Abstract
This review considers current knowledge surrounding species boundaries of the Armillaria root-rot pathogens and their distribution. In addition, a phylogenetic tree using translation elongation factor subunit 1-alpha (tef-1α) from isolates across the globe are used to present a global phylogenetic framework for the genus. Defining species boundaries based on DNA sequence-inferred phylogenies has been a central focus of contemporary mycology. The results of such studies have in many cases resolved the biogeographic history of species, mechanisms involved in dispersal, the taxonomy of species and how certain phenotypic characteristics have evolved throughout lineage diversification. Such advances have also occurred in the case of Armillaria spp. that include important causal agents of tree root rots. This commenced with the first phylogeny for Armillaria that was based on IGS-1 (intergenic spacer region one) DNA sequence data, published in 1992. Since then phylogenies were produced using alternative loci, either as single gene phylogenies or based on concatenated data. Collectively these phylogenies revealed species clusters in Armillaria linked to their geographic distributions and importantly species complexes that warrant further research.
1. Introduction
Armillaria spp. (Basidiomycota, Agaricales, Physalacriaceae) are amongst the best known and most important pathogens of forest trees but are also beneficial to horticulture and growth of edible fungi. They are found on almost every continent, have resulted in serious losses in sustainably managed natural forests as well as in plantations of non-native tree species [1]. Some species are important mycorrhizal symbionts of orchids [2,3,4,5,6]. Other species are symbionts of edible fungi such as Polyporus umbellatus and necessary for their production [7,8,9,10]. Most species have a saprophytic lifestyle during which they contribute to the decomposition of organic material in forest ecosystems and they become pathogens when environmental conditions are favourable for infection. Armillaria spp. have also become notoriously interesting to biologists associated with the fact that they include some of the world’s largest and oldest organisms [11,12].
The genus Armillaria has a long and controversial taxonomic history. This dates back to the 1700’s and the description of Agaricus melleus Vahl, Fl. dan.: t. (now A. mellea) by Danish Botanist Martin Vahl [13]. Subsequently, a number of taxonomic revisions were published and the valid name for the genus was often in dispute [14]. Armillaria Fr. Staude is now the broadly accepted generic name with Armillaria mellea (Vahl) P. Kumm. representing the type of the genus [14,15] (see [1] for characteristics of the genus). At least 129 species are currently recognised in Armillaria (MycoBank: html://http://www.mycobank.org/). In addition to these species, a number of biological species and phylogenetic groups have also been identified but these have yet to be formally described (e.g., [6]).
Phylogenetic studies have revolutionised our understanding of species boundaries in Armillaria. The first study to apply DNA sequence data to this question was conducted by Anderson and Stasovski [16]. This resulted in a phylogenetic concept for the relationships among Armillaria species from North America. This ground-breaking work was followed by a number of studies respectively dealing with species from Africa [17], Europe [18,19], Australasia [20,21], South America [22,23] and South-east Asia [6,24,25].
There have been few studies that have considered the phylogenetic relationships of Armillaria spp. at a global scale [26,27,28,29]. Maphosa et al. [26] and later Coetzee et al. [27] showed that that species of Armillaria form three distinct geographical clades. These include the African clade, the Holarctic clade (sensu [30]) and a non-Holarctic clade including several regions in the Holotropical and Austral kingdoms as defined by Morrone [30]. This subdivision was supported more recently by Koch et al. [29]. Using a concatenated dataset comprising six gene regions (rDNA large subunit [LSU], translation elongation factor subunit 1-alpha [tef-1α], RNA polymerase subunit II [rpb2], β-tubulin [TUB], Glyceraldehyde-3-phosphate dehydrogenase [gpd] and actin-1), the phylogenetic tree generated by these authors grouped Armillaria species from Africa, the Northern Hemisphere and Australian/temperate South America into clades representing those regions. In contrast to the results of Maphosa et al. [26] and Coetzee et al. [27], isolates of A. mellea formed a clade sister to the northern hemisphere—Australasian—temperate South America clades and the exannulated species (A. ectypa and A. tabescens) formed a monophyletic group basal to all Armillaria species. The placement of A. ectypa and A. tabescens basal to other Armillaria spp. and the fact that they are the only exannulated species provided sufficient support for the description of the new genus Desarmillaria [29]. Following the work of Koch et al. [29], Klopfenstein et al. [28] identified “superclades” in a global phylogeny of Armillaria spp. These were referred to as the Gallica, Solidipes/ostoyae, Mellea and Socialis/tabescens superclades.
The age of the most recent ancestor of Armillaria (including Desarmillaria) was dated within the Early Paleogene. Coetzee et al. [27] estimated the most recent ancestor of Armilllaria at 54 million years ago (MYA). This was supported by Koch et al. [29] who dated the most common ancestor of Armillaria at 41 MYA. Consequently, the age of the ancestral lineage of Armillaria post-dates the continental break up of Gondwana. This suggests that extant species attained their current biogeographical distribution through long-distance dispersal rather than by vicariance events as was previously hypothesised by [31].
This review considers species boundaries of the Armillaria root-rot pathogens and their distribution. For this purpose, we interrogated current knowledge regarding species clusters and phylogenetic relationships of species or taxa (biological species or lineages) based on morphological cohesion, phenotypic characteristics and phylogenetic analyses. A phylogenetic tree based on tef-1α DNA sequences was constructed using the dataset from Klopfenstein et al. [28] with additional sequences of taxa not included in their study and serves as phylogenetic framework for Armillaria in this review. A curated DNA sequence database was also constructed to aid in future studies.
2. Genes and Genomic Regions Employed in Phylogenetic Studies of Armillaria Species
Several genes and genomic regions have been employed in phylogenetic studies of Armillaria species (see Table S1 for a summary). Most early phylogenetic studies on Armillaria have incorporated the non-coding regions of the nuclear rRNA cistron to construct phylogenetic trees. Among these regions, the intergenic spacer 1 region (IGS-1), located between the nuclear ribosomal large subunit (nLSU, 28S gene) and the 5S gene, was first used to determine the relationships among North America Armillaria species [16]. In subsequent studies, the internally transcribed spacer regions (ITS-1 and ITS-2) together with the 5.8S gene (situated between the ITS-1 and ITS-2) were employed for phylogenetic inference [18,20,21]. Although these regions were successfully used in various studies, it was recognised that they fail to provide sufficient resolution to differentiate certain Armillaria species. Moreover, intra-strain nucleotide heterogeneity was reported for the ITS and IGS-1 regions in some species [32,33], making it impossible to sequence these regions without cloning. An additional complication was the fact that the 5S gene is inverted for the African Armillaria species relative to other species [17,34]. Consequently, the IGS-1 region cannot be used in a global phylogeny of Armillaria species. Thus, alternative loci were considered for phylogenetic purposes.
The protein-coding genes, or regions of these genes, most commonly used in the fungal phylogenetic analyses include the tef-1α and rpb2 genes (see [35] for a detailed account). Following the study on Phellinus nigrolimitatus by Kauserud et al. [36] and using the primers developed by these authors, Maphosa et al. [26] published the first phylogeny of Armillaria species based on sequences from a region of the tef-1α gene. Baumgartner et al. [37] showed that tef-1α occurs as a single copy gene in A. mellea. However, Guo et al. [6] discovered some diploid strains of Armillaria that contained intra-strain heterozygous sites in the tef-1α region and suggested this might be due to hybridisation events. Despite this, several studies have shown that this region in general provides the necessary resolution to infer species relationships with confidence, especially if phylogenies of species from the same geographic region are considered [6,19,25,26,28,38,39,40].
Matheny et al. [35] showed that rpb2 sequences can be used to resolve the phylogeny between closely related species in the Basidiomycota. This led to the research of Brazee et al. [39] to evaluate partial tef-1α, rpb2, and nLSU sequences for the identification of isolates representing A. calvescens and A. gallica from north-eastern North America and the application of this gene in a phylogeny of six Armillaria species from that geographic region. Armillaria calvescens and A. gallica are morphologically very similar with some differences in microscopic characters of their basidiocarps [41]. Brazee et al. [39] showed that sequences from the rpb2 gene cannot discriminate between A. calvescens and A. gallica. However, they found that these species and the four other species included in their study (A. gemina, A. mellea, A. sinapina, and A. solidipes) could be separated based on tef-1α gene sequences.
Recently, Guo et al. [6] utilised β–tubulin, in addition to ITS and tef-1α, sequence to study the phylogenetic relationships of Chinese biological species of Armillaria. β–tubulin and tef-1α gene trees generated in that study conflicted in the monophyly of strains. Strains that formed a monophyletic group in the tef-1α gene tree formed sub-clades distant to each other in the β–tubulin gene tree. β–tubulin genes underwent multiple independent duplications and losses in fungal lineages [42]. It is thus possible that paralogous copies were retained in different strains of Armillaria yielding paraphyletic groups in phylogenetic analyses. Sequences for β–tubulin have consequently not been used in more recent phylogenetic studies and it is doubtful if the gene carries adequate phylogenetic signal to provide a robust phylogeny for Armillaria species.
In addition to the loci described above, anonymous sequences generated through Sequencing With Arbitrary Primer Pairs (SWAPP) were used to determine the phylogeny of North American biological species [43]. Phylogenetic analysis of a matrix in which these sequences were combined yielded a phylogenetic tree that resolved the phylogenetic history of the North American species [43]. This method, however, has not been employed in subsequent phylogenetic studies of Armillaria.
