In Silico Identification of Novel Immunogenic Secreted Proteins of Mycoplasma bovis from Secretome Data and Experimental Verification
Abstract
1. Introduction
2. Results
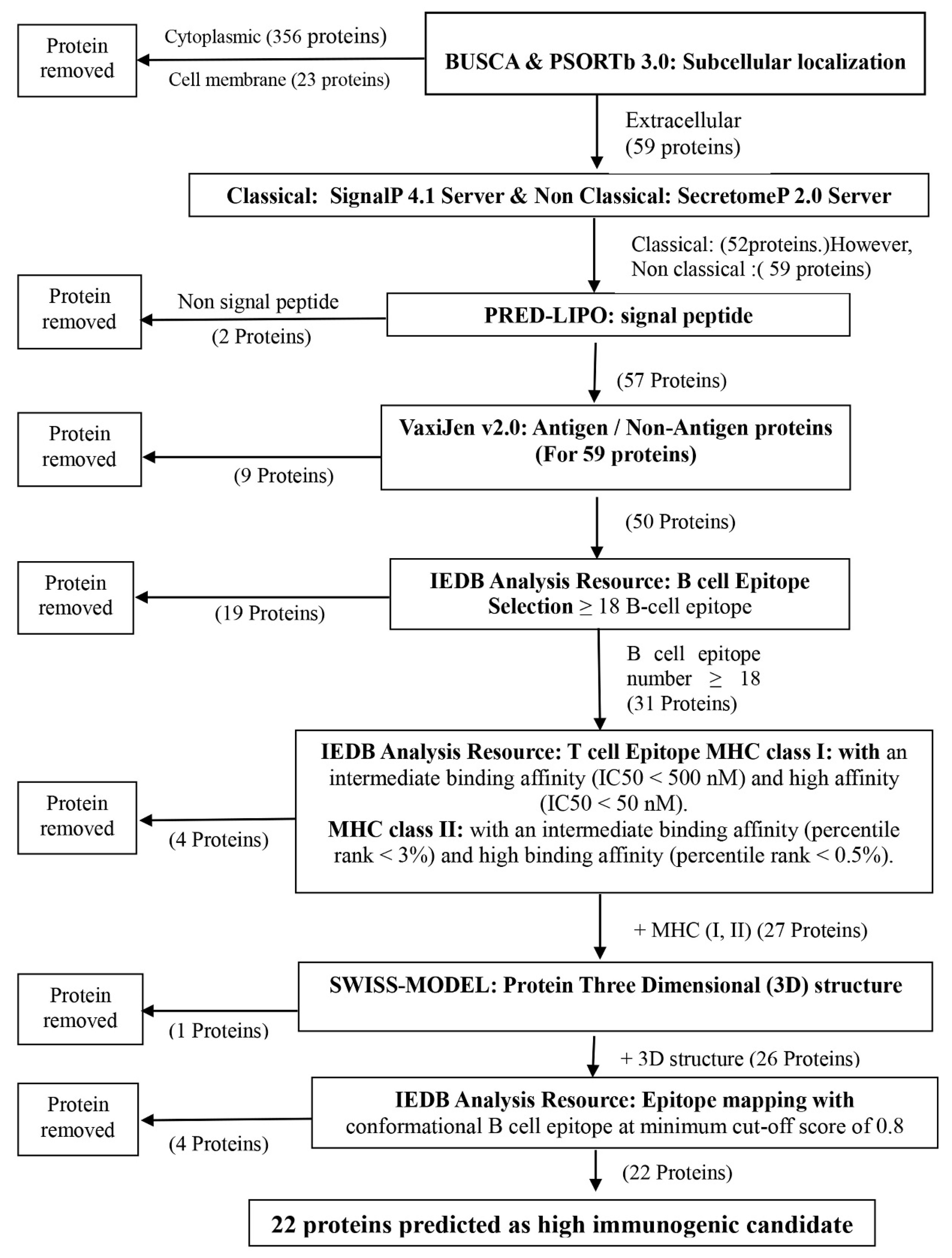
2.1. Prediction of Protein Subcellular Localization
2.2. Prediction of Secreted Proteins
2.3. Analysis of Linear B-Cell and MHC Class I and Class II T-Cell Epitopes
2.4. Overlapping Residues of MHC Class I and II T-cell Epitopes
2.5. 3D Structural Modeling and Conformational B-Cell Epitopes
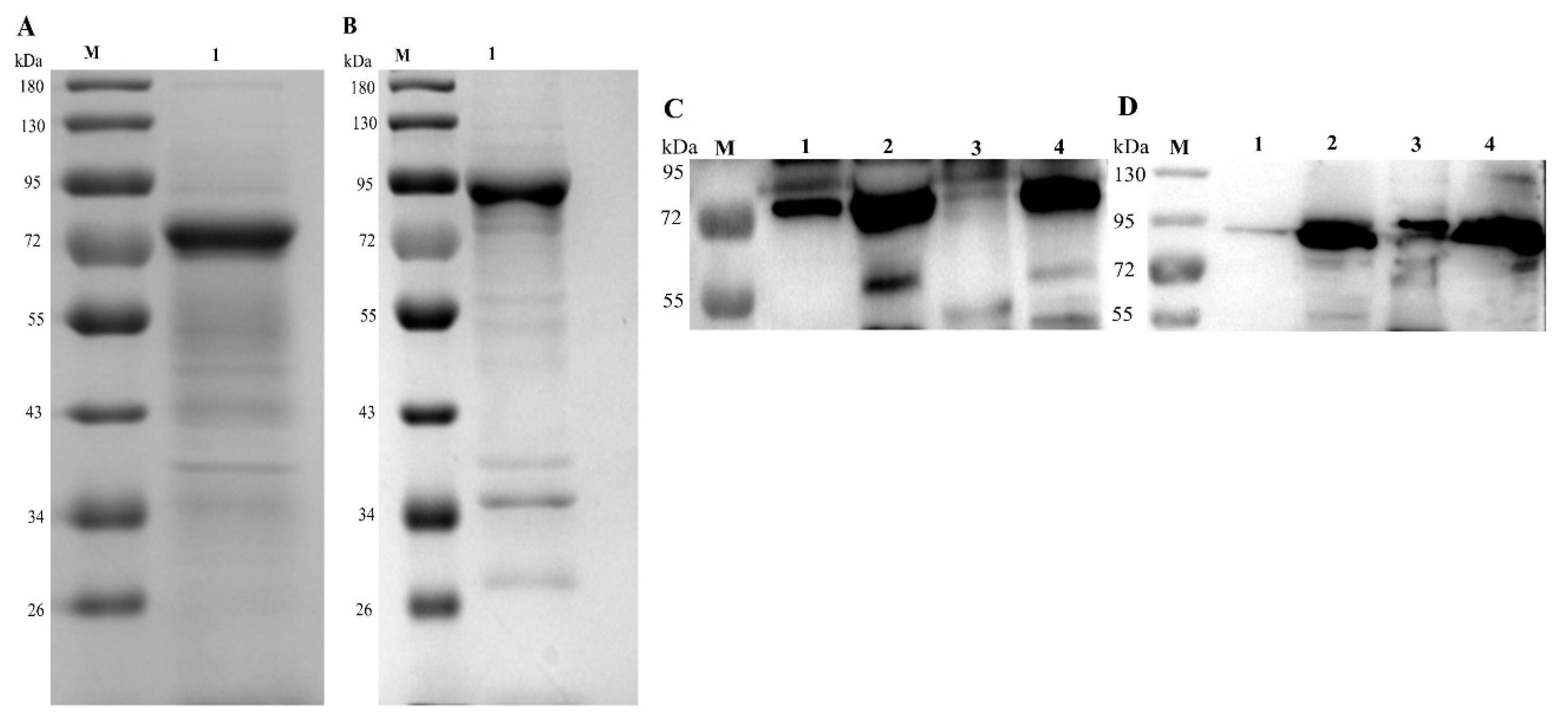
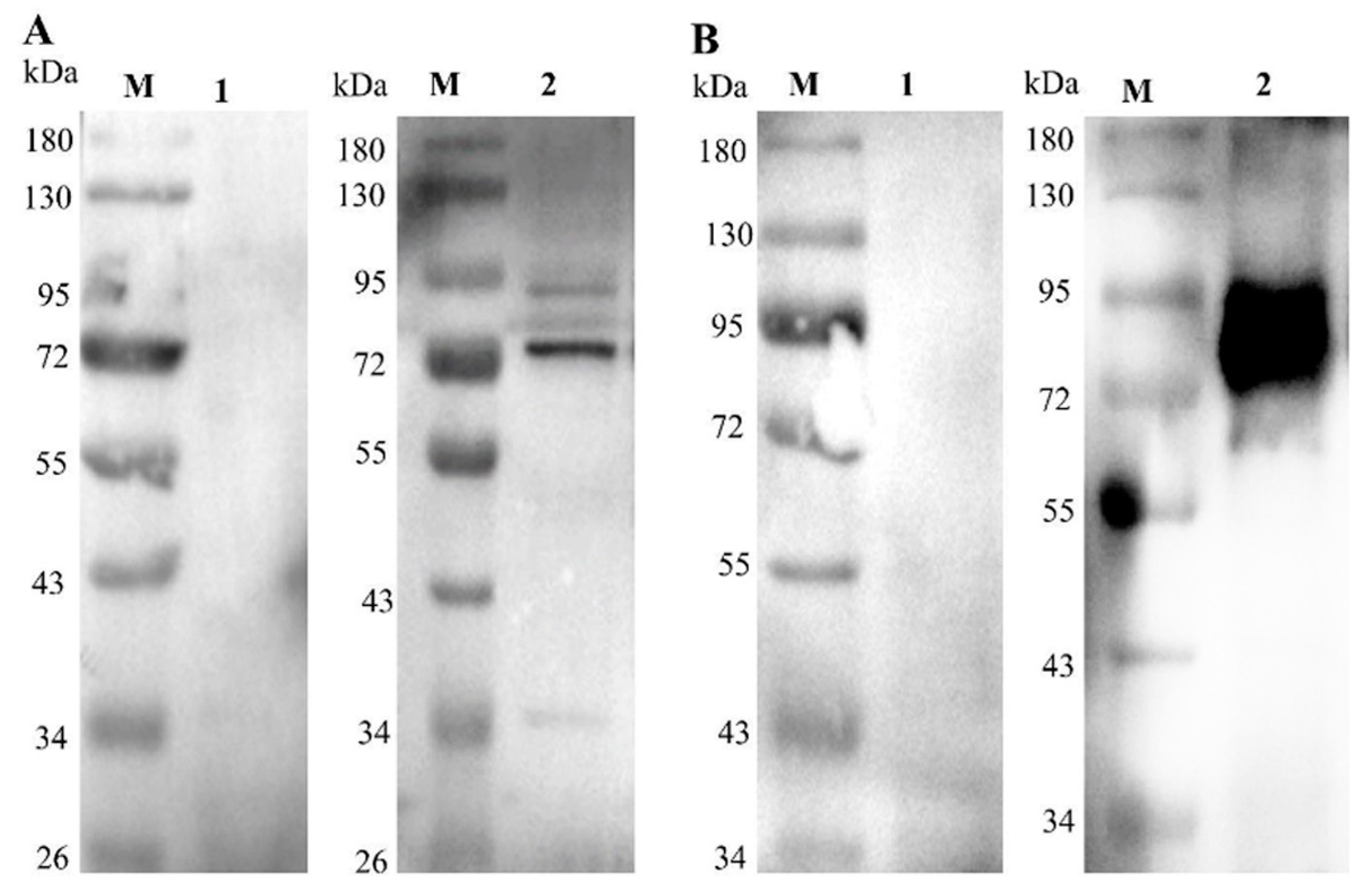
2.6. Verification of Secretion and Immunogenicity of Recombinant Proteins
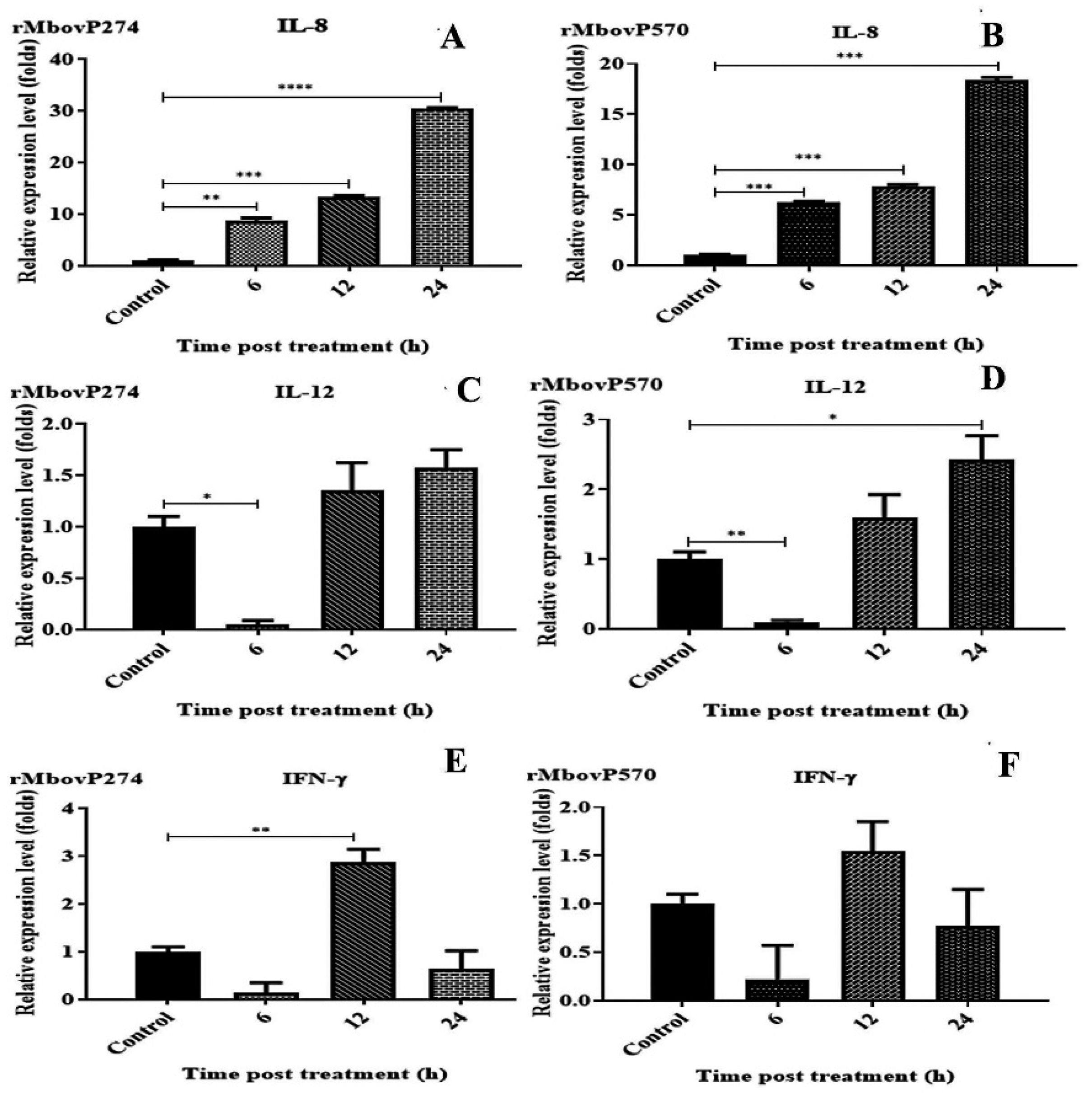
2.7. Inflammatory Cytokines Induced by Secreted rMbovP274 and rMbovP570
3. Discussion
3.1. Twenty-Two Secreted Immunogenic Proteins Identified
3.2. Secreted Proteins Differentially Expressed in Virulent and Attenuated M. bovis Strains
3.3. MbovP274 and MbovP570 Are Potential Protective Antigens
4. Materials and Methods
4.1. Ethics Statement
4.2. Secretome Data for M. bovis Strains Used in this Study
4.3. Prediction of Subcellular Localization of the Secretomes
4.4. Prediction of Protein Secretion and Antigenic Proteins
4.5. Prediction of B and T-Cell Epitopes
4.6. Prediction of Conformational B-Cell Epitopes Based on 3D Structure
4.7. Cloning and Expression of Immunogenic Candidates Proteins for Validation
4.8. Production of Polyclonal Antibodies against Recombinant Proteins
4.9. Validation of Secretion and Immunogenicity of the Recombinant Proteins
4.10. Quantitative PCR (qPCR) Detection of Inflammatory Cytokines Induced by Proteins
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Mustafa, R.; Qi, J.; Ba, X.; Chen, Y.; Hu, C.; Liu, X.; Tu, L.; Peng, Q.; Chen, H.; Guo, A. In vitro quinolones susceptibility analysis of Chinese Mycoplasma bovis isolates and their phylogenetic scenarios based upon QRDRs of DNA topoisomerases revealing a unique transition in ParC. Pak. Vet. J. 2013, 33, 364–369. [Google Scholar]
- Qi, J.; Guo, A.; Cui, P.; Chen, Y.; Mustafa, R.; Ba, X.; Hu, C.; Bai, Z.; Chen, X.; Shi, L.; et al. Comparative geno-plasticity analysis of Mycoplasma bovis HB0801 (Chinese isolate). PLoS ONE 2012, 7, e38239. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Han, X.; Chen, Y.; Mustafa, R.; Qi, J.; Chen, X.; Hu, C.; Chen, H.; Guo, A. Attenuated Mycoplasma bovis strains provide protection against virulent infection in calves. Vaccine 2014, 32, 3107–3114. [Google Scholar] [CrossRef]
- Nicholas, R.A.; Ayling, R.; McAuliffe, L. Vaccines for Mycoplasma diseases in animals and man. J. Comp. Pathol. 2009, 140, 85–96. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, S.; Gangopadhay, K.; Mukherjee, S.B. Identification of potential new vaccine candidates in Salmonella typhi using reverse vaccinology and subtractive genomics-based approach. bioRxiv 2019, 521518. [Google Scholar] [CrossRef]
