The Diverse Roles of the Global Transcriptional Regulator PhoP in the Lifecycle of Yersinia pestis
Abstract
1. Introduction
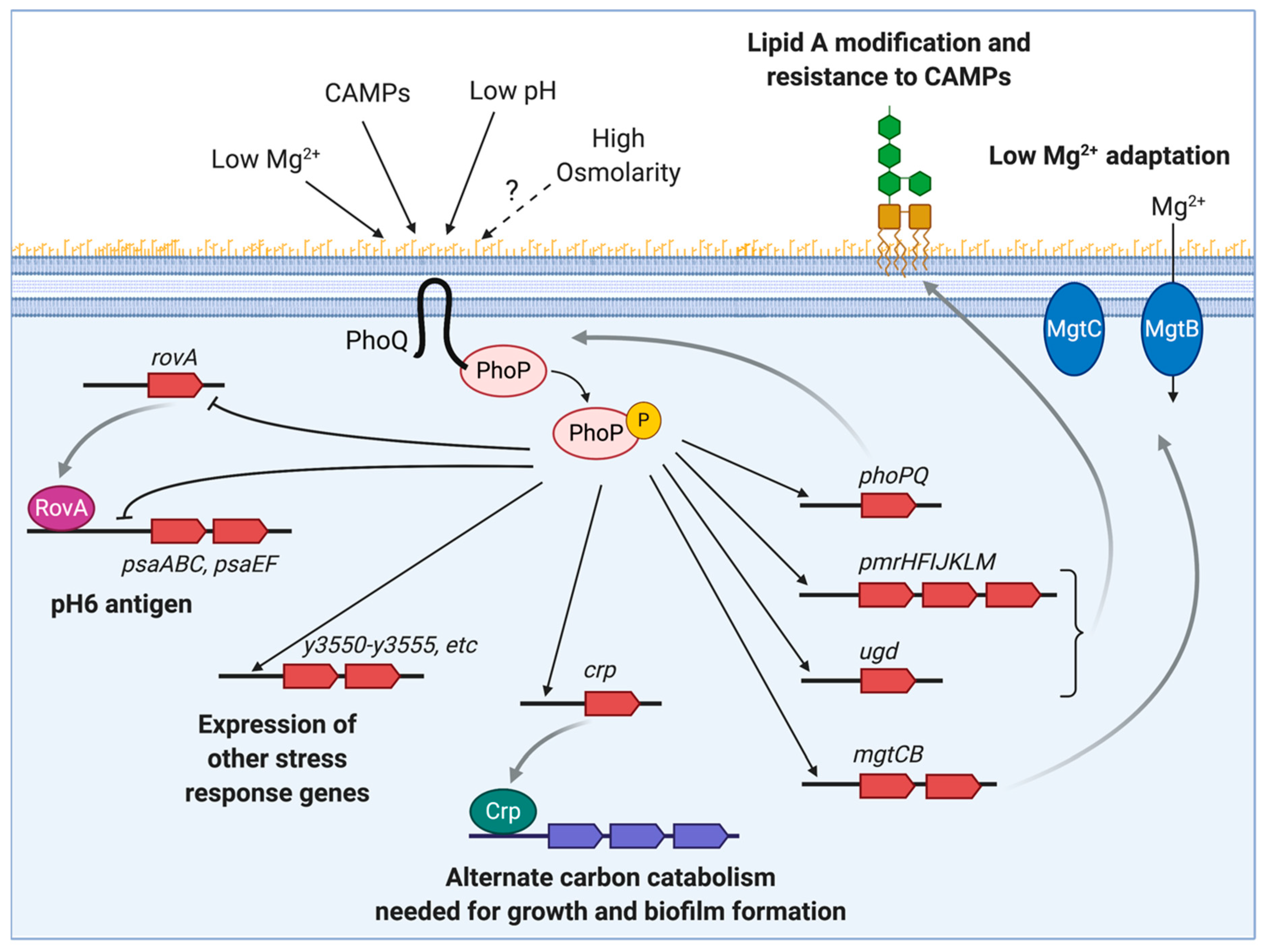
2. Y. pestis PhoP Regulatory Networks
3. Role of phoP in Intracellular Replication in Mammalian Hosts
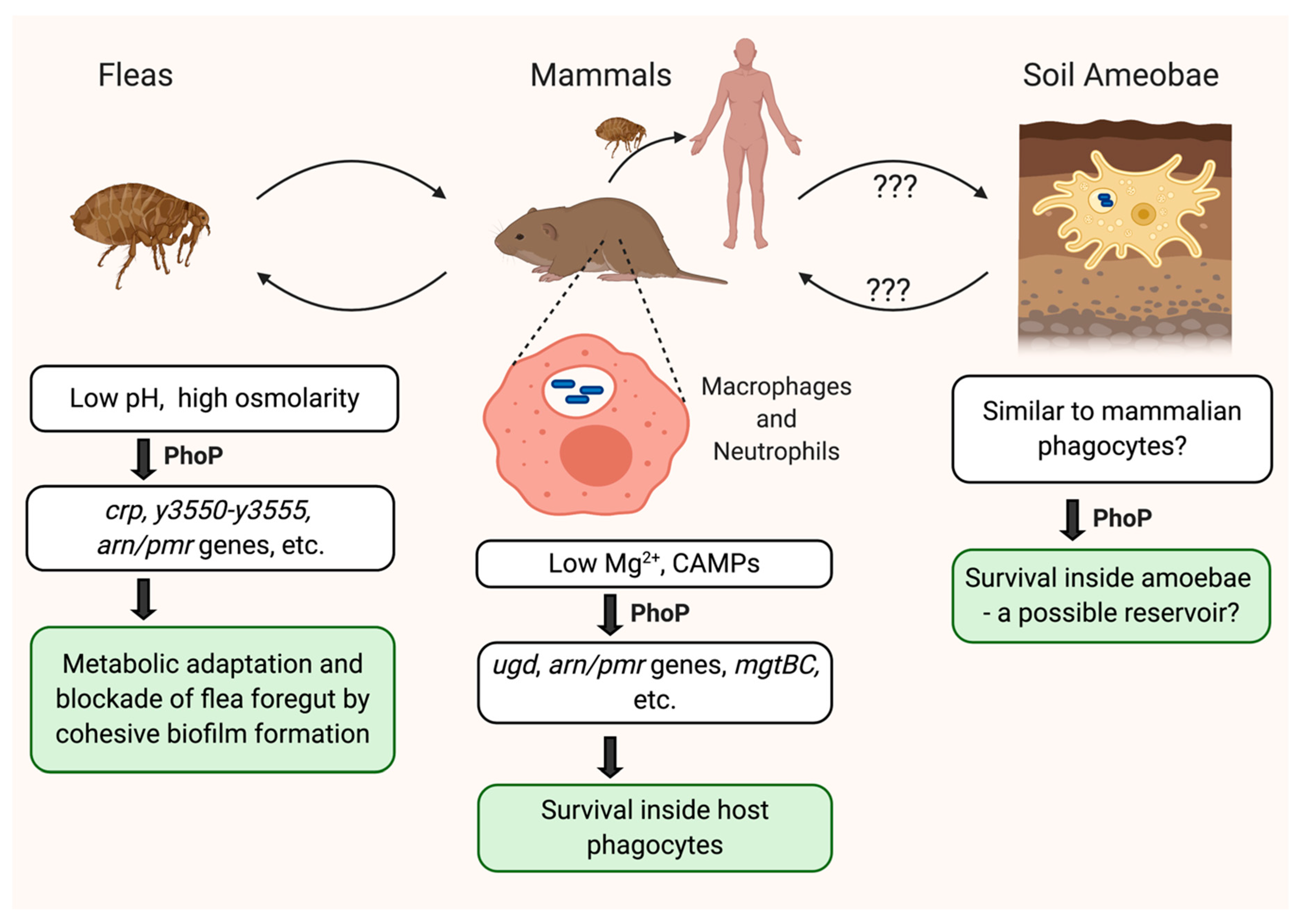
4. Biofilm and Flea Colonization
5. Amoeba as a Potential Host While the Pathogen Is Quiescent
6. The Effects of a phoP SNP on the PhoP Function and the Evolution of Y. pestis Virulence
7. Summary and Future Perspective
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Achtman, M.; Zurth, K.; Morelli, G.; Torrea, G.; Guiyoule, A.; Carniel, E. Yersinia pestis, the cause of plague, is a recently emerged clone of Yersinia pseudotuberculosis. Proc. Natl. Acad. Sci. USA 1999, 96, 14043–14048. [Google Scholar] [CrossRef]
- Rasmussen, S.; Allentoft, M.E.; Nielsen, K.; Orlando, L.; Sikora, M.; Sjogren, K.G.; Pedersen, A.G.; Schubert, M.; Van Dam, A.; Kapel, C.M.; et al. Early divergent strains of Yersinia pestis in Eurasia 5000 years ago. Cell 2015, 163, 571–582. [Google Scholar] [CrossRef]
- Rascovan, N.; Sjogren, K.G.; Kristiansen, K.; Nielsen, R.; Willerslev, E.; Desnues, C.; Rasmussen, S. Emergence and spread of basal lineages of Yersinia pestis during the Neolithic decline. Cell 2019, 176, 295–305. [Google Scholar] [CrossRef]
- Spyrou, M.A.; Tukhbatova, R.I.; Wang, C.C.; Valtuena, A.A.; Lankapalli, A.K.; Kondrashin, V.V.; Tsybin, V.A.; Khokhlov, A.; Kuhnert, D.; Herbig, A.; et al. Analysis of 3800-year-old Yersinia pestis genomes suggests Bronze Age origin for bubonic plague. Nat. Commun. 2018, 9, 2234. [Google Scholar] [CrossRef]
- Chain, P.S.; Carniel, E.; Larimer, F.W.; Lamerdin, J.; Stoutland, P.O.; Regala, W.M.; Georgescu, A.M.; Vergez, L.M.; Land, M.L.; Motin, V.L.; et al. Insights into the evolution of Yersinia pestis through whole-genome comparison with Yersinia pseudotuberculosis. Proc. Natl. Acad. Sci. USA 2004, 101, 13826–13831. [Google Scholar] [CrossRef]
- Chouikha, I.; Hinnebusch, B.J. Yersinia-flea interactions and the evolution of the arthropod-borne transmission route of plague. Curr. Opin. Microbiol. 2012, 15, 239–246. [Google Scholar] [CrossRef]
- Perry, R.D.; Fetherston, J.D. Yersinia pestis—Etiologic agent of plague. Clin. Microbiol. Rev. 1997, 10, 35–66. [Google Scholar] [CrossRef]
- Ayyadurai, S.; Houhamdi, L.; Lepidi, H.; Nappez, C.; Raoult, D.; Drancourt, M. Long-term persistence of virulent Yersinia pestis in soil. Microbiology 2008, 154, 2865–2871. [Google Scholar] [CrossRef]