3. Curation of Sequences from GenBank and Phylogenetic Analyses
A large number of DNA sequences for Armillaria species are available in GenBank. As is the case with many other fungal sequence deposits, many of these are not linked to published papers. The authenticity and origin of these sequences are often questionable. Another problem is that strain numbers can be linked to deposited sequences where the same strain has been assigned different strain numbers by researchers in different laboratories. Utilisation of these data can consequently lead to an artificial inflation of the taxon sampling in phylogenetic studies.
A database that curates high quality published sequence data linked to strain and the biogeographic information is currently not available. Therefore, we obtained sequences for the ITS, IGS-1, tef-1α and LSU regions from GenBank using published accession numbers rather than blast or keyword searches. A “bionumber” was assigned that linked individual strains to their available DNA sequences, their different strain numbers published by different researchers, biogeographic information and publication references. This information is available in a Microsoft® Excel Spreadsheet accessible at: http://www.fabinet.up.ac.za/mcoetzee (Table S2) and is updated when sequences in new publications become available.
A phylogenetic tree based on tef-1α sequence data and trees based on published phylogenies was generated for this review (Figure 1 and Figure S1). The tef-1α data set comprised the sequence dataset of Klopfenstein et al. [28] and additional sequences obtained from GenBank (Table S3). In total the dataset included 94 taxa, of which 47 Armillaria sequences were new additions. Sequences were re-aligned using the online version of MAFFT version 7 [44] with default settings. The phylogenetic tree was constructed based on Bayesian inference of phylogenies using BEAST version 1.10 [45] and following the protocol described in Klopfenstein et al. [28]. Guyanagaster necrorhizus was used as the outgroup species to root the tree. Published phylogenetic trees were redrawn because tef-1α sequence data were not available for all taxa or species. This also made it possible to indicate alternative phylogenetic positions of taxa or species where loci other than tef-1α were used for phylogenetic analyses. For this purpose, manually edited trees were generated in Mesquite version 3.5 [46] with the phylogenetic position of taxa based on their published relationship with other Armillaria species (Figure S1).
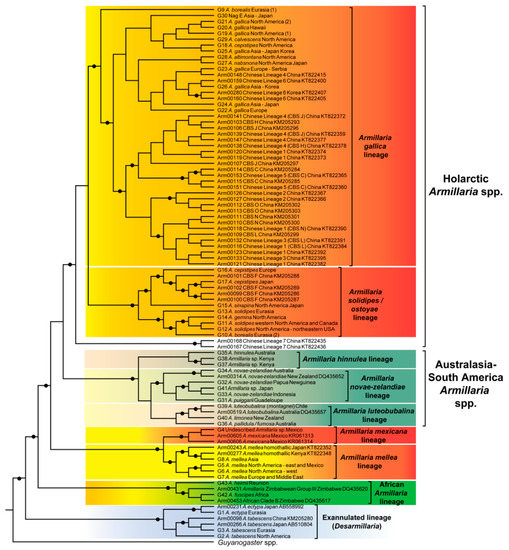
Figure 1.
Phylogenetic tree generated from tef-1α DNA sequence data showing the phylogenetic relationships of Armillaria species and lineages. Circles at nodes indicate posterior probability values > 0.95. Group numbers from Klopfenstein et al. [28] are shown at the species names where applicable. Arm numbers are bionumbers link to published sequences of Armillaria strains and for which information is provided in Table S2.
4. Relatives of Armillaria
Armillaria was originally assigned to the Tricholomataceae, one of the largest families in the Agaricales that included a diverse assemblage of genera. Molecular systematic studies revealed the polyphyly of the family [47,48,49,50]. Several genera previously residing in the Tricholomataceae were transferred to other families by Matheny et al. [49] based on their monophyletic groupings. In that study, Armillaria formed a monophyletic group together with other genera that were assigned to the Physalacriaceae. This family together with several other families that also include the Marasmiaceae constitute the Marasimioid clade in the Agaricales [49].
The question regarding the closest relative to Armillaria is intriguing. Currently, the phylogenetically closest known relatives of Armillaria are Guyanagaster necrorhizus and G. lucianii [29,51], sequestrate fungi that are known only from the neotropical rainforests of the Guiana Shield (Pakaraima Mountains, Guyana). Phylogenetic analyses that included various genera in the Agaricales and using DNA sequences for five loci (18S, ITS, and 28S rDNA, rpb2, and tef-1α) placed G. necrorhizus sister to A. tabescens and A. mellea [51]. Subsequent to this study, Koch et al. [29] inferred the phylogenetic relationship of Guyanagaster relative to the global collection of Armillaria species. That study showed that G. necrorhizus and G. lucianii is a monophyletic group sister to Armillaria and the newly erected genus Desarmillaria.
For many years it was suggested that A. tabescens, which was referred to as A. socialis in Europe [52], and A. ectypa should be moved to discrete genera [52,53]. This notion was based on the fact that these are the only known Armillaria species that do not produce an annulus on the stipe at maturity. It was further thought that A. socialis (A. tabescens) is more thermophilic than other Armillaria species residing in the Northern Hemisphere [52,54]. Pegler [53] treated A. tabescens and A. ectypa in the section Desarmillaria, but without making a formal transfer to that genus.
The placement of A. ectypa and A. tabescens in a genus different to Armillaria is contentious because both genera have morphological features typical of Armillaria spp., including basidiospore morphology, the production of rhizomorphs in culture and diploid secondary mycelium. Furthermore, phylogenetic trees generated by Maphosa et al. [26] and Coetzee et al. [27] (Figure S1a) clustered the exannulated species in a monophyletic group that included Armillaria species from the Northern Hemisphere. Recently, phylogenetic analysis (Figure S1b) based on six gene regions by Koch et al. [29], however, showed that A. tabescens and A. ectypa reside in a monophyletic group sister to a clade that includes all known Armillaria lineages. Based on their morphological differences and phylogenetic position relative to other Armillaria spp. the authors formally erected the genus Desarmillaria and introduced the new combinations D. tabescens and D. ectypa to accommodate the two exannulated species of “Armillaria”. Desarmillaria consequently represents the phylogenetically closest known genus to Armillaria. In the present review, A. tabescens and A. ectypa are treated as Armillaria due to their taxonomic history and similarity to other Armillaria species.
5. Species Clusters and Phylogenetic Relationships Based on Morphological Cohesion, Phenotypic Characteristics and Phylogenetic Analyses
Most early studies on Armillaria focused on relationships among those species residing in the Holarctic region. But more recently, there has been work on species from the non-Holarctic regions, mainly focusing on species from sub-Sahara Africa, Australia and New Zealand [17,20,21]. It was only relatively recently that efforts were made to resolve phylogenetic relationships among species in South America [23]. Collectively, these studies suggest that Armillaria species reside in clusters, which we refer to here as lineages, that reflect their origin, morphological similarity and ecological characteristics. It has also been shown that the Australasian-South American species form a monophyletic group sister to the Holarctic species (with the exception of A. mellea and A. mexicana) and that species from Africa are a basal group in Armillaria as illustrated in the present study and supported by [28,29]. These different lineages are discussed below.
5.1. Species Lineages from the Holarctic
In this review, five Holarctic lineages are recognised (Figure 1). These are referred to as the A. gallica, A. solidipes/ostoyae, A. mellea, A. mexicana and Exannulated lineages based on results from Coetzee et al. [27] and Klopfenstein et al. [28] (although the latter publication referred to these lineages as superclades) as well as the analysis conducted for this review. Armillaria tabescens and A. ectypa are collectively referred to in the present review as the Exannulated lineage. Results of phylogenetic analyses by Coetzee et al. [27] and Klopfenstein et al. [28] suggested that the Armillaria gallica lineage forms a sister group with the A. solidipes/ostoyae lineage. The phylogeny produced by Coetzee et al. [27] indicated that the A. mellea and the Exannulated lineages are basal to the Holarctic species. However, this was not supported in the phylogenetic trees of Klopfenstein et al. [28], Koch et al. [29] and neither in the phylogenetic analyses emerging from the present study. Rather, the remainder of the Northern Hemisphere lineages were shown to be more closely related to lineages from the Southern Hemisphere than to the A. mellea and Exannulated lineages. An in-depth discussion dealing with the northern hemisphere lineages is provided in Klopfenstein et al. [28], therefore only the main issues relating the phylogeny and taxonomy of species in these lineages are provided below.
5.1.1. The Armillaria solidipes/ostoyae Lineage
Species in the Armillaria solidipes/ostoyae lineage include: A. ostoyae, A. borealis and A. gemina, A. sinapina (G15, North America and Japan) and A. cepistipes (G16 from Europe and G17 from Japan) and Chinese Biological Species (CBS) F (proposed as being A. cepistipes by [6]) based on phylogenetic trees (Figure 1) generated from tef-1α DNA sequence data [25,28]. Within this group, A. ostoyae, A. borealis and A. gemina, are morphologically similar in having a thick anulus, stipes that are more or less equal in shape and the presence of distinct dark scales [55]. Armillaria ostoyae (proposed by [56] as A. solidipes because this is the older name, but not always accepted [57]) has a transcontinental distribution, occurring in Europe, North America and Asia (China, Japan and South Korea), A. borealis occurs in both Europe and Asia (China) while A. gemina is confined to north-eastern North America [31,58,59,60] and as shown in Figure 2 (Table S4).