- Mukherjee, P.; Mani, S. Methodologies to decipher the cell secretome. Biochim. Biophys. Acta (BBA) Proteins Proteom. 2013, 1834, 2226–2232. [Google Scholar] [CrossRef]
- Huntley, J.F.; Conley, P.G.; Hagman, K.E.; Norgard, M.V. Characterization of Francisella tularensis outer membrane proteins. J. Bacteriol. 2007, 189, 561–574. [Google Scholar] [CrossRef]
- Vafina, G.; Zainutdinova, E.; Bulatov, E.; Filimonova, M.N. Endonuclease from Gram-negative bacteria Serratia marcescens is as effective as pulmozyme in the hydrolysis of DNA in sputum. Front. Pharmacol. 2018, 9, 114. [Google Scholar] [CrossRef]
- Trajković, V. The role of mycobacterial secretory proteins in immune response in tuberculosis. Med. Pregl. 2004, 57, 25–28. [Google Scholar]
- Wang, Y.; Wang, X.; Ali, F.; Li, Z.; Fu, Y.; Yang, X.; Lin, W.; Lin, X.-M. Comparative extracellular proteomics of Aeromonas hydrophila reveals iron-regulated secreted proteins as potential vaccine candidates. Front. Immunol. 2019, 10, 256. [Google Scholar] [CrossRef]
- Vivek-Ananth, R.P.; Mohanraj, K.; Vandanashree, M.; Jhingran, A.; Craig, J.P.; Samal, A. Comparative systems analysis of the secretome of the opportunistic pathogen Aspergillus fumigatus and other Aspergillus species. Sci. Rep. 2018, 8, 6617. [Google Scholar] [CrossRef]
- Satchidanandam, V.; Kumar, N.; Biswas, S.; Jumani, R.S.; Jain, C.; Rani, R.; Aggarwal, B.; Singh, J.; Kotnur, M.R.; Sridharan, A. The secreted protein Rv1860 of Mycobacterium tuberculosis stimulates human polyfunctional CD8+ T cells. Clin. Vaccine Immunol. 2016, 23, 282–293. [Google Scholar] [CrossRef] [PubMed]
- Khan, F.A.; Faisal, M.; Chao, J.; Liu, K.; Chen, X.; Zhao, G.; Menghwar, H.; Zhang, H.; Zhu, X.; Rasheed, M.A.; et al. Immunoproteomic identification of MbovP579, a promising diagnostic biomarker for serological detection of Mycoplasma bovis infection. Oncotarget 2016, 7, 39376–39395. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhao, G.; Guo, Y.; Menghwar, H.; Chen, Y.; Chen, H.; Guo, A. Mycoplasma bovis MBOV_RS02825 encodes a secretory nuclease associated with cytotoxicity. Int. J. Mol. Sci. 2016, 17, 628. [Google Scholar] [CrossRef] [PubMed]
- Rasheed, M.A.; Qi, J.; Zhu, X.; Chenfei, H.; Menghwar, H.; Khan, F.A.; Zhao, G.; Zubair, M.; Hu, C.; Chen, Y.; et al. Comparative genomics of Mycoplasma bovis strains reveals that decreased virulence with increasing passages might correlate with potential virulence-related factors. Front. Cell. Infect. Microbiol. 2017, 7, 177. [Google Scholar] [CrossRef] [PubMed]
- Gondaira, S.; Higuchi, H.; Iwano, H.; Nakajima, K.; Kawai, K.; Hashiguchi, S.; Konnai, S.; Nagahata, H. Cytokine mRNA profiling and the proliferative response of bovine peripheral blood mononuclear cells to Mycoplasma bovis. Vet. Immunol. Immunopathol. 2015, 165, 45–53. [Google Scholar] [CrossRef]
