Evaluation of Biocontrol Activities of Streptomyces spp. against Rice Blast Disease Fungi
Abstract
1. Introduction
2. Results
2.1. Isolation of Rhizospheric Actinomycetes
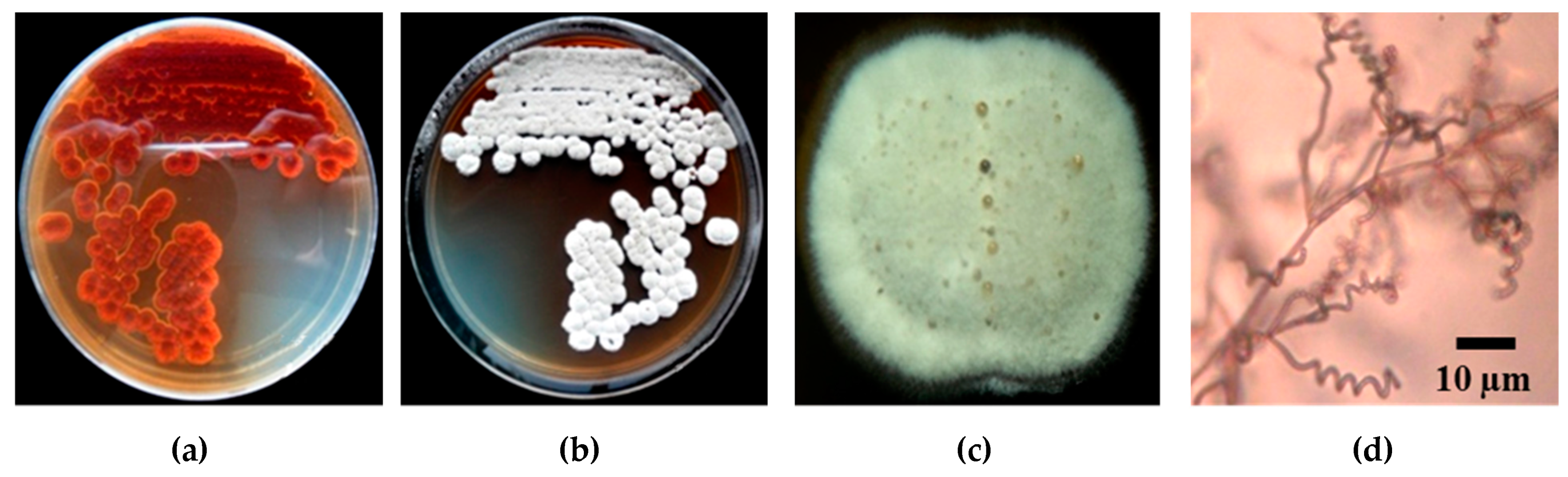
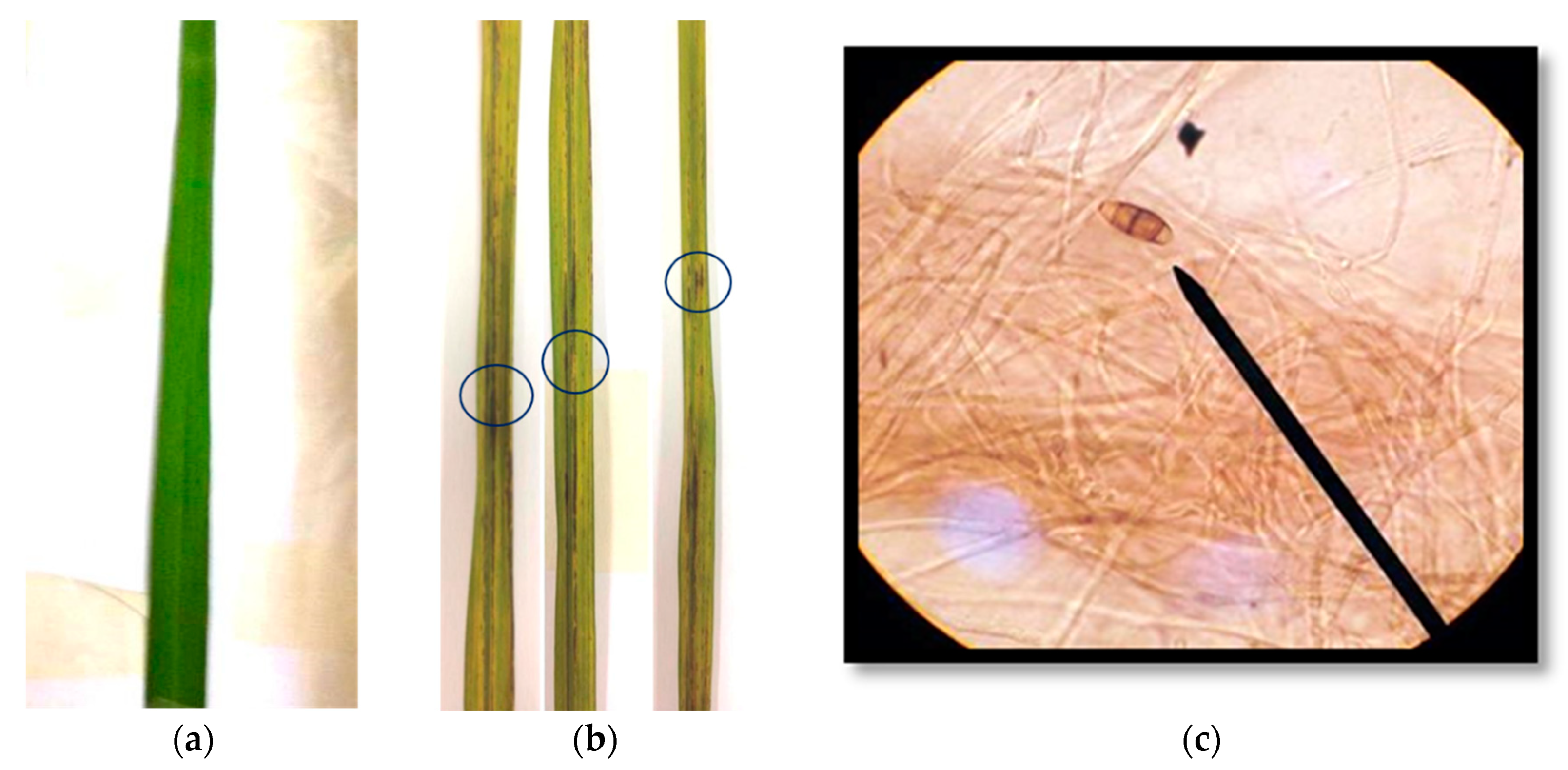
2.2. Isolation of Blast Disease Fungi from Rice Leaves and Pathogenicity Test
2.3. Plant Growth Properties of the Actinomycete Isolates
2.4. Greenhouse Experiment
2.4.1. Pathogenicity Test of Selected Streptomyces
2.4.2. Evaluation of Streptomyces against Pyricularia sp. under Greenhouse Condition
2.4.3. Plant Growth Promotion effect
2.5. Rhizospheric Colonization by Streptomyces
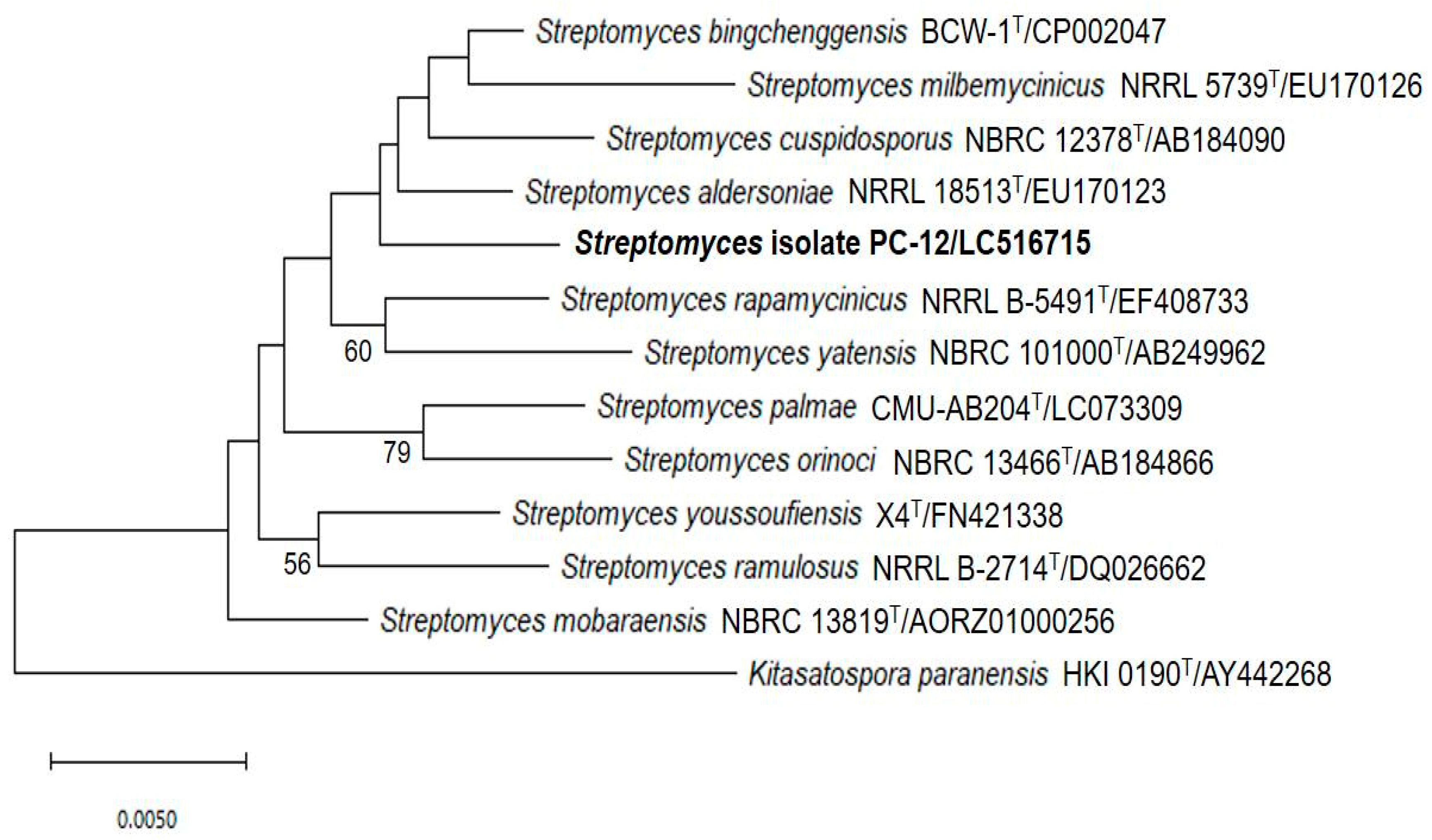