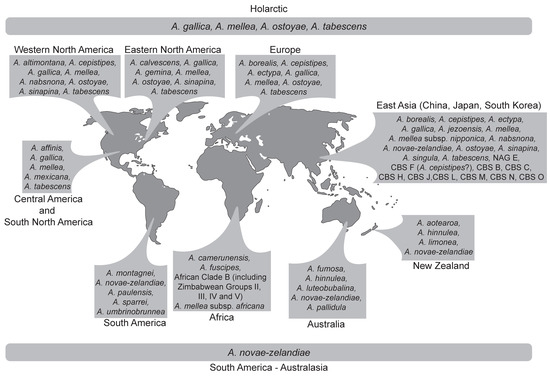
Figure 2.
Map showing the geographic distribution of Armillaria species and biological species not assigned to morphological species.
Phylogenetic studies have revealed that A. ostoyae, A. borealis and A. gemina are phylogenetically more closely related to each other than they are to the other Armillaria species from the Holarctic region (Figure 1). Anderson et al. [61] showed, using rDNA Restriction Fragment Length Polymorphism (RFLP) profiles, that the three species harbour distinct rDNA classes. This was later reflected in their grouping based on ITS and IGS-1 sequence phylogenies that separated them from other Holarctic Armillaria species [16,18]. More recent research has, however, shown that strains representing A. borealis and A. ostoyae, based on their identification using mating tests, have polyphyletic origins [6,19,25,28,62]. Strains of A. ostoyae reside in two paraphyletic lineages based on IGS-1 and tef-1α sequences (Figure 1), with one lineage grouping within a monophyletic group with A. borealis and A. gemina [25,28]. IGS-1 sequences group strains of A. borealis in a monophyletic group [25], however, phylogenetic trees generated from tef-1α gene sequences (Figure 1) place them in paraphyletic groups [25,28]. One group forms a monophyletic group with A. gemina and A. ostoyae, while strains from the second group cluster in a monophyletic group distant to the A. solidipes/ostoyae lineage. Overall the results of these studies suggest that A. ostoyae and A. borealis represent species complexes with cryptic species that cannot be detected using mating tests.
Phylogenetic studies based on ITS, IGS-1 or a combination of these regions together with tef-1α sequences have shown that isolates of A. sinapina and A. cepistipes are closely related to Armillaria species in the A. gallica lineage [25,63,64]. However, phylogenetic trees generated by different authors based on tef-1α sequence data alone suggested that some isolates representing these species are closely related to A. borealis, A. gemina and A. ostoyae [6,64]. Most recently, Klopfenstein et al. [28] showed that A. sinapina (Group 15, from North America and Japan) and two groups of A. cepistipes (Group 16 from Europe and Group 17 from Japan) constitute a monophyletic group sister to A. borealis—A. solidipes/ostoyae—A. gemina with strong statistical support. This was also the case in the current analysis (Figure 1).
5.1.2. The Armillaria gallica Lineage
The Armillaria gallica lineage represents the largest and most complex group of Holarctic species. The cluster includes A. altimontana, A. calvescens, A. cepistipes, A. gallica, A. jezoensis, A. nabsnona, A. sinapina and A. singula (Figure 1 and Figure 2, Table S4). Several biological species and phylogenetic lineages that are not yet linked to morphological species have been identified and reside in this lineage (Figure 1, Table S4). Species in this lineage can generally be distinguished by their thin delicate annuli and more bulbose or clavate stipes [55]. The species residing in this cluster differ in their geographical distributions, some being confined to certain continents while others have a transcontinental distribution (Figure 2, Table S4).
Determining the phylogenetic position of species in the A. gallica lineage is complicated by gene trees in which isolates of the same species occur at different positions, isolates supposed to represent the same species being placed distant to each other in phylogenetic trees generated from the same locus and overall low statistical support for nodes [25,28]. The taxonomic status of A. calvescens, A. nabsnona, A. altimontana and NAG E is supported by the results of several studies that showed that isolates representing these species formed distinct monophyletic clades [6,25,28] (Figure 1). However, their phylogenetic position relative to each other and the remaining Armillaria species in this cluster is not clear due to low statistical support in the published phylogenetic trees [6,25,28] and in our analyses for this review.
Within the A. gallica lineage, the taxonomy of A. gallica and A. cepistipes is problematic since phylogenetic studies have shown that these species are polyphyletic [6,25,28]. DNA sequence analyses revealed that European and Northern American A. gallica are phylogenetically distinct [62] and that A. gallica from East-Asia is genetically different to European A. gallica [6]. It is also known that A. gallica constitutes up to eight clades [28] with multiple phylogenetic groups being present in North America [65], Europe [66] and Asia [6,25]. Guo et al. [6] suggested that A. gallica is restricted to Europe and “A. gallica” from East Asia and North America should be treated as a different species. Armillaria cepistipes from North America resides in the A. gallica lineage while A. cepistipes from Europe and Japan cluster with species in the A. solidipes/ostoyae lineage based on tef-1α sequence data [25,28] (Figure 1). But, phylogenetic trees based on sequences for other loci group A. cepistipes from the different regions in the A. gallica lineage [25,38,63]. Collectively these studies have shown that A. gallica and A. cepistipes represent species complexes with several cryptic species that warrant further taxonomic investigation.
Several biological species that are restricted to China (Figure 2, Table S4) and that were later assigned to phylogenetic lineages [6] reside in the A. gallica lineage [25]. The Chinese biological species (CBS) in this lineage include CBS C, J, H, O, M and L (Figure 1). The phylogenetic relationships among these biological species are presently not resolved because the loci used in phylogenetic analyses do not provide sufficient phylogenetic signal for a robust phylogeny [6,25]. This is further complicated by the fact that single gene genealogies or concatenated data yield phylogenetic trees in which a phylogenetic lineage includes more than one biological species, while isolates identified as being the same biological species may reside in more than one phylogenetic lineage [6] (Table S4).
5.1.3. The Armillaria mellea Lineage
Armillaria mellea is the only species accommodated in this lineage (Figure 1). Representatives of this lineage are characterised by the lack of clamp connections at the base of their basidia, the presence of a distinct annulus, honey coloured caps and robust basidiocarps [67]. Isolates in the lineage have a transcontinental distribution occurring in eastern, western and central North America [31,65,68,69], Europe [58], Middle East [70] and Asia [59,71] (Figure 2). Armillaria mellea is also known from South Africa [72], Tanzania, Ethiopia, Sao Tome and Kenya [73,74,75] and as illustrated in Figure 2 (Table S4). However, these isolates represent examples of accidental introductions into new environments [72,76,77,78].
Members of the A. mellea lineage display considerable intraspecific variation at the DNA sequence level. Phylogenetic studies based on ITS, IGS-1 and tef-1α sequence data have revealed geographic clades within the lineage, distinguished by their origins. These include clades from Europe, western and eastern North America and Asia [26,79]. Population differentiation between western and eastern North America was later supported by microsatellite analysis [69]. The phylogenetic relationships among the clades are, however, not clear since the phylogenetic position of the clades based on different loci are not congruent. In view of the intraspecific variation in A. mellea, it was suggested that geographically separated populations of this species are in the process of speciation as a result of genetic isolation [79]. Based on DNA sequence variation between the geographical clusters, we hypothesise that the A. mellea lineage represents a species complex that is comprised of different cryptic species.
Members of the A. mellea lineage display intraspecific variation in their sexual systems. Similar to most Armillaria species, isolates that have been identified as A. mellea from Europe and North America and the majority of isolates from China are heterothallic. Homothallic forms were reported from Japan and China [59,80] as well as from the African countries of Ethiopia, Kenya, Tanzania, and Sao Tome [75,81]. The homothallic mating system was shown to be a unique form of secondary homothallism, and is therefore referred to as non-heterothallic [80]. The non-heterothallic isolates of A. mellea from Africa and Japan have been recognised as A. mellea ssp. africana [31] and A. mellea ssp. nipponica [82], respectively. Ota et al. [81] found that the Japanese population has high genetic diversity while African A. mellea isolates were identical to Japanese somatic incompatibility group A isolates based on somatic compatibility tests, isozyme and Random Amplification of Polymorphic DNA (RAPD) analyses. The authors therefore hypothesised that A. mellea spp. africana originated from East Asia. Phylogenetic analyses of sequences for tef-1α and ITS showed that the non-heterothallic and heterothallic forms from Asia and Africa are closely related in a clade [6,25]. Consequently, they cannot be differentiated based on the currently available DNA sequence data. However, Ota et al. [81] separated the two mating forms based on RAPDs, but isolates from China were not included (only isolates from Japan) in their study.
5.1.4. The Armillaria mexicana Lineage
Armillaria mexicana is the only species accommodated in this lineage (Figure 1) and is known only from Mexico (Figure 2, Table S4). The species was placed with A. mellea in the “Mellea superclade” as denoted by Klopfenstein et al. [28]. However, with the exception of fibulae that are absent in the basidiomata, this species shows no other morphological similarity with A. mellea [83]. Isolates representing this species form a monophyletic group sister to A. mellea [28,83] (Figure 1). Furthermore, this species is unique in having the largest ITS region thus far reported for fungi with an ITS1 of 1299 bp and an ITS2 of 582 bp [83]. By virtue of these characteristics, A. mexicana is considered to represent a well-defined lineage in this review.
5.1.5. The Exannulated Lineage of Species
Armillaria tabescens (A. socialis [52]) and A. ectypa (now referred to as Desarmillaria tabescens and D. ectypa [29]) are characterised by the lack of an annulus on their stipes. The two species differ in terms of their ecology and distribution (Table S4). Armillaria ectypa has a narrow distribution, being reported only from Europe (see [84] for an account of the distribution and references), Turkey [85], China ([86] cited in [59]) and Japan [87,88]. In Europe it is an extremely rare species and in many countries is considered to be endangered. Armillaria ectypa occurs as a saprophyte on woody material found in peat bogs and marshy habitats (e.g., [54]). Armillaria tabescens has a much broader distribution than A. ectypa being reported from southern Europe [52,58], North and Central America [89,90,91], South Korea [60,92,93], southern Japan [71] and China [59,94]. In some of these regions they are considered important pathogens on woody shrubs and trees (e.g., [91]).