- Gurung, R.B.; Purdie, A.C.; Begg, D.J.; Whittington, R.J. In silico identification of epitopes in Mycobacterium avium subsp. paratuberculosis proteins that were upregulated under stress conditions. Clin. Vaccine Immunol. 2012, 19, 855–864. [Google Scholar] [CrossRef]
- Ribas-Aparicio, R.M.; Arturo, C.J.; Alicia, J.A.; Paulina, M.G.; Gerardo, A.O. The impact of bioinformatics on vaccine design and development. In Vaccines; InTech: Rijeka, Croatia, 2017. [Google Scholar]
- Chen, X.; Huang, J.; Zhu, H.; Guo, Y.; Khan, F.A.; Menghwar, H.; Zhao, G.; Guo, A. P27 (MBOV_RS03440) is a novel fibronectin binding adhesin of Mycoplasma bovis. Int. J. Med. Microbiol. 2018, 308, 848–857. [Google Scholar] [CrossRef]
- Zhao, G.; Zhang, H.; Chen, X.; Zhu, X.; Guo, Y.; He, C.; Khan, F.A.; Chen, Y.; Hu, C.; Chen, H.; et al. Mycoplasma bovis NADH oxidase functions as both a NADH oxidizing and O 2 reducing enzyme and an adhesin. Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef]
- Zubair, M.; Muhamed, S.A.; Khan, F.A.; Zhao, G.; Menghwar, H.; Faisal, M.; Zhang, H.; Zhu, X.; Rasheed, M.A.; Chen, Y.; et al. Identification of 60 secreted proteins for Mycoplasma bovis with secretome assay. Microb. Pathog. 2020, 104135. [Google Scholar] [CrossRef]
- Wawegama, N.K.; Browning, G.F.; Kanci, A.; Marenda, M.S.; Markham, P.F. Development of a recombinant protein-based enzyme-linked immunosorbent assay for diagnosis of Mycoplasma bovis infection in cattle. Clin. Vaccine Immunol. 2014, 21, 196–202. [Google Scholar] [CrossRef]
- Chong, P.; Huang, J.-H.; Leng, C.-H.; Liu, S.-J.; Chen, H.-W. Recombinant lipoproteins as novel vaccines with intrinsic adjuvant. In Advances in Protein Chemistry and Structural Biology; Elsevier: Amsterdam, The Netherlands, 2015. [Google Scholar]
- Park, H.; Do, E.; Kim, M.; Park, H.-J.; Lee, J.; Han, S.-W. A LysR-type transcriptional regulator LcrX is involved in virulence, biofilm formation, swimming motility, siderophore secretion, and growth in sugar sources in Xanthomonas axonopodis pv. glycines. Front. Plant. Sci. 2019, 10, 1657. [Google Scholar] [CrossRef] [PubMed]
- Gondaira, S.; Higuchi, H.; Iwano, H.; Nishi, K.; Nebu, T.; Nakajima, K.; Nagahata, H. Innate immune response of bovine mammary epithelial cells to Mycoplasma bovis. J. Vet. Sci. 2018, 19, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Chao, J.; Han, X.; Liu, K.; Li, Q.; Peng, Q.; Lu, S.; Zhao, G.; Zhu, X.; Hu, G.; Dong, Y.; et al. Calves infected with virulent and attenuated Mycoplasma bovis strains have upregulated Th17 inflammatory and Th1 protective responses, respectively. Genes 2019, 10, 656. [Google Scholar] [CrossRef] [PubMed]
