SARS-CoV-2 and Coronavirus Disease 2019: What We Know So Far
Abstract
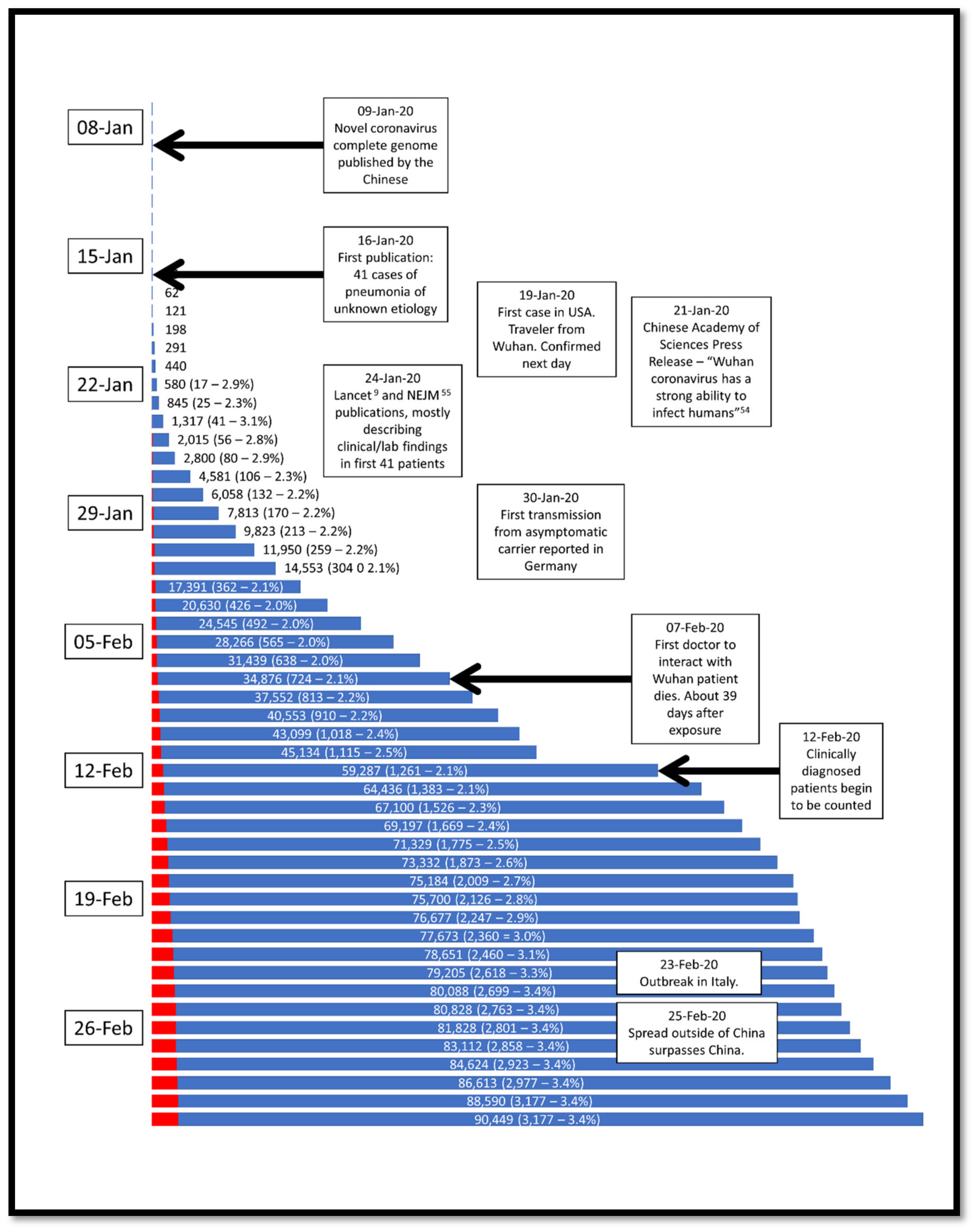
1. Introduction
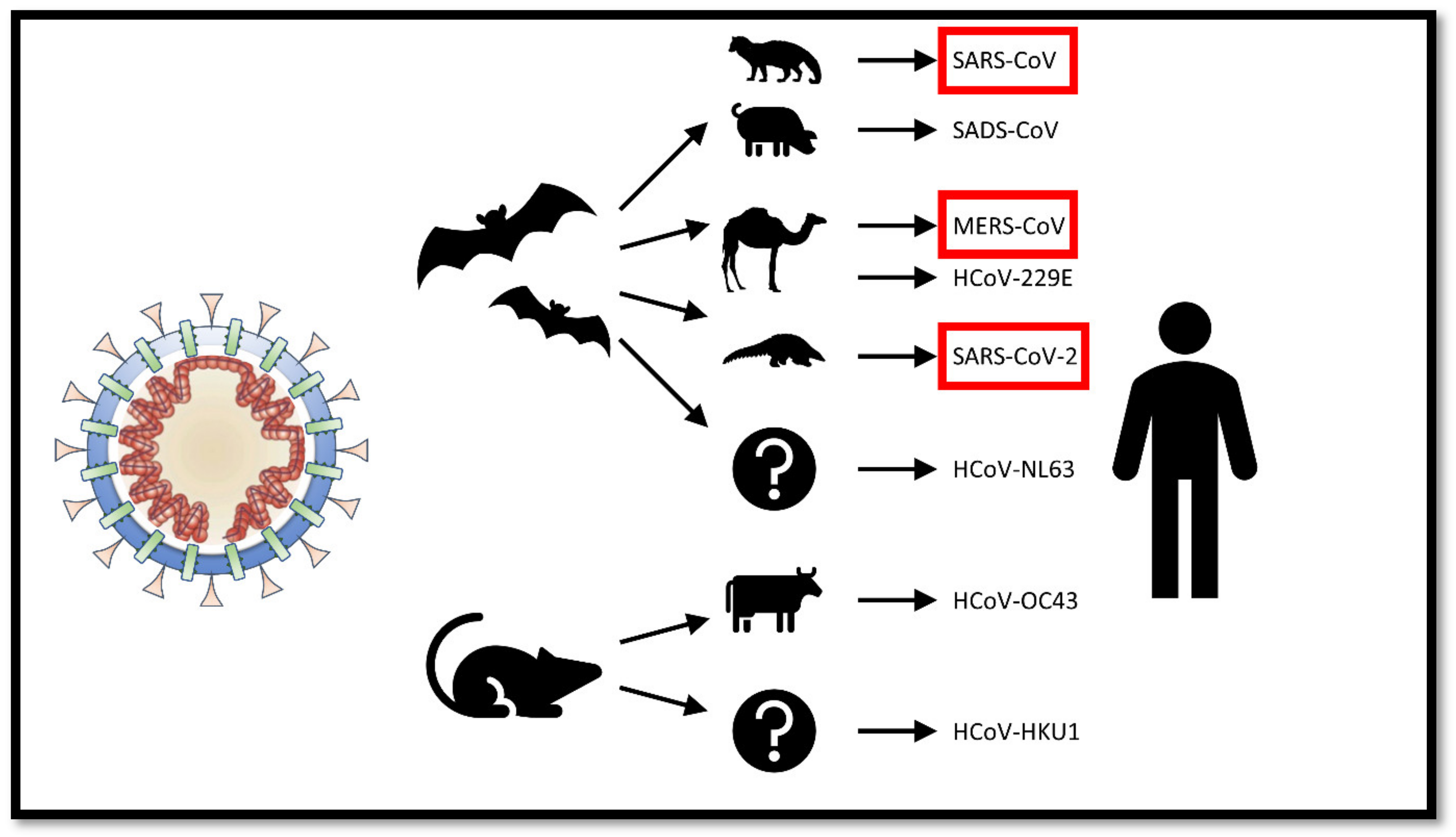
2. The Origin of SARS-CoV-2
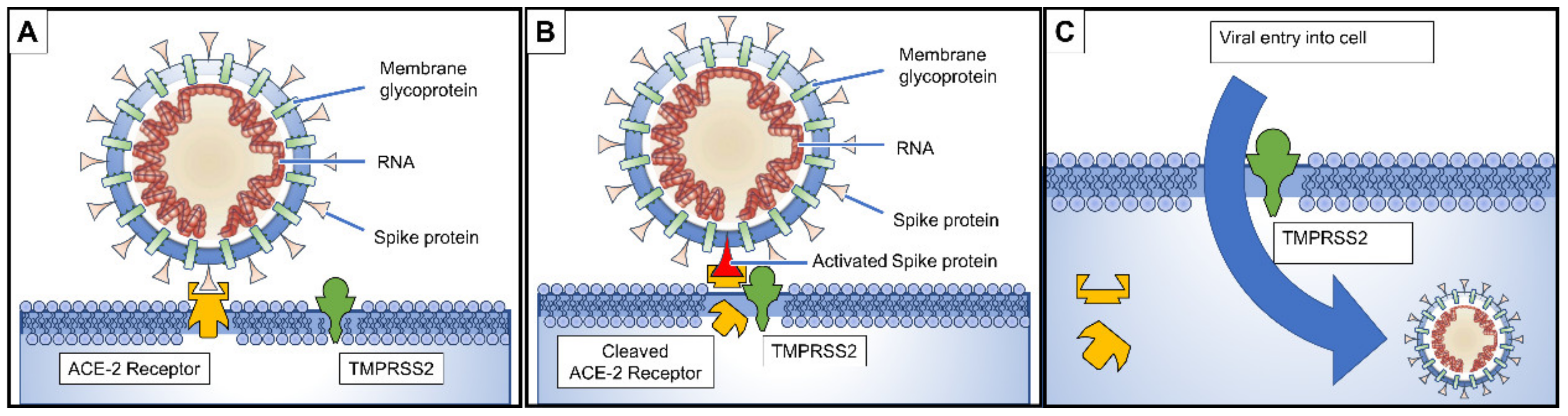
3. Pathogenesis and Clinical Presentation
4. Epidemiology
4.1. Transmission Rate
4.2. Incubation Period
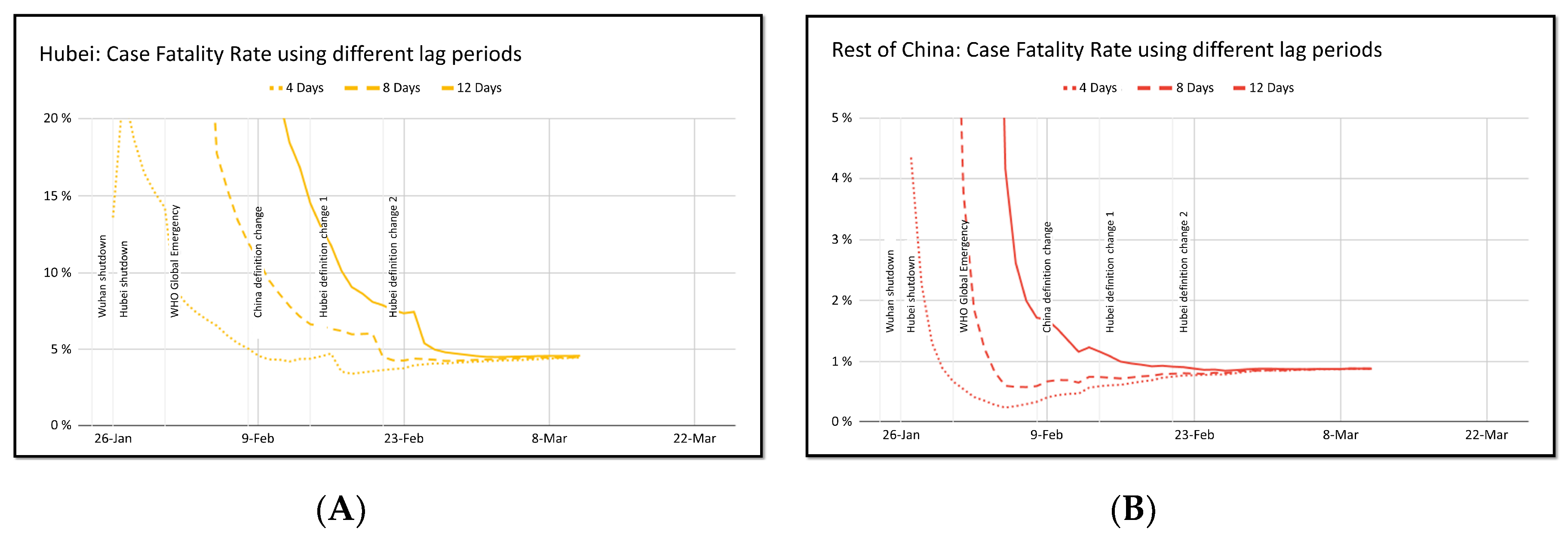
4.3. Case Fatality Rate
4.4. Asymptomatic Transmission
5. Risk Factors for Mortality
6. Treatment
7. Vaccine Development
Author Contributions
Funding
Conflicts of Interest
References
- Coronaviridae—Positive Sense RNA Viruses—Positive Sense RNA Viruses. 2011. Available online: https://talk.ictvonline.org/ictv-reports/ictv_9th_report/positive-sense-rna-viruses-2011/w/posrna_viruses/222/coronaviridae (accessed on 15 March 2020).
