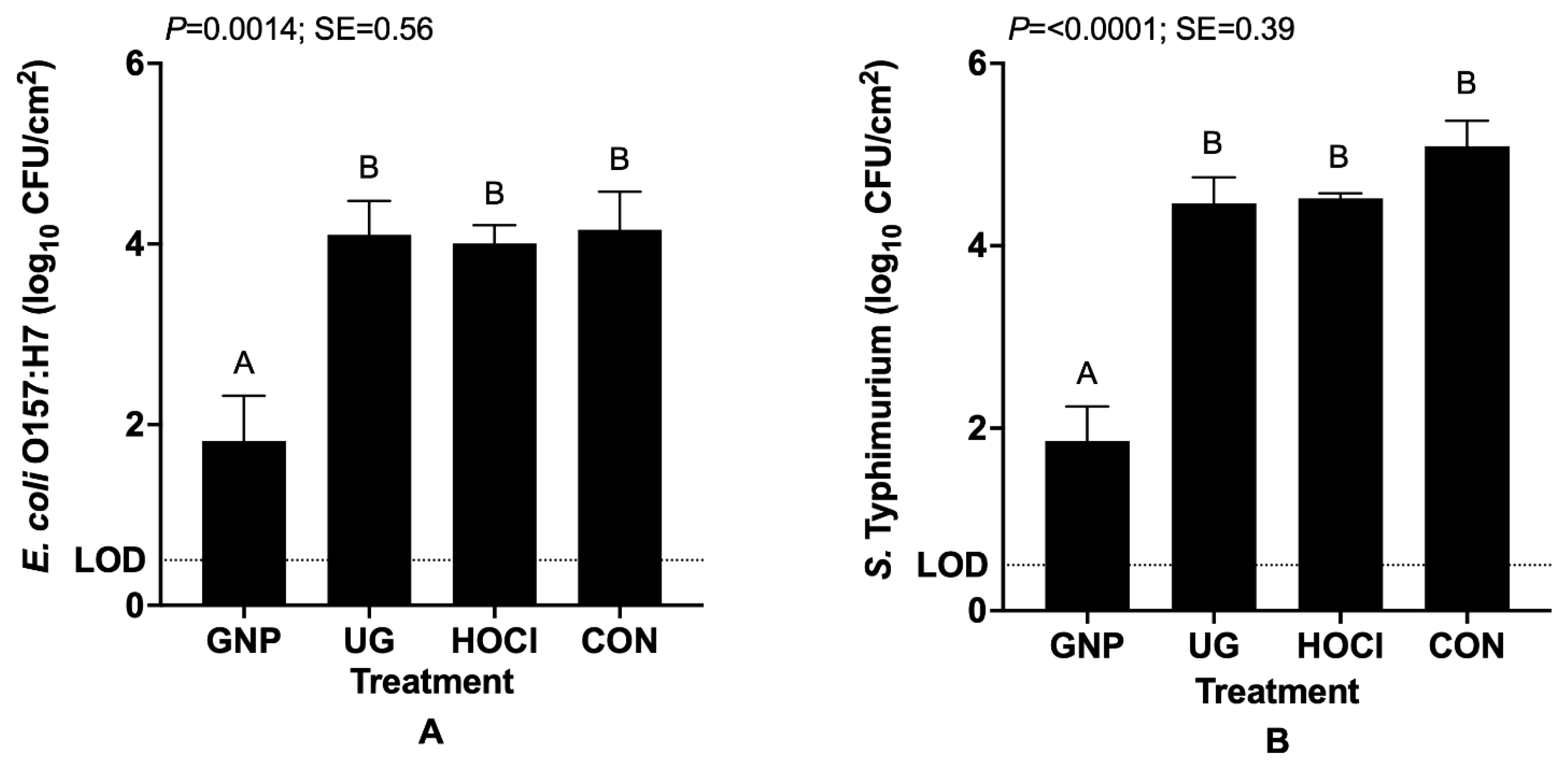
3.1.1. Pathogen Contamination on Tomatoes Precedes Sanitization Treatment (Scenario 1)
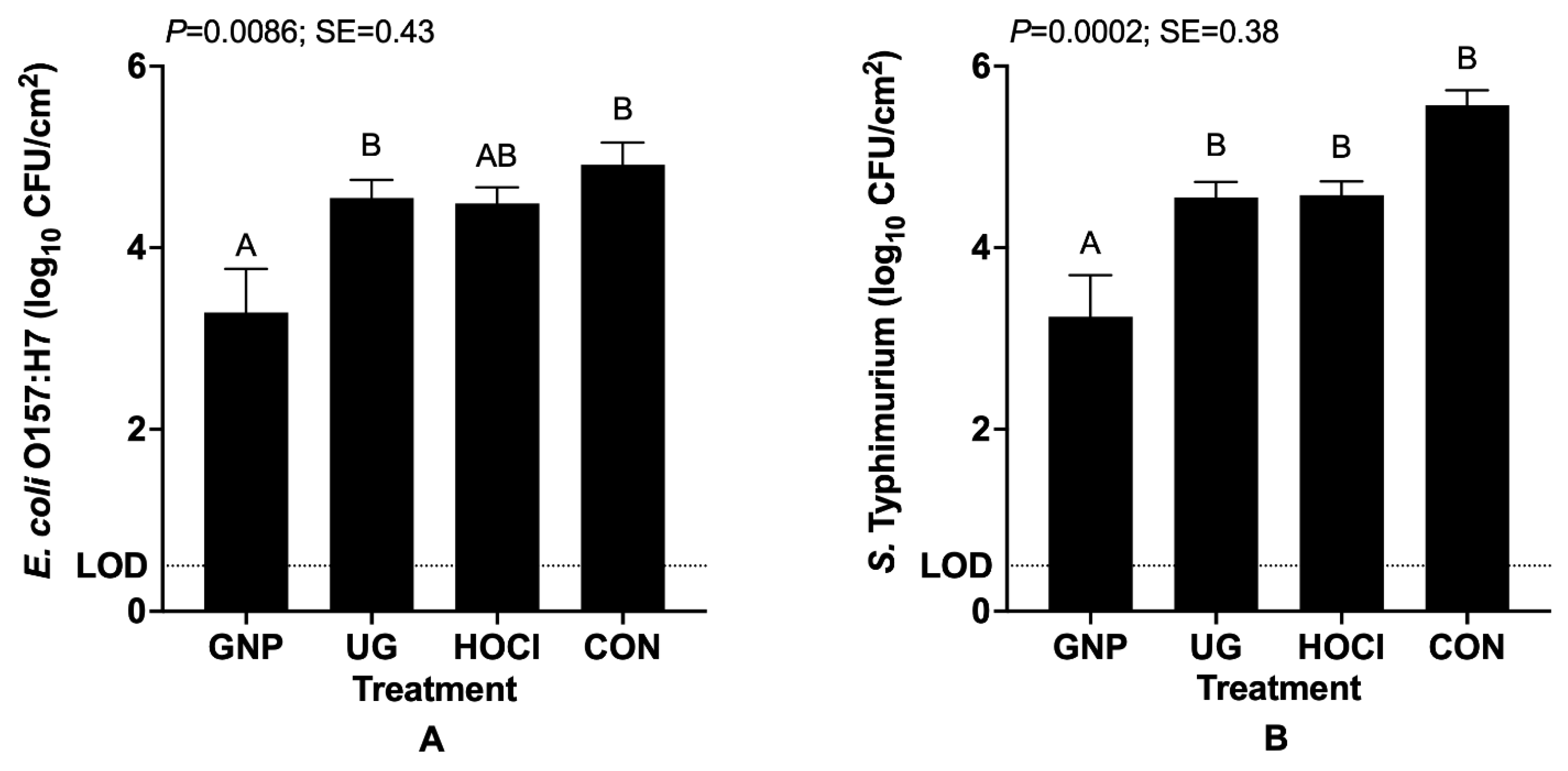
3.1.2. Pathogen Survival When Contamination of Tomatoes Both Precedes and Follows Sanitization Treatment (Scenario 2)
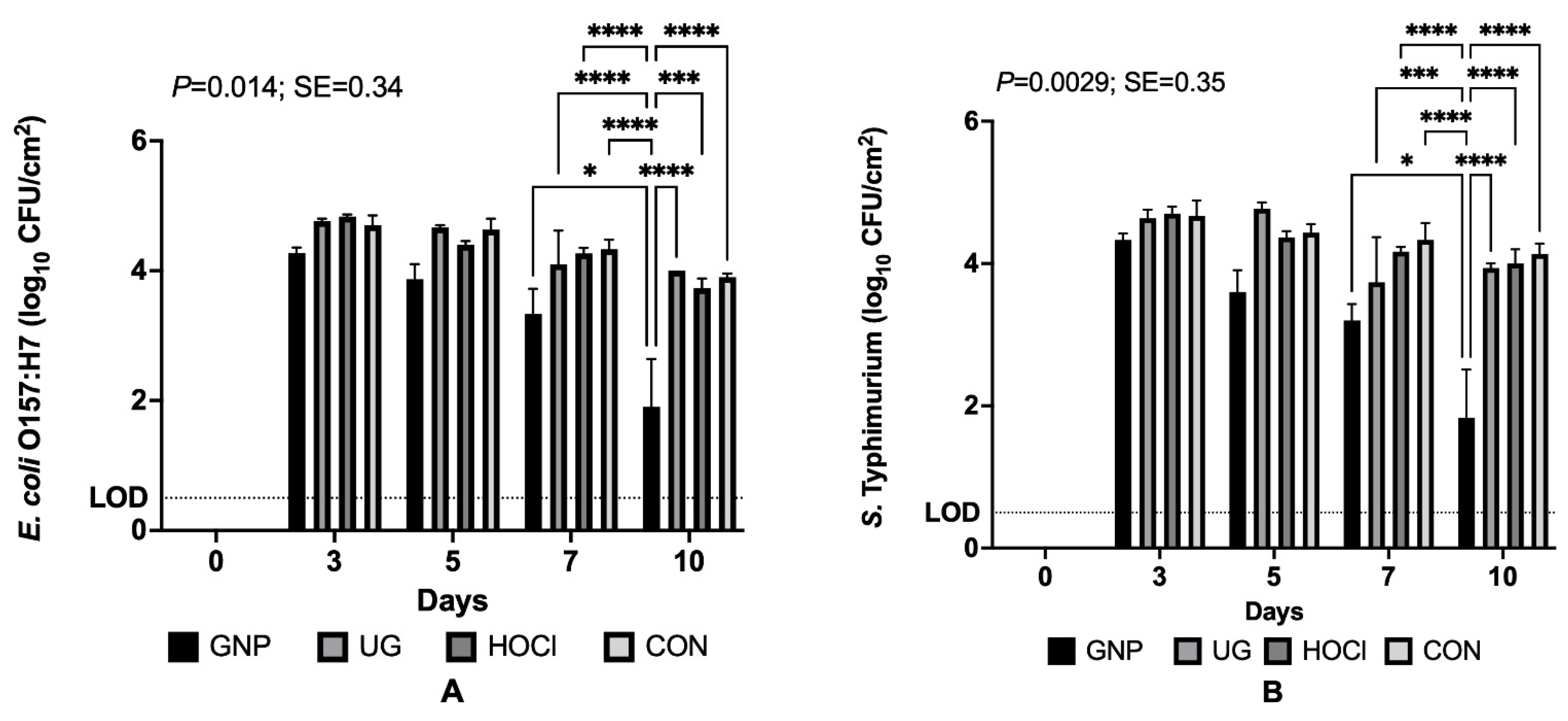