2.6. Identification of Streptomyces PC 12
3. Discussion
4. Materials and Methods
4.1. Isolation of Rhizospheric Actinomycetes
4.2. Isolation of Rice Blast Disease Fungi
4.3. Pathogenicity Assay
4.4. In vitro Antifungal Assay
4.5. Actinomycetes Identification
4.5.1. Cultural and Morphological Characterizations
4.5.2. Cell Wall Composition Analysis
4.5.3. Molecular Characterization of Potential Antagonistic Isolate
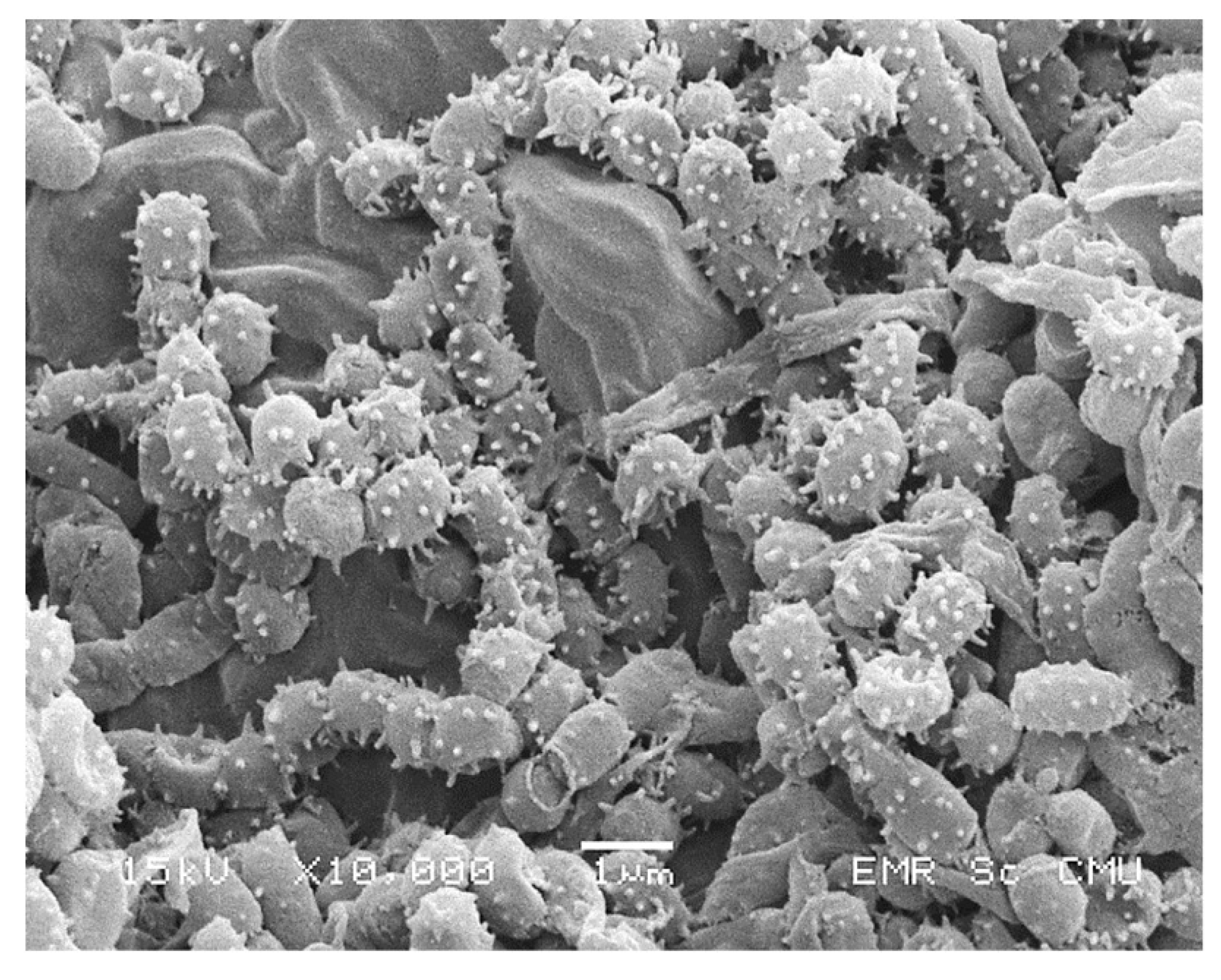
4.5.4. Scanning Electron Microscopy
4.6. Screening of Plant Growth Promoting Activities
4.6.1. Siderophore Production
4.6.2. HCN Production
4.6.3. Hydrolytic Enzyme Production
4.7. Greenhouse Experiments
- T 1 = Healthy control (no fungus, no actinomycetes);
- T 2 = Healthy rice plant inoculate with Streptomyces strain W 1;
- T 3 = Rice plant inoculate with Streptomyces strain W 1+ Pyricularia sp.;
- T 4 = Healthy rice plant inoculate with Streptomyces strain D 4.3;
- T 5 = Rice plant inoculate with Streptomyces strain D 4.3 + Pyricularia sp.;
- T 6 = Healthy rice plant inoculate with Streptomyces strain PC 12;
- T 7 = Rice plant inoculate with Streptomyces strain PC 12 + Pyricularia sp.;
- T 8 = Healthy rice plant inoculate with Streptomyces strain D 4.1;
- T 9 = Rice plant inoculate with Streptomyces strain D 4.1+ Pyricularia sp.;
- T 10 = Disease control (Pyricularia sp. alone);
- T 11 = Rice plant inoculate with Fungicide + Pyricularia sp.
