Whole-Transcriptome Sequencing Reveals Characteristics of Cancer Microbiome in Korean Patients with GI Tract Cancer: Fusobacterium nucleatum as a Therapeutic Target
Abstract
1. Introduction
2. Materials and Methods
2.1. Public Whole Transcriptome Dataset Analysis
2.2. Bacterial Culture Growth and Supernatants
2.3. Cell Culture Growth, siRNA Transfection, and Treatment of Bacterial Supernatants
2.4. Total RNA Extraction and qRT-PCR Analysis
2.5. WST-1 Assay
2.6. Statistical Analysis
3. Results and Discussion
3.1. Fusobacterium nucleatum Is Enriched in Gastrointestinal Tract Cancer Tissues
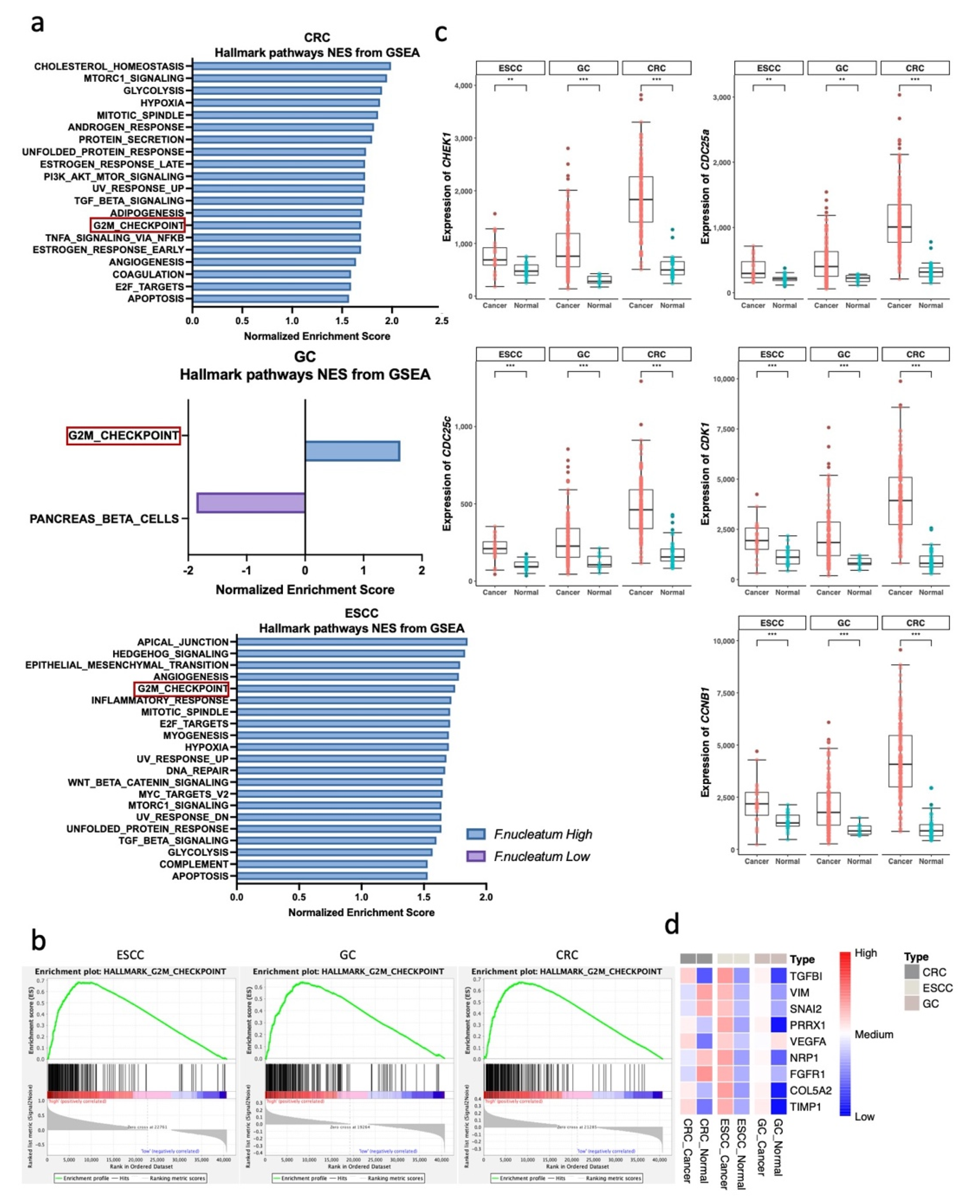
3.2. G2M Checkpoint Pathway Is Associated with F. nucleatum Abundance in GI Tract Cancer
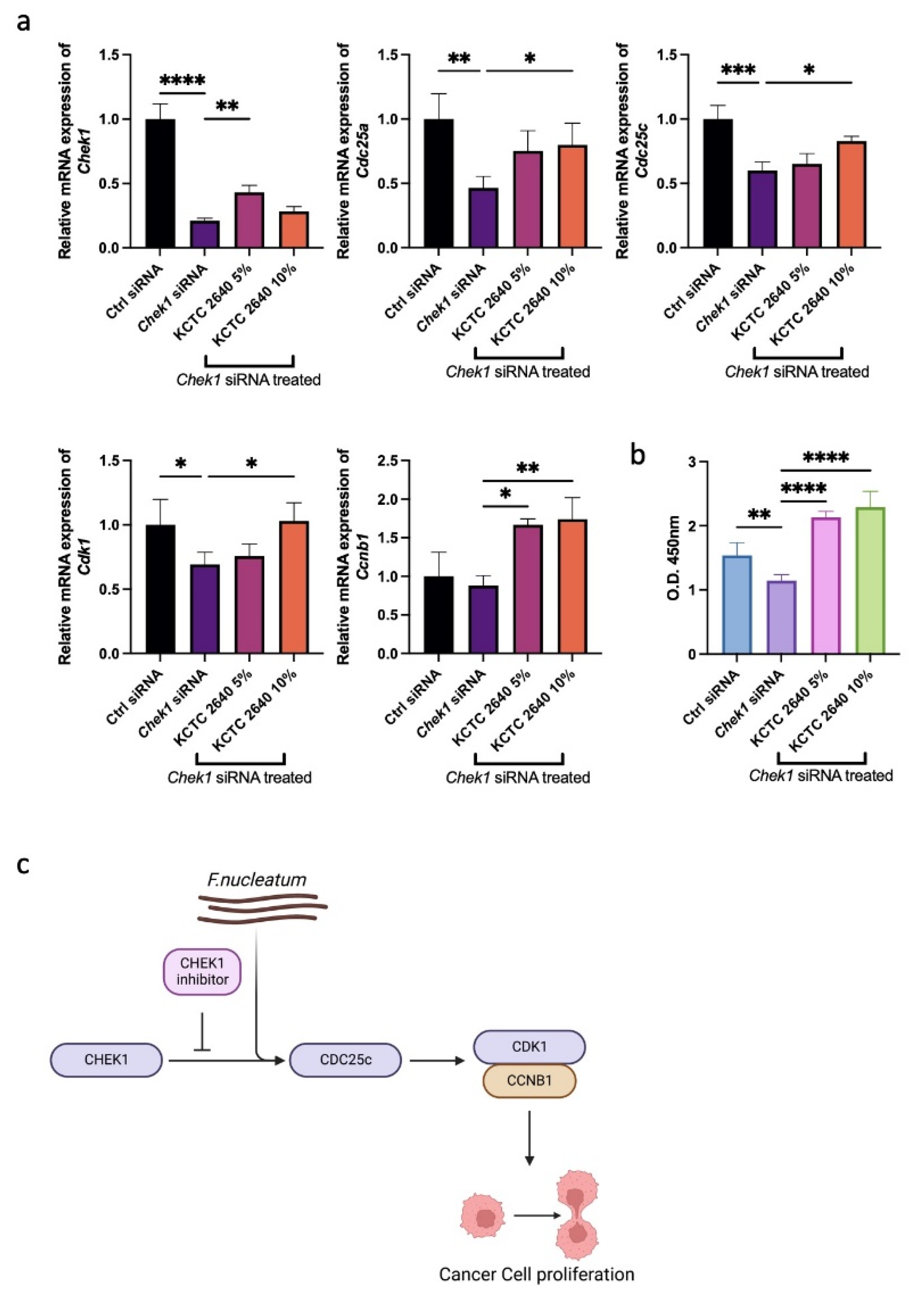
3.3. F. nucleatum Promotes Cancer Cell Proliferation through the G2M Checkpoint Pathway
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bhatt, A.P.; Redinbo, M.R.; Bultman, S.J. The role of the microbiome in cancer development and therapy. CA A Cancer J. Clin. 2017, 67, 326–344. [Google Scholar] [CrossRef] [PubMed]
- Hong, M.; Tao, S.; Zhang, L.; Diao, L.-T.; Huang, X.; Huang, S.; Xie, S.-J.; Xiao, Z.-D.; Zhang, H. RNA sequencing: New technologies and applications in cancer research. J. Hematol. Oncol. 2020, 13, 166. [Google Scholar] [CrossRef] [PubMed]
- Allen, J.; Sears, C.L. Impact of the gut microbiome on the genome and epigenome of colon epithelial cells: Contributions to colorectal cancer development. Genome Med. 2019, 11, 11. [Google Scholar] [CrossRef]
- Cullin, N.; Azevedo Antunes, C.; Straussman, R.; Stein-Thoeringer, C.K.; Elinav, E. Microbiome and cancer. Cancer Cell 2021, 39, 1317–1341. [Google Scholar] [CrossRef] [PubMed]
- Doorakkers, E.; Lagergren, J.; Engstrand, L.; Brusselaers, N. Eradication of Helicobacter pylori and Gastric Cancer: A Systematic Review and Meta-analysis of Cohort Studies. J. Natl. Cancer Inst. 2016, 108. [Google Scholar] [CrossRef] [PubMed]