- Cui, J.; Li, F.; Shi, Z.-L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Fan, H.; Lan, T.; Yang, X.-L.; Shi, W.-F.; Zhang, W.; Zhu, Y.; Zhang, Y.-W.; Xie, Q.-M.; Mani, S.; et al. Fatal swine acute diarrhoea syndrome caused by an HKU2-related coronavirus of bat origin. Nature 2018, 556, 255–258. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Gorbalenya, A.E.; Baker, S.C.; Baric, R.S.; de Groot, R.J.; Drosten, C.; Gulyaeva, A.A.; Haagmans, B.L.; Lauber, C.; Leontovich, A.M.; Neuman, B.W.; et al. Severe acute respiratory syndrome-related coronavirus: The species and its viruses—A statement of the Coronavirus Study Group. Microbiology 2020. [Google Scholar] [CrossRef]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef]
- Lu, H.; Stratton, C.W.; Tang, Y. Outbreak of pneumonia of unknown etiology in Wuhan, China: The mystery and the miracle. J. Med. Virol. 2020, 92, 401–402. [Google Scholar] [CrossRef]
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: A descriptive study. Lancet 2020, 395, 507–513. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Tan, W.; Zhao, X.; Ma, X.; Wang, W.; Niu, P.; Xu, W.; Gao, G.F.; Wu, G. A Novel Coronavirus Genome Identified in a Cluster of Pneumonia Cases—Wuhan, China 2019−2020. China CDC Wkly. 2020, 2, 61–62. [Google Scholar]
- Johns Hopkins University Coronavirus COVID-19 (2019-nCoV). Available online: https://gisanddata.maps.arcgis.com/apps/opsdashboard/index.html#/bda7594740fd40299423467b48e9ecf6 (accessed on 25 February 2020).
