Motility, Biofilm Formation and Antimicrobial Efflux of Sessile and Planktonic Cells of Achromobacter xylosoxidans
Abstract
1. Introduction
2. Results and Discussion
2.1. Differentially Expressed Genes
2.1.1. Increased Efflux Pump Activity During Sessile Growth
2.1.2. Motility, Stress Response and Quorum Sensing
2.1.3. Sulfur Metabolism
2.1.4. Constituents of the Extracellular Matrix
2.2. Examination of Inactivated Mutants
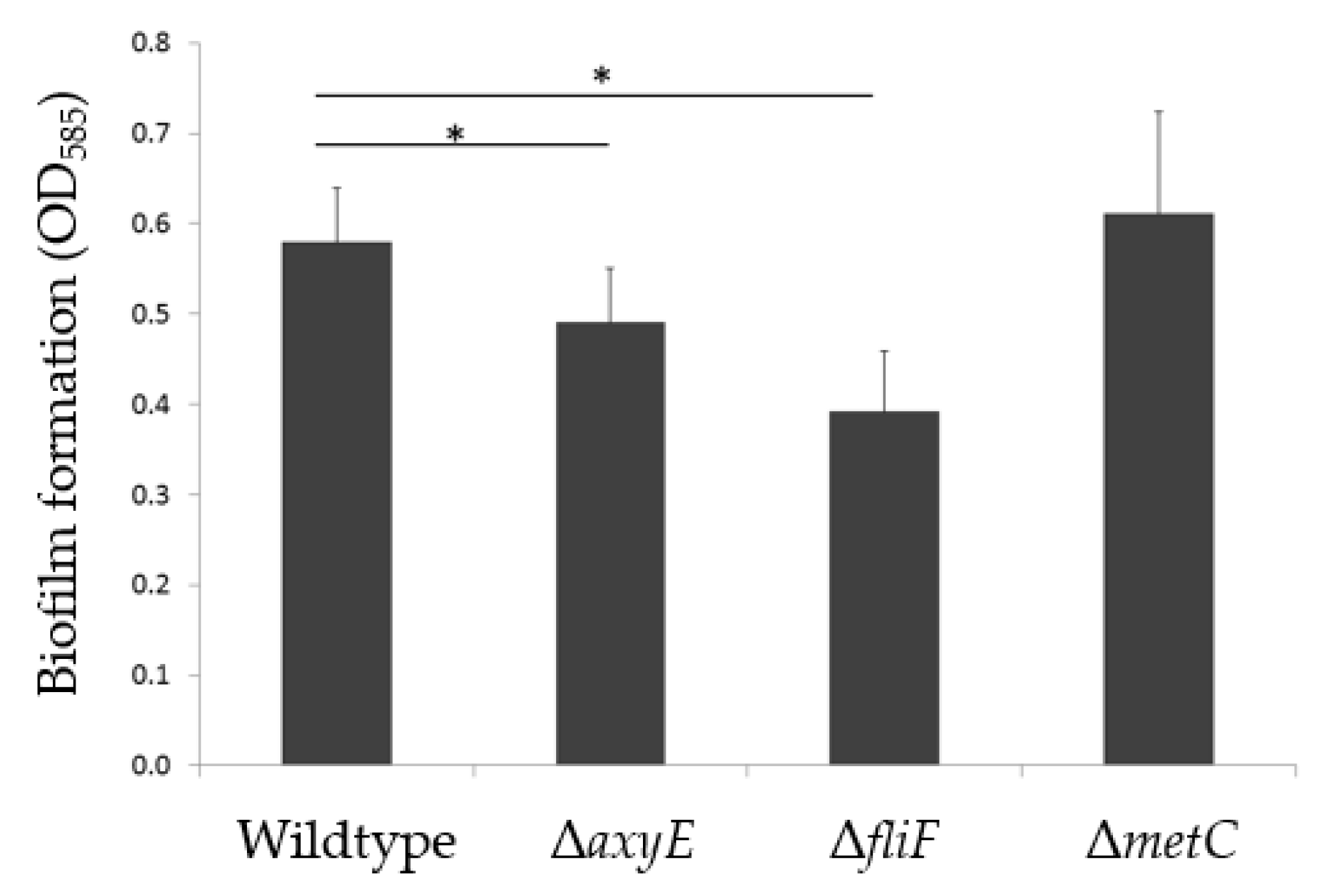
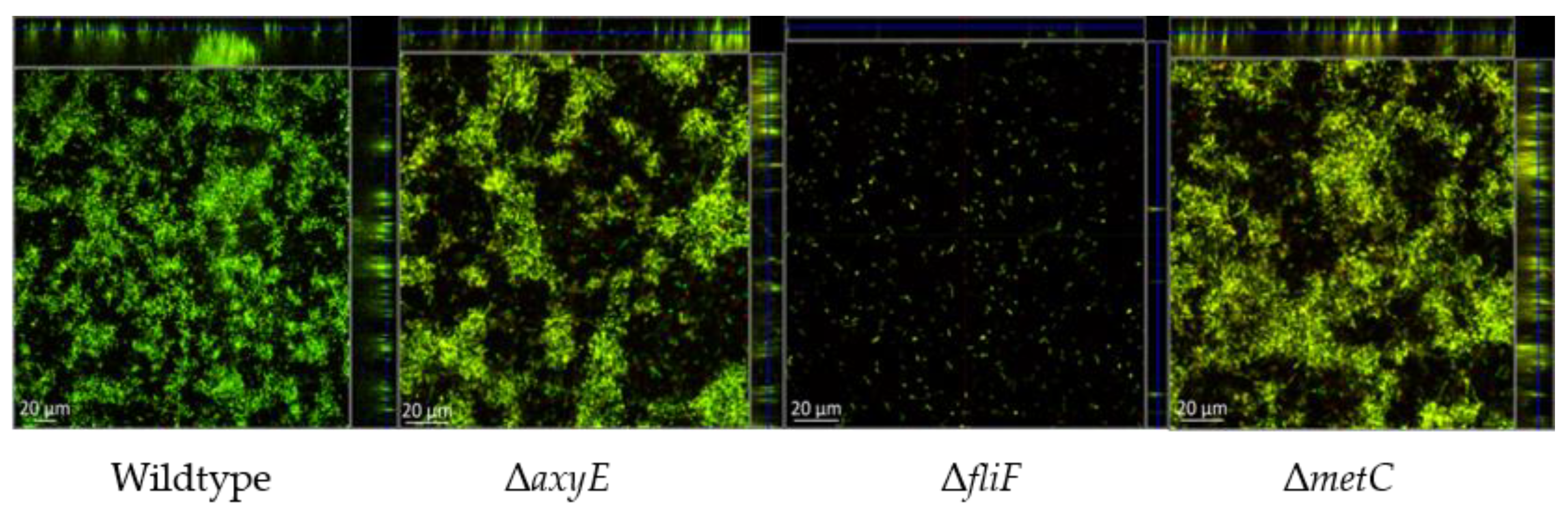
2.2.1. RND-Type Efflux Pump AxyEF-OprN Affects Antimicrobial Susceptibility and Biofilm Formation
2.2.2. Motility Impairment Affects Biofilm Formation
2.2.3. Methionine Biosynthesis
3. Materials and Methods
3.1. Clinical Strain of A. xylosoxidans
3.2. Culture at Separate Growth Phases and RNA Extraction
3.3. Sequencing and Data Processing
3.4. Gene Inactivation
3.5. Visualization and Quantitation of Biofilm Formation
3.6. Antimicrobial Susceptibility
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Davies, J.C.; Rubin, B.K. Emerging and unusual gram-negative infections in cystic fibrosis. Semin. Respir. Crit. Care Med. 2007, 28, 312–321. [Google Scholar] [CrossRef] [PubMed]
- De Baets, F.; Schelstraete, P.; Van Daele, S.; Haerynck, F.; Vaneechoutte, M. Achromobacter xylosoxidans in cystic fibrosis: Prevalence and clinical relevance. J. Cyst. Fibros. 2007, 6, 75–78. [Google Scholar] [CrossRef] [PubMed]
- Dupont, C.; Michon, A.L.; Jumas-Bilak, E.; Norskov-Lauritsen, N.; Chiron, R.; Marchandin, H. Intrapatient diversity of Achromobacter spp. involved in chronic colonization of Cystic Fibrosis airways. Infect. Genet. Evol. 2015, 32, 214–223. [Google Scholar] [CrossRef] [PubMed]
- Rolston, K.V.; Messer, M. The in-vitro susceptibility of Alcaligenes denitrificans subsp. xylosoxidans to 40 antimicrobial agents. J. Antimicrob. Chemother. 1990, 26, 857–860. [Google Scholar] [CrossRef]
- Raso, T.; Bianco, O.; Grosso, B.; Zucca, M.; Savoia, D. Achromobacter xylosoxidans respiratory tract infections in cystic fibrosis patients. Eur. J. Clin. Microbiol. Infect. Dis. 2008, 116, 837–841. [Google Scholar]