Phylogenetic trees generated from different loci have suggested that A. ectypa and A. tabescens speciated from a common ancestor. In a study by Ota et al. [95], isolates representing A. ectypa and A. tabescens, respectively, formed sister groups. However, this relationship did not receive statistical support through bootstrap analysis and an outgroup to polarise the tree was not included in their study. The sister relationship between A. ectypa and A. tabescens was strongly supported in the phylogenetic trees presented in the studies of Coetzee et al. [27], Koch et al. [29] and Klopfenstein et al. [28].
The taxonomy of A. tabescens from North America, Europe and Asia is controversial due to inconclusive results emerging from sexual compatibility and phylogenetic studies. Sexual compatibility between isolates from North America and one isolate from Italy suggested that these isolates represent the same species [96]. In contrast, Guillaumin et al. [58] reported that A. tabescens from Europe is compatible with isolates from Asia but not with isolates from North America. Kile et al. [31] consequently suggested that A. tabescens from North America should be assigned to A. monodelpha, but this name was considered illegitimate by Volk et al. [14]. Ota et al. [71] found that strains from Japan were interfertile with European strains but intersterile with one isolate from North America. Later, Qin et al. [59] reported that strains from China (CBS I) are sexually compatible with strains of A. tabescens from Europe. In a more recent phylogenetic study, Tsykun et al. [64] showed that European and Japanese isolates of A. tabescens form separate clades when sequences from the IGS-1, ITS, and tef-1α regions are combined. That study, however, included a limited number of samples of A. tabescens and the grouping of Japanese isolates received marginal statistical support.
In view of the discrepancies in sexual compatibility between isolates of A. tabescens from the different regions, Coetzee et al. [25] attempted to resolve the taxonomic uncertainty following a phylogenetic approach. Results of their study supported the notion that A. tabescens from North America, Europe and Asia should be treated as a single taxon. The analyses presented by Coetzee et al. [25] was based on sequences for only two loci that are linked (ITS and IGS-1) and limited strain sampling. There is consequently a need for further studies before a definitive conclusion can be reached regarding the taxonomic treatment of A. tabescens from Europe, North America and Asia.
5.2. Species Lineages from Australasia-South America
In comparison to the Holarctic species, studies focusing on the phylogenetic relationships of Armillaria species from the non-Holarctic regions, excluding sub-Sahara Africa, are limited in number. To date, the most comprehensive work has been conducted on species from Australia and New Zealand [20,21] and the Patagonian Andes [23]. Results of these studies showed that the Australasian-South American Armillaria species represent a very divergent group of fungi in terms of their DNA sequence characteristics. The IGS-1 ranged between 400 to 1500 bp among different species [20], in comparison to the narrow size range (845–920 bp) among the Holarctic species [64,66,97]. Intra-strain sequence heterogeneity of the IGS-1 copies [20] and heterogenic multi copies of the tef-1α gene within the genomes of certain species [23] have been reported. Due to this variation, phylogenetic studies focussed mainly on ITS and LSU sequence data to infer phylogenetic relationships. Collectively, results of these studies showed that the Armillaria species from South America and Australasia reside in clades that we refer to as the Armillaria hinnulea lineage, Armillaria novae-zelandiae lineage and Armillaria luteobubalina lineage.
5.2.1. The Armillaria hinnulea Lineage
The Armillaria hinnulea lineage includes A. hinnulea, A. aotearoa, A. umbrinobrunnea and A. sparrei (Figure 1 and Figure S1e). Armillaria hinnulea is found in Tasmania, south-eastern Australia and Nothofagus forests in the north-western part of the South Island of New Zealand [98,99,100] (Figure 2, Table S4). Armillaria aotearoa has a restricted distribution known only from the North and South Islands of New Zealand and is morphologically similar to A. hinnulea ([100] and references therein). Armillaria umbrinobrunnea and A. sparrei are restricted to South America ([101] and references therein) (Figure 2, Table S4).
Phylogenetic inference based on ITS sequences has revealed that A. hinnulea and A. aotearoa, previously referred to as “unknown species” [20,26], are more closely related to Armillaria species from the Holarctic than to those from Australia, New Zealand and South America [20] (Figure S1d). However, phylogenetic trees generated from tef-1α sequences (Figure 1) placed A. hinnulea sister to A. fumosa and A. pallidula that resides in the Armillaria novae-zelandiae lineage [26]. It was suggested that the incongruence in the gene tree topologies was due to the rapid evolution of the ITS region and that the tef-1α gene of A. hinnulea retained the ancestral character states of species within the non-Holarctic group [26].
The close affinity of A. hinnulea with species from the Holarctic is supported by morphological characteristics. Similar to species in the A. gallica and A. ostoyae clusters, A. hinnulea produces clamp connections in sub-hymenial layers of the basidiocarp [98]. These findings are also consistent with the view of Kile and Watling [98] that A. hinnulea resembles the Holarctic species A. cepistipes.
5.2.2. The Armillaria novae-zelandiae Lineage
The Armillaria novae-zelandiae lineage accommodates A. novae-zelandiae and one isolate of A. puiggarii from Guadeloupe (Figure 1). Among the species occurring in the South American and Australasia regions, A. novae-zelandiae has the widest geographic distribution, occurring in Australia, New Zealand, South America, Papua New Guinea, Indonesia, Malaysia and Amami-Oshima, which is a subtropical island of Japan [22,95,98,102] (Figure 2, Table S4).
Comparison of published phylogenetic trees and those emerging from the present study (Figure 1) suggest four separate lineages for isolates of A. novae-zelandiae [20,22,23,26,95,101]. These also reflect their geographic origins: Australia, New Zealand, South America (Argentina and Chile) and Asia (Indonesia, Malaysia and Amami-Oshimi). Isolates from Australia and New Zealand are reciprocally monophyletic but considered conspecific by virtue of their similar basidiocarp morphology, vegetative growth characteristics and sexual compatibility [98,103]. The South American lineage is sister to the Australasian clade [22,23], while isolates from Asia are a basal monophyletic lineage within A. novae-zelandiae [22,26,95]. Ota et al. [95] reported that the basidiocarp morphology of collections from Amami-Oshimi closely resembles that of A. fuscipes from Africa. However, it is phylogenetically distantly related to African species and could therefore represent a newly discovered species. The lineages within A. novae-zelandiae suggest that the separate populations are in the process of allopatric speciation due to geographic isolation.
5.2.3. The Armillaria luteobubalina Lineage
The Armillaria luteobubalina lineage is represented by several species that have different distribution patterns in the southern hemisphere (Figure 1 and Figure 2, Table S4). This group includes A. fumosa, A. pallidula, A. luteobubalina, A. limonea, A. montagnei and A. paulensis. Armillaria pallidula, A. fumosa and A. luteobubalina are restricted to Australia [104]. Armillaria montagnei has been identified in Chile and Argentina [22,101]. Armillaria limonea was reported from New Zealand and South America (Chile and Argentina) [105,106,107]. Armillaria paulensis is known from only one location in the south of São Paulo City [108].
The relationships among species within the A. luteobubalina lineage are not yet clear. This is due to topological differences among single locus trees and lack of nodal support for certain species groupings. However, comparisons of published phylogenetic trees have revealed a clustering of certain species that are supported by phenotypic traits.
Armillaria pallidula and A. fumosa together with A. luteobubalina and A. montagnei are pairs of sibling species based on individual ITS, tef-1α and LSU phylogenetic trees [20,23,26] (Figure 1 and Figure S1a,e). Armillaria pallidula and A. fumosa cannot be differentiated based on ITS or tef-1α DNA sequences [20,26] and they are similar in some morphological features [104]. However, they were shown to be distinct biological species based on interfertility tests [104]. The sister relationship between A. luteobubalina and A. montagnei is supported by pileus characteristics and an apparently “unpleasant” flavour [101,109,110]. Phylogenetic trees generated from ITS sequence data showed that A. paulensis is a close evolutionary relative of the A. luteobubalina–A. montagnei group [23,108].
5.3. The African Armillaria Lineage
The African lineage includes two major clades [17,26,111], referred to as A. fuscipes (=A. heimii) and African Clade B [17]. A disguising feature of this cluster is the fact that the 5S gene is inverted relative to other Armillaria species [17,34]. In addition, all isolates included in the studies of several authors [75,112,113] lack a tetrapolar (bifactorial) heterothallic mating system with some being bipolar (unifactorial) heterothallic while others are homothallic.
A number of studies have revealed sub-groups within the two major African clades. Two sister sub-groups were recognised in the A. fuscipes clade represented by isolates from Ethiopia and Kenya, respectively, based on ITS and IGS-1 sequence data [17,114]. Clade B is the most diverse group based on ITS-1 and IGS-1 sequences as well as AFLP and somatic compatibility studies. Together with other isolates from Africa, this clade includes sub-groups referred to as Zimbabwean Groups II, III, IV and V [115,116]. Collectively, results of these studies suggest that the different sub-groups in the major clades possibly represent distinct species. But, Pérez-Sierra et al. [111] were of the view that the major clades and their sub-groups were a single species comprising different molecular groups.