- Savojardo, C.; Martelli, P.L.; Fariselli, P.; Profiti, G.; Casadio, R. BUSCA: An integrative web server to predict subcellular localization of proteins. Nucleic Acids Res. 2018, 46, W459–W466. [Google Scholar] [CrossRef] [PubMed]
- Yu, N.Y.; Wagner, J.R.; Laird, M.R.; Melli, G.; Rey, S.; Lo, R.; Dao, P.; Sahinalp, S.C.; Ester, M.; Foster, L.J.; et al. PSORTb 3.0: Improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 2010, 26, 1608–1615. [Google Scholar] [CrossRef]
- Petersen, T.N.; Brunak, S.; Von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef]
- Bendtsen, J.D.; Jensen, L.J.; Blom, N.S.; Von Heijne, G.; Brunak, S. Feature-based prediction of non-classical and leaderless protein secretion. Protein Eng. Des. Sel. 2004, 17, 349–356. [Google Scholar] [CrossRef]
- Bagos, P.G.; Tsirigos, K.D.; Liakopoulos, T.D.; Hamodrakas, S.J. Prediction of lipoprotein signal peptides in Gram-positive bacteria with a Hidden Markov Model. J. Proteome Res. 2008, 7, 5082–5093. [Google Scholar] [CrossRef]
- Toseland, C.P.; Doytchinova, I.; Zaharieva, N.; Dimitrov, I. Immunogenicity prediction by VaxiJen: A ten year overview. J. Proteom. Bioinform. 2017, 10, 298–310. [Google Scholar]
- Doytchinova, I.; Toseland, C.P. VaxiJen: A server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 2007, 8, 4. [Google Scholar] [CrossRef] [PubMed]
- Singh, H.; Ansari, H.R.; Raghava, G.P. Improved method for linear B-cell epitope prediction using antigen’s primary sequence. PLoS ONE 2013, 8, e62216. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, M.; Lundegaard, C.; Worning, P.; Lauemøller, S.L.; Lamberth, K.; Buus, S.; Brunak, S.; Lund, O. Reliable prediction of T-cell epitopes using neural networks with novel sequence representations. Protein Sci. 2003, 12, 1007–1017. [Google Scholar] [CrossRef] [PubMed]
- Tenzer, S.; Peters, B.; Bulik, S.; Schoor, O.; Lemmel, C.; Schatz, M.M.; Kloetzel, P.-M.; Rammensee, H.-G.; Schild, H.; Holzhütter, H.-G. Modeling the MHC class I pathway by combining predictions of proteasomal cleavage, TAP transport and MHC class I binding. Cell. Mol. Life Sci. CMLS 2005, 62, 1025–1037. [Google Scholar] [CrossRef] [PubMed]
- Stabel, J.R.; Stabel, T.J. Immortalization and characterization of bovine peritoneal macrophages transfected with SV40 plasmid DNA. Vet. Immunol. Immunopathol. 1995, 45, 211–220. [Google Scholar] [CrossRef]
- O’Gorman, G.M.; Park, S.D.E.; Hill, E.W.; Meade, K.G.; Mitchell, L.C.; Agaba, M.; Gibson, J.P.; Hanotte, O.; Naessens, J.; Kemp, S.J.; et al. Cytokine mRNA profiling of peripheral blood mononuclear cells from trypanotolerant and trypanosusceptible cattle infected with Trypanosoma congolense. Physiol. Genom. 2006, 28, 53–61. [Google Scholar]

| No. | Proteins 1 | Signal Peptide 2 | Classical 3 | Non Classical 4 | Antigenic/Non-Antigenic 5 | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Mnemonic | Accession No. | GO Ids | GO Terms | Score | Overall Protective Antigen Prediction Score | Antigen/Non-Antigen | ||||