- 0. no lesions observed;
- 1. small brown specks of pin-point size without sporulating centre;
- 2. larger brown specks but less than 1 mm in diameter;
- 3. small roundish to slightly elongated, necrotic grey spots 1– 2 mm;
- 4. typical susceptible blast lesions (spindle-shaped) 3 mm or longer, infecting less than 4.0% of the leaf area;
- 5. typical blast lesions infecting 4.1–10.0% of the leaf area;
- 6. typical blast lesions infecting 10.1–25.0% of the leaf area;
- 7. typical blast lesions infecting 25.1–50.0% of the leaf area;
- 8. typical blast lesions infecting 50.1–75.0% of the leaf area;
- 9. typical blast lesions infecting more than 75.1% of the leaf area.
4.8. Effect of Streptomyces Application on Plant Growth Promotion under Greenhouse Conditions
4.8.1. Measurement of Shoot and Root Weight and Tiller Number
4.8.2. Rhizospheric Colonization by Streptomyces
4.9. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Chaiharn, M.; Chunhaleuchanon, S.; Lumyong, S. Screening siderophore producing bacteria as potential biological control agent for fungal rice pathogens in Thailand. World J. Microbiol. Biotechnol. 2009, 25, 1919–1928. [Google Scholar] [CrossRef]
- Asibi, A.E.; Chai, Q.; Coulter, J.A. Rice blast: A disease with implications for global food security. Agronomy 2019, 9, 451. [Google Scholar] [CrossRef]
- Kato, H. Rice blast disease. Pestic. Outlook. 2001, 12, 23–25. [Google Scholar] [CrossRef]
- Hamer, J.E.; Howard, R.J.; Chumley, F.G.; Valent, B. A mechanism for surface attachment in spores of a plant pathogenic fungus. Science 1988, 239, 288–290. [Google Scholar] [CrossRef]
- Vidhyasekaran, P. Fungal pathogenesis in plants and crops. In Molecular Biology and Host Defense Mechanisms, 2th ed.; Vidhyasekaran, P., Ed.; CRC Press: New York, NY, USA, 2008; pp. 1–53. [Google Scholar]
- Bae, C.Y.; Kim, S.; Choi, W.; Lee, Y.H. Involvement of extracellular matrix and integrin-like proteins on conidal adhesion and appressorium differentiation in Magnaporthe oryzae. J. Microbiol. Biotechnol. 2007, 17, 1198–1203. [Google Scholar] [PubMed]
- Inoue, K.; Suzuki, T.; Ikeda, K.; Jiang, S.; Hosogi, N.; Hyon, G.S.; Hida, S.; Yamada, T.; Park, P. Extracellular matrix of Magnaporthe oryzae may have a role in host adhesion during fungal penetration and is digested by matrix metalloproteinases. J. Gen. Plant Pathol. 2007, 73, 388–398. [Google Scholar] [CrossRef]
- Wilson, R.A.; Talbot, N.J. Under pressure: Investigating the biology of plant infection by Magnaporthe oryzae. Nat. Rev. Microbiol. 2009, 7, 185–195. [Google Scholar] [CrossRef] [PubMed]
- Sesma, A.; Osbourn, A.E. The rice leaf blast pathogen undergoes developmental processes typical of root-infecting fungi. Nature 2004, 431, 582–586. [Google Scholar] [CrossRef]