- Almuzara, M.; Limansky, A.; Ballerini, V.; Galanternik, L.; Famiglietti, A.; Vay, C. In vitro susceptibility of Achromobacter spp. isolates: Comparison of disk diffusion, Etest and agar dilution methods. Int. J. Antimicrob. Agents 2010, 35, 68–71. [Google Scholar] [CrossRef] [PubMed]
- Amoureux, L.; Bador, J.; Siebor, E.; Taillefumier, N.; Fanton, A.; Neuwirth, C. Epidemiology and resistance of Achromobacter xylosoxidans from cystic fibrosis patients in Dijon, Burgundy: First French data. J. Cyst. Fibros. 2013, 12, 170–176. [Google Scholar] [CrossRef] [PubMed]
- Hansen, C.R.; Pressler, T.; Nielsen, K.G.; Jensen, P.O.; Bjarnsholt, T.; Hoiby, N. Inflammation in Achromobacter xylosoxidans infected cystic fibrosis patients. J. Cyst. Fibros. 2010, 9, 51–58. [Google Scholar] [CrossRef]
- Jakobsen, T.H.; Hansen, M.A.; Jensen, P.O.; Hansen, L.; Riber, L.; Cockburn, A.; Kolpen, M.; Ronne Hansen, C.; Ridderberg, W.; Eickhardt, S.; et al. Complete genome sequence of the cystic fibrosis pathogen Achromobacter xylosoxidans NH44784-1996 complies with important pathogenic phenotypes. PLoS ONE 2013, 8, e68484. [Google Scholar] [CrossRef]
- Nielsen, S.M.; Norskov-Lauritsen, N.; Bjarnsholt, T.; Meyer, R.L. Achromobacter Species Isolated from Cystic Fibrosis Patients Reveal Distinctly Different Biofilm Morphotypes. Microorganisms 2016, 4, 33. [Google Scholar] [CrossRef]
- De la Fuente-Nunez, C.; Reffuveille, F.; Fernandez, L.; Hancock, R.E. Bacterial biofilm development as a multicellular adaptation: Antibiotic resistance and new therapeutic strategies. Curr. Opin. Microbiol. 2013, 16, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Hall-Stoodley, L.; Costerton, J.W.; Stoodley, P. Bacterial biofilms: From the natural environment to infectious diseases. Nat. Rev. Microbiol. 2004, 2, 95–108. [Google Scholar] [CrossRef] [PubMed]
- Costerton, J.W.; Stewart, P.S.; Greenberg, E.P. Bacterial biofilms: A common cause of persistent infections. Science 1999, 284, 1318–1322. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.C.; Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 2010, 8, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Stoodley, P.; Sauer, K.; Davies, D.G.; Costerton, J.W. Biofilms as complex differentiated communities. Annu. Rev. Microbiol. 2002, 56, 187–209. [Google Scholar] [CrossRef] [PubMed]