| 1 | Mbov_0016 a,b | AFM51394_1 | GO:0005615 | Extracellular | 0.92 | L | Y | Y | 0.5093 | A |
| 2 | Mbov_0037 a,b | AFM51415_2 | GO:0005615 | Extracellular | 0.87 | L | Y | Y | 0.3970 | NA |
| 3 | Mbov_0038 a,b | AFM51416_1 | GO:0005615 | Extracellular | 0.73 | T | N | Y | 0.4219 | A |
| 4 | Mbov_0049 a,b | AFM51424_1 | GO:0005615 | Extracellular | 0.88 | L | Y | Y | 0.5480 | A |
| 5 | Mbov_0111 a,b | AFM51486_1 | GO:0005615 | Extracellular | 0.77 | L | Y | Y | 0.5117 | A |
| 6 | Mbov_0154 a,b | AFM51527_1 | GO:0005615 | Extracellular | 0.96 | T | N | Y | 0.5622 | A |
| 7 | Mbov_0156 a,b | AFM51529_1 | GO:0005615 | Extracellular | 0.92 | L | Y | Y | 0.5078 | A |
| 8 | Mbov_0211 b | AFM51580_1 | GO:0005615 | Extracellular | 0.96 | L | Y | Y | 0.3222 | NA |
| 9 | Mbov_0217 a,b | AFM51586_1 | GO:0005615 | Extracellular | 0.96 | L | Y | Y | 0.4783 | A |
| 10 | Mbov_0274 a | AFM51642_1 | GO:0005615 | Extracellular | 0.91 | L | Y | Y | 0.5003 | A |
| 11 | Mbov_0283 b | AFM51651_1 | GO:0005615 | Extracellular | 0.96 | S | Y | Y | 0.9083 | A |
| 12 | Mbov_0290 b | AFM51658_1 | GO:0005615 | Extracellular | 0.91 | L | Y | Y | 0.4128 | A |
| 13 | Mbov_0296 b | AFM51664_1 | GO:0005615 | Extracellular | 0.96 | S | Y | Y | 0.4878 | A |
| 14 | Mbov_0326 b | AFM51694_1 | GO:0005615 | Extracellular | 0.96 | L | Y | Y | 0.3961 | NA |
| 15 | Mbov_0339 b | AFM51707_1 | GO:0005615 | Extracellular | 0.91 | L | Y | Y | 1.2671 | A |
| 16 | Mbov_0341 b | AFM51709_1 | GO:0005615 | Extracellular | 0.91 | S | Y | Y | 0.2139 | NA |
| 17 | Mbov_0350 a,b | AFM51716_1 | GO:0005615 | Extracellular | 0.97 | S | Y | Y | 0.4786 | A |
| 18 | Mbov_0364 a,b | AFM51726_1 | GO:0005615 | Extracellular | 0.96 | S | Y | Y | 0.4827 | A |
| 19 | Mbov_0368 b | AFM51730_1 | GO:0005615 | Extracellular | 0.97 | S | Y | Y | 0.5245 | A |
| 20 | Mbov_0374 b | AFM51736_1 | GO:0005615 | Extracellular | 0.97 | L | Y | Y | 0.4516 | A |
| 21 | Mbov_0393 a,b | AFM51750_1 | GO:0005615 | Extracellular | 0.96 | T | Y | Y | 0.3484 | NA |
| 22 | Mbov_0449 a,b | AFM51806_2 | GO:0005615 | Extracellular | 0.95 | S | Y | Y | 1.0254 | A |
| 23 | Mbov_0458 a,b | AFM51815_1 | GO:0005615 | Extracellular | 0.96 | L | Y | Y | 1.0635 | A |
| 24 | Mbov_0461 a,b | AFM51818_1 | GO:0005615 | Extracellular | 0.96 | S | Y | Y | 0.5130 | A |
| 25 | Mbov_0462 a,b | AFM51819_1 | GO:0005615 | Extracellular | 0.97 | L | Y | Y | 1.1146 | A |
| 26 | Mbov_0467 b | AFM51823_1 | GO:0005615 | Extracellular | 0.93 | T | N | Y | 0.3328 | NA |
| 27 | Mbov_0468 a,b | AFM51824_1 | GO:0005615 | Extracellular | 0.97 | L | Y | Y | 0.4890 | A |
| 28 | Mbov_0469 a,b | AFM51825_1 | GO:0005615 | Extracellular | 0.97 | L | Y | Y | 1.0196 | A |
| 29 | Mbov_0471 a,b | AFM51827_1 | GO:0005615 | Extracellular | 0.92 | S | Y | Y | 0.5445 | A |