- Buc, E.; Dubois, D.; Sauvanet, P.; Raisch, J.; Delmas, J.; Darfeuille-Michaud, A.; Pezet, D.; Bonnet, R. High prevalence of mucosa-associated E. coli producing cyclomodulin and genotoxin in colon cancer. PLoS ONE 2013, 8, e56964. [Google Scholar] [CrossRef]
- Haghi, F.; Goli, E.; Mirzaei, B.; Zeighami, H. The association between fecal enterotoxigenic B. fragilis with colorectal cancer. BMC Cancer 2019, 19, 1–4. [Google Scholar] [CrossRef]
- Scanu, T.; Spaapen, R.M.; Bakker, J.M.; Pratap, C.B.; Wu, L.-E.; Hofland, I.; Broeks, A.; Shukla, V.K.; Kumar, M.; Janssen, H. Salmonella manipulation of host signaling pathways provokes cellular transformation associated with gallbladder carcinoma. Cell Host Microbe 2015, 17, 763–774. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Alcoholado, L.; Ramos-Molina, B.; Otero, A.; Laborda-Illanes, A.; Ordóñez, R.; Medina, J.A.; Gómez-Millán, J.; Queipo-Ortuño, M.I. The Role of the Gut Microbiome in Colorectal Cancer Development and Therapy Response. Cancers 2020, 12, 1406. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Wang, X.; Fu, L.; Yang, K. Fusobacterium nucleatum: A new player in regulation of cancer development and therapeutic response. Cancer Drug Resist. 2022, 5, 436–450. [Google Scholar] [CrossRef] [PubMed]
- Suraya, R.; Nagano, T.; Kobayashi, K.; Nishimura, Y. Microbiome as a Target for Cancer Therapy. Integr. Cancer Ther. 2020, 19, 1534735420920721. [Google Scholar] [CrossRef]
- Kim, S.K.; Kim, H.J.; Park, J.L.; Heo, H.; Kim, S.Y.; Lee, S.I.; Song, K.S.; Kim, W.H.; Kim, Y.S. Identification of a molecular signature of prognostic subtypes in diffuse-type gastric cancer. Gastric. Cancer 2020, 23, 473–482. [Google Scholar] [CrossRef]
- You, B.H.; Yoon, J.H.; Kang, H.; Lee, E.K.; Lee, S.K.; Nam, J.W. HERES, a lncRNA that regulates canonical and noncanonical Wnt signaling pathways via interaction with EZH2. Proc. Natl. Acad. Sci. USA 2019, 116, 24620–24629. [Google Scholar] [CrossRef] [PubMed]