- McDougald, D.; Rice, S.A.; Barraud, N.; Steinberg, P.D.; Kjelleberg, S. Should we stay or should we go: Mechanisms and ecological consequences for biofilm dispersal. Nat. Rev. Microbiol. 2011, 10, 39–50. [Google Scholar] [CrossRef] [PubMed]
- Dotsch, A.; Eckweiler, D.; Schniederjans, M.; Zimmermann, A.; Jensen, V.; Scharfe, M.; Geffers, R.; Haussler, S. The Pseudomonas aeruginosa transcriptome in planktonic cultures and static biofilms using RNA sequencing. PLoS ONE 2012, 7, e31092. [Google Scholar] [CrossRef]
- Dingemans, J.; Monsieurs, P.; Yu, S.H.; Crabbe, A.; Forstner, K.U.; Malfroot, A.; Cornelis, P.; Van Houdt, R. Effect of Shear Stress on Pseudomonas aeruginosa Isolated from the Cystic Fibrosis Lung. MBio 2016, 7. [Google Scholar] [CrossRef]
- Nielsen, S.; Meyer, R.; Nørskov-Lauritsen, N. Differences in gene expression profiles between early and late isolates in monospecies Achromobacter biofilm. Pathogens 2017, 6, 20. [Google Scholar] [CrossRef]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef]
- Van Acker, H.; Coenye, T. The Role of Efflux and Physiological Adaptation in Biofilm Tolerance and Resistance. J. Biol. Chem. 2016, 291, 12565–12572. [Google Scholar] [CrossRef] [PubMed]
- Baugh, S.; Ekanayaka, A.S.; Piddock, L.J.; Webber, M.A. Loss of or inhibition of all multidrug resistance efflux pumps of Salmonella enterica serovar Typhimurium results in impaired ability to form a biofilm. J. Antimicrob. Chemother. 2012, 67, 2409–2417. [Google Scholar] [CrossRef] [PubMed]
- Lamers, R.P.; Cavallari, J.F.; Burrows, L.L. The efflux inhibitor phenylalanine-arginine beta-naphthylamide (PAbetaN) permeabilizes the outer membrane of gram-negative bacteria. PLoS ONE 2013, 8, e60666. [Google Scholar] [CrossRef] [PubMed]
- Kvist, M.; Hancock, V.; Klemm, P. Inactivation0 of efflux pumps abolishes bacterial biofilm formation. Appl. Environ. Microbiol. 2008, 74, 7376–7382. [Google Scholar] [CrossRef] [PubMed]