In addition to the taxa described above and A. mellea ssp. africana, an additional lineage occurring on tea (Camellia sinensis) in Kenya has been recognised. The lineage is referred to as Kenyan Group III [117], SIG (somatic incompatibility group) II [118], SIG III [119] and Kenyan Group II [120] by different authors. Different from other African taxa, this group is more similar to northern hemisphere species in that the 5S gene is not inverted [117,120]. Furthermore, it is phylogenetically more closely related to northern hemisphere species [120], and formed a sister group with A. hinnulea based on IGS-1 [117] and tef-1α sequences [28] (Figure 1). We postulate that the fungus was most likely introduced into Africa.
6. Conclusions and Future Prospects
Substantial progress has been made during the past 26 years towards understanding the identity and phylogenetic relationships of Armillaria root pathogens. Using DNA sequence data and phylogenetic methods, new species are regularly being discovered or recognised. However, there is still much to be learned regarding the species boundaries between closely related taxa, especially those in the A. gallica lineage. But, questions relating to species boundaries of these taxa will be resolved only by molecular phylogenies based on DNA or amino acid sequences from multiple independent evolving loci.
Genealogical Concordance Phylogenetic Species Recognition (GCPSR) using the concordance among several gene trees [121,122] to delineate species has become standard in fungal taxonomy. But, with the exception of a few studies (e.g., [6,64]) this taxonomic method has not yet been widely implemented in Armillaria taxonomy. The limited application of GCPSR in Armillaria taxonomic studies can be ascribed mainly to the absence of a standard set of informative unlinked single copy genes. This warrants a concerted effort to identify an assemblage of genes that will facilitate GCPSR in Armillaria taxonomy.
Phylogenomics based on a large collection of orthologous genes or transcriptomes provides a promising method for future research that focuses on Armillaria species boundaries, constructing robust phylogenetic frameworks, and to diagnose species. Earlier work by Rokas et al. [123] showed that at least 20 unlinked genes or 8000 randomly selected orthologous nucleotides distributed over the sampled genomes are required to construct a robust species phylogeny and this will probably be the case for Armillaria phylogenomics. Using a phylogenomic approach has already lead to progress in fungal systematics and the diagnoses of species new to science [124]. It is evident that the reduction in cost associated with whole genomes sequencing [125], advances in whole genome sequence technology and bioinformatic pipelines will make it feasible for research groups to conduct phylogenomic studies on large collections of Armillaria isolates. Even if whole genomes cannot be sequenced, several genome sampling methods are available to obtain phylogenetic informative characters that can be used in phylogenomic studies at various scales (see [126]). Sequences of the genomes of key species [127] are already providing exciting prospects to study the evolution and systematics of Armillaria. They are certain to lead to important breakthroughs regarding not only the taxonomy but the biology and ecology of these fascinating fungi in the future.
Supplementary Materials
The following are available online at http://www.mdpi.com/2076-0817/7/4/83/s1. Table S1: Summary of genes and genomic regions employed in phylogenetic studies of Armillaria species. Table S2: Curated database of published DNA sequences from GenBank for Armillaria species. Table S3: Additional tef-1α DNA sequences from Armillaria species or taxa included in this study. Table S4: Distribution of Armillaria species and taxa from phylogenetic studies and their associated biological species designation. Figure S1. Phylogenetic trees generated from publications for species for which tef-1α DNA are not available or that had conflicting phylogenetic positions based on genomic regions other than tef-1α.
Author Contributions
Methodology, M.P.A.C.; writing—original draft preparation, M.P.A.C.; writing—review and editing, M.P.A.C., B.D.W. and M.J.W.
Funding
This research was funded by the Tree Protection Co-operative Programme (TPCP), Department of Science and Technology (DST)—National Research Foundation (NRF) Centre of Excellence in Tree Health Biotechnology (CTHB).
Acknowledgments
We thank the reviewers for their valuable and insightful suggestions to improve the original manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References and Note
- Shaw, C.G.; Kile, G.A. Armillaria Root Disease. Agriculture Handbook No. 691; Forest Service, United States Department of Agriculture: Washington, DC, USA, 1991.
- Cha, J.Y.; Igarashi, T. Armillaria species associated with Gastrodia elata in Japan. Eur. J. For. Pathol. 1995, 25, 319–326. [Google Scholar] [CrossRef]
- Terashita, T. Biological species of Armillaria symbiotic with Galeola septentrionalis. Nippon Kingakukai Kaiho 1996, 37, 45–49. [Google Scholar]
- Cha, J.Y.; Igarashi, T. Armillaria jezoensis, a new symbiont of Galeola septentrionalis (Orchidaceae) in Hokkaido. Mycoscience 1996, 37, 21–24. [Google Scholar] [CrossRef]
- Sekizaki, H.; Kuninaga, S.; Yamamoto, M.; Asazu, S.N.; Sawa, S.; Kojoma, M.; Yokosawa, R.; Yoshida, N. Identification of Armillaria nabsnona in gastrodia tubers. Biol. Pharm. Bull. 2008, 31, 1410–1414. [Google Scholar] [CrossRef] [PubMed]
- Guo, T.; Wang, H.C.; Xue, W.Q.; Zhao, J.; Yang, Z.L. Phylogenetic analyses of Armillaria reveal at least 15 phylogenetic lineages in China, seven of which are associated with cultivated Gastrodia elata. PLoS ONE 2016, 11, e0154794. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.X.; Xu, J.T. Nutrient source of sclerotia of Grifola umbellata and its relationship to Armillaria mellea. Acta Bot. Sin. 1992, 34, 576–580. [Google Scholar]
- Kikuchi, G.; Yamaji, H. Identification of Armillaria species associated with Polyporus umbellatus using ITS sequences of nuclear ribosomal DNA. Mycoscience 2010, 51, 366–372. [Google Scholar] [CrossRef]
- Lee, M.W.; Chang, K.C.; Shin, D.B.; Lee, K.R.; Im, K.H.; Jin, G.H.; Shin, P.G.; Xing, Y.M.; Chen, J.; Guo, S.X.; et al. The culture conditions for mycelial growth and sclerotial formation of Polyporus umbellatus. J. Mushroom Sci. Prod. 2013, 11, 194–200. [Google Scholar] [CrossRef]
- Xing, X.; Men, J.; Guo, S. Phylogenetic constrains on Polyporus umbellatus-Armillaria associations. Sci. Rep. 2017, 7, 4226. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.L.; Bruhn, J.N.; Anderson, J.B. The fungus Armillaria bulbosa is among the largest and oldest living organisms. Nature 1992, 356, 428–431. [Google Scholar] [CrossRef]
- Ferguson, B.A.; Dreisbach, T.A.; Parks, C.G.; Filip, G.M.; Schmitt, C.L. Coarse-scales population structure of pathogenic Armillaria species in a mixed-conifer forest in the Blue Mountains of northeast Oregon. Can. J. For. Res. 2003, 33, 612–623. [Google Scholar] [CrossRef]
- Vahl, M. (1787–1799). Flora Danica. Fasc. 16–21, Tab. 901–1260.—(1794). Nogle iagttagelser ved en reise giennem Norge til dets nordlige dele (2). - Skr. Naturhist. Selsk.