- China CDC Weekly. Available online: http://weekly.chinacdc.cn/news/TrackingtheEpidemic.htm (accessed on 25 February 2020).
- Worldometer: Coronavirus Update (Live). Available online: https://www.worldometers.info/coronavirus/ (accessed on 25 February 2020).
- Klassen, D. Updated COVID-19 Statistics. Available online: https://nucleuswealth.com/articles/updated-coronavirus-statistics-cases-deaths-mortality-rate/ (accessed on 3 March 2020).
- WHO Director-General’s Opening Remarks at the Media Briefing on COVID-19—11 March 2020. Available online: https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020 (accessed on 16 March 2020).
- Fan, Y.; Zhao, K.; Shi, Z.-L.; Zhou, P. Bat Coronaviruses in China. Viruses 2019, 11, 210. [Google Scholar] [CrossRef]
- Cyranoski, D. Did pangolins spread the China coronavirus to people? Nature 2020. [Google Scholar] [CrossRef]
- Ksiazek, T.G.; Erdman, D.; Goldsmith, C.S.; Zaki, S.R.; Peret, T.; Emery, S.; Tong, S.; Urbani, C.; Comer, J.A.; Lim, W.; et al. A Novel Coronavirus Associated with Severe Acute Respiratory Syndrome. N. Engl. J. Med. 2003, 348, 1953–1966. [Google Scholar] [CrossRef] [PubMed]
- Andersen, K.; Rambaut, A.; Lipkin, W.I.; Holmes, E.C.; Garry, R.F. The Proximal Origin of SARS-CoV-2. Available online: http://virological.org/t/the-proximal-origin-of-sars-cov-2/398 (accessed on 25 February 2020).
- Kuba, K.; Imai, Y.; Rao, S.; Gao, H.; Guo, F.; Guan, B.; Huan, Y.; Yang, P.; Zhang, Y.; Deng, W.; et al. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus–induced lung injury. Nat. Med. 2005, 11, 875–879. [Google Scholar] [CrossRef] [PubMed]
- Glowacka, I.; Bertram, S.; Muller, M.A.; Allen, P.; Soilleux, E.; Pfefferle, S.; Steffen, I.; Tsegaye, T.S.; He, Y.; Gnirss, K.; et al. Evidence that TMPRSS2 Activates the Severe Acute Respiratory Syndrome Coronavirus Spike Protein for Membrane Fusion and Reduces Viral Control by the Humoral Immune Response. J. Virol. 2011, 85, 4122–4134. [Google Scholar] [CrossRef]
- Heurich, A.; Hofmann-Winkler, H.; Gierer, S.; Liepold, T.; Jahn, O.; Pohlmann, S. TMPRSS2 and ADAM17 Cleave ACE2 Differentially and Only Proteolysis by TMPRSS2 Augments Entry Driven by the Severe Acute Respiratory Syndrome Coronavirus Spike Protein. J. Virol. 2014, 88, 1293–1307. [Google Scholar] [CrossRef]
- Shulla, A.; Heald-Sargent, T.; Subramanya, G.; Zhao, J.; Perlman, S.; Gallagher, T. A Transmembrane Serine Protease Is Linked to the Severe Acute Respiratory Syndrome Coronavirus Receptor and Activates Virus Entry. J. Virol. 2011, 85, 873–882. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. Discovery of a novel coronavirus associated with the recent pneumonia outbreak in humans and its potential bat origin. Microbiology 2020. [Google Scholar] [CrossRef]
- Schnirring, L. More Outbreak Details Emerge as COVID-19 Cases top 70,000. Available online: http://www.cidrap.umn.edu/news-perspective/2020/02/more-outbreak-details-emerge-covid-19-cases-top-70000 (accessed on 25 February 2020).
- The Epidemiological Characteristics of an Outbreak of 2019 Novel Coronavirus Diseases (COVID-19)—China 2020. Available online: http://weekly.chinacdc.cn/en/article/id/e53946e2-c6c4-41e9-9a9b-fea8db1a8f51 (accessed on 25 February 2020).
- PM Abe Asks All of Japan Schools to Close Over coronavirus. Reuters. 2020. Available online: https://www.reuters.com/article/us-china-health-japan-idUSKCN20L0BI (accessed on 27 February 2020).