- Ashkani, S.; Yusop, M.R.; Shabanimofrad, M.; Haron, A.R.; Sahebi, M.; Latif, M.A. Genetic analysis of resistance to rice blast: A study on the inheritance of resistance to the blast disease pathogen in an F3 population of rice. J. Phytopathol. 2015, 163, 300–309. [Google Scholar] [CrossRef]
- Pooja, K.; Katoch, A. Past, present and future of rice blast management. Plant Sci. Today 2014, 1, 165–173. [Google Scholar] [CrossRef]
- Law, J.W.; Ser, H.L.; Khan, T.M.; Chuah, L.H.; Pusparajah, P.; Chan, K.G.; Goh, B.H.; Lee, L.H. The potential of Streptomyces as biocontrol agents against the rice blast fungus, Magnaporthe oryzae (Pyricularia oryzae). Front. Microbiol. 2017, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Yoon, M.Y.; Cha, B.; Kim, J.C. Recent trends in studies on botanical fungicides in agriculture. Plant Pathol. J. 2013, 29, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, S.; Meisner, C.; Wheeler, D.; Xuyen, K.; Lam, N.T. Pesticide poisoning of farm workers-implications of blood test results from Vietnam. Int. J. Hyg. Environ. Health 2007, 210, 121–132. [Google Scholar] [CrossRef] [PubMed]
- Fattahi, F.; Mousavi Moghadam, M.; Khanbabaei, R. The effect of tricyclazole on testosterone changes and testicular structure in mice. J. Babol. Univ. Med. Sci. 2015, 17, 43–49. [Google Scholar] [CrossRef]
- Newitt, J.T.; Prudence, S.M.M.; Hutchings, M.I.; Worsley, S.F. Biocontrol of cereal crop diseases using streptomycetes. Pathogens 2019, 8, 78. [Google Scholar] [CrossRef]
- Copping, L.G.; Duke, S.O. Natural products that have been used commercially as crop protection agents. Pest Manag. Sci. 2007, 63, 524–554. [Google Scholar] [CrossRef]
- Yang, P.; Li, M.; Zhao, J.; Zhu, M.; Shan, H.; Li, J. Oligomycins A and C, major secondary metabolites isolated from the newly isolated strain Streptomyces diastaticus. Folia Microbiol. 2010, 55, 10–16. [Google Scholar] [CrossRef]
- Boukaew, S.; Prasertsan, P. Suppression of rice sheath blight disease using a heat stable culture filtrate from Streptomyces philanthi RM-1-138. Crop Protect. 2014, 61, 1–10. [Google Scholar] [CrossRef]
- Ningthoujam, S.; Sanasam, S.; Tamreihao, K.; Nimaich, S. Antagonistic activities of local actinomycete isolates against rice fungal pathogens. Afr. J. Microbiol. Res. 2009, 3, 737–742. [Google Scholar]
- Awla, H.K.; Kadir, J.; Othman, R.; Rashid, T.S.; Hamid, S.; Wong, M.-Y. Plant growth-promoting abilities and biocontrol efficacy of Streptomyces sp. UPMRS4 against Pyricularia oryzae. Biol. Control. 2017, 112, 55–63. [Google Scholar] [CrossRef]
- Prabavathy, V.R.; Mathivanan, N.; Muragesan, K. Control of blast and sheath blight disease of rice using antifungal metabolites produced by Streptomyces sp. PM 5. Biol. Control. 2006, 3, 313–319. [Google Scholar] [CrossRef]