- Boegler, K.A.; Graham, C.B.; Montenieri, J.A.; MacMillan, K.; Holmes, J.L.; Petersen, J.M.; Gage, K.L.; Eisen, R.J. Evaluation of the infectiousness to mice of soil contaminated with Yersinia pestis-infected blood. Vector Borne Zoonotic Dis. 2012, 12, 948–952. [Google Scholar] [CrossRef]
- Eisen, R.J.; Gage, K.L. Adaptive strategies of Yersinia pestis to persist during inter-epizootic and epizootic periods. Vet. Res. 2009, 40, 1. [Google Scholar] [CrossRef]
- Marceau, M. Transcriptional regulation in Yersinia: An update. Curr. Issues Mol. Biol. 2005, 7, 151–177. [Google Scholar]
- Sun, Y.C.; Hinnebusch, B.J.; Darby, C. Experimental evidence for negative selection in the evolution of a Yersinia pestis pseudogene. Proc. Natl. Acad. Sci. USA 2008, 105, 8097–8101. [Google Scholar] [CrossRef]
- Groisman, E.A. The pleiotropic two-component regulatory system PhoP-PhoQ. J. Bacteriol. 2001, 183, 1835–1842. [Google Scholar] [CrossRef]
- Bader, M.W.; Sanowar, S.; Daley, M.E.; Schneider, A.R.; Cho, U.; Xu, W.; Klevit, R.E.; Le Moual, H.; Miller, S.I. Recognition of antimicrobial peptides by a bacterial sensor kinase. Cell 2005, 122, 461–472. [Google Scholar] [CrossRef]
- Prost, L.R.; Daley, M.E.; Le Sage, V.; Bader, M.W.; Le Moual, H.; Klevit, R.E.; Miller, S.I. Activation of the bacterial sensor kinase PhoQ by acidic pH. Mol. Cell 2007, 26, 165–174. [Google Scholar] [CrossRef]
- Yuan, J.; Jin, F.; Glatter, T.; Sourjik, V. Osmosensing by the bacterial PhoQ/PhoP two-component system. Proc. Natl. Acad. Sci. USA 2017, 114, E10792–E10798. [Google Scholar] [CrossRef]
- Lippa, A.M.; Goulian, M. Perturbation of the oxidizing environment of the periplasm stimulates the PhoQ/PhoP system in Escherichia coli. J. Bacteriol. 2012, 194, 1457–1463. [Google Scholar] [CrossRef]
- Zhou, D.; Han, Y.; Qin, L.; Chen, Z.; Qiu, J.; Song, Y.; Li, B.; Wang, J.; Guo, Z.; Du, Z.; et al. Transcriptome analysis of the Mg2+-responsive PhoP regulator in Yersinia pestis. FEMS Microbiol. Lett. 2005, 250, 85–95. [Google Scholar] [CrossRef][Green Version]
- Grabenstein, J.P.; Fukuto, H.S.; Palmer, L.E.; Bliska, J.B. Characterization of phagosome trafficking and identification of PhoP-regulated genes important for survival of Yersinia pestis in macrophages. Infect. Immun. 2006, 74, 3727–3741. [Google Scholar] [CrossRef]
- Li, Y.; Gao, H.; Qin, L.; Li, B.; Han, Y.; Guo, Z.; Song, Y.; Zhai, J.; Du, Z.; Wang, X.; et al. Identification and characterization of PhoP regulon members in Yersinia pestis biovar Microtus. BMC Genom. 2008, 9, 143. [Google Scholar] [CrossRef]
- Winfield, M.D.; Latifi, T.; Groisman, E.A. Transcriptional regulation of the 4-amino-4-deoxy-L-arabinose biosynthetic genes in Yersinia pestis. J. Biol. Chem. 2005, 280, 14765–14772. [Google Scholar] [CrossRef] [PubMed]