| 30 | Mbov_0473 a,b | AFM51829_1 | GO:0005615 | Extracellular | 0.91 | L | Y | Y | 0.6593 | A |
| 31 | Mbov_0505 a,b | AFM51861_1 | GO:0005615 | Extracellular | 0.78 | S | Y | Y | 0.6022 | A |
| 32 | Mbov_0515 a,b | AFM51869_1 | GO:0005615 | Extracellular | 0.97 | L | Y | Y | 0.5203 | A |
| 33 | Mbov_0516 a,b | AFM51870_1 | GO:0005615 | Extracellular | 0.96 | S | Y | Y | 0.4863 | A |
| 34 | Mbov_0517 a,b | AFM51871_1 | GO:0005615 | Extracellular | 0.95 | S | Y | Y | 0.4955 | A |
| 35 | Mbov_0518 b | AFM51872_1 | GO:0005615 | Extracellular | 0.96 | S | Y | Y | 0.5014 | A |
| 36 | Mbov_0519 a,b | AFM51873_1 | GO:0005615 | Extracellular | 0.79 | T | Y | Y | 0.4463 | A |
| 37 | Mbov_0536 a,b | AFM51890_1 | GO:0005615 | Extracellular | 0.87 | L | Y | Y | 0.4537 | A |
| 38 | Mbov_0537 a,b | AFM51891_1 | GO:0005615 | Extracellular | 0.93 | L | Y | Y | 0.4844 | A |
| 39 | Mbov_0570 a,b | AFM51924_1 | GO:0005615 | Extracellular | 0.9 | L | Y | Y | 0.5094 | A |
| 40 | Mbov_0579 a,b | AFM51933_1 | GO:0005615 | Extracellular | 0.96 | L | Y | Y | 0.4661 | A |
| 41 | Mbov_0580 a,b | AFM51934_1 | GO:0005615 | Extracellular | 0.97 | S | Y | Y | 0.4848 | A |
| 42 | Mbov_0585 a,b | AFM51939_1 | GO:0005615 | Extracellular | 0.96 | S | Y | Y | 0.4742 | A |
| 43 | Mbov_0654 a,b | AFM52005_1 | GO:0005615 | Extracellular | 0.8 | L | Y | Y | 0.8413 | A |
| 44 | Mbov_0656 a,b | AFM52007_1 | GO:0005615 | Extracellular | 0.8 | NS | N | Y | 1.0586 | A |
| 45 | Mbov_0658 a,b | AFM52009_1 | GO:0005615 | Extracellular | 0.92 | S | Y | Y | 0.4564 | A |
| 46 | Mbov_0674 a,b | AFM52024_1 | GO:0005615 | Extracellular | 0.87 | T | Y | Y | 0.5553 | A |
| 47 | Mbov_0675 a,b | AFM52025_1 | GO:0005615 | Extracellular | 0.97 | L | Y | Y | 0.5009 | A |
| 48 | Mbov_0693 a,b | AFM52042_1 | GO:0005615 | Extracellular | 0.95 | T | N | Y | 0.2052 | NA |
| 49 | Mbov_0696 a,b | AFM52045_1 | GO:0005615 | Extracellular | 0.96 | S | Y | Y | 0.8248 | A |
| 50 | Mbov_0732 a | AFM52081_1 | GO:0005737 | Extracellular | 8.91 | NS | N | Y | 0.2937 | NA |
| 51 | Mbov_0739 a,b | AFM52087_1 | GO:0005615 | Extracellular | 0.96 | L | Y | Y | 0.5234 | A |
| 52 | Mbov_0743 a,b | AFM52091_1 | GO:0005615 | Extracellular | 0.82 | T | N | Y | 0.5336 | A |
| 53 | Mbov_0768 a,b | AFM52116_1 | GO:0005615 | Extracellular | 0.97 | S | Y | Y | 0.5745 | A |
| 54 | Mbov_0793 a,b | AFM52141_1 | GO:0005615 | Extracellular | 0.9 | L | Y | Y | 1.1056 | A |
| 55 | Mbov_0794 a,b | AFM52142_1 | GO:0005615 | Extracellular | 0.9 | L | Y | Y | 0.8247 | A |
| 56 | Mbov_0795 a | AFM52143_1 | GO:0005615 | Extracellular | 0.93 | L | Y | Y | 0.3425 | NA |
| 57 | Mbov_0796 a,b | AFM52144_1 | GO:0005615 | Extracellular | 0.95 | L | Y | Y | 0.8931 | A |
| 58 | Mbov_0797 a,b | AFM52145_1 | GO:0005615 | Extracellular | 0.96 | L | Y | Y | 1.1257 | A |