- Aeschlimann, J.R. The role of multidrug efflux pumps in the antibiotic resistance of Pseudomonas aeruginosa and other gram-negative bacteria. Insights from the Society of Infectious Diseases Pharmacists. Pharmacotherapy 2003, 23, 916–924. [Google Scholar] [CrossRef] [PubMed]
- Bador, J.; Amoureux, L.; Duez, J.M.; Drabowicz, A.; Siebor, E.; Llanes, C.; Neuwirth, C. First description of an RND-type multidrug efflux pump in Achromobacter xylosoxidans, AxyABM. Antimicrob. Agents Chemother. 2011, 55, 4912–4914. [Google Scholar] [CrossRef] [PubMed]
- Bador, J.; Amoureux, L.; Blanc, E.; Neuwirth, C. Innate aminoglycoside resistance of Achromobacter xylosoxidans is due to AxyXY-OprZ, an RND-type multidrug efflux pump. Antimicrob. Agents Chemother. 2013, 57, 603–605. [Google Scholar] [CrossRef]
- Li, X.Z.; Nikaido, H. Efflux-mediated drug resistance in bacteria: An update. Drugs 2009, 69, 1555–1623. [Google Scholar] [CrossRef]
- Schroeder, M.; Brooks, B.D.; Brooks, A.E. The Complex Relationship between Virulence and Antibiotic Resistance. Genes 2017, 8, 39. [Google Scholar] [CrossRef]
- Soto, S.M. Role of efflux pumps in the antibiotic resistance of bacteria embedded in a biofilm. Virulence 2013, 4, 223–229. [Google Scholar] [CrossRef]
- Nachin, L.; Nannmark, U.; Nystrom, T. Differential roles of the universal stress proteins of Escherichia coli in oxidative stress resistance, adhesion, and motility. J. Bacteriol. 2005, 187, 6265–6272. [Google Scholar] [CrossRef] [PubMed]
- Valentini, M.; Filloux, A. Biofilms and Cyclic di-GMP (c-di-GMP) Signaling: Lessons from Pseudomonas aeruginosa and Other Bacteria. J. Biol. Chem. 2016, 291, 12547–12555. [Google Scholar] [CrossRef] [PubMed]
- Ha, D.G.; O’Toole, G.A. c-di-GMP and its Effects on Biofilm Formation and Dispersion: A Pseudomonas Aeruginosa Review. Microbiol. Spectr. 2015, 3. MB-0003-2014. [Google Scholar] [CrossRef]
- Bergeron, J.R. Structural modeling of the flagellum MS ring protein FliF reveals similarities to the type III secretion system and sporulation complex. PeerJ 2016, 4, 1718. [Google Scholar] [CrossRef] [PubMed]