- Volk, T.J.; Burdsall, H.H. A Nomenclatural Study of Armillaria and Armillariella Species (Basidiomycotina, Tricholomataceae); Fungiflora: Førde, Norway, 1995; p. 121. [Google Scholar]
- Watling, R.; Kile, G.A.; Gregory, N.M. The genus Armillaria–nomenclature, typification, the identity of Armillaria mellea and species differentiation. Trans. Br. Mycol. Soc. 1982, 78, 271–285. [Google Scholar] [CrossRef]
- Anderson, J.B.; Stasovski, E. Molecular phylogeny of Northern Hemisphere species of Armillaria. Mycologia 1992, 84, 505–516. [Google Scholar] [CrossRef]
- Coetzee, M.P.; Wingfield, B.D.; Bloomer, P.; Wingfield, M.J. Phylogenetic analyses of DNA sequences reveal species partitions amongst isolates of Armillaria from Africa. Mycol. Res. 2005, 109, 1223–1234. [Google Scholar] [CrossRef] [PubMed]
- Chillali, M.; Wipf, D.; Guillaumin, J.-J.; Mohammed, C.; Botton, B. Delineation of the European Armillaria species based on the sequences of the internal transcribed spacer (ITS) of ribosomal DNA. New Phytol. 1998, 138, 553–561. [Google Scholar] [CrossRef]
- Mulholland, V.; MacAskill, G.A.; Laue, B.E.; Steele, H.; Kenyon, D.; Green, S. Development and verification of a diagnostic assay based on EF-1 a for the identification of Armillaria species in Northern Europe. For. Pathol. 2012, 42, 229–238. [Google Scholar] [CrossRef]
- Coetzee, M.P.A.; Wingfield, B.D.; Bloomer, P.; Ridley, G.S.; Kile, G.A.; Wingfield, M.J. Phylogenetic relationships of Australian and New Zealand Armillaria species. Mycologia 2001, 93, 887–896. [Google Scholar] [CrossRef]
- Dunne, C.P.; Glen, M.; Tommerup, I.C.; Shearer, B.L.; Hardy, G.E.S.J. Sequence variation in the rDNA ITS of Australian Armillaria species and intra-specific variation in A. luteobubalina. Australas. Plant Pathol. 2002, 31, 241–251. [Google Scholar] [CrossRef]
- Coetzee, M.P.A.; Wingfield, B.D.; Bloomer, P.; Ridley, G.S.; Wingfield, M.J. Molecular identification and phylogeny of Armillaria isolates from South America and Indo-Malaysia. Mycologia 2003, 95, 285–293. [Google Scholar] [CrossRef] [PubMed]
- Pildain, M.; Coetzee, M.; Rajchenberg, M.; Petersen, R.; Wingfield, M.; Wingfield, B. Molecular phylogeny of Armillaria from the Patagonian Andes. Mycol. Prog. 2009, 8, 181–194. [Google Scholar] [CrossRef]
- Terashima, K.; Cha, J.Y.; Yajima, T.; Igarashi, T.; Miura, K. Phylogenetic analysis of Japanese Armillaria based on the intergenic spacer (IGS) sequences of their ribosomal DNA. Eur. J. For. Pathol. 1998, 28, 11–19. [Google Scholar] [CrossRef]
- Coetzee, M.P.A.; Wingfield, B.D.; Zhao, J.; van Coller, S.J.; Wingfield, M.J. Phylogenetic relationships among biological species of Armillaria from China. Mycoscience 2015, 56, 530–541. [Google Scholar] [CrossRef]
- Maphosa, L.; Wingfield, B.D.; Coetzee, M.P.A.; Mwenje, E.; Wingfield, M.J. Phylogenetic relationships among Armillaria species inferred from partial elongation factor 1-alpha DNA sequence data. Australas. Plant Pathol. 2006, 35, 513–520. [Google Scholar] [CrossRef]
- Coetzee, M.P.A.; Bloomer, P.; Wingfield, M.J.; Wingfield, B.D. Paleogene radiation of a plant pathogenic mushroom. PLoS ONE 2011, 6, e28545. [Google Scholar] [CrossRef] [PubMed]
- Klopfenstein, N.B.; Stewart, J.E.; Ota, Y.; Hanna, J.W.; Richardson, B.A.; Ross-Davis, A.L.; Elías-Román, R.D.; Korhonen, K.; Keča, N.; Iturritxa, E.; et al. Insights into the phylogeny of Northern Hemisphere Armillaria: Neighbor-net and Bayesian analyses of translation elongation factor 1-α gene sequences. Mycologia 2017, 109, 75–91. [Google Scholar] [CrossRef] [PubMed]
- Koch, R.A.; Wilson, A.W.; Séné, O.; Henkel, T.W.; Aime, M.C. Resolved phylogeny and biogeography of the root pathogen Armillaria and its gasteroid relative, Guyanagaster. BMC Evol. Biol. 2017, 17, 33. [Google Scholar] [CrossRef] [PubMed]
- Morrone, J.J. Biogeographical regionalisation of the world: A reappraisal. Aust. Syst. Bot. 2015, 28, 81–90. [Google Scholar] [CrossRef]
- Kile, G.A.; Guillaumin, J.-J.; Mohammed, C.; Watling, R. Biogeography and pathology of Armillaria. In Proceedings of the Eigth International Conference on Root and Butt Rots, Wik, Sweden; Haikko, Finland, 9–16 August 1993; Swedish University of Agricultural Science: Uppsala, Sweden, 1994; pp. 411–436. [Google Scholar]
- Coetzee, M.P.A.; Wingfield, B.D.; Kirisits, T.; Chhetri, D.B.; Bloomer, P.; Wingfield, M.J. Identification of Armillaria isolates from Bhutan based on DNA sequence comparisons. Plant Pathol. 2005, 54, 36–45. [Google Scholar] [CrossRef]
- Hanna, J.W.; Klopfenstein, N.B.; Kim, M.S.; McDonald, G.I.; Moore, J.A. Phylogeographic patterns of Armillaria ostoyae in the western United States. For. Pathol. 2007, 37, 192–216. [Google Scholar] [CrossRef]
- Coetzee, M.P.A.; Wingfield, B.D.; Coutinho, T.A.; Wingfield, M.J. Identification of the causal agent of Armillaria root rot of Pinus species in South Africa. Mycologia 2000, 92, 777–785. [Google Scholar] [CrossRef]
- Matheny, P.B.; Wang, Z.; Binder, M.; Curtis, J.M.; Lim, Y.W.; Nilsson, R.H.; Hughes, K.W.; Hofstetter, V.; Ammirati, J.F.; Schoch, C.; et al. Contributions of rpb2 and tef1 to the phylogeny of mushrooms and allies (Basidiomycota, Fungi). Mol. Phylogen. Evol. 2007, 43, 430–451. [Google Scholar] [CrossRef] [PubMed]
- Kauserud, H.; Schumacher, T. Outcrossing or inbreeding: DNA markers provide evidence for type of reproductive mode in Phellinus nigrolimitatus (Basidiomycota). Mycol. Res. 2001, 105, 676–683. [Google Scholar] [CrossRef]
- Baumgartner, K.; Bhat, R.; Fujiyoshi, P. A rapid infection assay for Armillaria and real-time PCR quantitation of the fungal biomass in planta. Fungal Biol 2010, 114, 107–119. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, E.; Ota, Y.; Hattori, T.; Kikuchi, T. Sequence-based identification of Japanese Armillaria species using the elongation factor-1 alpha gene. Mycologia 2010, 102, 890–910. [Google Scholar] [CrossRef]
- Brazee, N.J.; Hulvey, J.P.; Wick, R.L. Evaluation of partial tef1, rpb2, and nLSU sequences for identification of isolates representing Armillaria calvescens and Armillaria gallica from northeastern North America. Fungal Biol. 2011, 115, 741–749. [Google Scholar] [CrossRef] [PubMed]
- Ross-Davis, A.L.; Hanna, J.W.; Klopfenstein, N.B.; Kim, M.-S. Advances toward DNA-based identification and phylogeny of North American Armillaria species using elongation factor-1 alpha gene. Mycoscience 2012, 53, 161–165. [Google Scholar] [CrossRef]
- Burdsall, H.H.; Volk, T.J. The state of taxonomy of the genus Armillaria. McIlvainea 1993, 11, 4–12. [Google Scholar]
- Zhao, Z.; Liu, H.; Luo, Y.; Zhou, S.; An, L.; Wang, C.; Jin, Q.; Zhou, M.; Xu, J.-R. Molecular evolution and functional divergence of tubulin superfamily in the fungal tree of life. Sci. Rep. 2014, 4, 6746. [Google Scholar] [CrossRef] [PubMed]
- Piercey-Normore, M.D.; Egger, K.N.; Bérubé, J.A. Molecular phylogeny and evolutionary divergence of North American Biological Species of Armillaria. Mol. Phylogen. Evol. 1998, 10, 49–66. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2017, bbx108. [Google Scholar] [CrossRef] [PubMed]
- Suchard, M.A.; Lemey, P.; Baele, G.; Ayres, D.L.; Drummond, A.J.; Rambaut, A. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol. 2018, 4, vey016. [Google Scholar] [CrossRef] [PubMed]
- Maddison, W.P.; Maddison, D.R. Mesquite: A Modular System for Evolutionary Analysis. Version 3.5. Available online: http://www.mesquiteproject.org (accessed on 21 October 2018).