- O’Loughlin, J.L.; Spinner, J.L.; Minnich, S.A.; Kobayashi, S.D. Yersinia pestis two-component gene regulatory systems promote survival in human neutrophils. Infect. Immun. 2010, 78, 773–782. [Google Scholar] [CrossRef] [PubMed]
- Bishop, R.E. The lipid A palmitoyltransferase PagP: Molecular mechanisms and role in bacterial pathogenesis. Mol. Microbiol. 2005, 57, 900–912. [Google Scholar] [CrossRef]
- Chandler, C.E.; Harberts, E.M.; Pelletier, M.R.; Thaipisuttikul, I.; Jones, J.W.; Hajjar, A.M.; Sahl, J.W.; Goodlett, D.R.; Pride, A.C.; Rasko, D.A.; et al. Early evolutionary loss of the lipid A modifying enzyme PagP resulting in innate immune evasion in Yersinia pestis. Proc. Natl. Acad. Sci. USA 2020, 117, 22984–22991. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wang, L.; Han, Y.; Yan, Y.; Tan, Y.; Zhou, L.; Cui, Y.; Du, Z.; Wang, X.; Bi, Y.; et al. Autoregulation of PhoP/PhoQ and positive regulation of the cyclic AMP receptor protein-cyclic AMP complex by PhoP in Yersinia pestis. J. Bacteriol. 2013, 195, 1022–1030. [Google Scholar] [CrossRef] [PubMed]
- Ritzert, J.T.; Minasov, G.; Embry, R.; Schipma, M.J.; Satchell, K.J.F. The cyclic AMP receptor protein regulates quorum sensing and global gene expression in Yersinia pestis during planktonic growth and growth in biofilms. mBio 2019, 10. [Google Scholar] [CrossRef]
- Zhang, Y.; Gao, H.; Wang, L.; Xiao, X.; Tan, Y.; Guo, Z.; Zhou, D.; Yang, R. Molecular characterization of transcriptional regulation of rovA by PhoP and RovA in Yersinia pestis. PLoS ONE 2011, 6, e25484. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, L.; Fang, N.; Qu, S.; Tan, Y.; Guo, Z.; Qiu, J.; Zhou, D.; Yang, R. Reciprocal regulation of pH 6 antigen gene loci by PhoP and RovA in Yersinia pestis biovar Microtus. Future Microbiol. 2013, 8, 271–280. [Google Scholar] [CrossRef]
- Bijlsma, J.J.; Groisman, E.A. The PhoP/PhoQ system controls the intramacrophage type three secretion system of Salmonella enterica. Mol. Microbiol. 2005, 57, 85–96. [Google Scholar] [CrossRef]
- Palmer, A.D.; Kim, K.; Slauch, J.M. PhoP-mediated repression of the SPI1 type 3 secretion system in Salmonella enterica serovar Typhimurium. J. Bacteriol. 2019, 201, e00264-19. [Google Scholar] [CrossRef]
- Perez, J.C.; Groisman, E.A. Transcription factor function and promoter architecture govern the evolution of bacterial regulons. Proc. Natl. Acad. Sci. USA 2009, 106, 4319–4324. [Google Scholar] [CrossRef] [PubMed]
- Viboud, G.I.; Bliska, J.B. Yersinia outer proteins: Role in modulation of host cell signaling responses and pathogenesis. Annu. Rev. Microbiol. 2005, 59, 69–89. [Google Scholar] [CrossRef]