- Park, D.Y.; Choi, C.; Shin, E.; Lee, J.H.; Kwon, C.H.; Jo, H.J.; Kim, H.R.; Kim, H.S.; Oh, N.; Lee, J.S.; et al. NTRK1 fusions for the therapeutic intervention of Korean patients with colon cancer. Oncotarget 2016, 7, 8399–8412. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- Wood, D.E.; Lu, J.; Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 2019, 20, 257. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2013, 30, 923–930. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef]
- Kolde, R. Pheatmap: Pretty heatmaps. R Package Version 2012, 1, 726. [Google Scholar]
- Chen, H.; Boutros, P.C. VennDiagram: A package for the generation of highly-customizable Venn and Euler diagrams in R. BMC Bioinform. 2011, 12, 35. [Google Scholar] [CrossRef]
- Wickham, H. Data analysis. In ggplot2; Springer: Germany, 2016; pp. 189–201. [Google Scholar]
- Yamamura, K.; Baba, Y.; Nakagawa, S.; Mima, K.; Miyake, K.; Nakamura, K.; Sawayama, H.; Kinoshita, K.; Ishimoto, T.; Iwatsuki, M.; et al. Human Microbiome Fusobacterium Nucleatum in Esophageal Cancer Tissue Is Associated with Prognosis. Clin. Cancer Res. 2016, 22, 5574–5581. [Google Scholar] [CrossRef]
- Irfan, M.; Delgado, R.Z.R.; Frias-Lopez, J. The Oral Microbiome and Cancer. Front. Immunol. 2020, 11, 591088. [Google Scholar] [CrossRef] [PubMed]
- Mitsuhashi, K.; Nosho, K.; Sukawa, Y.; Matsunaga, Y.; Ito, M.; Kurihara, H.; Kanno, S.; Igarashi, H.; Naito, T.; Adachi, Y.; et al. Association of Fusobacterium species in pancreatic cancer tissues with molecular features and prognosis. Oncotarget 2015, 6, 7209–7220. [Google Scholar] [CrossRef] [PubMed]
- Boehm, E.T.; Thon, C.; Kupcinskas, J.; Steponaitiene, R.; Skieceviciene, J.; Canbay, A.; Malfertheiner, P.; Link, A. Fusobacterium nucleatum is associated with worse prognosis in Lauren’s diffuse type gastric cancer patients. Sci. Rep. 2020, 10, 16240. [Google Scholar] [CrossRef]
- Khatoon, J.; Rai, R.P.; Prasad, K.N. Role of Helicobacter pylori in gastric cancer: Updates. World J. Gastrointest. Oncol. 2016, 8, 147–158. [Google Scholar] [CrossRef]