- CNN World; Yeung, J.; Marsh, J.; Kottasová, I.; Vera, A. March 15 Coronavirus News. Available online: https://www.cnn.com/world/live-news/coronavirus-outbreak-2-03-15-20-intl-hnk/index.html (accessed on 16 March 2020).
- CDC Coronavirus Disease 2019 (COVID-19)—Resources for K-12 Schools and Childcare Programs. Available online: https://www.cdc.gov/coronavirus/2019-ncov/community/schools-childcare/index.html (accessed on 16 March 2020).
- Delamater, P.L.; Street, E.J.; Leslie, T.F.; Yang, Y.T.; Jacobsen, K.H. Complexity of the Basic Reproduction Number (R0). Emerg. Infect. Dis. J. 2019, 25. [Google Scholar] [CrossRef]
- Zhao, S.; Lin, Q.; Ran, J.; Musa, S.S.; Yang, G.; Wang, W.; Lou, Y.; Gao, D.; Yang, L.; He, D.; et al. Preliminary estimation of the basic reproduction number of novel coronavirus (2019-nCoV) in China, from 2019 to 2020: A data-driven analysis in the early phase of the outbreak. Int. J. Infect. Dis. 2020, 92, 214–217. [Google Scholar] [CrossRef]
- WHO. Novel Coronavirus—China. Available online: http://www.who.int/csr/don/12-january-2020-novel-coronavirus-china/en/ (accessed on 25 February 2020).
- CDC Coronavirus Disease 2019 (COVID-19). Available online: https://www.cdc.gov/coronavirus/2019-ncov/about/transmission.html (accessed on 3 March 2020).
- Guerra, F.M.; Bolotin, S.; Lim, G.; Heffernan, J.; Deeks, S.L.; Li, Y.; Crowcroft, N.S. The basic reproduction number (R0) of measles: A systematic review. Lancet Infect. Dis. 2017, 17, e420–e428. [Google Scholar] [CrossRef]
- Marangi, L.; Mirinaviciute, G.; Flem, E.; Tomba, G.S.; Guzzetta, G.; Blasio, B.F.; de Manfredi, P. The natural history of varicella zoster virus infection in Norway: Further insights on exogenous boosting and progressive immunity to herpes zoster. PLoS ONE 2017, 12, e0176845. [Google Scholar] [CrossRef] [PubMed]
- Kampf, G.; Todt, D.; Pfaender, S.; Steinmann, E. Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. J. Hosp. Infect. 2020, 104, 246–251. [Google Scholar] [CrossRef] [PubMed]
- van Doremalen, N.; Bushmaker, T.; Karesh, W.B.; Munster, V.J. Stability of Middle East respiratory syndrome coronavirus in milk. Emerg. Infect. Dis. 2014, 20, 1263–1264. [Google Scholar] [CrossRef] [PubMed]
- Warnes, S.L.; Little, Z.R.; Keevil, C.W. Human Coronavirus 229E Remains Infectious on Common Touch Surface Materials. mBio 2015, 6, e01697-15. [Google Scholar] [CrossRef] [PubMed]
- Yeung, J. China is Literally Laundering its Money to Contain the Coronavirus—CNN. Available online: https://edition.cnn.com/2020/02/17/asia/china-is-disinfecting-cash-coronavirus-intl-hnk-scli/index.html (accessed on 25 February 2020).
- Zou, L.; Ruan, F.; Huang, M.; Liang, L.; Huang, H.; Hong, Z.; Yu, J.; Kang, M.; Song, Y.; Xia, J.; et al. SARS-CoV-2 Viral Load in Upper Respiratory Specimens of Infected Patients. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.M.; Lau, E.H.Y.; Wong, J.Y.; et al. Early Transmission Dynamics in Wuhan, China, of Novel Coronavirus–Infected Pneumonia. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- Guan, W.; Ni, Z.; Hu, Y.; Liang, W.; Ou, C.; He, J.; Liu, L.; Shan, H.; Lei, C.; Hui, D.S.; et al. Clinical characteristics of 2019 novel coronavirus infection in China. Respir. Med. 2020. [Google Scholar] [CrossRef]
- Backer, J.A.; Klinkenberg, D.; Wallinga, J. Incubation period of 2019 novel coronavirus (2019-nCoV) infections among travellers from Wuhan, China, 20–28 January 2020. Eurosurveillance 2020, 25, 2000062. [Google Scholar] [CrossRef]
- Coronavirus: Wuhan Speeds Up Testing and Treatment of Patients, East Asia News & Top Stories—The Straits Times. Available online: https://www.straitstimes.com/asia/east-asia/coronavirus-wuhan-speeds-up-testing-and-treatment-of-patients (accessed on 15 March 2020).
