Molecular Detection of Spotted-Fever Group Rickettsiae in Ticks Collected from Domestic and Wild Animals in Corsica, France
Abstract
1. Introduction
2. Results
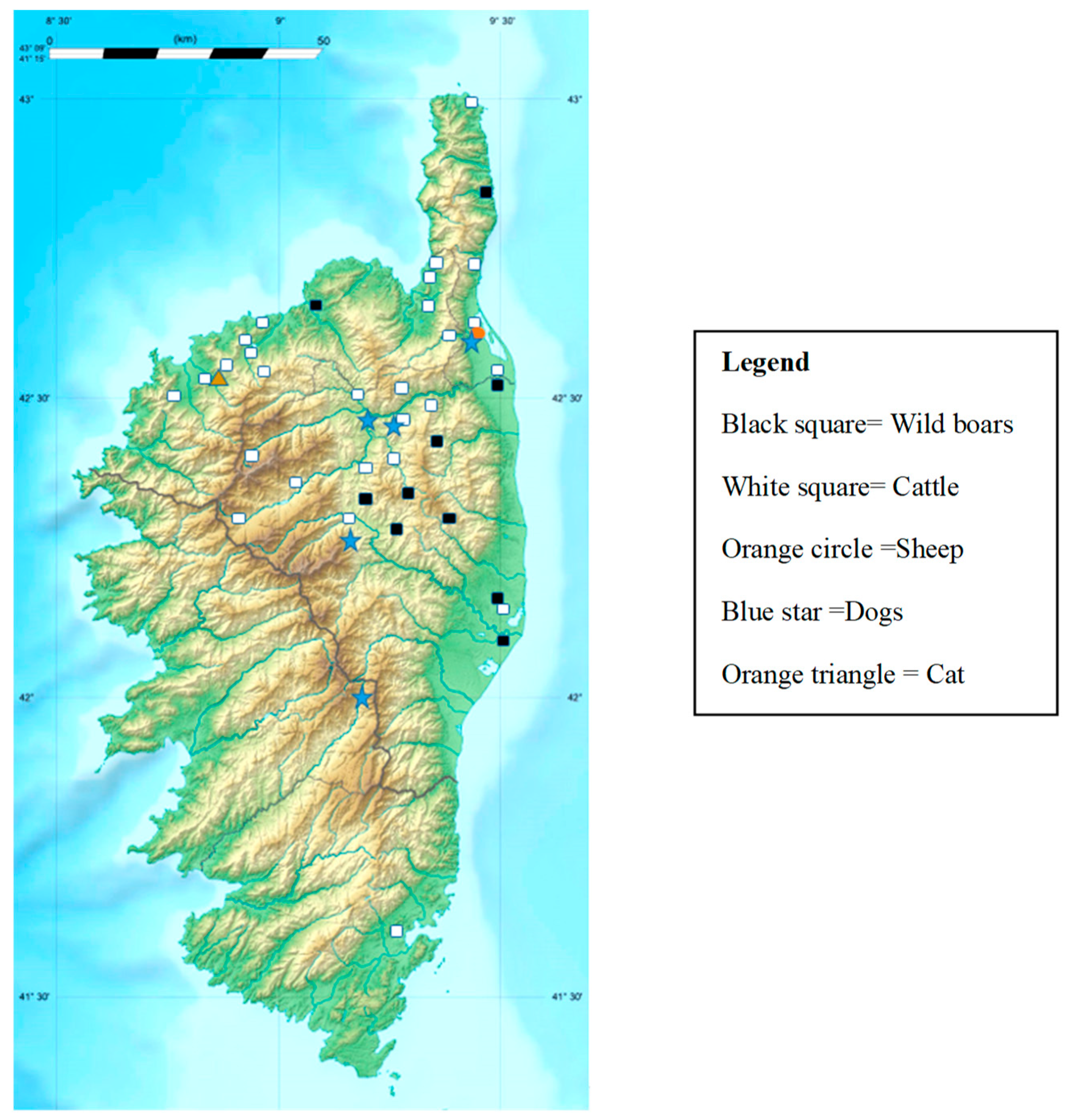
2.1. Sampling and Identification of Ticks
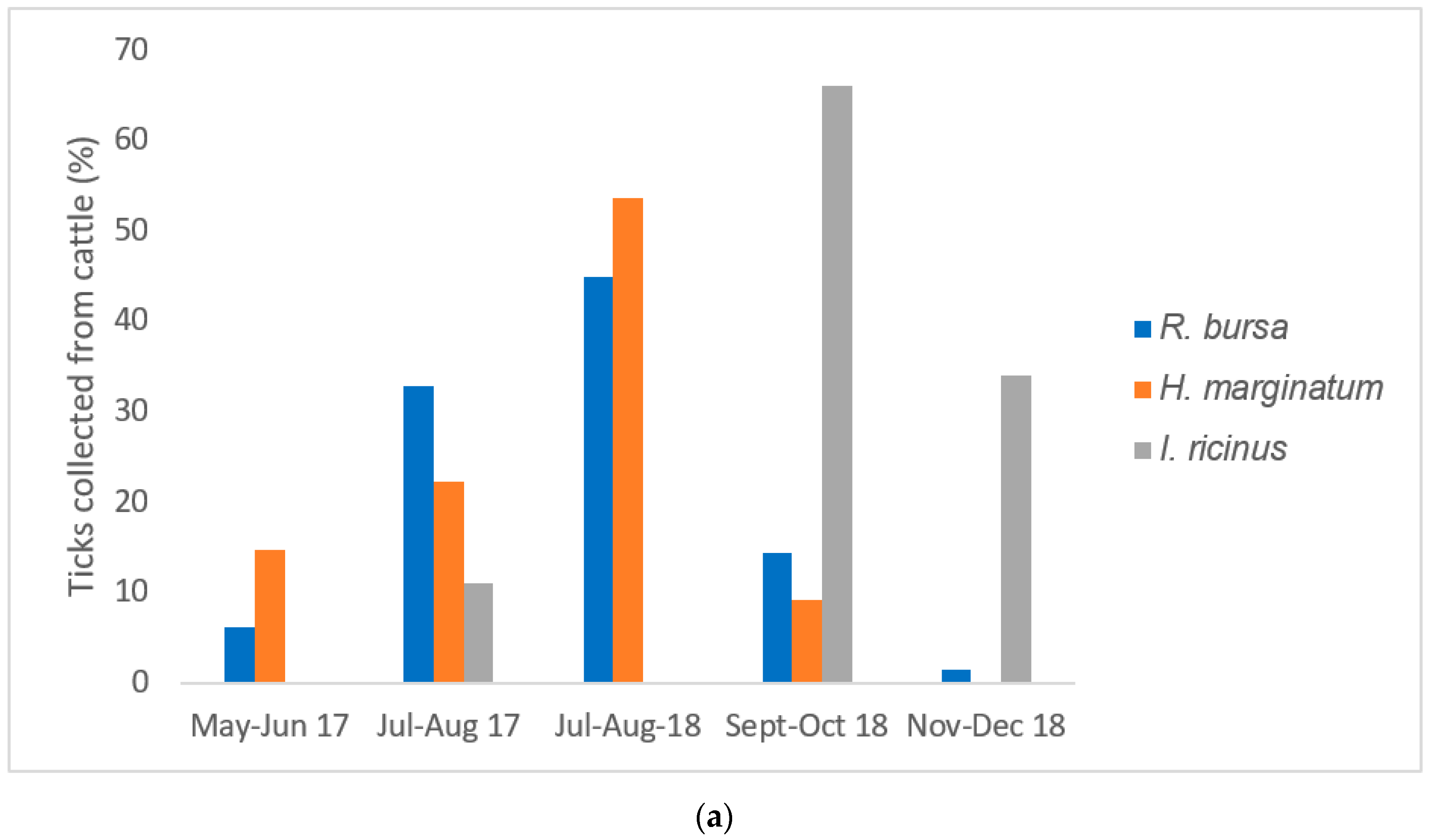
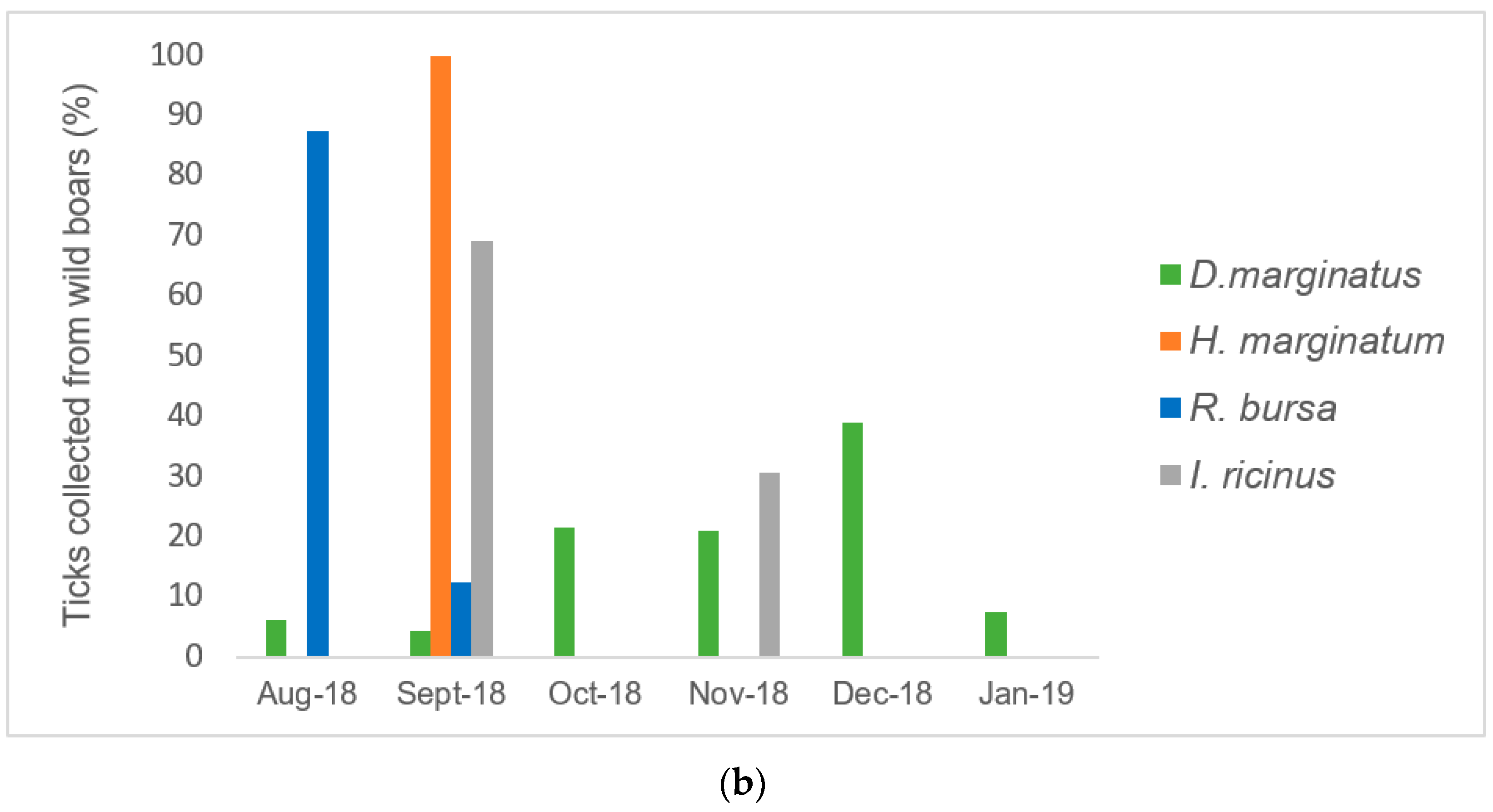
2.2. Infestation Rate in Cattle, Sheep, and Wild Boars
2.3. Detection of Rickettsial DNA
2.4. Rickettsia Species Identification Based on Sequence Analysis
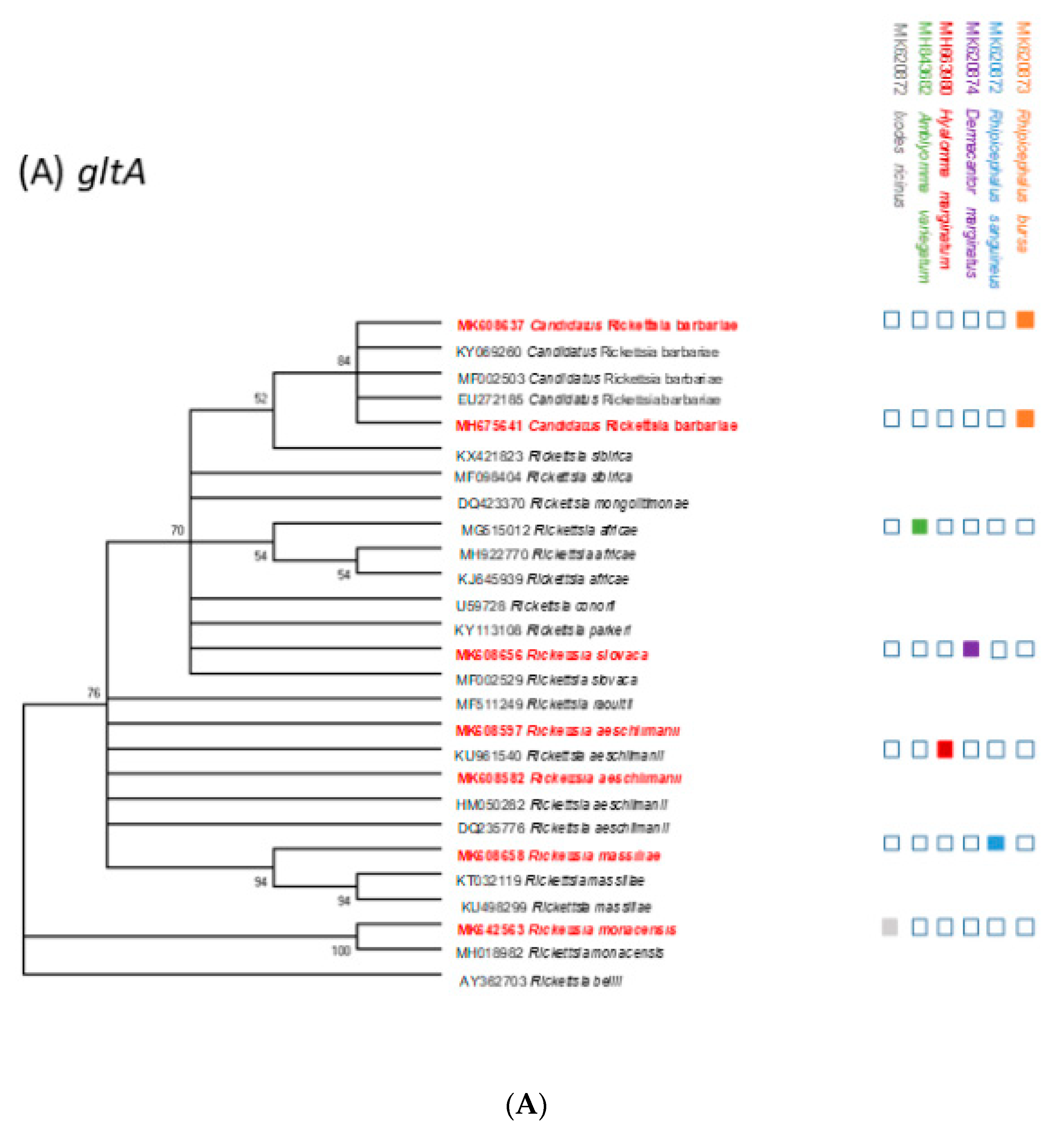
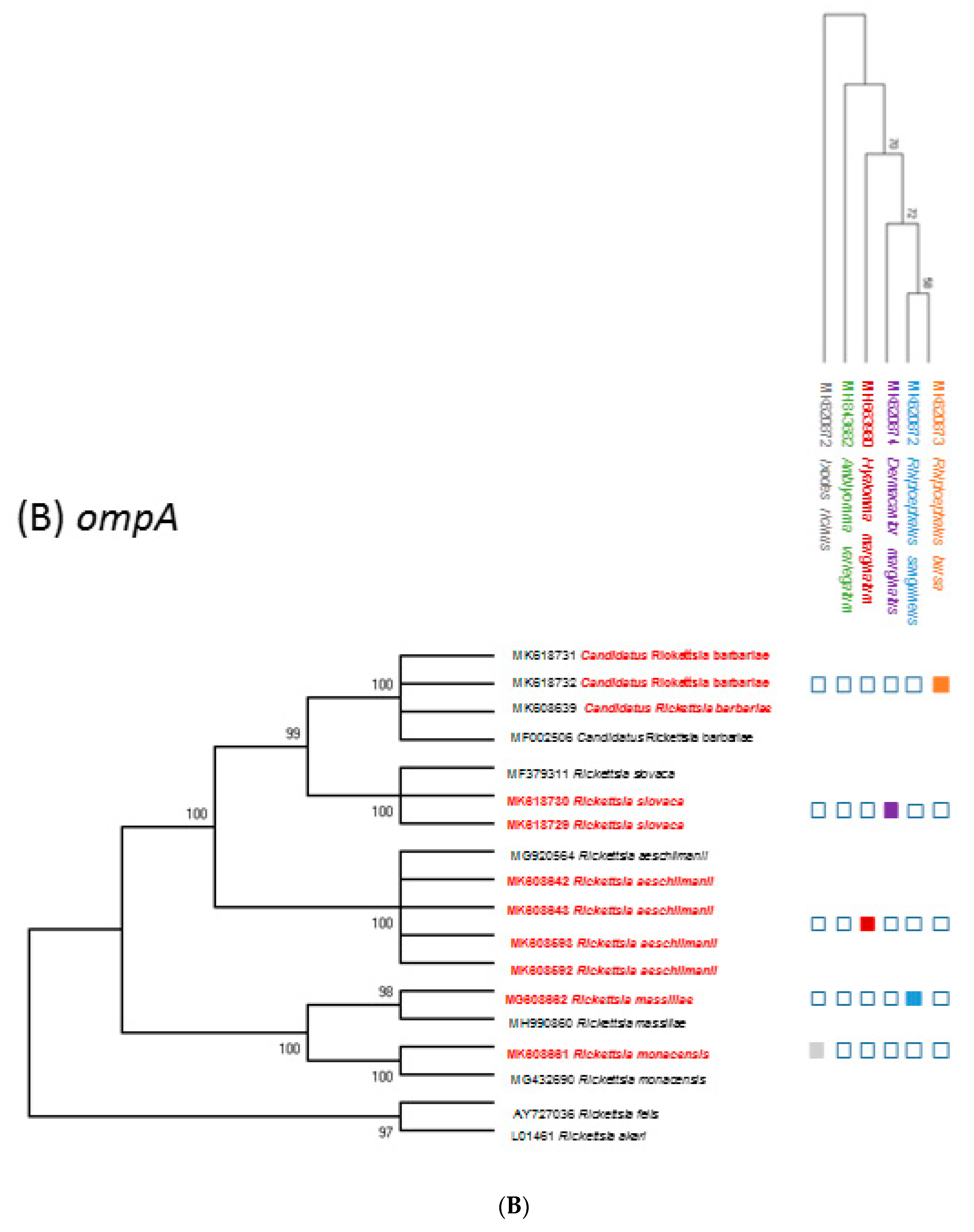
2.5. Phylogenetic Analysis
3. Discussion
4. Methods
4.1. Study Area and Ticks
4.2. Morphologic Identification of Ticks at the Species Level
4.3. DNA Extraction
4.4. Molecular Identification of Ticks at the Species Level
4.5. PCR Detection of Rickettsia
4.6. Statistical Analysis
5. Conclusions
6. Declarations
6.1. Ethics Approval and Consent to Participate
6.2. Availability of Data and Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| DEBONEL | Dermacantor-borne necrosis lymphadenopathy |
| gltA | citrate synthase |
| MLE | maximum likelihood estimation |
| MSF | Mediterranean spotted fever |
| ompA | outer membrane proteins A |
| PCR | polymerase chain reaction |
| SFG | spotted fever group |
| TIBOLA | tick-borne lymphadenopathy |
References
- Parola, P.; Paddock, C.D.; Socolovschi, C.; Labruna, M.B.; Mediannikov, O.; Kernif, T.; Abdad, M.Y.; Stenos, J.; Bitam, I.; Fournier, P.E.; et al. Update on tick-borne rickettsioses around the world: A geographic approach. Clin. Microbiol. Rev. 2013, 26, 657–702. [Google Scholar] [CrossRef]
- Merhej, V.; Angelakis, E.; Socolovschi, C.; Raoult, D. Rickettsial evolution in the light of comparative genomics. Biol. Rev. Camb. Philos. Soc. 2011, 86, 379–405. [Google Scholar] [CrossRef]
- Thu, M.J.; Qiu, Y.; Matsuno, K.; Kajihara, M.; Mori-Kajihara, A.; Omori, R.; Monma, N.; Chiba, K.; Seto, J.; Gokuden, M.; et al. Diversity of spotted fever group rickettsiae and their association with host ticks in Japan. Sci. Rep. 2019, 9, 1500. [Google Scholar] [CrossRef]
- Merhej, V.; Angelakis, E.; Socolovschi, C.; Raoult, D. Genotyping, evolution and epidemiological findings of Rickettsia species. Infect. Genet. Evol. 2014, 25, 122–137. [Google Scholar] [CrossRef]
- Hornok, S.; Csorgo, T.; de la Fuente, J.; Gyuranecz, M.; Privigyei, C.; Meli, M.L.; Kreizinger, Z.; Gonczi, E.; Fernandez de Mera, I.G.; Hofmann-Lehmann, R. Synanthropic birds associated with high prevalence of tick-borne rickettsiae and with the first detection of Rickettsia aeschlimannii in Hungary. Vector Borne Zoonotic Dis. 2013, 13, 77–83. [Google Scholar] [CrossRef]