- Moncalvo, J.-M.; Lutzoni, F.M.; Rhener, S.A.; Johnson, J.; Vilgalys, R. Phylogenetic relationships of agaric fungi based on nuclear large subunit ribosomal DNA sequences. Syst. Biol. 2000, 49, 278–305. [Google Scholar] [CrossRef] [PubMed]
- Moncalvo, J.-M.; Vilgalys, R.; Redhead, S.A.; Johnson, J.E.; James, T.Y.; Catherine Aime, M.; Hofstetter, V.; Verduin, S.J.W.; Larsson, E.; Baroni, T.J.; et al. One hundred and seventeen clades of euagarics. Mol. Phylogen. Evol. 2002, 23, 357–400. [Google Scholar] [CrossRef]
- Matheny, P.B.; Curtis, J.M.; Hofstetter, V.; Aime, M.C.; Moncalvo, J.-M.; Ge, Z.-W.; Yang, Z.-L.; Slot, J.C.; Ammirati, J.F.; Baroni, T.J.; et al. Major clades of Agaricales: A multilocus phylogenetic overview. Mycologia 2006, 98, 982–995. [Google Scholar] [CrossRef] [PubMed]
- Garnica, S.; Weiss, M.; Walther, G.; Oberwinkler, F. Reconstructing the evolution of agarics from nuclear gene sequences and basidiospore ultrastructure. Mycol. Res. 2007, 111, 1019–1029. [Google Scholar] [CrossRef] [PubMed]
- Henkel, T.W.; Smith, M.E.; Aime, M.C. Guyanagaster, a new wood-decaying sequestrate fungal genus related to Armillaria (Physalacriaceae, Agaricales, Basidiomycota). Am. J. Bot. 2010, 97, 1471–1484. [Google Scholar] [CrossRef] [PubMed]
- Antonín, V.; Jankovský, L.; Lochman, J.; Tomšovský, M. Armillaria socialis morphological anatomical and ecological characteristics, pathology, distribution in the Czech Republic and Europe and remarks on its genetic variation. Czech Mycol. 2006, 58, 209–224. [Google Scholar]
- Pegler, D.N. Taxonomy, nomenclature and description of Armillaria. In Armillaria Root Rot: Biology and Control of Honey Fungus; Fox, R.T.V., Ed.; Intercept Limited: Andover, UK, 2000; pp. 81–93. [Google Scholar]
- Zolciak, A.; Bouteville, R.J.; Tourvieille, J.; Roeckel-Drevet, P.; Nicolas, P.; Guillaumin, J.J. Occurrence of Armillaria ectypa (Fr.) Lamoure in peat bogs of the Auvergne—The reproduction system of the species. Cryptogam. Mycol. 1997, 18, 299–313. [Google Scholar]
- Korhonen, K. Armillaria since Elias Fries. Acta Universitatis Upsaliensis Symbolae Botanicae Upsalienses 1995, 30, 153–161. [Google Scholar]
- Burdsall, H.H.; Volk, T.J. Armillaria solidipes, an older name for the fungus called Armillaria ostoyae. N. Am. Fungi 2008, 3, 261–267. [Google Scholar] [CrossRef]
- Hunt, R.S.; Morrison, D.J.; Bérubé, J. Armillaria solidipes is not a replacement name for A. ostoyae. For. Pathol. 2011, 41, 253–254. [Google Scholar] [CrossRef]
- Guillaumin, J.J.; Mohammed, C.; Anselmi, N.; Courtecuisse, R.; Gregory, S.C.; Holdenrieder, O.; Intini, M.; Lung, B.; Marxmuller, H.; Morrison, D.; et al. Geographical distribution and ecology of the Armillaria species in western Europe. Eur. J. For. Pathol. 1993, 23, 321–341. [Google Scholar] [CrossRef]
- Qin, G.F.; Zhao, J.; Korhonen, K. A study on intersterility groups of Armillaria in China. Mycologia 2007, 99, 430–441. [Google Scholar] [CrossRef] [PubMed]
- Park, K.H.; Oh, S.Y.; Park, M.S.; Kim, M.S.; Klopfenstein, N.B.; Kim, N.K.; Park, J.Y.; Kim, J.J.; Han, S.K.; Lee, J.K.; et al. Re-evaluation of Armillaria and Desarmillaria in South Korea based on ITS/tef1 sequences and morphological characteristics. For. Pathol. 2018, e12447. [Google Scholar] [CrossRef]
- Anderson, J.B.; Bailey, S.S.; Pukkila, P.J. Variation in ribosomal DNA among biological species of Armillaria, a genus of root-infecting fungi. Evolution 1989, 43, 1652–1662. [Google Scholar] [PubMed]
- Antonín, V.; Tomšovský, M.; Sedlák, P.; Májek, T.; Jankovský, L. Morphological and molecular characterization of the Armillaria cepistipes—A. gallica complex in the Czech Republic and Slovakia. Mycol. Prog. 2009, 8, 259–271. [Google Scholar] [CrossRef]
- Kim, M.S.; Klopfenstein, N.B.; Hanna, J.W.; McDonald, G.I. Characterization of North American Armillaria species: Genetic relationships determined by ribosomal DNA sequences and AFLP markers. For. Pathol. 2006, 36, 145–164. [Google Scholar] [CrossRef]
- Tsykun, T.; Rigling, D.; Prospero, S. A new multilocus approach for a reliable DNA-based identification of Armillaria species. Mycologia 2013, 105, 1059–1076. [Google Scholar] [CrossRef] [PubMed]
- Elías-Román, R.D.; Guzmán-Plazola, R.A.; Klopfenstein, N.B.; Alvarado-Rosales, D.; Calderón-Zavala, G.; Mora-Aguilera, J.A.; Kim, M.S.; García-Espinosa, R. Incidence and phylogenetic analyses of Armillaria spp. associated with root disease in peach orchards in the State of Mexico, Mexico. For. Pathol. 2013, 43, 390–401. [Google Scholar] [CrossRef]
- Keča, N.; Klopfenstein, N.B.; Kim, M.S.; Solheim, H.; Woodward, S. Initial characterization of an unidentified Armillaria isolate from Serbia using LSU-IGS1 and TEF-1-α genes. For. Pathol. 2015, 45, 120–126. [Google Scholar] [CrossRef]
- Motta, J.J.; Korhonen, K. A note on Armilaria mellea and Armillaria bulbosa from the Middle Atlantic States. Mycologia 1986, 78, 471–474. [Google Scholar] [CrossRef]
- Elias-Roman, R.D.; Guzman-Plazola, R.A.; Alvarado-Rosales, D.; Calderon-Zavala, G.; Mora-Aguilera, J.A.; Garcia-Espinosa, R.; Kim, M.S.; Ross-Davis, A.L.; Hanna, J.W.; Klopfenstein, N.B. Armillaria root disease in peach orchards of the state of Mexico, Mexico: Characterization of Armillaria species and assessment of disease impact. Phytopathology 2013, 103, 39. [Google Scholar]
- Baumgartner, K.; Travadon, R.; Bruhn, J.; Bergemann, S.E. Contrasting patterns of genetic diversity and population structure of Armillaria mellea sensu stricto in the eastern and western United States. Phytopathology 2010, 100, 708–718. [Google Scholar] [CrossRef] [PubMed]
- Asef, M.R.; Mohammadi Goltapeh, E.; Alizadeh, A. Identification of Armillaria biological species in Iran. Fungal Divers. 2003, 14, 51–60. [Google Scholar]
- Ota, Y.; Matsushita, N.; Nagasawa, E.; Terashita, T.; Fukuda, K.; Suzuki, K. Biological species of Armillaria in Japan. Plant Dis. 1998, 82, 537–543. [Google Scholar] [CrossRef]
- Coetzee, M.P.A.; Musasira, N.Y.; Roux, J.; Roets, F.; van der Merwe, N.A.; Wingfield, M.J. Armillaria root rot spreading into a natural woody ecosystem in South Africa. Plant Pathol. 2018, 67, 883–891. [Google Scholar] [CrossRef]
- Guillaumin, J.-J.; Mohammed, C.; Abomo-Ndongo, S. Vegetative incompatibility and sexual systems of Armillaria isolates from tropical Africa. In Proceedings of the Eighth International Conference on Root and Butt Rots, Wik, Sweden; Haikko, Finland, 9–16 August 1993; Swedish University of Agricultural Science: Uppsala, Sweden, 1994; pp. 349–354. [Google Scholar]
- Mohammed, C.; Guillaumin, J.-J.; Botton, B.; Intini, M. Species of Armillaria in tropical Africa. In Proceedings of the Eight International Conference on Root and Butt Rots, Wik, Sweden; Haikko, Finland, 9–16 August 1993; Swedish University of Agricultural Science: Uppsala, Sweden, 1994; pp. 402–410. [Google Scholar]
- Abomo-Ndongo, S.; Mohammed, C.; Guillaumin, J.-J. Sexual behaviour of Armillaria heimii and A. mellea isolates from Africa. Eur. J. For. Pathol. 1997, 27, 207–224. [Google Scholar] [CrossRef]
- Coetzee, M.P.A.; Wingfield, B.D.; Harrington, T.C.; Steimel, J.; Coutinho, T.A.; Wingfield, M.J. The root rot fungus Armillaria mellea introduced into South Africa by early Dutch settlers. Mol. Ecol. 2001, 10, 387–396. [Google Scholar] [CrossRef] [PubMed]
- Coetzee, M.P.A.; Wingfield, B.D.; Roux, J.; Crous, P.W.; Denman, S.; Wingfield, M.J. Discovery of two northern hemisphere Armillaria species on Proteaceae in South Africa. Plant Pathol. 2003, 52, 604–612. [Google Scholar] [CrossRef]
- Wingfield, M.J.; Coetzee, M.P.A.; Crous, P.W.; Six, D.; Wingfield, B.D. Fungal phoenix rising from the ashes? IMA Fungus 2011, 1, 149–153. [Google Scholar] [CrossRef]
- Coetzee, M.P.A.; Wingfield, B.D.; Harrington, T.C.; Dalevi, D.; Coutinho, T.A.; Wingfield, M.J. Geographical diversity of Armillaria mellea s. s. based on phylogenetic analysis. Mycologia 2000, 92, 105–113. [Google Scholar] [CrossRef]
- Ota, Y.; Fukuda, K.; Suzuki, K. The nonheterothallic life cycle of Japanese Armillaria mellea. Mycologia 1998, 90, 396–405. [Google Scholar] [CrossRef]
- Ota, Y.; Intini, M.; Hattori, T. Genetic characterization of heterothallic and non-heterothallic Armillaria mellea sensu stricto. Mycol. Res. 2000, 104, 1046–1054. [Google Scholar] [CrossRef]
- Cha, J.Y.; Igarashi, T. A note on Armillaria mellea subsp. nipponica subsp. nov. in Japan. Mycoscience 1995, 36, 143–146. [Google Scholar] [CrossRef]