3.1.3. Pathogen Survival on Tomatoes When Contamination Follows Sanitization Treatment (Scenario 3)
3.2. Antimicrobial Impacts of Sanitization Treatment by Experimental Scenario for Pathogen-Inoculated Tomatoes
3.3. Antimicrobial Impacts of Sanitizer Treatments on Tomato Hygiene Indicator Bacteria Groups for Tomato Samples Not Inoculated with Pathogens
3.3.1. Pathogen Contamination on Tomatoes Precedes Sanitization Treatment
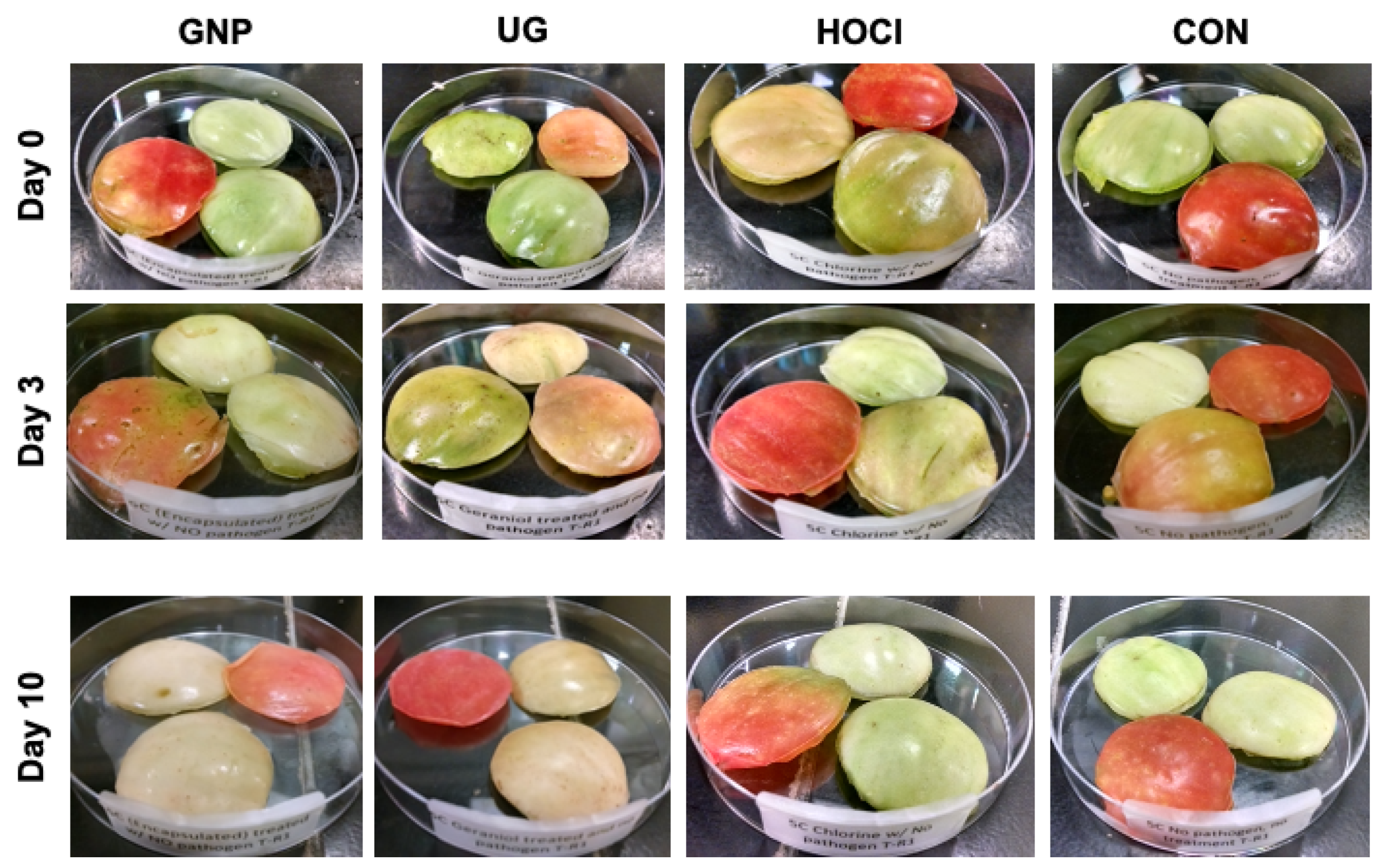
3.3.2. Impact of Sanitization Treatment on Tomato Skin Appearance during Storage
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- USDA Economic Research Service (ERS). Annual Number of Foodborne Illnesses Associated with Outbreaks in Tomatoes Has Generally Decreased Since Their High in 2001. Available online: https://www.ers.usda.gov/data-products/chart-gallery/gallery/chart-detail/?chartId=93465 (accessed on 21 December 2021).