- Ibarra, V.; Oteo, J.A.; Portillo, A.; Santibanez, S.; Blanco, J.R.; Metola, L.; Eiros, J.M.; Perez-Martinez, L.; Sanz, M. Rickettsia slovaca infection: DEBONEL/TIBOLA. Ann. N. Y. Acad. Sci. 2006, 1078, 206–214. [Google Scholar] [CrossRef]
- Parola, P.; Rovery, C.; Rolain, J.M.; Brouqui, P.; Davoust, B.; Raoult, D. Rickettsia slovaca and R. raoultii in tick-borne Rickettsioses. Emerg. Infect. Dis. 2009, 15, 1105–1108. [Google Scholar] [CrossRef]
- Cicculli, V.; Capai, L.; Quilichini, Y.; Masse, S.; Fernandez-Alvarez, A.; Minodier, L.; Bompard, P.; Charrel, R.; Falchi, A. Molecular investigation of tick-borne pathogens in ixodid ticks infesting domestic animals (cattle and sheep) and small rodents (black rats) of Corsica, France. Ticks Tick Borne Dis. 2019, 10, 606–610. [Google Scholar] [CrossRef]
- Grech-Angelini, S.; Stachurski, F.; Lancelot, R.; Boissier, J.; Allienne, J.F.; Marco, S.; Maestrini, O.; Uilenberg, G. Ticks (Acari: Ixodidae) infesting cattle and some other domestic and wild hosts on the French Mediterranean island of Corsica. Parasit. Vectors 2016, 9, 582. [Google Scholar] [CrossRef]
- Chisu, V.; Leulmi, H.; Masala, G.; Piredda, M.; Foxi, C.; Parola, P. Detection of Rickettsia hoogstraalii, Rickettsia helvetica, Rickettsia massiliae, Rickettsia slovaca and Rickettsia aeschlimannii in ticks from Sardinia, Italy. Ticks Tick Borne Dis. 2017, 8, 347–352. [Google Scholar] [CrossRef]
- Belkahia, H.; Ben Said, M.; Ghribi, R.; Selmi, R.; Ben Asker, A.; Yahiaoui, M.; Bousrih, M.; Daaloul-Jedidi, M.; Messadi, L. Molecular detection, genotyping and phylogeny of Anaplasma spp. in Rhipicephalus ticks from Tunisia. Acta Trop. 2019, 191, 38–49. [Google Scholar] [CrossRef]
- Pereira, A.; Parreira, R.; Cotao, A.J.; Nunes, M.; Vieira, M.L.; Azevedo, F.; Campino, L.; Maia, C. Tick-borne bacteria and protozoa detected in ticks collected from domestic animals and wildlife in central and southern Portugal. Ticks Tick Borne Dis. 2018, 9, 225–234. [Google Scholar] [CrossRef]
- Mierzejewska, E.J.; Alsarraf, M.; Behnke, J.M.; Bajer, A. The effect of changes in agricultural practices on the density of Dermacentor reticulatus ticks. Vet. Parasitol. 2015, 211, 259–265. [Google Scholar] [CrossRef]
- Mura, A.; Masala, G.; Tola, S.; Satta, G.; Fois, F.; Piras, P.; Rolain, J.M.; Raoult, D.; Parola, P. First direct detection of rickettsial pathogens and a new rickettsia, ‘Candidatus Rickettsia barbariae’, in ticks from Sardinia, Italy. Clin. Microbiol. Infect. 2008, 14, 1028–1033. [Google Scholar]