- Bai, Y.; Yao, L.; Wei, T.; Tian, F.; Jin, D.-Y.; Chen, L.; Wang, M. Presumed Asymptomatic Carrier Transmission of COVID-19. JAMA 2020. [Google Scholar] [CrossRef]
- Rothe, C.; Schunk, M.; Sothmann, P.; Bretzel, G.; Froeschl, G.; Wallrauch, C.; Zimmer, T.; Thiel, V.; Janke, C.; Guggemos, W.; et al. Transmission of 2019-nCoV Infection from an Asymptomatic Contact in Germany. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- Lipsitch, M.; Swerdlow, D.L.; Finelli, L. Defining the Epidemiology of Covid-19—Studies Needed. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Zhao, Z.; Wang, Y.; Zhou, Y.; Ma, Y.; Zuo, W. Single-cell RNA expression profiling of ACE2, the putative receptor of Wuhan 2019-nCov. Bioinformatics 2020. [Google Scholar] [CrossRef]
- Cai, G. Tobacco-Use Disparity in Gene Expression of ACE2, the Receptor of 2019-nCov. Life Sci. 2020. [Google Scholar] [CrossRef]
- Liu, S.; Zhang, M.; Yang, L.; Li, Y.; Wang, L.; Huang, Z.; Wang, L.; Chen, Z.; Zhou, M. Prevalence and patterns of tobacco smoking among Chinese adult men and women: Findings of the 2010 national smoking survey. J. Epidemiol. Community Health 2017, 71, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Wei, M.; Yuan, J.; Liu, Y.; Fu, T.; Yu, X.; Zhang, Z.-J. Novel Coronavirus Infection in Hospitalized Infants Under 1 Year of Age in China. JAMA 2020. [Google Scholar] [CrossRef]
- Wax, R.S.; Christian, M.D. Practical recommendations for critical care and anesthesiology teams caring for novel coronavirus (2019-nCoV) patients. Can. J. Anesth. Can. Anesth. 2020. [Google Scholar] [CrossRef]
- Chu, C.M. Role of lopinavir/ritonavir in the treatment of SARS: Initial virological and clinical findings. Thorax 2004, 59, 252–256. [Google Scholar] [CrossRef]
- Sheahan, T.P.; Sims, A.C.; Graham, R.L.; Menachery, V.D.; Gralinski, L.E.; Case, J.B.; Leist, S.R.; Pyrc, K.; Feng, J.Y.; Trantcheva, I.; et al. Broad-spectrum antiviral GS-5734 inhibits both epidemic and zoonotic coronaviruses. Sci. Transl. Med. 2017, 9, eaal3653. [Google Scholar] [CrossRef]
- Mulangu, S.; Dodd, L.E.; Davey, R.T.; Tshiani Mbaya, O.; Proschan, M.; Mukadi, D.; Lusakibanza Manzo, M.; Nzolo, D.; Tshomba Oloma, A.; Ibanda, A.; et al. A Randomized, Controlled Trial of Ebola Virus Disease Therapeutics. N. Engl. J. Med. 2019, 381, 2293–2303. [Google Scholar] [CrossRef]
- Wang, M.; Cao, R.; Zhang, L.; Yang, X.; Liu, J.; Xu, M.; Shi, Z.; Hu, Z.; Zhong, W.; Xiao, G. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020, 30, 269–271. [Google Scholar] [CrossRef]
- Holshue, M.L.; DeBolt, C.; Lindquist, S.; Lofy, K.H.; Wiesman, J.; Bruce, H.; Spitters, C.; Ericson, K.; Wilkerson, S.; Tural, A.; et al. First Case of 2019 Novel Coronavirus in the United States. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- NIH (National Institute of Allergy and Infectious Diseases). NIH Clinical Trial of Remdesivir to Treat COVID-19 Begins. Available online: https://www.niaid.nih.gov/news-events/nih-clinical-trial-remdesivir-treat-covid-19-begins (accessed on 27 February 2020).
- Gilead Sciences Initiates Two Phase 3 Studies of Investigational Antiviral Remdesivir for the Treatment of COVID-19. Available online: https://www.gilead.com/news-and-press/press-room/press-releases/2020/2/gilead-sciences-initiates-two-phase-3-studies-of-investigational-antiviral-remdesivir-for-the-treatment-of-covid-19 (accessed on 27 February 2020).
