Comparison of Potato and Asian Citrus Psyllid Adult and Nymph Transcriptomes Identified Vector Transcripts with Potential Involvement in Circulative, Propagative Liberibacter Transmission
Abstract
:1. Introduction
2. Results and Discussion
2.1. Assembly and Annotation of Potato Psyllid Transcriptomes
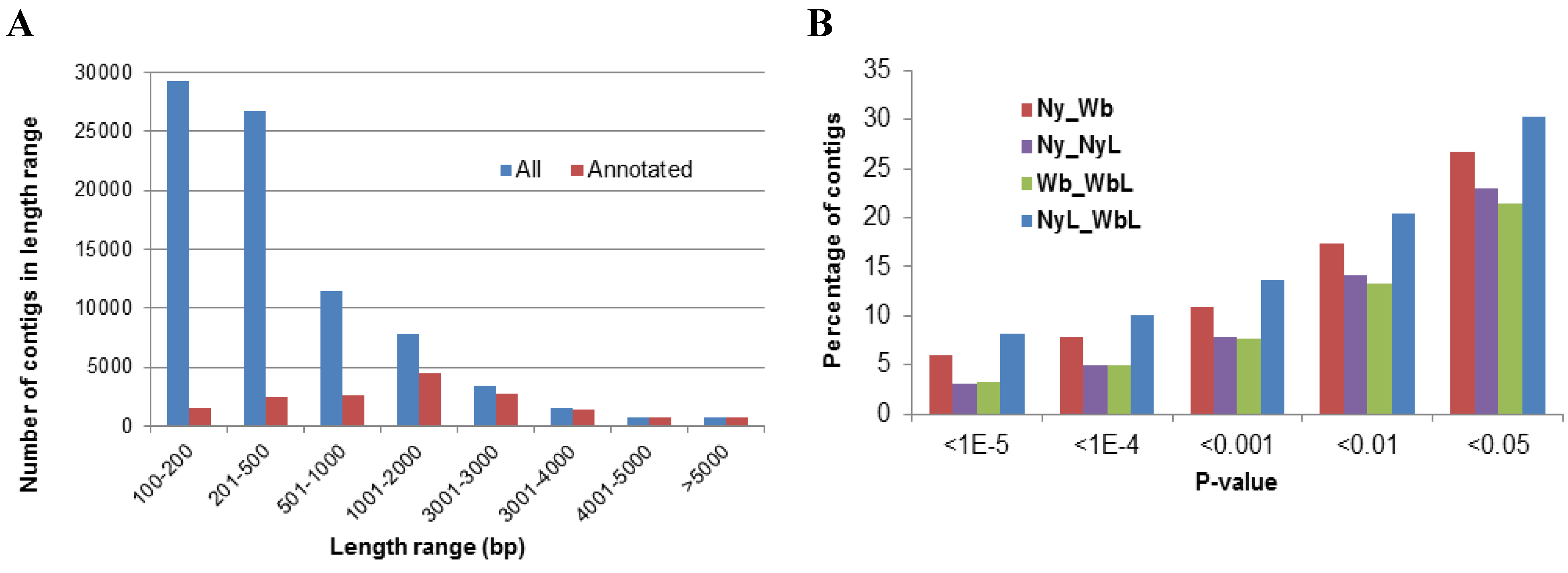
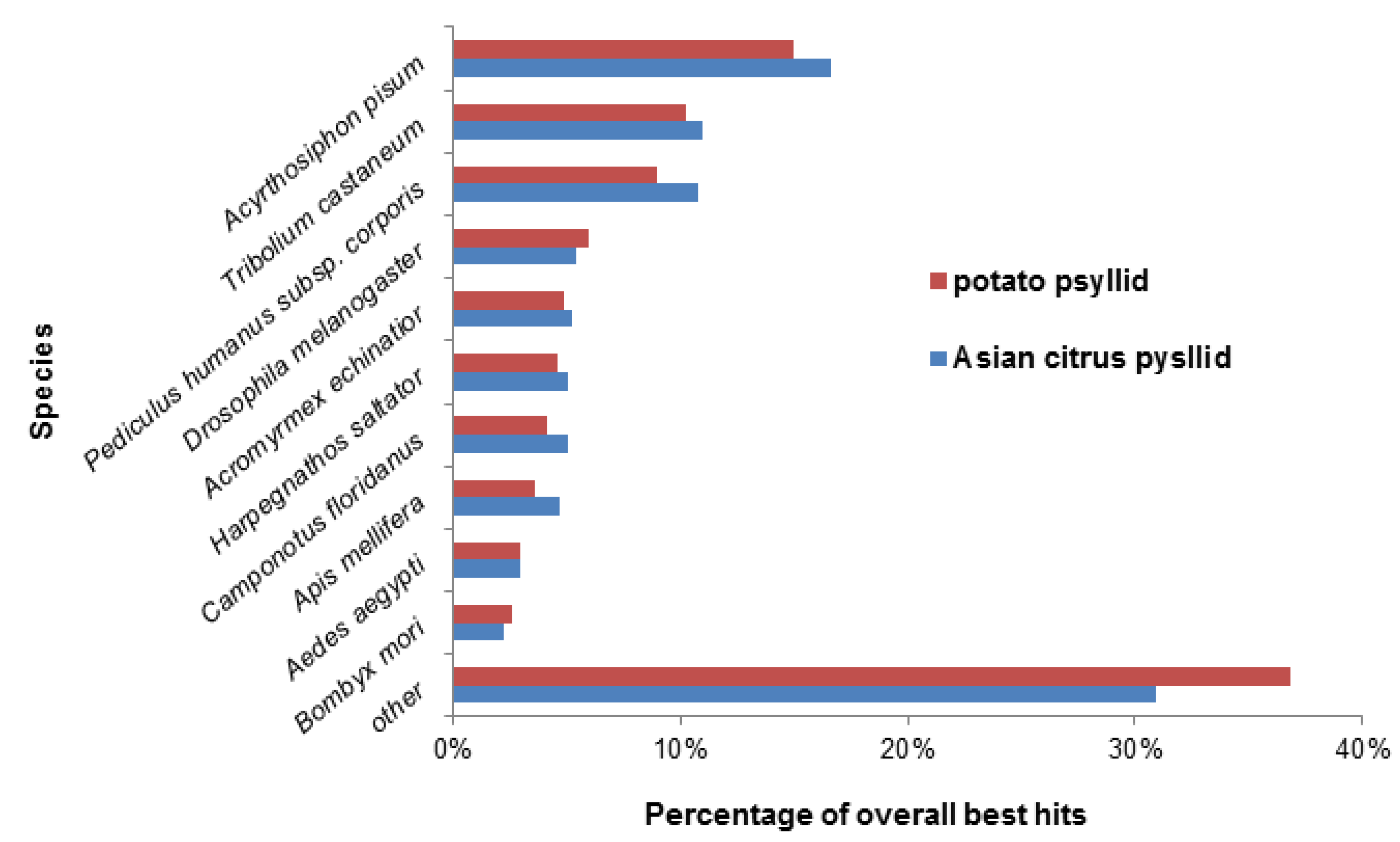
2.2. Effects of Potato Psyllid Adult and Nymph Life Stages, and Liberibacter Infection on Gene Expression
| Expression Response | Biological Process | Cellular Process | Molecular Function | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PoP | ACP | PoP | ACP | PoP | ACP | |||||||
| Adults | Nymphs | Adults | Nymphs | Adults | Nymphs | Adults | Nymphs | Adults | Nymphs | Adults | Nymphs | |