- Prentice, M.B.; Rahalison, L. Plague. Lancet 2007, 369, 1196–1207. [Google Scholar] [CrossRef]
- Vadyvaloo, V.; Jarrett, C.; Sturdevant, D.E.; Sebbane, F.; Hinnebusch, B.J. Transit through the flea vector induces a pretransmission innate immunity resistance phenotype in Yersinia pestis. PLoS Pathog. 2010, 6, e1000783. [Google Scholar] [CrossRef] [PubMed]
- Pujol, C.; Bliska, J.B. Turning Yersinia pathogenesis outside in: Subversion of macrophage function by intracellular yersiniae. Clin. Immunol. 2005, 114, 216–226. [Google Scholar] [CrossRef] [PubMed]
- Finegold, M.J. Pneumonic plague in monkeys. An. electron microscopic study. Am. J. Pathol. 1969, 54, 167–185. [Google Scholar]
- Lukaszewski, R.A.; Kenny, D.J.; Taylor, R.; Rees, D.G.; Hartley, M.G.; Oyston, P.C. Pathogenesis of Yersinia pestis infection in BALB/c mice: Effects on host macrophages and neutrophils. Infect. Immun. 2005, 73, 7142–7150. [Google Scholar] [CrossRef]
- Bosio, C.M.; Goodyear, A.W.; Dow, S.W. Early interaction of Yersinia pestis with APCs in the lung. J. Immunol. 2005, 175, 6750–6756. [Google Scholar] [CrossRef]
- Cavanaugh, D.C.; Randall, R. The role of multiplication of Pasteurella pestis in mononuclear phagocytes in the pathogenesis of flea-borne plague. J. Immunol. 1959, 83, 348–363. [Google Scholar]
- Charnetzky, W.T.; Shuford, W.W. Survival and growth of Yersinia pestis within macrophages and an effect of the loss of the 47-megadalton plasmid on growth in macrophages. Infect. Immun. 1985, 47, 234–241. [Google Scholar] [CrossRef]
- Shannon, J.G.; Hasenkrug, A.M.; Dorward, D.W.; Nair, V.; Carmody, A.B.; Hinnebusch, B.J. Yersinia pestis subverts the dermal neutrophil response in a mouse model of bubonic plague. mBio 2013, 4, e00170-13. [Google Scholar] [CrossRef] [PubMed]
- Shannon, J.G.; Bosio, C.F.; Hinnebusch, B.J. Dermal neutrophil, macrophage and dendritic cell responses to Yersinia pestis transmitted by fleas. PLoS Pathog. 2015, 11, e1004734. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, R.J.; Lane, M.C.; Wagner, N.J.; Weening, E.H.; Miller, V.L. Dissemination of a highly virulent pathogen: Tracking the early events that define infection. PLoS Pathog. 2015, 11, e1004587. [Google Scholar] [CrossRef] [PubMed]
- Oyston, P.C.; Dorrell, N.; Williams, K.; Li, S.R.; Green, M.; Titball, R.W.; Wren, B.W. The response regulator PhoP is important for survival under conditions of macrophage-induced stress and virulence in Yersinia pestis. Infect. Immun. 2000, 68, 3419–3425. [Google Scholar] [CrossRef]