- Vincent, M.J.; Bergeron, E.; Benjannet, S.; Erickson, B.R.; Rollin, P.E.; Ksiazek, T.G.; Seidah, N.G.; Nichol, S.T. Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virol. J. 2005, 2, 69. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Vedantham, P.; Lu, K.; Agudelo, J.; Carrion, R.; Nunneley, J.W.; Barnard, D.; Pöhlmann, S.; McKerrow, J.H.; Renslo, A.R.; et al. Protease inhibitors targeting coronavirus and filovirus entry. Antiviral Res. 2015, 116, 76–84. [Google Scholar] [CrossRef] [PubMed]
- Yamawaki, H.; Futagami, S.; Kaneko, K.; Agawa, S.; Higuchi, K.; Murakami, M.; Wakabayashi, M.; Sakasegawa, N.; Kodaka, Y.; Ueki, N.; et al. Camostat Mesilate, Pancrelipase, and Rabeprazole Combination Therapy Improves Epigastric Pain in Early Chronic Pancreatitis and Functional Dyspepsia with Pancreatic Enzyme Abnormalities. Digestion 2019, 99, 283–292. [Google Scholar] [CrossRef] [PubMed]
- Ramsey, M.L.; Nuttall, J.; Hart, P.A. TACTIC Investigative Team a phase 1/2 trial to evaluate the pharmacokinetics, safety, and efficacy of NI-03 in patients with chronic pancreatitis: Study protocol for a randomized controlled trial on the assessment of camostat treatment in chronic pancreatitis (TACTIC). Trials 2019, 20, 501. [Google Scholar] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Krüger, N.; Müller, M.; Drosten, C.; Pöhlmann, S. The novel coronavirus 2019 (2019-nCoV) uses the SARS-coronavirus receptor ACE2 and the cellular protease TMPRSS2 for entry into target cells. bioRxiv 2020. [Google Scholar] [CrossRef]
- Kraft, C.S.; Hewlett, A.L.; Koepsell, S.; Winkler, A.M.; Kratochvil, C.J.; Larson, L.; Varkey, J.B.; Mehta, A.K.; Lyon, G.M.; Friedman-Moraco, R.J.; et al. The Use of TKM-100802 and Convalescent Plasma in 2 Patients with Ebola Virus Disease in the United States. Clin. Infect. Dis. 2015, 61, 496–502. [Google Scholar] [CrossRef]
- Walker, L.M.; Burton, D.R. Passive immunotherapy of viral infections: “super-antibodies” enter the fray. Nat. Rev. Immunol. 2018, 18, 297–308. [Google Scholar] [CrossRef]
- Wong, S.K.; Li, W.; Moore, M.J.; Choe, H.; Farzan, M. A 193-Amino Acid Fragment of the SARS Coronavirus S Protein Efficiently Binds Angiotensin-converting Enzyme 2. J. Biol. Chem. 2004, 279, 3197–3201. [Google Scholar] [CrossRef]
- Li, W.; Moore, M.J.; Vasilieva, N.; Sui, J.; Wong, S.K.; Berne, M.A.; Somasundaran, M.; Sullivan, J.L.; Luzuriaga, K.; Greenough, T.C.; et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 2003, 426, 450–454. [Google Scholar] [CrossRef]
- Elshabrawy, H.A.; Coughlin, M.M.; Baker, S.C.; Prabhakar, B.S. Human Monoclonal Antibodies against Highly Conserved HR1 and HR2 Domains of the SARS-CoV Spike Protein Are More Broadly Neutralizing. PLoS ONE 2012, 7, e50366. [Google Scholar] [CrossRef] [PubMed]
- Arbabi-Ghahroudi, M. Camelid Single-Domain Antibodies: Historical Perspective and Future Outlook. Front. Immunol. 2017, 8, 1589. [Google Scholar] [CrossRef] [PubMed]
- Moore, M.J.; Dorfman, T.; Li, W.; Wong, S.K.; Li, Y.; Kuhn, J.H.; Coderre, J.; Vasilieva, N.; Han, Z.; Greenough, T.C.; et al. Retroviruses Pseudotyped with the Severe Acute Respiratory Syndrome Coronavirus Spike Protein Efficiently Infect Cells Expressing Angiotensin-Converting Enzyme 2. J. Virol. 2004, 78, 10628–10635. [Google Scholar] [CrossRef] [PubMed]
- Imai, Y.; Kuba, K.; Rao, S.; Huan, Y.; Guo, F.; Guan, B.; Yang, P.; Sarao, R.; Wada, T.; Leong-Poi, H.; et al. Angiotensin-converting enzyme 2 protects from severe acute lung failure. Nature 2005, 436, 112–116. [Google Scholar] [CrossRef] [PubMed]
- Reilly, J.; Calfee, C.; Christie, J. Acute Respiratory Distress Syndrome Phenotypes. Semin. Respir. Crit. Care Med. 2019, 40, 019–030. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.; Benthin, C.; Zeno, B.; Albertson, T.E.; Boyd, J.; Christie, J.D.; Hall, R.; Poirier, G.; Ronco, J.J.; Tidswell, M.; et al. A pilot clinical trial of recombinant human angiotensin-converting enzyme 2 in acute respiratory distress syndrome. Crit. Care 2017, 21, 234. [Google Scholar] [CrossRef] [PubMed]
- Maxmen, A. More than 80 clinical trials launch to test coronavirus treatments. Nature 2020, 578, 347–348. [Google Scholar] [CrossRef]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.-L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef]
- Wolff, J.A.; Malone, R.W.; Williams, P.; Chong, W.; Acsadi, G.; Jani, A.; Felgner, P.L. Direct gene transfer into mouse muscle in vivo. Science 1990, 247, 1465–1468. [Google Scholar] [CrossRef]
- Sahin, U.; Karikó, K.; Türeci, Ö. mRNA-based therapeutics—Developing a new class of drugs. Nat. Rev. Drug Discov. 2014, 13, 759–780. [Google Scholar] [CrossRef]
- Sahin, U.; Derhovanessian, E.; Miller, M.; Kloke, B.-P.; Simon, P.; Löwer, M.; Bukur, V.; Tadmor, A.D.; Luxemburger, U.; Schrörs, B.; et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 2017, 547, 222–226. [Google Scholar] [CrossRef] [PubMed]
- Sebastian, M.; Schröder, A.; Scheel, B.; Hong, H.S.; Muth, A.; von Boehmer, L.; Zippelius, A.; Mayer, F.; Reck, M.; Atanackovic, D.; et al. A phase I/IIa study of the mRNA-based cancer immunotherapy CV9201 in patients with stage IIIB/IV non-small cell lung cancer. Cancer Immunol. Immunother. CII 2019, 68, 799–812. [Google Scholar] [CrossRef] [PubMed]
- Diken, M.; Kranz, L.M.; Kreiter, S.; Sahin, U. mRNA: A Versatile Molecule for Cancer Vaccines. Curr. Issues Mol. Biol. 2017, 22, 113–128. [Google Scholar] [CrossRef] [PubMed]
- Safety and Immunogenicity Study of 2019-nCov Vaccine (mRNA-1273) to Treat Novel Coronavirus—Full Text View—ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/show/NCT04283461 (accessed on 27 February 2020).