- Socolovschi, C.; Reynaud, P.; Kernif, T.; Raoult, D.; Parola, P. Rickettsiae of spotted fever group, Borrelia valaisiana, and Coxiella burnetii in ticks on passerine birds and mammals from the Camargue in the south of France. Ticks Tick Borne Dis. 2012, 3, 355–360. [Google Scholar] [CrossRef]
- Chochlakis, D.; Ioannou, I.; Sandalakis, V.; Dimitriou, T.; Kassinis, N.; Papadopoulos, B.; Tselentis, Y.; Psaroulaki, A. Spotted fever group Rickettsiae in ticks in Cyprus. Microb. Ecol. 2012, 63, 314–323. [Google Scholar] [CrossRef]
- Vanegas, A.; Keller, C.; Kruger, A.; Manchang, T.K.; Hagen, R.M.; Frickmann, H.; Veit, A.; Achukwi, M.D.; Krucken, J.; Poppert, S. Molecular detection of spotted fever group rickettsiae in ticks from Cameroon. Ticks Tick Borne Dis. 2018, 9, 1049–1056. [Google Scholar] [CrossRef]
- Abdelkadir, K.; Palomar, A.M.; Portillo, A.; Oteo, J.A.; Ait-Oudhia, K.; Khelef, D. Presence of Rickettsia aeschlimannii, ‘Candidatus Rickettsia barbariae’ and Coxiella burnetii in ticks from livestock in Northwestern Algeria. Ticks Tick Borne Dis. 2019, 10, 924–928. [Google Scholar] [CrossRef]
- Fernandez de Mera, I.G.; Blanda, V.; Torina, A.; Dabaja, M.F.; El Romeh, A.; Cabezas-Cruz, A.; de la Fuente, J. Identification and molecular characterization of spotted fever group rickettsiae in ticks collected from farm ruminants in Lebanon. Ticks Tick Borne Dis. 2018, 9, 104–108. [Google Scholar] [CrossRef]
- Ereqat, S.; Nasereddin, A.; Al-Jawabreh, A.; Azmi, K.; Harrus, S.; Mumcuoglu, K.; Apanaskevich, D.; Abdeen, Z. Molecular Detection and Identification of Spotted Fever Group Rickettsiae in Ticks Collected from the West Bank, Palestinian Territories. PLoS Negl. Trop. Dis. 2016, 10, e0004348. [Google Scholar] [CrossRef] [PubMed]
- Papa, A.; Xanthopoulou, K.; Kotriotsiou, T.; Papaioakim, M.; Sotiraki, S.; Chaligiannis, I.; Maltezos, E. Rickettsia species in human-parasitizing ticks in Greece. Trans. R. Soc. Trop. Med. Hyg. 2016, 110, 299–304. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, K.; Parola, P.; Brouqui, P.; Raoult, D. Rickettsia aeschlimannii in Hyalomma ticks from Corsica. Eur. J. Clin. Microbiol. Infect. Dis. 2004, 23, 732–734. [Google Scholar] [CrossRef] [PubMed]
- Punda-Polic, V.; Petrovec, M.; Trilar, T.; Duh, D.; Bradaric, N.; Klismanic, Z.; Avsic-Zupanc, T. Detection and identification of spotted fever group rickettsiae in ticks collected in southern Croatia. Exp. Appl. Acarol. 2002, 28, 169–176. [Google Scholar] [CrossRef] [PubMed]