- Xu, T.; Cao, L.; Zeng, J.; Franco, C.M.M.; Yang, Y.; Hu, X.; Liu, Y.; Wang, X.; Gao, Y.; Bu, Z.; et al. The antifungal action mode of the rice endophyte Streptomyces hygroscopicus OsiSh-2 as a potential biocontrol agent against the rice blast pathogen. Pestic Biochem. Phys. 2019, 160, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Nonomura, H. Key for classification and identification of 458 species of the Streptomycetes included in ISP. J. Ferment. Technol. 1974, 52, 78–92. [Google Scholar]
- Mew, T.W.; Gonzales, P. A Handbook of Rice Seed Borne Fungi. Los Banos, Laguna: IRRI; Enfield, N.H., Ed.; Science Publishers, Inc.: New York, NY, USA, 2012; pp. 27–31. [Google Scholar]
- Bussaban, B.; Lumyong, S.; Lumyong, P.; Seelanan, T.; Park, D.C.; McKenzie, E.H.C.; Hyde, K.D. Molecular and morphological characterization of Pyricularia and allied genera. Mycologia 2005, 97, 1002–1011. [Google Scholar] [CrossRef] [PubMed]
- Chaiharn, C.; Sujada, N.; Pathom-aree, W.; Lumyong, S. Biological control of Rigidoporus microporus the casue of white root disease in rubber using PGPRs in vivo. Chiang Mai J. Sci. 2019, 46, 850–866. [Google Scholar]
- Hata, E.M.; Sijam, K.; Ahmad, Z.A.M.; Yusof, M.T.; Azman, N.A. In vitro antimicrobial assay of actinomycetes in rice against Xanthomonas oryzae pv. oryzicola and as potential plant growth promoter. Braz. Arch. Biol. Technol. 2015, 58, 821–832. [Google Scholar] [CrossRef]
- Banchio, E.; Bogino, P.C.; Zygadlo, J.; Giordano, W. Plant growth promoting rhizobacteria improve growth and essential oil yield in Origanum majorana L. Biochem. Syst. Ecol. 2008, 36, 766–771. [Google Scholar] [CrossRef]
- Kim, M.; Oh, H.-S.; Park, S.-C.; Chun, J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int. J. Syst. Evol. Microbiol. 2014, 64, 346–351. [Google Scholar] [CrossRef]
- Loqman, S.; Barka, E.E.A.; Clement, C.E.; Ouhdouch, E.Y. Antagonistic actinomycetes from Moroccan soil to control the grapevine gray mold. World J. Microbiol. Biotechnol. 2009, 25, 81–91. [Google Scholar] [CrossRef]
- Schulz, B.; Wanke, U.; Draeger, S. Endophytes from herbaceous plants and shrubs: Effectiveness of surface sterilization methods. Mycol. Res. 1993, 97, 1447–1450. [Google Scholar] [CrossRef]
- Plodpai, P.; Chuenchitt, S.; Petcharat, V.; Chakthong, S.; Piyawan, S.; Voravuthikunchai, S.S. Anti-Rhizoctonia solani activity by Desmos chinensis extracts and its mechanism of action. Crop Prot. 2013, 43, 65–71. [Google Scholar] [CrossRef]
- Hamdia, Z.A.; Kalaivani, N. Evaluating efficacy of Trichoderma isolates and Bacillus subtilis as biological control agents against Rhizoctonia solani. Res. J. Appl. Sci. 2013, 8, 72–81. [Google Scholar]
- Alfredo, M.S.; Aleli Cornelia, R.P. Biological control of sheath blight of upland rice with Trichoderma species. J. Trop. Plant Pathol. 2011, 69, 1–9. [Google Scholar]
- Lee, J.M.; Tan, W.S.; Ting, A.S.Y. Revealing the antimicrobial and enzymatic potentials of culturable fungal endophytes from tropical pitcher plants (Nepenthes spp.). Mycosphere 2014, 5, 364–377. [Google Scholar] [CrossRef]