| 59 | Mbov_0798 a,b | AFM52146_1 | GO:0005615 | Extracellular | 0.96 | L | Y | Y | 0.8682 | A |
| No. | Proteins | Epitopes | ||||
|---|---|---|---|---|---|---|
| Mnemonic | Accession No. | Protein | Over All B Cell Epitopes Numbers | B-Cells | T-Cell (MHC Class I & II) | |
| 1 | Mbov_0016 a,b | AFM51394_1 | P48-like surface lipoprotein | 22 | + | +(I,II) |
| 2 | Mbov_0038 a,b | AFM51416_1 | putative transmembrane protein | 161 | + | +(I,II) |
| 3 | Mbov_0049 a,b | AFM51424_1 | putative lipoprotein | 34 | + | +(I,II) |
| 4 | Mbov_0111 a,b | AFM51486_1 | putative lipoprotein | 53 | + | +(I,II) |
| 5 | Mbov_0154 a,b | AFM51527_1 | putative transmembrane protein | 24 | + | +(I,II) |
| 6 | Mbov_0217 a,b | AFM51586_1 | putative lipoprotein | 21 | + | +(I,II) |
| 7 | Mbov_0274 a | AFM51642_1 | putative lipoprotein | 35 | + | +(I,II) |
| 8 | Mbov_0290 b | AFM51658_1 | putative lipoprotein | 26 | + | - (I,II) |
| 9 | Mbov_0296 b | AFM51664_1 | putative lipoprotein | 30 | + | +(I,II) |
| 10 | Mbov_0350 a,b | AFM51716_1 | putative lipoprotein | 36 | + | - (I,II) |
| 11 | Mbov_0364 a,b | AFM51726_1 | putative membrane lipoprotein | 25 | + | - (I,II) |
| 12 | Mbov_0374 b | AFM51736_1 | putative lipoprotein | 38 | + | - (I,II) |
| 13 | Mbov_0468 a,b | AFM51824_1 | putative lipoprotein | 24 | + | +(I,II) |
| 14 | Mbov_0471 a,b | AFM51827_1 | Periplasmic protease | 30 | + | +(I,II) |
| 15 | Mbov_0505 a,b | AFM51861_1 | putative lipoprotein | 38 | + | +(I,II) |
| 16 | Mbov_0515 a,b | AFM51869_1 | putative lipoprotein | 52 | + | +(I,II) |
| 17 | Mbov_0516 a,b | AFM51870_1 | Putative transmembrane protein | 46 | + | +(I,II) |
| 18 | Mbov_0517 a,b | AFM51871_1 | Putative transmembrane protein | 42 | + | +(I,II) |
| 19 | Mbov_0518 b | AFM51872_1 | putative lipoprotein | 45 | + | +(I,II) |
| 20 | Mbov_0519 a,b | AFM51873_1 | Putative transmembrane protein | 41 | + | +(I,II) |
| 21 | Mbov_0536 a,b | AFM51890_1 | putative lipoprotein | 19 | + | +(I,II) |
| 22 | Mbov_0570 a,b | AFM51924_1 | putative lipoprotein | 34 | + | +(I,II) |
| 23 | Mbov_0579 a,b | AFM51933_1 | membrane lipoprotein P81 | 39 | + | +(I,II) |
| 24 | Mbov_0580 a,b | AFM51934_1 | nuclease | 18 | + | +(I,II) |
| 25 | Mbov_0585 a,b | AFM51939_1 | putative lipoprotein | 18 | + | +(I,II) |
| 26 | Mbov_0658 a,b | AFM52009_1 | Periplasmic protease | 29 | + | +(I,II) |
| 27 | Mbov_0674 a,b | AFM52024_1 | putative lipoprotein | 18 | + | +(I,II) |
| 28 | Mbov_0675 a,b | AFM52025_1 | 5′nucleotidase | 36 | + | +(I,II) |
| 29 | Mbov_0739 a,b | AFM52087_1 | putative lipoprotein | 38 | + | +(I,II) |
| 30 | Mbov_0743 a,b | AFM52091_1 | putative transmembrane protein | 48 | + | +(I,II) |
| 31 | Mbov_0798 a,b | AFM52146_1 | variable surface lipoprotein VspHB0801-6 | 22 | + | +(I,II) |