- Kolpen, M.; Hansen, C.R.; Bjarnsholt, T.; Moser, C.; Christensen, L.D.; van Gennip, M.; Ciofu, O.; Mandsberg, L.; Kharazmi, A.; Döring, G.; et al. Polymorphonuclear leucocytes consume oxygen in sputum from chronic Pseudomonas aeruginosa pneumonia in cystic fibrosis. Thorax 2010, 65, 57–62. [Google Scholar] [CrossRef] [PubMed]
- Kolpen, M.; Kragh, K.N.; Bjarnsholt, T.; Line, L.; Hansen, C.R.; Dalbøge, C.S.; Hansen, N.; Kühl, M.; Høiby, N.; Jensen, P.Ø. Denitrification by cystic fibrosis pathogens—Stenotrophomonas maltophilia is dormant in sputum. Int. J. Med. Microbiol. 2015, 305, 1–10. [Google Scholar] [CrossRef]
- Clausen, T.; Huber, R.; Prade, L.; Wahl, M.C.; Messerschmidt, A. Crystal structure of Escherichia coli cystathionine gamma-synthase at 1.5 A resolution. EMBO J. 1998, 17, 6827–6838. [Google Scholar] [CrossRef]
- Ejim, L.J.; D’Costa, V.M.; Elowe, N.H.; Loredo-Osti, J.C.; Malo, D.; Wright, G.D. Cystathionine beta-lyase is important for virulence of Salmonella enterica serovar Typhimurium. Infect. Immun. 2004, 72, 3310–3314. [Google Scholar] [CrossRef]
- Shah, D.H.; Shringi, S.; Desai, A.R.; Heo, E.J.; Park, J.H.; Chae, J.S. Effect of metC mutation on Salmonella Gallinarum virulence and invasiveness in 1-day-old White Leghorn chickens. Vet. Microbiol. 2007, 119, 352–357. [Google Scholar] [CrossRef]
- Mayer, C.; Moritz, R.; Kirschner, C.; Borchard, W.; Maibaum, R.; Wingender, J.; Flemming, H.C. The role of intermolecular interactions: Studies on model systems for bacterial biofilms. Int. J. Biol. Macromol. 1999, 26, 3–16. [Google Scholar] [CrossRef]
- Whitfield, C. Biosynthesis and assembly of capsular polysaccharides in Escherichia coli. Annu. Rev. Biochem. 2006, 75, 39–68. [Google Scholar] [CrossRef] [PubMed]
- Hentzer, M.; Teitzel, G.M.; Balzer, G.J.; Heydorn, A.; Molin, S.; Givskov, M.; Parsek, M.R. Alginate overproduction affects Pseudomonas aeruginosa biofilm structure and function. J. Bacteriol. 2001, 183, 5395–5401. [Google Scholar] [CrossRef] [PubMed]
- The European Committee on Antimicrobial Susceptibility Testing—EUCAST. Available online: http://www.eucast.org/clinical_breakpoints/ (accessed on 9 March 2018).