- Gurtler, J.B.; Harlee, N.A.; Smelser, A.M.; Schneider, K.R. Salmonella enterica contamination of market fresh tomatoes: A review. J. Food Prot. 2018, 81, 1193–1213. [Google Scholar] [CrossRef] [PubMed]
- Colombe, S.; Jernberg, C.; Löf, E.; Angervall, A.L.; Mellström-Dahlgren, H.; Dotevall, L.; Bengnér, M.; Hall, I.; Sundqvist, L.; Kühlmann-Berenzon, S.; et al. Outbreak of unusual H2S-negative monophasic Salmonella Typhimurium strain likely associated with small tomatoes, Sweden, August to October 2019. Eurosurveillance 2019, 24, 1900643. [Google Scholar] [CrossRef] [PubMed]
- Deering, A.J.; Jack, D.R.; Pruitt, R.E.; Mauer, L.J. Movement of Salmonella serovar Typhimurium and E. coli O157:H7 to ripe tomato fruit following various routes of contamination. Microorganisms 2015, 3, 809–825. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhou, D.; Cao, Y.; Zhang, Y.; Xiao, X.; Liu, F.; Yu, Y. Synergistic inactivation of Escherichia coli O157:H7 and Staphylococcus aureus by gallic acid and thymol and its potential application on fresh-cut tomatoes. Food Microbiol. 2022, 102, 103925. [Google Scholar] [CrossRef] [PubMed]
- Ruengvisesh, S.; Oh, J.K.; Kerth, C.R.; Akbulut, M.; Taylor, T.M. Inhibition of bacterial human pathogens on tomato skin surfaces using eugenol-loaded surfactant micelles during refrigerated and abuse storage. J. Food Saf. 2019, 39, e12598. [Google Scholar] [CrossRef]
- He, Q.; Guo, M.; Jin, T.Z.; Arabi, S.A.; Liu, D. Ultrasound improves the decontamination effect of thyme essential oil nanoemulsions against Escherichia coli O157:H7 on cherry tomatoes. Int. J. Food Microbiol. 2021, 337, 108936. [Google Scholar] [CrossRef]
- Kirkland, E.; Green, L.R.; Stone, C.; Reimann, D.; Nicholas, D.; Mason, R.; Frick, R.; Coleman, S.; Bushnell, L.; Blade, H.; et al. Tomato handling practices in restaurants. J. Food Prot. 2009, 72, 1692–1698. [Google Scholar] [CrossRef][Green Version]
- Perez-Lewis, K.L.; Yegin, Y.; Oh, J.K.; Castillo, A.; Cisneros-Zevallos, L.; Kerth, C.R.; Scholar, E.; Taylor, T.M. Encapsulated plant-derived antimicrobial reduces enteric bacterial pathogens on melon surfaces during differing contamination and sanitization treatment scenarios. Appl. Microbiol. 2021, 1, 460–470. [Google Scholar] [CrossRef]
- Yegin, Y.; Perez-Lewis, K.L.; Liu, S.; Kerth, C.R.; Cisneros-Zevallos, L.; Castillo, A.; Akbulut, M.; Taylor, T.M. Antimicrobial-loaded polymeric micelles inhibit enteric bacterial pathogens on spinach leaf surfaces during multiple simulated pathogen contamination events. Front. Sus. Food Syst. 2021, 5, 646980. [Google Scholar] [CrossRef]
- Perez-Lewis, K.L.; Yegin, Y.; Cisneros-Zevallos, L.; Castillo, A.; Kerth, C.R.; Akbulut, M.; Taylor, T.M. Geraniol-loaded polymeric nanoparticles inhibit enteric pathogens on spinach during posttreatment refrigerated and temperature abuse storage. Front. Sustain. Food Syst. 2018, 2, 4. [Google Scholar] [CrossRef]
- Yegin, Y.; Perez-Lewis, K.L.; Zhang, M.; Akbulut, M.; Taylor, T.M. Development and characterization of geraniol-loaded polymeric nanoparticles with antimicrobial activity against foodborne bacterial pathogens. J. Food Eng. 2016, 170, 64–71. [Google Scholar] [CrossRef]