- Chitimia-Dobler, L.; Schaper, S.; Riess, R.; Bitterwolf, K.; Frangoulidis, D.; Bestehorn, M.; Springer, A.; Oehme, R.; Drehmann, M.; Lindau, A.; et al. Imported Hyalomma ticks in Germany in 2018. Parasit. Vectors 2019, 12, 134. [Google Scholar] [CrossRef] [PubMed]
- Parola, P.; Paddock, C.D.; Raoult, D. Tick-borne rickettsioses around the world: Emerging diseases challenging old concepts. Clin. Microbiol. Rev. 2005, 18, 719–756. [Google Scholar] [CrossRef] [PubMed]
- Raoult, D.; Fournier, P.E.; Abboud, P.; Caron, F. First documented human Rickettsia aeschlimannii infection. Emerg. Infect. Dis. 2002, 8, 748–749. [Google Scholar] [CrossRef] [PubMed]
- Rukhkian, M. Vertical migrations of hungry imagoes of the tick Dermacentor marginatus. Parazitologiia 1987, 21, 680–683. [Google Scholar]
- Masala, G.; Chisu, V.; Satta, G.; Socolovschi, C.; Raoult, D.; Parola, P. Rickettsia slovaca from Dermacentor marginatus ticks in Sardinia, Italy. Ticks Tick Borne Dis. 2012, 3, 393–395. [Google Scholar] [CrossRef]
- Martello, E.; Mannelli, A.; Grego, E.; Ceballos, L.A.; Ragagli, C.; Stella, M.C.; Tomassone, L. Borrelia burgdorferi sensu lato and spotted fever group rickettsiae in small rodents and attached ticks in the Northern Apennines, Italy. Ticks Tick Borne Dis. 2019, 10, 862–867. [Google Scholar] [CrossRef]
- Turebekov, N.; Abdiyeva, K.; Yegemberdiyeva, R.; Dmitrovsky, A.; Yeraliyeva, L.; Shapiyeva, Z.; Amirbekov, A.; Oradova, A.; Kachiyeva, Z.; Ziyadina, L.; et al. Prevalence of Rickettsia species in ticks including identification of unknown species in two regions in Kazakhstan. Parasit. Vectors 2019, 12, 197. [Google Scholar] [CrossRef] [PubMed]
- Raoult, D.; Lakos, A.; Fenollar, F.; Beytout, J.; Brouqui, P.; Fournier, P.E. Spotless rickettsiosis caused by Rickettsia slovaca and associated with Dermacentor ticks. Clin. Infect. Dis. 2002, 34, 1331–1336. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, K.; Ogawa, M.; Brouqui, P.; Raoult, D.; Parola, P. Transmission of Rickettsia massiliae in the tick, Rhipicephalus turanicus. Med. Vet. Entomol. 2005, 19, 263–270. [Google Scholar] [CrossRef] [PubMed]
- Beeler, E.; Abramowicz, K.F.; Zambrano, M.L.; Sturgeon, M.M.; Khalaf, N.; Hu, R.; Dasch, G.A.; Eremeeva, M.E. A focus of dogs and Rickettsia massiliae-infected Rhipicephalus sanguineus in California. Am. J. Trop. Med. Hyg. 2011, 84, 244–249. [Google Scholar] [CrossRef] [PubMed]
- Estrada-Pena, A.; Roura, X.; Sainz, A.; Miro, G.; Solano-Gallego, L. Species of ticks and carried pathogens in owned dogs in Spain: Results of a one-year national survey. Ticks Tick Borne Dis. 2017, 8, 443–452. [Google Scholar] [CrossRef] [PubMed]