- Fleurbaaij, F.; Henneman, A.A.; Corver, J.; Knetsch, C.W.; Smits, W.K.; Nauta, S.T.; Giera, M.; Dragan, I.; Kumar, N.; Lawley, T.D.; et al. Proteomic identification of Axc, a novel beta-lactamase with carbapenemase activity in a meropenem-resistant clinical isolate of Achromobacter xylosoxidans. Nat. Sci. Rep. 2018, 8, 8181. [Google Scholar] [CrossRef] [PubMed]
- Bador, J.; Neuwirth, C.; Grangier, N.; Muniz, M.; Germé, L.; Bonnet, J.; Pillay, V.; Llanes, C.; de Curraize, C.; Amoureux, L. Role of AxyZ transcriptional regulator in overproduction of AxyXY OprZ multidrug efflux system in Achromobacter species mutants selected by tobramycin. Antimicrob. Agents Chemother. 2017, 61, 8. [Google Scholar] [CrossRef] [PubMed]
- Whiteley, M.; Bangera, M.G.; Bumgarner, R.E.; Parsek, M.R.; Teitzel, G.M.; Lory, S.; Greenberg, E.P. Gene expression in Pseudomonas aeruginosa biofilms. Nature 2001, 413, 860–864. [Google Scholar] [CrossRef]
- Guttenplan, S.B.; Kearns, D.B. Regulation of flagellar motility during biofilm formation. FEMS Microbiol. Rev. 2013, 37, 849–871. [Google Scholar] [CrossRef] [PubMed]
- Soutourina, O.; Poupel, O.; Coppee, J.Y.; Danchin, A.; Msadek, T.; Martin-Verstraete, I. CymR, the master regulator of cysteine metabolism in Staphylococcus aureus, controls host sulphur source utilization and plays a role in biofilm formation. Mol. Microbiol. 2009, 73, 194–211. [Google Scholar] [CrossRef] [PubMed]
- Allesen-Holm, M.; Barken, K.B.; Yang, L.; Klausen, M.; Webb, J.S.; Kjelleberg, S.; Molin, S.; Givskov, M.; Tolker-Nielsen, T. A characterization of DNA release in Pseudomonas aeruginosa cultures and biofilms. Mol. Microbiol. 2006, 59, 1114–1128. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, G.A. Microtiter dish biofilm formation assay. J. Vis. Exp. 2011, 47, 2437. [Google Scholar] [CrossRef]
- Harrison, J.J.; Stremick, C.A.; Turner, R.J.; Allan, N.D.; Olson, M.E.; Ceri, H. Microtiter susceptibility testing of microbes growing on peg lids: A miniaturized biofilm model for high-throughput screening. Nat. Protoc. 2010, 5, 1236–1254. [Google Scholar] [CrossRef]
| Gene Name | Function | Fold Change | |
|---|---|---|---|
| RND efflux system membrane fusion protein AxyA | Virulence | Up | 7.4 |
| Flagellar basal-body rod modification protein FlgD | Flagellar motility | Down | −7.3 |
| Flagellar basal-body rod protein FlgC | Flagellar motility | Down | −5.1 |
| Flagellar biosynthesis protein FlhB | Flagellar motility | Down | −5.1 |
| Flagellar biosynthesis protein FliC | Flagellar motility | Down | −8.5 |
| Flagellar biosynthesis protein FliL | Flagellar motility | Down | −6.7 |
| Flagellar biosynthesis protein FliQ | Flagellar motility | Down | −5.2 |
| Flagellar biosynthesis protein FliT | Flagellar motility | Down | −5.2 |
| Flagellar hook-associated protein FlgK | Flagellar motility | Down | −5.2 |
| Flagellar hook-associated protein FlgL | Flagellar motility | Down | −5.3 |
| Flagellar hook-associated protein FliD | Flagellar motility | Down | −6.0 |
| Flagellar hook-basal body complex protein FliE | Flagellar motility | Down | −5.4 |
| Flagellar L-ring protein FlgH | Flagellar motility | Down | −5.0 |
| Flagellar motor switch protein FliM | Flagellar motility | Down | −5.7 |