- Harris, L.J.; Farber, J.N.; Beuchat, L.R.; Parish, M.E.; Suslow, T.V.; Garrett, E.H.; Busta, F.F. Outbreaks associated with fresh produce: Incidence, growth, and survival of pathogens in fresh and fresh-cut produce. Comp. Rev. Food Sci. Food Saf. 2003, 2, 78–141. [Google Scholar] [CrossRef]
- Wang, H.; Feng, H.; Luo, Y.; Zhang, A. Produce surface characteristics affect product quality and safety. Acta Hort. 2007, 746, 131–138. [Google Scholar] [CrossRef]
- Lu, Y.; Wu, C. Reduction of Salmonella enterica contamination on grape tomatoes by washing with thyme oil, thymol, and carvacrol as compared to chlorine treatment. J. Food Prot. 2010, 73, 2270–2275. [Google Scholar] [CrossRef] [PubMed]
- Shu, X.; Singh, M.; Karampundi, N.B.R.; Bridges, D.F.; Kitazumi, A.; Wu, V.C.H.; De los Reyes, B.G. Respones of Escherichia coli and Listeria monocytogenes to ozone treatment on non-host tomato: Efficacy of intervention and evidence of induced acclimation. PLoS ONE 2021, 16, e0256324. [Google Scholar] [CrossRef]
- Afari, G.K.; Hung, Y.-C. A meta-analysis on the effectiveness of electrolyzed water treatments in reducing foodborne pathogens on different foods. Food Cont. 2018, 93, 150–164. [Google Scholar] [CrossRef]
- Abuladze, T.; Li, M.; Menetrez, M.Y.; Dean, T.; Senecal, A.; Sulakvelidze, A. Bacteriophages reduce experimental contamination of hard surfaces, tomato, spinach, broccoli, and ground beef by Escherichia coli O157:H7. Appl. Environ. Microbiol. 2008, 74, 6230–6238. [Google Scholar] [CrossRef]
- Wei, W.; Wang, X.; Xie, Z.; Wang, W.; Xu, J.; Liu, Y.; Gao, H.; Zhou, Y. Evaluation of sanitizing methods for reducing microbial contamination on fresh strawberry, cherry tomato, and red bayberry. Front. Microbiol. 2017, 8, 2397. [Google Scholar] [CrossRef]
- Gündüz, G.T.; Gönül, Ş.A.; Karapinar, M. Efficacy of sumac and oregano in the inactivation of Salmonella Typhimurium on tomatoes. Int. J. Food Microbiol. 2010, 141, 39–44. [Google Scholar] [CrossRef]
- Mattson, T.E.; Johny, A.K.; Amalaradjou, M.A.R.; More, K.; Schreiber, D.T.; Venkitanarayanan, K. Inactivation of Salmonella spp. on tomatoes by plant molecules. Int. J. Food Microbiol. 2011, 144, 464–468. [Google Scholar] [CrossRef]
- Lu, Y.; Joerger, R.; Wu, C. Similar reduction of Salmonella enterica Typhimurium on grape tomatoes and its cross-contamination in wash water by washing with natural antimicrobials as compared with chlorine treatment. Food Bioproc. Technol. 2014, 7, 661–760. [Google Scholar] [CrossRef]
- Wang, H.; Zhou, B.; Feng, H. Surface characteristics of fresh produce and their impact on attachment and removal of human pathogens on produce surfaces. In Decontamination of Fresh and Minimally Processed Produce, 1st ed.; Gómez-López, V.M., Ed.; John Wiley & Sons, Inc.: Ames, IA, USA, 2012; pp. 43–57. [Google Scholar]
- Yaron, S.; Römling, U. Biofilm formation by enteric pathogens and its role in plant colonization and persistence. Microb. Biotechnol. 2014, 7, 496–516. [Google Scholar] [CrossRef] [PubMed]
- Murray, K.; Wu, F.; Shi, J.; Xue, S.J.; Warriner, K. Challenges in the microbiological food safety of fresh produce: Limitations of post-harvest washing and the need for alternative interventions. Food Qual. Saf. 2017, 1, 289–301. [Google Scholar] [CrossRef]