- Lauzi, S.; Maia, J.P.; Epis, S.; Marcos, R.; Pereira, C.; Luzzago, C.; Santos, M.; Puente-Payo, P.; Giordano, A.; Pajoro, M.; et al. Molecular detection of Anaplasma platys, Ehrlichia canis, Hepatozoon canis and Rickettsia monacensis in dogs from Maio Island of Cape Verde archipelago. Ticks Tick Borne Dis. 2016, 7, 964–969. [Google Scholar] [CrossRef] [PubMed]
- Estrada-Pena, A.; Gray, J.S.; Kahl, O.; Lane, R.S.; Nijhof, A.M. Research on the ecology of ticks and tick-borne pathogens—Methodological principles and caveats. Front. Cell. Infect. Microbiol. 2013, 3, 29. [Google Scholar] [CrossRef]
- Estrada-Pena, A.; Bouattour, A.; Camicas, J.L.; Walker, A.R. Ticks of Veterinary and Medical Importance: The Mediterranean Basin; A guide of identification of species; University of Zaragoza Press, Zaragoza, Ed.; University of Zaragoza Press: Zaragoza, Spain, 2004. [Google Scholar]
- Estrada-Pena, A.; Nava, S.; Petney, T. Description of all the stages of Ixodes inopinatus n. sp. (Acari: Ixodidae). Ticks Tick Borne Dis. 2014, 5, 734–743. [Google Scholar] [CrossRef]
- Black, W.C.T.; Piesman, J. Phylogeny of hard-and soft-tick taxa (Acari: Ixodida) based on mitochondrial 16S rDNA sequences. Proc. Natl. Acad. Sci. USA 1994, 91, 10034–10038. [Google Scholar] [CrossRef]
- Roux, V.; Rydkina, E.; Eremeeva, M.; Raoult, D. Citrate synthase gene comparison, a new tool for phylogenetic analysis, and its application for the rickettsiae. Int. J. Syst. Bacteriol. 1997, 47, 252–261. [Google Scholar] [CrossRef]
- Mediannikov, O.Y.; Sidelnikov, Y.; Ivanov, L.; Mokretsova, E.; Fournier, P.E.; Tarasevich, I.; Raoult, D. Acute tick-borne rickettsiosis caused by Rickettsia heilongjiangensis in Russian Far East. Emerg. Infect. Dis. 2004, 10, 810–817. [Google Scholar] [CrossRef] [PubMed]
- Labruna, M.B.; Whitworth, T.; Horta, M.C.; Bouyer, D.H.; McBride, J.W.; Pinter, A.; Popov, V.; Gennari, S.M.; Walker, D.H. Rickettsia species infecting Amblyomma cooperi ticks from an area in the state of Sao Paulo, Brazil, where Brazilian spotted fever is endemic. J. Clin. Microbiol. 2004, 42, 90–98. [Google Scholar] [CrossRef] [PubMed]
- Regnery, R.L.; Spruill, C.L.; Plikaytis, B.D. Genotypic identification of rickettsiae and estimation of intraspecies sequence divergence for portions of two rickettsial genes. J. Bacteriol. 1991, 173, 1576–1589. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schaffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Biggerstaff, B. PooledInfRate software. Vector Borne Zoonotic Dis. 2005, 5, 420–421. [Google Scholar] [CrossRef] [PubMed]