- Könönen, E.; Gursoy, U.K. Oral Prevotella Species and Their Connection to Events of Clinical Relevance in Gastrointestinal and Respiratory Tracts. Front. Microbiol. 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Visconti, R.; Della Monica, R.; Grieco, D. Cell cycle checkpoint in cancer: A therapeutically targetable double-edged sword. J. Exp. Clin. Cancer Res. 2016, 35, 153. [Google Scholar] [CrossRef]
- Zhang, Y.; Hunter, T. Roles of Chk1 in cell biology and cancer therapy. Int. J. Cancer 2014, 134, 1013–1023. [Google Scholar] [CrossRef]
- Qiu, Z.; Oleinick, N.L.; Zhang, J. ATR/CHK1 inhibitors and cancer therapy. Radiother. Oncol. 2018, 126, 450–464. [Google Scholar] [CrossRef] [PubMed]
- Albiges, L.; Goubar, A.; Scott, V.; Vicier, C.; Lefèbvre, C.; Alsafadi, S.; Commo, F.; Saghatchian, M.; Lazar, V.; Dessen, P.; et al. Chk1 as a new therapeutic target in triple-negative breast cancer. Breast 2014, 23, 250–258. [Google Scholar] [CrossRef]
- Fang, Y.; Yu, H.; Liang, X.; Xu, J.; Cai, X. Chk1-induced CCNB1 overexpression promotes cell proliferation and tumor growth in human colorectal cancer. Cancer Biol. Ther. 2014, 15, 1268–1279. [Google Scholar] [CrossRef]
- Ribatti, D. Epithelial-mesenchymal transition in morphogenesis, cancer progression and angiogenesis. Exp. Cell Res. 2017, 353, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Newsome, R.C.; Yang, Y.; Jobin, C. The microbiome, gastrointestinal cancer, and immunotherapy. J. Gastroenterol. Hepatol. 2022, 37, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.A.; Liu, F.; Riordan, S.M.; Lee, C.S.; Zhang, L. Global Investigations of Fusobacterium nucleatum in Human Colorectal Cancer. Front. Oncol. 2019, 9, 566. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.H.; Cho, Y.; Kim, D.H.; Woo, H.J.; Yang, J.Y.; Kwon, H.J.; Yeon, M.J.; Park, M.; Kim, S.H.; Moon, C.; et al. Menadione induces G2/M arrest in gastric cancer cells by down-regulation of CDC25C and proteasome mediated degradation of CDK1 and cyclin B1. Am. J. Transl. Res. 2016, 8, 5246–5255. [Google Scholar]
- Jung, K.-W.; Won, Y.-J.; Kong, H.-J.; Lee, E.S. Cancer Statistics in Korea: Incidence, Mortality, Survival, and Prevalence in 2015. Cancer Res. Treat. 2018, 50, 303–316. [Google Scholar] [CrossRef] [PubMed]