- Elías-Román, R.D.; Medel-Ortiz, R.; Alvarado-Rosales, D.; Hanna, J.W.; Ross-Davis, A.L.; Kim, M.-S.; Klopfenstein, N.B. Armillaria mexicana, a newly described species from Mexico. Mycologia 2018, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Stasińska, M. Armillaria ectypa, a rare fungus of mire in Poland. Acta Mycol. 2015, 50, 1–6. [Google Scholar] [CrossRef]
- Sesli, E.; Denchev, C.M. Checklists of the myxomycetes, larger ascomycetes, and larger basidiomycetes in Turkey. Mycotaxon 2008, 106, 65–67. [Google Scholar]
- Zhang, M. The Fungi in the Region of Hengduan Mountains; Science Press: Beijing, China, 1996. [Google Scholar]
- Kudo, S.; Nagasawa, E. Armillaria ectypa rediscovered in Aomori Prefecture, northern Japan. Rep. Tottori Mycol. Inst. 2003, 41, 26–34, (In Japanese with English Abstract). [Google Scholar]
- Ito, S. Occurrence of Armillaria ectypa in Aomori prefecture. Newsl. Mycol. Soc. Jpn. 2004, 4, 4–5. (In Japanese) [Google Scholar]
- Bruhn, J.N.; Wetteroff, J.J., Jr.; Mihail, J.D.; Kabrick, J.M.; Pickens, J.B. Distribution of Armillaria species in upland Ozark Mountain forests with respect to site, overstory species composition and oak decline. For. Pathol. 2000, 30, 43–60. [Google Scholar] [CrossRef]
- Schnabel, G.; Ash, J.S.; Bryson, P.K. Identification and characterization of Armillaria tabescens from the southeastern United States. Mycol. Res. 2005, 109, 1208–1222. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.S.; Klopfenstein, N.B.; Hanna, J.W.; Cannon, P.; Medel, R.; López, A. First report of armillaria root disease caused by Armillaria tabescens on Araucaria araucana in Veracruz, Mexico. Plant Dis. 2010, 94, 784. [Google Scholar] [CrossRef]
- Cha, J.Y.; Lee, S.Y.; Chun, K.W.; Lee, S.Y.; Ohga, S. Armillaria root rot caused by Armillaria tabescens on Prunus salicina in a Korean garden. J. Fac. Agric. Kyushu Univ. 2009, 54, 273–277. [Google Scholar]
- Lee, S.K.; Seo, S.T. First report of Armillaria root disease caused by Armillaria tabescens on Carpinus tschonoskii in South Korea. Plant Dis. 2016, 100, 213. [Google Scholar] [CrossRef]
- Dai, Y.C.; Cui, B.K.; Yuan, H.S.; Li, B.D. Pathogenic wood-decaying fungi in China. For. Pathol. 2007, 37, 105–120. [Google Scholar] [CrossRef]
- Ota, Y.; Kim, M.-S.; Neda, H.; Klopfenstein, N.B.; Hasegawa, E. The phylogenetic position of an Armillaria species from Amami-Oshima, a subtropical island of Japan, based on elongation factor and ITS sequences. Mycoscience 2011, 52, 53–58. [Google Scholar] [CrossRef]
- Darmono, T.W.; Burdsall, H.H.; Volk, T.J. Interfertility among isolates of Armillaria tabescens in North America. Sydowia 1992, 42, 105–116. [Google Scholar]
- Harrington, T.C.; Wingfield, B.D. A PCR-based identification method for species of Armillaria. Mycologia 1995, 87, 280–288. [Google Scholar] [CrossRef]
- Kile, G.A.; Watling, R. Armillaria species from south-eastern Australia. Trans. Br. Mycol. Soc. 1983, 81, 129–140. [Google Scholar] [CrossRef]
- Ramsfield, T.D.; Power, M.W.P.; Ridley, O.S. A comparison of populations of Armillaria hinnulea in New Zealand and Australia. N. Z. Plant Prot. 2008, 61, 41–47. [Google Scholar]
- Hood, I.A.; Ramsfield, T.D. Armillaria aotearoa species nova. N. Z. J. For. Sci. 2016, 46, 1–11. [Google Scholar] [CrossRef]
- Pildain, M.B.; Coetzee, M.P.A.; Wingfield, B.D.; Wingfield, M.J.; Rajchenberg, M. Taxonomy of Armillaria in the Patagonian forests of Argentina. Mycologia 2010, 102, 392–403. [Google Scholar] [CrossRef] [PubMed]
- Guillaumin, J.-J.; Mohammed, C.; Berthelay, S. Armillaria novae-zelandiae in Papua New Guinea. Mycol. Res. 1992, 96, 278–280. [Google Scholar] [CrossRef]
- Stevenson, G. The Agaricales of New Zealand V. Tricholomataceae. Kew Bull. 1964, 19, 1–59. [Google Scholar] [CrossRef]
- Kile, G.A.; Watling, R. Identification and occurrence of Australian Armillaria species, including A. pallidula sp. nov. and comparative studies between them and non-Australian tropical and Indian Armillaria. Trans. Br. Mycol. Soc. 1988, 91, 305–315. [Google Scholar] [CrossRef]
- Singer, R. Mycoflora Australis. Beihefte zur Nova Hedwigia 1969, 29, 40–49. [Google Scholar]
- Horak, E. Flora criptogámica de Tierra del Fuego. Fungi: Basidiomycetes Agaricales y Gasteromycetes Secotioides; CONICET-FECIC: Buenos Aires, Argentina, 1979.
- Boesewinkel, H.J. New plant disease records in New Zealand: Records in the period 1969–1976. N. Z. J. Agric. Res. 1977, 20, 583–589. [Google Scholar] [CrossRef]
- Lima, M.L.A.; Asai, T.; Capelari, M. Armillaria paulensis: A new South American species. Mycol. Res. 2008, 112, 1122–1128. [Google Scholar] [CrossRef] [PubMed]
- Singer, R. The Armillariella mellea group. Lloydia 1956, 19, 176–187. [Google Scholar]
- Podger, F.D.; Kile, G.A.; Watling, R.; Fryer, J. Spread and effects of Armillaria luteobubalina sp. nov. in an Australian Eucaluptus regnans plantation. Trans. Br. Mycol. Soc. 1978, 71, 77–87. [Google Scholar] [CrossRef]
- Pérez-Sierra, A.; Guillaumin, J.-J.; Spooner, B.M.; Bridge, P.D. Characterization of Armillaria heimii from Africa. Plant Pathol. 2004, 53, 220–230. [Google Scholar] [CrossRef]
- Abomo-Ndongo, S.; Tourvieille, J.; Guillaumin, J.J. The Buller phenomenon in Armillaria heimii Pegler, a bipolar diploid basidiomycete. Cryptogam. Mycol. 2002, 23, 335–347. [Google Scholar]
- Guillaumin, J.-J.; Legrand, P. Armillaria root rots. In Infectious Forest Diseases; Gonthier, P., Nicolotti, G., Eds.; CABI Publishing: Oxfordshire, UK, 2013; pp. 159–177. [Google Scholar]
- Gezahgne, A.; Coetzee, M.P.A.; Wingfield, B.D.; Wingfield, M.J.; Roux, J. Identification of the Armillaria root rot pathogen in Ethiopian plantations. For. Pathol. 2004, 34, 133–145. [Google Scholar] [CrossRef]
- Mwenje, E.; Wingfield, B.D.; Coetzee, M.P.; Wingfield, M.J. Molecular characterisation of Armillaria species from Zimbabwe. Mycol. Res. 2003, 107, 291–296. [Google Scholar] [CrossRef] [PubMed]
- Wingfield, B.D.; Maphosa, L.; Coetzee, M.P.A.; Mwenje, E.; Wingfield, M.J. Characterisation of Zimbabwean Armillaria using IGS-1 sequences and AFLP analysis. Fungal Divers. 2009, 34, 187–196. [Google Scholar]
- Mwenje, E.; Wingfield, B.D.; Coetzee, M.P.A.; Nemato, H.; Wingfield, M.J. Armillaria species on tea in Kenya identified using isozyme and DNA sequence comparisons. Plant Pathol. 2006, 55, 343–350. [Google Scholar] [CrossRef]
- Mwangi, L.M.; Lin, D.; Hubbes, M. Identification of Kenyan Armillaria isolates by cultural morphology intersterility tests and analysis of isozyme profiles. Eur. J. For. Pathol. 1989, 19, 399–406. [Google Scholar] [CrossRef]
- Agustian, A.; Mohammed, C.; Guillaumin, J.-J.; Botton, B. Discrimination of some African Armillaria species by isozyme electrophoretic analysis. New Phytol. 1994, 128, 135–143. [Google Scholar] [CrossRef]
- Otieno, W.; Pérez Sierra, A.; Termorshuizen, A. Characterization of Armillaria isolates from tea (Camellia sinensis) in Kenya. Mycologia 2003, 95, 160–175. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.W.; Jacobson, D.J.; Kroken, S.; Kasuga, T.; Geiser, D.M.; Hibbett, D.S.; Fisher, M.C. Phylogenetic species recognition and species concepts in fungi. Fungal Genet. Biol. 2000, 31, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Dettman, J.R.; Jacobson, D.J.; Taylor, J.W. A multilocus genealogical approach to phylogenetic species recognition in the model eukaryote Neurospora. Evolution 2003, 57, 2703–2720. [Google Scholar] [CrossRef] [PubMed]
- Rokas, A.; Williams, B.L.; King, N.; Carroll, S.B. Genome-scale approaches to resolving incongruence in molecular phylogenies. Nature 2003, 425, 798–804. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Luo, J.; Bhattacharya, D. Chapter Eight—Advances in fungal phylogenomics and their impact on fungal systematics. In Advances in Genetics; Townsend, J.P., Wang, Z., Eds.; Academic Press: Cambridge, MA, USA, 2017; Volume 100, pp. 309–328. [Google Scholar]
- Wetterstrand, K. DNA Sequencing Costs: Data from the NHGRI Genome Sequencing Program (GSP). Available online: http//:www.genome.gov/sequencingcostsdata (accessed on 21 October 2018).
- Lemmon, E.M.; Lemmon, A.R. High-throughput genomic data in systematics and phylogenetics. Annu. Rev. Ecol. Evol. Syst. 2013, 44, 99–121. [Google Scholar] [CrossRef]
- Sipos, G.; Prasanna, A.N.; Walter, M.C.; O’Connor, E.; Bálint, B.; Krizsán, K.; Kiss, B.; Hess, J.; Varga, T.; Slot, J.; et al. Genome expansion and lineage-specific genetic innovations in the forest pathogenic fungi Armillaria. Nat. Ecol. Evol. 2017, 1, 1931–1941. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).