- Team, R.D.C. A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2012); R Foundation for Statistical Computing: Vienna, Austria, 2015. [Google Scholar]

| Host (n of Hosts Inspected; Infestation Rate (%)) | Tick Species | n Ticks (%) | Male n (%) | Female n (%) | Nymph n (%) |
|---|---|---|---|---|---|
| Cattle (n = 381; 37.7%) | R. bursa | 608 (73.0) | 350 (57.6) | 225 (37.0) | 33 (5.4) |
| H. marginatum | 216 (25.8) | 163 (75.4) | 53 (24.6) | 0 (0.0) | |
| I. ricinus | 10 (1.2) | 3 (3.0) | 7 (7.0) | 0 (0.0) | |
| Total | 834 | 516 (62.0) | 285 (34.0) | 33 (4.0) | |
| Sheep (n = 500; 2.0%) | R. bursa | 10 (100) | 10 (100.0) | 0 | 0 |
| Total | 10 | 10 | |||
| Wild boars (n = 119; 38.6%) | D. marginatus | 223 (91.0) | 144 (64.5) | 79 (35.5) | 0 (0.0) |
| I. ricinus | 13 (5.3) | 1 (8.0) | 12 (92.0) | 0 (0.0) | |
| R. bursa | 8 (3.3) | 1 (12.5) | 7 (87.5) | 0 (0.0) | |
| H. marginatum | 1 (0.4) | 0 (0.0) | 1 (100) | 0 (0.0) | |
| Total | 245 | 146 (60.0) | 99 (40.0) | 0 (0.0) | |
| Dogs (n = 13 * §) | R. sanguineus s.l. | 24 (89.0) | 2 (8.3) | 14 (58.3) | 8(33.3) |
| I. ricinus | 3 (11.0) | 0 | 3 (100) | 0 | |
| Total | 27 | 2 (7.4) | 17 (63.0) | 8 (29.6) | |
| Cat (n = 1 * §) | D. marginatus | 1 (100) | 0 | 0 | 1(100) |
| Total (n = 1014) | All species | 1117 | 674 (60.4) | 401 (35.8) | 42 (3.8) |
| Number of Individual Ticks or Ticks per Pool (n) | Number of Pools with n Ticks | Positive Rickettsia spp. n s(%) | ||||||
|---|---|---|---|---|---|---|---|---|
| 2017 (July–August) | H. marginatum | R. bursa | I. ricinus | Total | H.marginatum | R.bursa | I. ricinus | Total |
| 1 | 15 | 19 | 1 | 35 | 11 (73) | 6 (32) | 0 (0) | 17 (49) |
| 2 | 13 | 9 | 0 | 22 | 11 (85) | 1 (11) | 0 (0) | 12 (55) |
| 3 | 6 | 6 | 0 | 12 | 4 (67) | 1 (17) | 0 (0) | 5 (42) |
| 4 | 1 | 7 | 0 | 8 | 1 (100) | 3 (43) | 0 (0) | 4 (50) |
| 5 | 1 | 8 | 0 | 9 | 1 (100) | 5 (63) | 0 (0) | 6 (67) |
| 6 | 2 | 19 | 0 | 21 | 1 (50) | 8 (42) | 0 (0) | 9 (43) |
| Total pools | 38 | 68 | 1 | 107 | 29 (76) | 24 (35) | 0 (0) | 53 (50) |
| MLE (95% CI) | 50.5% (37.0–64.4) | 12.2% (8.1–17.4) | 0 | 20.7% (16.0–26.1) | ||||
| 2018 (July–December) | H. marginatum | R. bursa | I. ricinus | Total | H. marginatum | R. bursa | I. ricinus | Total |
| 1 | 25 | 12 | 1 | 38 | 24 (96) | 5 (42) | 0 (0) | 29 (76) |
| 2 | 9 | 7 | 1 | 17 | 5 (55) | 5 (71) | 0 (0) | 10 (62) |
| 3 | 14 | 8 | 1 | 23 | 13 (93) | 4 (50) | 1 (100) | 18 (78) |