- Straley, S.C.; Harmon, P.A. Growth in mouse peritoneal macrophages of Yersinia pestis lacking established virulence determinants. Infect. Immun. 1984, 45, 649–654. [Google Scholar] [CrossRef]
- Hitchen, P.G.; Prior, J.L.; Oyston, P.C.; Panico, M.; Wren, B.W.; Titball, R.W.; Morris, H.R.; Dell, A. Structural characterization of lipo-oligosaccharide (LOS) from Yersinia pestis: Regulation of LOS structure by the PhoPQ system. Mol. Microbiol. 2002, 44, 1637–1650. [Google Scholar] [CrossRef]
- Fukuto, H.S.; Svetlanov, A.; Palmer, L.E.; Karzai, A.W.; Bliska, J.B. Global gene expression profiling of Yersinia pestis replicating inside macrophages reveals the roles of a putative stress-induced operon in regulating type III secretion and intracellular cell division. Infect. Immun. 2010, 78, 3700–3715. [Google Scholar] [CrossRef]
- Klein, K.A.; Fukuto, H.S.; Pelletier, M.; Romanov, G.; Grabenstein, J.P.; Palmer, L.E.; Ernst, R.; Bliska, J.B. A transposon site hybridization screen identifies galU and wecBC as important for survival of Yersinia pestis in murine macrophages. J. Bacteriol. 2012, 194, 653–662. [Google Scholar] [CrossRef]
- Ford, D.C.; Joshua, G.W.P.; Wren, B.W.; Oyston, P.C.F. The importance of the magnesium transporter MgtB for virulence of Yersinia pseudotuberculosis and Yersinia pestis. Microbiology 2014, 160 Pt 12, 2710–2717. [Google Scholar] [CrossRef]
- Pujol, C.; Klein, K.A.; Romanov, G.A.; Palmer, L.E.; Cirota, C.; Zhao, Z.; Bliska, J.B. Yersinia pestis can reside in autophagosomes and avoid xenophagy in murine macrophages by preventing vacuole acidification. Infect. Immun. 2009, 77, 2251–2261. [Google Scholar] [CrossRef]
- Connor, M.G.; Pulsifer, A.R.; Chung, D.; Rouchka, E.C.; Ceresa, B.K.; Lawrenz, M.B. Yersinia pestis targets the host endosome recycling pathway during the biogenesis of the Yersinia-containing vacuole to avoid killing by macrophages. mBio 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Connor, M.G.; Pulsifer, A.R.; Price, C.T.; Abu Kwaik, Y.; Lawrenz, M.B. Yersinia pestis requires host Rab1b for survival in macrophages. PLoS Pathog. 2015, 11, e1005241. [Google Scholar] [CrossRef] [PubMed]
- Spinner, J.L.; Winfree, S.; Starr, T.; Shannon, J.G.; Nair, V.; Steele-Mortimer, O.; Hinnebusch, B.J. Yersinia pestis survival and replication within human neutrophil phagosomes and uptake of infected neutrophils by macrophages. J. Leukoc. Biol. 2014, 95, 389–398. [Google Scholar] [CrossRef] [PubMed]
- Bliska, J.B. (Geisel School of Medicine at Dartmouth, Hanover, NH, USA). Personal communication, 2020.
- Bozue, J.; Mou, S.; Moody, K.L.; Cote, C.K.; Trevino, S.; Fritz, D.; Worsham, P. The role of the phoPQ operon in the pathogenesis of the fully virulent CO92 strain of Yersinia pestis and the IP32953 strain of Yersinia pseudotuberculosis. Microb. Pathog. 2011, 50, 314–321. [Google Scholar] [CrossRef] [PubMed]