- Shirling, E.B.; Gottlieb, D. Methods for characterization of Streptomyces species. Int. J. Syst. Bacteriol. 1966, 16, 313–340. [Google Scholar] [CrossRef]
- Hasegawa, T.; Takizawa, M.; Tanida, S. A rapid analysis for chemical grouping of aerobic actinomycetes. J. Gen. Appl. Microbiol. 1983, 29, 319–322. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogeny: An appropriate use of the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Tamura, K.; Nei, M.; Kumar, S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc. Nat. Acad. Sci. USA 2004, 101, 11030–11035. [Google Scholar] [CrossRef]
- Schwyn, B.; Neilands, J.B. Universal chemical assay for the detection and determination of siderophores. Anal. Biochem. 1987, 160, 47–56. [Google Scholar] [CrossRef]
- International Rice Research Institute. Standards Evaluation System for Rice; IRRI: Manila, Philippines, 2013. [Google Scholar]
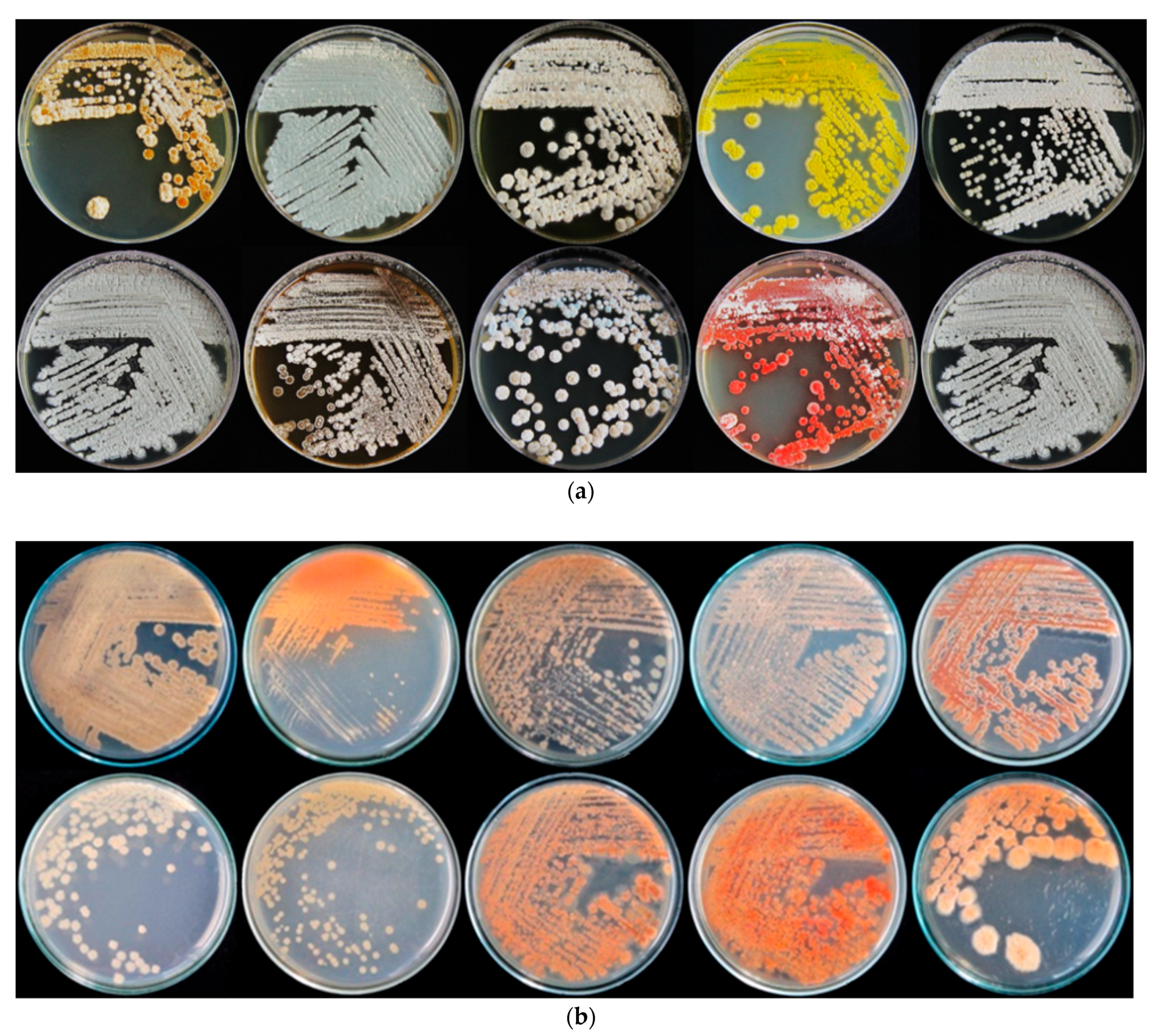


| No. | Color of Aerial Mycelium | Soluble Pigment | Number of Isolates | Percentage of all Isolates (%) |
|---|---|---|---|---|
| 1 | Gray | Brown | 41 | 36.6 |
| 2 | Brown | Brown | 23 | 20.5 |
| 3 | White-Gray | Brown | 14 | 12.5 |
| 4 | White | Brown | 12 | 10.7 |
| 5 | Yellow | - | 10 | 8.9 |
| 6 | Green | - | 3 | 2.7 |
| 7 | Orange | Orange | 3 | 2.7 |
| 8 | Red | Brown | 3 | 2.7 |
| 9 | Pink | - | 2 | 1.8 |
| 10 | Black | - | 1 | 0.9 |
| Total | 112 | 100 |
| Isolate. | Inhibition Zone (mm) | HCN Production | ||||
|---|---|---|---|---|---|---|
| Antagonistic to Pyricularia sp. (%) | Siderophore Production | Protease Production | Cellulase Production | Chitinase Production | ||
| PC-1 | 39.5 ± 2.7 f | 11.4 ± 0.9 e | 51.3 ± 1.9 b | - | - | + |
| PC-5 | 41.3 ± 1.8 e | 8.9 ± 0.7 f | 50.6 ± 1.5 b | - | - | + |
| PC-7 | 29.7 ± 3.5 g | 10.4 ± 0.5 e | 40.7 ± 1.1 c | - | - | + |
| PC-9 | 22.6 ± 2.9 g | 12.5 ± 0.9 e | 43.2 ± 1.3 c | - | - | + |
| PC-12 | 87.3 ± 1.8 b | 24.5 ± 0.9 d | 25.7 ± 1.3 e | 57.2 ± 3.1 c | 40.3 ± 2.7 c | + |
| PC-17 | 45.6 ± 3.1 e | 18.5 ± 1.1 e | 28.6 ± 1.5 e | 43.1 ± 2.7 d | 35.7 ± 2.1 d | + |
| PC-23 | 31.2 ± 4.9 f | 17.3 ± 1.3 e | 33.5 ± 1.3 d | 27.9 ± 1.7 e | - | + |
| PC-24 | 71.6 ± 1.4 bc | 32.4 ± 1.3 c | 73.7 ± 1.5 a | - | - | + |
| PC-37 | 38.4 ± 7.8 f | 28.9 ± 1.5 d | 38.4 ± 1.7 d | - | - | + |
| PC-39 | 51.7 ± 1.9 d | 17.5 ± 1.1 e | 34.7 ± 1.5 d | - | - | + |
| PC-42 | 66.0 ± 3.5 c | 32.8 ± 1.7 c | 46.2 ± 1.1 c | - | - | + |
| PC-47 | 33.5 ± 3.1 f | 36.7 ± 1.9 c | 51.5 ± 1.9 b | 40.5 ± 3.7 d | 21.4 ± 1.3 e | + |
| PC-55 | 27.7 ± 7.5 g | 41.0 ± 1.7 b | 44.3 ± 1.7 c | 64.2 ± 2.7 b | 27.9 ± 2.7 e | + |
| PC-63 | 69.3 ± 3.7 c | 40.5 ± 1.3 b | 52.4 ± 1.5 b | - | - | + |
| PC-71 | 21.0 ± 1.7 h | 34.6 ± 1.1 c | 47.2 ± 1.9 c | - | - | + |
| PC-73 | 38.6 ± 2.1 f | 43.4 ± 1.3 b | 24.2 ± 1.3 e | - | - | + |
| PC-88 | 26.1 ± 0.8 g | 42.8 ± 1.5 b | 35.7 ± 1.7 d | - | - | + |
| PC-94 | 71.3 ± 1.4 bc | 50.2 ± 1.3 a | 28.9 ± 1.5 e | - | - | + |