| 4 | 5 | 15 | 0 | 20 | 4 (80) | 7 (46) | 0 (0) | 11 (55) |
| 5 | 5 | 14 | 0 | 19 | 5 (100) | 10 (66) | 0 (0) | 15 (79) |
| 6 | 1 | 30 | 0 | 31 | 1 (100) | 20 (64) | 0 (0) | 21 (68) |
| Total pools | 59 | 86 | 3 | 148 | 52 (88) | 51 (58) | 1 (33) | 104 (70) |
| MLE (95% CI) | 65.3% (52.8–77.3) | 20.1% (15.3–25.6) | 2.8% (0.1–11.9) | 30.7% (25.6–36.2) | ||||
| 2017 and 2018 | H. marginatum | R. bursa | I. ricinus | Total | H. marginatum | R. bursa | I. ricinus | Total |
| 1 | 40 | 31 | 2 | 73 | 35 (87) | 11 (35) | 0 (0) | 46 (63) |
| 2 | 22 | 16 | 1 | 39 | 16 (73) | 6 (37) | 0 (0) | 22 (56) |
| 3 | 20 | 14 | 1 | 35 | 17 (85) | 5 (36) | 1 (100) | 23 (66) |
| 4 | 6 | 22 | 0 | 28 | 5 (83) | 10 (45) | 0 (0) | 15 (53) |
| 5 | 6 | 22 | 0 | 28 | 6 (100) | 15 (68) | 0 (0) | 21 (75) |
| 6 | 3 | 49 | 0 | 52 | 2(67) | 28 (56) | 0 (0) | 30 (58) |
| Total pools | 97 | 154 | 4 | 255 | 81 (83) | 75 (48) | 1 (25) | 157 (61) |
| MLE (95% CI) | 58% (49.4–67.9) | 16.6% (13.3–20.4) | 2.7% (0.1–11.6) | 26.3% (22.7–30.1) | ||||
| Number of Individual Ticks or Ticks per Pool (n) | Number of Pools with n Ticks | Positive Rickettsia spp. n (%) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| D. marginatus | I. ricinus | R. bursa | H. marginatum | Total | D. marginatus | I. ricinus | R. bursa | H. marginatum | Total | |
| 1 | 15 | 1 | 3 | 1 | 20 | 8 (53) | 1 (100) | 2 (67) | 1 (100) | 12 (60) |
| 2 | 10 | 2 | 1 | 0 | 13 | 7 (70) | 1 (50) | 1(100) | 0 (0) | 9 (62) |
| 3 | 11 | 0 | 1 | 0 | 12 | 7 (64) | 0 (0) | 1(100) | 0 (0) | 8 (67) |
| 4 | 11 | 2 | 0 | 0 | 13 | 8 (73) | 0 (0) | 0 (0) | 0 (0) | 8 (62) |
| 5 | 9 | 0 | 0 | 0 | 9 | 3 (34) | 0 (0) | 0 (0) | 0 (0) | 3 (33) |
| 6 | 11 | 0 | 0 | 0 | 11 | 6 (56) | 0 (0) | 0 (0) | 0 (0) | 6 (55) |
| Total pools | 67 | 5 | 5 | 1 | 78 | 39 (58) | 2 (40) | 4 (80) | 1 (100) | 46 (59) |
| MLE (95% CI) | 23.0% (16.9–30.0) | 4.9% (0.0–14.4) | 10.9% (3.5–23.7) | 2.4% (0.0–10.2) | 24.4% (18.5–31.1) | |||||
| Host | Tick Species | No. of Pools Positive for Rickettsiae | No of gltA Sequences | No of ompA Sequences | Identified Rickettsia Species |
|---|---|---|---|---|---|
| 2017 | |||||
| Cattle | H. marginatum | 29 | 24 | 5 | Ri.aeschlimanii |
| R. bursa | 24 | 5 | 2 | Candidatus Ri. barbariae | |
| Sheep | R. bursa | 2 | 2 | 1 | Candidatus Ri. barbariae |
| Dogs | R.sanguineuss.l. | 1 | 1 | 1 | Ri. massiliae |
| I. ricinus | 1 | 1 | 1 | Ri. monacensis | |
| Cat | D. marginatus | 1 | 1 | 0 | Ri. slovaca |
| Total pools 2017 | 58 | 34 | 10 | ||
| 2018 | |||||
| Cattle | H. marginatum | 52 | 31 | 13 | Ri.aeschlimanii |