- Grabenstein, J.P.; Marceau, M.; Pujol, C.; Simonet, M.; Bliska, J.B. The response regulator PhoP of Yersinia pseudotuberculosis is important for replication in macrophages and for virulence. Infect. Immun. 2004, 72, 4973–4984. [Google Scholar] [CrossRef] [PubMed]
- Fields, P.I.; Groisman, E.A.; Heffron, F. A Salmonella locus that controls resistance to microbicidal proteins from phagocytic cells. Science 1989, 243 Pt 1, 1059–1062. [Google Scholar] [CrossRef]
- Galan, J.E.; Curtiss, R., 3rd. Virulence and vaccine potential of phoP mutants of Salmonella typhimurium. Microb. Pathog. 1989, 6, 433–443. [Google Scholar]
- Miller, S.I.; Kukral, A.M.; Mekalanos, J.J. A two-component regulatory system (phoP phoQ) controls Salmonella typhimurium virulence. Proc. Natl. Acad. Sci. USA 1989, 86, 5054–5058. [Google Scholar] [CrossRef]
- Pechous, R.D.; Sivaraman, V.; Price, P.A.; Stasulli, N.M.; Goldman, W.E. Early host cell targets of Yersinia pestis during primary pneumonic plague. PLoS Pathog. 2013, 9, e1003679. [Google Scholar] [CrossRef]
- St John, A.L.; Ang, W.X.G.; Huang, M.N.; Kunder, C.A.; Chan, E.W.; Gunn, M.D.; Abraham, S.N. S1P-Dependent trafficking of intracellular Yersinia pestis through lymph nodes establishes Buboes and systemic infection. Immunity 2014, 41, 440–450. [Google Scholar] [CrossRef]
- Ye, Z.; Kerschen, E.J.; Cohen, D.A.; Kaplan, A.M.; van Rooijen, N.; Straley, S.C. Gr1+ cells control growth of YopM-negative Yersinia pestis during systemic plague. Infect. Immun. 2009, 77, 3791–3806. [Google Scholar] [CrossRef] [PubMed]
- Rebeil, R.; Jarrett, C.O.; Driver, J.D.; Ernst, R.K.; Oyston, P.C.; Hinnebusch, B.J. Induction of the Yersinia pestis PhoP-PhoQ regulatory system in the flea and its role in producing a transmissible infection. J. Bacteriol. 2013, 195, 1920–1930. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Vadyvaloo, V.; Viall, A.K.; Jarrett, C.O.; Hinz, A.K.; Sturdevant, D.E.; Joseph Hinnebusch, B. Role of the PhoP-PhoQ gene regulatory system in adaptation of Yersinia pestis to environmental stress in the flea digestive tract. Microbiology 2015, 161, 1198–1210. [Google Scholar] [CrossRef] [PubMed]
- Willias, S.P.; Chauhan, S.; Lo, C.C.; Chain, P.S.; Motin, V.L. CRP-mediated carbon catabolite regulation of Yersinia pestis biofilm formation is enhanced by the carbon storage regulator protein, CsrA. PLoS ONE 2015, 10, e0135481. [Google Scholar] [CrossRef]