| PC-102 | 38.3 ± 4.0 f | 34.5 ± 1.5 c | 47.6 ± 1.3 c | 57.6 ± 4.1 c | - | + |
| PC-110 | 49.5 ± 2.3 e | 37.4 ± 1.3 c | 28.7 ± 1.7 e | - | - | + |
| PC-117 | 67.2 ± 4.2 c | 50.7 ± 1.1 a | 37.8 ± 1.9 d | - | - | + |
| PC-119 | 75.8 ± 3.7 b | 40.8 ± 1.5 b | 45.4 ± 1.5 c | - | - | + |
| PC-120 | 74.3 ± 4.2 b | 31.7 ± 1.3 c | 50.2 ± 1.9 b | - | - | + |
| D 4.1 | 82.2 ± 2.5 a | 52.3 ± 1.9 a | 53.5 ± 1.9 b | 70.8 ± 4.7 a | 57.4 ± 3.1 a | + |
| D 4.3 | 80.0 ± 2.7 a | 51.4 ± 1.1a | 52.2 ± 1.9 b | 69.5 ± 3.7 b | 48.5 ± 2.7 b | + |
| W 1 | 80.5 ± 1.7 a | 47.8 ± 1.5 c | 50.7 ± 1.5 b | 57.8 ± 1.9 c | 34.8 ± 2.1 d | + |
| W 9 | 76.4 ± 3.2 b | 41.5 ± 1.1 c | 34.7 ± 1.3 d | - | - | + |
| Treatment | Blast disease Parameters 60 Days | Plant growth Parameters60 Days | |||||
|---|---|---|---|---|---|---|---|
| Lesion Length (cm) | No. of Dried Leaves per Plant | Disease Severity (%) | Plant Height (cm) | Root Length (cm) | Root Dry Weight (g) | No of Tiller | |
| Streptomyces W1 | 1.19 ± 0.21 b | 3.67 ± 0.27 c | 51.9 ± 0.31 c | 45.08 ± 0.57 c | 29.43 ± 0.17 c | 2.13 ± 0.07 ab | 12.8 ± 0.31 bc |
| Streptomyces D 4.3 | 0.98 ± 0.19 b | 3.37 ± 0.21 c | 44.5 ± 0.37 b | 52.69 ± 0.47 b | 37.00 ± 0.19 b | 2.53 ± 0.11 a | 15.2 ± 0.27 b |
| Streptomyces PC 12 | 0.57 ± 0.19 c | 3.10 ± 0.19 c | 31.4 ± 0.37 a | 76.93 ± 0.59 a | 41.60 ± 0.27 a | 2.68 ± 0.17 a | 19.4 ± 0.45 a |
| Streptomyces D 4.1 | 1.07 ± 0.15 b | 4.50 ± 0.29 b | 42.8 ± 0.31 b | 43.57 ± 0.37 c | 31.89 ± 0.21 b | 2.05 ± 0.19 ab | 14.7 ± 0.21 b |
| Pathogen (treated control) | 6.78 ± 1.17 a | 8.38 ± 0.35 a | 87.5 ± 0.47 d | 36.59 ± 0.41 d | 26.17 ± 0.15 c | 1.88 ± 0.13 b | ND |
| Healthy (Untreated control) | ND | ND | ND | 40.70 ± 0.35 cd | 29.43 ± 0.29 c | 1.96 ± 0.17 b | 9.20 ± 0.31 d |
| Streptomyces Strains | Cell Number (CFU/mL) |
|---|---|
| Streptomyces W1 | 3.2 × 103 |
| Streptomyces D 4.3 | 3.4 × 104 |
| Streptomyces PC 12 | 4.5 × 104 |
| Streptomyces D 4.1 | 2.7 × 103 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chaiharn, M.; Theantana, T.; Pathom-aree, W. Evaluation of Biocontrol Activities of Streptomyces spp. against Rice Blast Disease Fungi. Pathogens 2020, 9, 126. https://doi.org/10.3390/pathogens9020126
Chaiharn M, Theantana T, Pathom-aree W. Evaluation of Biocontrol Activities of Streptomyces spp. against Rice Blast Disease Fungi. Pathogens. 2020; 9(2):126. https://doi.org/10.3390/pathogens9020126
Chicago/Turabian StyleChaiharn, Mathurot, Teerayut Theantana, and Wasu Pathom-aree. 2020. "Evaluation of Biocontrol Activities of Streptomyces spp. against Rice Blast Disease Fungi" Pathogens 9, no. 2: 126. https://doi.org/10.3390/pathogens9020126
APA StyleChaiharn, M., Theantana, T., & Pathom-aree, W. (2020). Evaluation of Biocontrol Activities of Streptomyces spp. against Rice Blast Disease Fungi. Pathogens, 9(2), 126. https://doi.org/10.3390/pathogens9020126