| R. bursa | 51 | 14 | 5 | Ri. aeschlimanii; Candidatus Ri. barbariae | |
| I. ricinus | 1 | 1 | 0 | Ri.aeschlimanii | |
| Wild boars | D. marginatus | 39 | 6 | 3 | Ri. slovaca |
| I. ricinus | 2 | 2 | 0 | Ri. slovaca | |
| R. bursa | 4 | 2 | 2 | Ri. slovaca | |
| H. marginatum | 1 | 1 | 0 | Ri.aeschlimanii | |
| Total pools 2018 | 150 | 57 | 23 | ||
| Total pools 2017–2018 | 208 | 91 | 33 | Ri. aeschlimanii; Candidatus Ri. barbariae; Ri. massiliae; Ri. monacensis; Ri. slovaca |
| Species | Target | Name | Sequence | Annealing Temperature (°C) | References |
|---|---|---|---|---|---|
| rPCR | |||||
| Rickettsia spp. | gltA | Rspp-F | GAGAGAAAATTATATCCAAATGTTGAT | 60 | |
| Rspp-R | AGGGTCTTCGTGCATTTCTT | ||||
| Rspp-P | CATTGTGCCATCCAGCCTACGGT | ||||
| Conventional PCR (Sequencing) | |||||
| Rickettsia spp. | gltA | CS2D | ATGACCAATGAAAATAATAAT | 54 | [40,41] |
| CSEndR | CTTATACTCTCTATGTACA | ||||
| 409D | CCTATGGCTATTATGCTTGC | 54 | |||
| 1258R | ATTGCAAAAAGTACAGTGAACA | ||||
| ompA | Rr190.70p | ATGGCGAATATTTCTCCAAAA | 48 | ||
| Rr190.602n | AGTGCAGCATTCGCTCCCCCT | ||||
| Ticks | 16S rDNA | 16S + 1 | CTGCTCAATGATTTTTTAAATTGCTGTGG | 48 and 54 | [39] |
| 16S − 1 | CCGGTCTGAACTCAGATCAAGT | ||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cicculli, V.; Maestrini, O.; Casabianca, F.; Villechenaud, N.; Charrel, R.; de Lamballerie, X.; Falchi, A. Molecular Detection of Spotted-Fever Group Rickettsiae in Ticks Collected from Domestic and Wild Animals in Corsica, France. Pathogens 2019, 8, 138. https://doi.org/10.3390/pathogens8030138
Cicculli V, Maestrini O, Casabianca F, Villechenaud N, Charrel R, de Lamballerie X, Falchi A. Molecular Detection of Spotted-Fever Group Rickettsiae in Ticks Collected from Domestic and Wild Animals in Corsica, France. Pathogens. 2019; 8(3):138. https://doi.org/10.3390/pathogens8030138
Chicago/Turabian StyleCicculli, Vincent, Oscar Maestrini, Francois Casabianca, Natacha Villechenaud, Remi Charrel, Xavier de Lamballerie, and Alessandra Falchi. 2019. "Molecular Detection of Spotted-Fever Group Rickettsiae in Ticks Collected from Domestic and Wild Animals in Corsica, France" Pathogens 8, no. 3: 138. https://doi.org/10.3390/pathogens8030138
APA StyleCicculli, V., Maestrini, O., Casabianca, F., Villechenaud, N., Charrel, R., de Lamballerie, X., & Falchi, A. (2019). Molecular Detection of Spotted-Fever Group Rickettsiae in Ticks Collected from Domestic and Wild Animals in Corsica, France. Pathogens, 8(3), 138. https://doi.org/10.3390/pathogens8030138