| Antigenic Proteins of M. bovis HB0801 with NCBI Protein ID | Protein Name | Domains | Name | Accession No. |
|---|---|---|---|---|
| Putative lipoprotein (MbovP049 a,b) AFM51424_1 | Putative lipoprotein | 330-636 | SMC_N super family | cl25732 |
| Putative lipoprotein (MbovP274 a) AFM51642_1 | Putative lipoprotein | 1-267 | UgpB super family | cl25886 |
| Putative lipoprotein (MbovP296 b) AFM51664_1 | Putative lipoprotein | 450-576 | Peptidase_S41 super family | cl02526 |
| Putative lipoprotein (MbovP570 a,b) AFM51924_1 | Putative lipoprotein | 458-624 | HemL super family | cl28400 |
| putative lipoprotein (MbovP585 a,b) AFM51939_1 | Putative lipoprotein | 364-499 | SMC_N super family | cl25732 |
| Gene Name | Primer Sequence (5′-3′) | Amplicon Size (bp) | Accession Number | Annealing Temperature (°C) | References |
|---|---|---|---|---|---|
| IL-1β | F: GTCATCTTCGAAACGTCCTCC R: TCCTCTCCTTGCACAAAGCTC | 191 | M37211 | 60 | [14] |
| IL-6 | F: ACCCCAGGCAGACTACTTCT R: CCCAGATTGGAAGCATCCGT | 195 | NM173923.2 | 60 | [14] |
| IL-12 | F: GCTTGGAGCACAGGGAGTAT R: AGTTGCAGGTTCTTGGGTGG | 151 | NM174356.1 | 60 | [14] |
| TNF- | F: CTCCATCAACAGCCCTCTGG R: GAGGGCATTGGCATACGAGT | 136 | NM173966 | 60 | [14] |
| IFN-γ | F: TCAAATTCCGGTGGATGATCTGC R: GACCATTACGTTGATGCTCTCCG | 150 | NM 174086.1 | 60 | [38] |
| IL-8 | F: GAAGAGAGCTGAGAAGCAAGATCC R: ACCCACACAGAACATGAGGC | 142 | NM173925.2 | 60 | [14] |
| β-Actin | F: AGCAAGCAGGAGTACGATGAG R: ATCCAACCGACTGCTGTCA | 241 | NM 173979.3 | 60 | [14] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shirani, I.; Zhang, H.; Zhao, G.; Lu, S.; Marawan, M.A.; Dawood, A.; Chen, Y.; Chen, X.; Chen, J.; Hu, C.; et al. In Silico Identification of Novel Immunogenic Secreted Proteins of Mycoplasma bovis from Secretome Data and Experimental Verification. Pathogens 2020, 9, 770. https://doi.org/10.3390/pathogens9090770
Shirani I, Zhang H, Zhao G, Lu S, Marawan MA, Dawood A, Chen Y, Chen X, Chen J, Hu C, et al. In Silico Identification of Novel Immunogenic Secreted Proteins of Mycoplasma bovis from Secretome Data and Experimental Verification. Pathogens. 2020; 9(9):770. https://doi.org/10.3390/pathogens9090770
Chicago/Turabian StyleShirani, Ihsanullah, Hui Zhang, Gang Zhao, Siyi Lu, Marawan A Marawan, Ali Dawood, Yingyu Chen, Xi Chen, Jianguo Chen, Changmin Hu, and et al. 2020. "In Silico Identification of Novel Immunogenic Secreted Proteins of Mycoplasma bovis from Secretome Data and Experimental Verification" Pathogens 9, no. 9: 770. https://doi.org/10.3390/pathogens9090770
APA StyleShirani, I., Zhang, H., Zhao, G., Lu, S., Marawan, M. A., Dawood, A., Chen, Y., Chen, X., Chen, J., Hu, C., Chen, H., & Guo, A. (2020). In Silico Identification of Novel Immunogenic Secreted Proteins of Mycoplasma bovis from Secretome Data and Experimental Verification. Pathogens, 9(9), 770. https://doi.org/10.3390/pathogens9090770