- Bontemps-Gallo, S.; Fernandez, M.; Dewitte, A.; Raphael, E.; Gherardini, F.C.; Elizabeth, P.; Koch, L.; Biot, F.; Reboul, A.; Sebbane, F. Nutrient depletion may trigger the Yersinia pestis OmpR-EnvZ regulatory system to promote flea-borne plague transmission. Mol. Microbiol. 2019, 112, 1471–1482. [Google Scholar] [CrossRef]
- Martinez-Chavarria, L.C.; Sagawa, J.; Irons, J.; Hinz, A.K.; Lemon, A.; Graca, T.; Downs, D.M.; Vadyvaloo, V. Putative horizontally acquired genes, highly transcribed during Yersinia pestis flea infection, are induced by hyperosmotic stress and function in aromatic amino acid metabolism. J. Bacteriol. 2020, 202. [Google Scholar] [CrossRef]
- Harari, O.; Park, S.Y.; Huang, H.; Groisman, E.A.; Zwir, I. Defining the plasticity of transcription factor binding sites by deconstructing DNA consensus sequences: The PhoP-binding sites among gamma/enterobacteria. PLoS Comput. Biol. 2010, 6, e1000862. [Google Scholar] [CrossRef]
- Ben-Efraim, S.; Aronson, M.; Bichowsky-Slomnicki, L. New antigenic component of Pasteurella pestis formed under specified conditions of pH and temperature. J. Bacteriol. 1961, 81, 704–714. [Google Scholar] [CrossRef]
- Aoyagi, K.L.; Brooks, B.D.; Bearden, S.W.; Montenieri, J.A.; Gage, K.L.; Fisher, M.A. LPS modification promotes maintenance of Yersinia pestis in fleas. Microbiology 2015, 161 Pt 3, 628–638. [Google Scholar] [CrossRef]
- Gerdes, K.; Christensen, S.K.; Løbner-Olesen, A. Prokaryotic toxin-antitoxin stress response loci. Nat. Rev. Microbiol. 2005, 3, 371–382. [Google Scholar] [CrossRef]
- Wood, T.K.; Knabel, S.J.; Kwan, B.W. Bacterial persister cell formation and dormancy. Appl. Environ. Microbiol. 2013, 79, 7116–7121. [Google Scholar] [CrossRef]
- Gollan, B.; Grabe, G.; Michaux, C.; Helaine, S. Bacterial persisters and infection: Past, present, and progressing. Annu. Rev. Microbiol. 2019, 73, 359–385. [Google Scholar] [CrossRef]
- Fukuto, H.S.; Vadyvaloo, V.; McPhee, J.B.; Poinar, H.N.; Holmes, E.C.; Bliska, J.B. A single amino acid change in the response regulator PhoP, acquired during Yersinia pestis evolution, affects PhoP target. Gene transcription and polymyxin B susceptibility. J. Bacteriol. 2018, 200. [Google Scholar] [CrossRef] [PubMed]
- Lemon, A.; Silva-Rohwer, A.; Sagawa, J.; Vadyvaloo, V. Co-infection assay to determine Yersinia pestis competitive fitness in fleas. Methods Mol. Biol. 2019, 2010, 153–166. [Google Scholar] [PubMed]
- Lemon, A.; Cherzan, N.; Vadyvaloo, V. Influence of temperature on development of Yersinia pestis foregut blockage in Xenopsylla cheopis (Siphonaptera: Pulicidae) and Oropsylla montana (Siphonaptera: Ceratophyllidae). J. Med. Entomol. 2020, 57, 1997–2007. [Google Scholar] [CrossRef] [PubMed]