- Smid, E.J.; Hendricks, L.; Boerringer, H.A.M.; Gorris, L.G.M. Surface disinfection of tomatoes using the natural plant compound trans-cinnamaldehyde. Postharvest Biol. Technol. 1996, 9, 343–350. [Google Scholar] [CrossRef]
- Keeratipibul, S.; Phewpan, A.; Lursinsap, C. Prediction of coliforms and Escherichia coli on tomato fruits and lettuce leaves after sanitizing by using artificial neural networks. LWT Food Sci. Technol. 2011, 44, 130–138. [Google Scholar] [CrossRef]
- Buendía-Moreno, L.; Sánchez-Martínez, M.J.; Antolinos, V.; Ros-Chumillas, M.; Navarro-Segura, L.; Soto-Jover, S.; Martínez-Hernández, G.B.; López-Gómez, A. Active cardboard box with a coating including essential oils entrapped within cyclodextrins and/or halloysite nanotubes. A case study for fresh tomato storage. Food Control 2020, 107, 106763. [Google Scholar] [CrossRef]

| Sanitizing Treatment 1 | Scenario 1 2 | Scenario 2 | Scenario 3 |
|---|---|---|---|
| GNP | 1.82 F 3 | 3.29 DE | 2.77 E |
| UG | 4.10 ABCD | 4.55 AB | 3.61 CDE |
| HOCl | 4.01 BCD | 4.49 AB | 3.55 CDE |
| CON | 4.15 ABC | 4.91 A | 3.61 CDE |
| p = 0.0006; Pooled SE = 0.18 |
| Sanitization Treatment 1 | Scenario 1 2 | Scenario 2 | Scenario 3 |
|---|---|---|---|
| GNP | 1.86 F 3 | 3.24 DE | 2.69 EF |
| UG | 4.47 BC | 4.55 B | 3.51 DE |
| HOCl | 4.52 BC | 4.58 B | 3.55 DE |
| CON | 5.09 AB | 5.57 A | 3.61 CD |
| p =< 0.0001; Pooled SE = 0.19 |
| Sanitization Treatment 1 | APC 2 | LAB | Coliforms |
|---|---|---|---|
| GNP | 3.61 A 3 | 3.39 A | 1.13 A |
| UG | 4.44 A | 4.28 B | 1.47 A |
| HOCl | 5.04 AB | 4.82 B | 2.23 AB |
| CON | 5.99 B | 5.77 B | 3.29 B |
| p =< 0.00001; SE = 0.45 | p = 0.0004; SE = 0.43 | p = 0.0054; SE = 0.55 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Perez-Lewis, K.L.; Yegin, Y.; Oh, J.K.; Castillo, A.; Cisneros-Zevallos, L.; Kerth, C.R.; Akbulut, M.; Taylor, T.M. Reduction of Bacterial Enteric Pathogens and Hygiene Indicator Bacteria on Tomato Skin Surfaces by a Polymeric Nanoparticle-Loaded Plant-Derived Antimicrobial. Microorganisms 2022, 10, 448. https://doi.org/10.3390/microorganisms10020448
Perez-Lewis KL, Yegin Y, Oh JK, Castillo A, Cisneros-Zevallos L, Kerth CR, Akbulut M, Taylor TM. Reduction of Bacterial Enteric Pathogens and Hygiene Indicator Bacteria on Tomato Skin Surfaces by a Polymeric Nanoparticle-Loaded Plant-Derived Antimicrobial. Microorganisms. 2022; 10(2):448. https://doi.org/10.3390/microorganisms10020448
Chicago/Turabian StylePerez-Lewis, Keila L., Yagmur Yegin, Jun K. Oh, Alejandro Castillo, Luis Cisneros-Zevallos, Chris R. Kerth, Mustafa Akbulut, and Thomas M. Taylor. 2022. "Reduction of Bacterial Enteric Pathogens and Hygiene Indicator Bacteria on Tomato Skin Surfaces by a Polymeric Nanoparticle-Loaded Plant-Derived Antimicrobial" Microorganisms 10, no. 2: 448. https://doi.org/10.3390/microorganisms10020448
APA StylePerez-Lewis, K. L., Yegin, Y., Oh, J. K., Castillo, A., Cisneros-Zevallos, L., Kerth, C. R., Akbulut, M., & Taylor, T. M. (2022). Reduction of Bacterial Enteric Pathogens and Hygiene Indicator Bacteria on Tomato Skin Surfaces by a Polymeric Nanoparticle-Loaded Plant-Derived Antimicrobial. Microorganisms, 10(2), 448. https://doi.org/10.3390/microorganisms10020448