| Flagellar motor switch protein FliN | Flagellar motility | Down | −7.3 |
| Flagellar M-ring protein FliF a | Flagellar motility | Down | −6.3 |
| Flagellar protein FliJ | Flagellar motility | Down | −5.9 |
| Universal stress protein family (tandem domain) | Stress response | Down | −9.2, −6.1 |
| Universal stress protein UspA | Stress response | Down | −6.6 |
| Diguanylate cyclase/ phosphodiesterase | Stress response | Up | 11.4 |
| Cystathionine beta-lyase, MetC a | Methionine biosynthesis | Up | 86 |
| Exopolysaccharide biosynthesis glycosyltransferase, EpsF | EPS biosynthesis | Up | 6.4 |
| ATP-binding proteins, KpsT, KpsE, KpsM | Cell wall and capsule | Up | 5.1 to 8.7 |
| AX08 Wildtype | AX08 ΔaxyE | |||
|---|---|---|---|---|
| Antibiotic (µg/mL) | MIC | S/R * | MIC | S/R * |
| Colistin | 1 | S | 1 | S |
| Polymyxin B | 1 | NI | 1 | NI |
| Piperacillin | 8 | S | 8 | S |
| Ticarcillin/Clavulanic Acid | ≤ 16 | S | ≤ 16 | S |
| Cefepime | 8 | S | 8 | S |
| Cefotaxime | > 32 | R | > 32 | R |
| Ceftazidime | 4 | S | 8 | S |
| Aztreonam | > 16 | R | > 16 | R |
| Ertapenem | 1 | NI | 0.5 | NI |
| Doripenem | 0.25 | S | 1 | S |
| Imipenem | 2 | S | ≤ 1 | S |
| Meropenem | ≤ 1 | S | ≤ 1 | S |
| Ciprofloxacin | 2 | R | 1 | R |
| Levofloxacin | 2 | R | ≤ 1 | S |
| Amikacin | > 32 | R | > 32 | R |
| Gentamicin | > 8 | R | > 8 | R |
| Tobramycin | > 8 | R | > 8 | R |
| Doxycycline | 8 | NI | 4 | NI |
| Minocycline | ≤ 2 | NI | ≤ 2 | NI |
| Tigecycline | 0.5 | NI | ≤ 0.25 | NI |
| Trimethoprim/ Sulfamethoxazole | ≤ 0.5 | NI | ≤ 0.5 | NI |
| Primer | Nucleotide Sequence (5’ – 3’) | Fragment Length (bp) | Target Gene |
|---|---|---|---|
| FliF_F | CGGTACCCGGGGATCGAACAGATCAACTACCAGCG | 771 | Flagellar M-ring gene, fliF |
| FliF_R | CGACTCTAGAGGATCTTGATGTGGCTGATGGTG | ||
| AxyE_F | CGGTACCCGGGGATCGACGTCAAGGAAAACCAG | 795 | RND-type multidrug resistance efflux pump gene, axyE |
| AxyE_R | CGACTCTAGAGGATCGGGCAACTGTTCGATCTT | ||
| MetC_F | CGGTACCCGGGGATCTGATGCAGGACAAGGAGT | 809 | Cystathionine beta-lyase gene, metC |
| MetC_R | CGACTCTAGAGGATCTAGGCGTAGGTGTCGTAG | ||
| M14Fa | CCAGGGTTTTCCCAGTCACGA | ||
| M14Ra | GCGGATAACAATTTCACACAGGA |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nielsen, S.M.; Penstoft, L.N.; Nørskov-Lauritsen, N. Motility, Biofilm Formation and Antimicrobial Efflux of Sessile and Planktonic Cells of Achromobacter xylosoxidans. Pathogens 2019, 8, 14. https://doi.org/10.3390/pathogens8010014
Nielsen SM, Penstoft LN, Nørskov-Lauritsen N. Motility, Biofilm Formation and Antimicrobial Efflux of Sessile and Planktonic Cells of Achromobacter xylosoxidans. Pathogens. 2019; 8(1):14. https://doi.org/10.3390/pathogens8010014
Chicago/Turabian StyleNielsen, Signe M., Line N. Penstoft, and Niels Nørskov-Lauritsen. 2019. "Motility, Biofilm Formation and Antimicrobial Efflux of Sessile and Planktonic Cells of Achromobacter xylosoxidans" Pathogens 8, no. 1: 14. https://doi.org/10.3390/pathogens8010014
APA StyleNielsen, S. M., Penstoft, L. N., & Nørskov-Lauritsen, N. (2019). Motility, Biofilm Formation and Antimicrobial Efflux of Sessile and Planktonic Cells of Achromobacter xylosoxidans. Pathogens, 8(1), 14. https://doi.org/10.3390/pathogens8010014