- Hinnebusch, B.J. The evolution of flea-borne transmission in Yersinia pestis. Curr. Issues Mol. Biol. 2005, 7, 197–212. [Google Scholar]
- Hinnebusch, B.J.; Bland, D.M.; Bosio, C.F.; Jarrett, C.O. Comparative ability of Oropsylla montana and Xenopsylla cheopis fleas to transmit Yersinia pestis by two different mechanisms. PLoS Negl. Trop. Dis. 2017, 11, e0005276. [Google Scholar] [CrossRef]
- Benavides-Montano, J.A.; Vadyvaloo, V. Yersinia pestis resists predation by Acanthamoeba castellanii and exhibits prolonged intracellular survival. Appl. Environ. Microbiol. 2017, 83. [Google Scholar] [CrossRef]
- Wren, B.W. The Yersiniae—A model genus to study the rapid evolution of bacterial pathogens. Nat. Rev. Microbiol. 2003, 1, 55–64. [Google Scholar] [CrossRef]
- Zimbler, D.L.; Schroeder, J.A.; Eddy, J.L.; Lathem, W.W. Early emergence of Yersinia pestis as a severe respiratory pathogen. Nat. Commun. 2015, 6, 7487. [Google Scholar] [CrossRef]
- Sun, Y.C.; Jarrett, C.O.; Bosio, C.F.; Hinnebusch, B.J. Retracing the evolutionary path that led to flea-borne transmission of Yersinia pestis. Cell Host Microbe 2014, 15, 578–586. [Google Scholar] [CrossRef] [PubMed]
- Bos, K.I.; Schuenemann, V.J.; Golding, G.B.; Burbano, H.A.; Waglechner, N.; Coombes, B.K.; McPhee, J.B.; DeWitte, S.N.; Meyer, M.; Schmedes, S.; et al. A draft genome of Yersinia pestis from victims of the Black Death. Nature 2011, 478, 506–510. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Yu, C.; Yan, Y.; Li, D.; Li, Y.; Jombart, T.; Weinert, L.A.; Wang, Z.; Guo, Z.; Xu, L.; et al. Historical variations in mutation rate in an epidemic pathogen, Yersinia pestis. Proc. Natl. Acad. Sci. USA 2013, 110, 577–582. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fukuto, H.S.; Viboud, G.I.; Vadyvaloo, V. The Diverse Roles of the Global Transcriptional Regulator PhoP in the Lifecycle of Yersinia pestis. Pathogens 2020, 9, 1039. https://doi.org/10.3390/pathogens9121039
Fukuto HS, Viboud GI, Vadyvaloo V. The Diverse Roles of the Global Transcriptional Regulator PhoP in the Lifecycle of Yersinia pestis. Pathogens. 2020; 9(12):1039. https://doi.org/10.3390/pathogens9121039
Chicago/Turabian StyleFukuto, Hana S., Gloria I. Viboud, and Viveka Vadyvaloo. 2020. "The Diverse Roles of the Global Transcriptional Regulator PhoP in the Lifecycle of Yersinia pestis" Pathogens 9, no. 12: 1039. https://doi.org/10.3390/pathogens9121039
APA StyleFukuto, H. S., Viboud, G. I., & Vadyvaloo, V. (2020). The Diverse Roles of the Global Transcriptional Regulator PhoP in the Lifecycle of Yersinia pestis. Pathogens, 9(12), 1039. https://doi.org/10.3390/pathogens9121039