- Vedeld, H.M.; Goel, A.; Lind, G.E. Epigenetic biomarkers in gastrointestinal cancers: The current state and clinical perspectives. In Proceedings of the Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2018; pp. 36–49. [Google Scholar]
- Bullman, S.; Pedamallu, C.S.; Sicinska, E.; Clancy, T.E.; Zhang, X.; Cai, D.; Neuberg, D.; Huang, K.; Guevara, F.; Nelson, T.; et al. Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science 2017, 358, 1443–1448. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ahn, H.; Min, K.; Lee, E.; Kim, H.; Kim, S.; Kim, Y.; Kim, G.; Cho, B.; Jeong, C.; Kim, Y.; et al. Whole-Transcriptome Sequencing Reveals Characteristics of Cancer Microbiome in Korean Patients with GI Tract Cancer: Fusobacterium nucleatum as a Therapeutic Target. Microorganisms 2022, 10, 1896. https://doi.org/10.3390/microorganisms10101896
Ahn H, Min K, Lee E, Kim H, Kim S, Kim Y, Kim G, Cho B, Jeong C, Kim Y, et al. Whole-Transcriptome Sequencing Reveals Characteristics of Cancer Microbiome in Korean Patients with GI Tract Cancer: Fusobacterium nucleatum as a Therapeutic Target. Microorganisms. 2022; 10(10):1896. https://doi.org/10.3390/microorganisms10101896
Chicago/Turabian StyleAhn, Hyeok, Kyungchan Min, Eulgi Lee, Hyun Kim, Sujeong Kim, Yunjae Kim, Gihyeon Kim, Beomki Cho, Chanyeong Jeong, Yeongmin Kim, and et al. 2022. "Whole-Transcriptome Sequencing Reveals Characteristics of Cancer Microbiome in Korean Patients with GI Tract Cancer: Fusobacterium nucleatum as a Therapeutic Target" Microorganisms 10, no. 10: 1896. https://doi.org/10.3390/microorganisms10101896
APA StyleAhn, H., Min, K., Lee, E., Kim, H., Kim, S., Kim, Y., Kim, G., Cho, B., Jeong, C., Kim, Y., & Park, H. (2022). Whole-Transcriptome Sequencing Reveals Characteristics of Cancer Microbiome in Korean Patients with GI Tract Cancer: Fusobacterium nucleatum as a Therapeutic Target. Microorganisms, 10(10), 1896. https://doi.org/10.3390/microorganisms10101896