| No. Cases (%) | Deaths (%) | CFR (%) | |
|---|---|---|---|
| Overall | 44,672 | 1023 | 2.3 |
| Age | |||
| 0–9 yrs | 416 (0.9) | - | - |
| 10–19 yrs | 549 (1.2) | 1 (0.1) | 0.2 |
| 20–29 yrs | 3619 (8.1) | 7 (0.7) | 0.2 |
| 30–39 yrs | 7600 (17.0) | 18 (1.8) | 0.2 |
| 40–49 yrs | 8571 (19.2) | 38 (3.7) | 0.4 |
| 50–59 yrs | 10,008 (22.4) | 130 (12.7) | 1.3 |
| 60–69 yrs | 8583 (19.2) | 309 (30.2) | 3.6 |
| 70–79 yrs | 3918 (8.8) | 312 (30.5) | 8.0 |
| ≥ 80 yrs | 1408 (3.2) | 208 (20.3) | 14.8 |
| Wuhan related exposure | |||
| Yes | 31,974 (85.8) | 853 (92.8) | 2.7 |
| No | 5295 (14.2) | 66 (7.2) | 1.2 |
| Case Severity | |||
| Mild | 36,180 (80.9) | - | - |
| Severe | 6168 (13.8) | - | - |
| Critical | 2087 (4.7) | 1023 (100) | 49.0 |
| Missing | 257 (0.6) | - | - |
| CFR (%) | |
|---|---|
| Age | |
| 0–9 yrs | - |
| 10–19 yrs | 0.2 |
| 20–29 yrs | 0.2 |
| 30–39 yrs | 0.2 |
| 40–49 yrs | 0.4 |
| 50–59 yrs | 1.3 |
| 60–69 yrs | 3.6 |
| 70–79 yrs | 8.0 |
| ≥ 80 yrs | 14.8 |
| Sex | |
| Male | 2.8 |
| Female | 1.7 |
| Preexisting Medical Condition | |
| Cardiovascular disease | 10.5 |
| Diabetes | 7.3 |
| Chronic respiratory disease | 6.3 |
| Hypertension | 6.0 |
| Cancer | 5.6 |
| No preexisting condition | 0.9 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rabi, F.A.; Al Zoubi, M.S.; Kasasbeh, G.A.; Salameh, D.M.; Al-Nasser, A.D. SARS-CoV-2 and Coronavirus Disease 2019: What We Know So Far. Pathogens 2020, 9, 231. https://doi.org/10.3390/pathogens9030231
Rabi FA, Al Zoubi MS, Kasasbeh GA, Salameh DM, Al-Nasser AD. SARS-CoV-2 and Coronavirus Disease 2019: What We Know So Far. Pathogens. 2020; 9(3):231. https://doi.org/10.3390/pathogens9030231
Chicago/Turabian StyleRabi, Firas A., Mazhar S. Al Zoubi, Ghena A. Kasasbeh, Dunia M. Salameh, and Amjad D. Al-Nasser. 2020. "SARS-CoV-2 and Coronavirus Disease 2019: What We Know So Far" Pathogens 9, no. 3: 231. https://doi.org/10.3390/pathogens9030231
APA StyleRabi, F. A., Al Zoubi, M. S., Kasasbeh, G. A., Salameh, D. M., & Al-Nasser, A. D. (2020). SARS-CoV-2 and Coronavirus Disease 2019: What We Know So Far. Pathogens, 9(3), 231. https://doi.org/10.3390/pathogens9030231