| Up-regulated | 7% | 4% | 14% | 28% | 7% | 3% | 14% | 29% | 9% | 4% | 14% | 28% |
| Down-regulated | 7% | 14% | 14% | 11% | 6% | 14% | 13% | 11% | 8% | 17% | 14% | 11% |
| Unchanged | 86% | 82% | 72% | 61% | 87% | 83% | 73% | 60% | 83% | 79% | 72% | 61% |
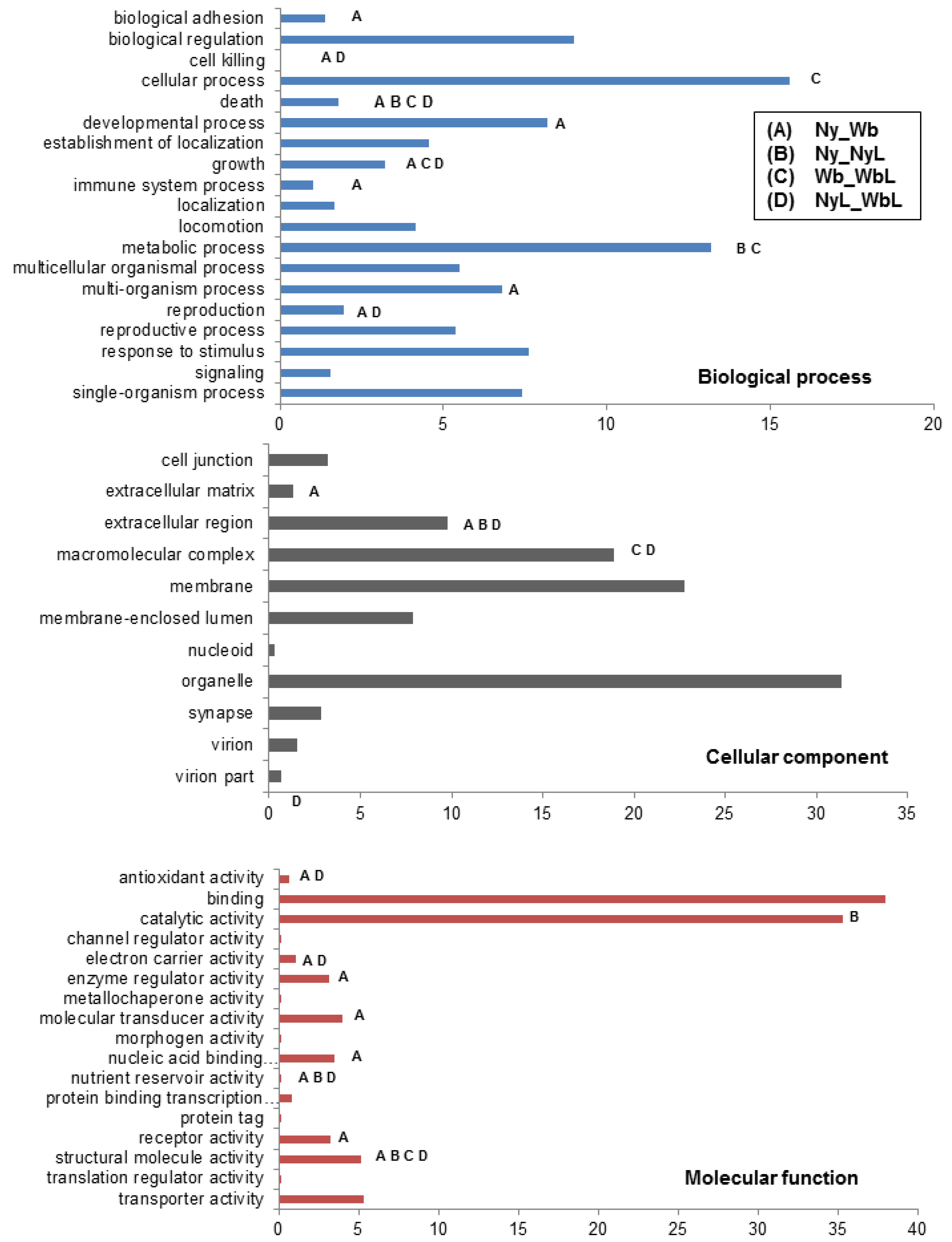
2.2.1. Adhesion-Related Transcripts
| Nymph | Fold Change * | Adult | Fold Change * |
|---|---|---|---|
| Putative salivary protein | 657.71 | Cuticular protein 100A | 49.29 |
| Abdominal-B | 91.35 | Cuticular protein PpolCPR68 | 46.07 |
| Cuticle protein 7 | 77.92 | Putative ribosomal protein S13e | −23.42 |
| Heat shock protein 70 A1 | −120.68 | Aconitate hydratase, mitochondrial | −120.89 |
| Heat shock 70 kDa protein IV | −337.04 | Cytochrome b561 | 17.63 |
| Ubiquitin | −256.48 | Senescence-associated protein, putative | 19.59 |
| 60S ribosomal protein L5 | −175.71 | ATP synthase subunit alpha, mitochondrial | −109.49 |
| Heat shock protein 70 | −67.87 | Arginine kinase | 30.16 |
| Putative ribosomal protein S13e | −69.98 | 60S ribosomal protein L5 | −20.22 |
| Probable RNA-directed DNA polymerase from transposon BS | −61.87 | Endonuclease-reverse transcriptase | −96.94 |
| Putative reverse transcriptase | −47.65 | Ubiquitin | −25.09 |
| FixR | 26.65 | Cathepsin D | −20.77 |
| Translationally-controlled tumor protein homolog | −92.8 | Reverse transcriptase-like protein | −27.8 |
| Cathepsin B preproprotein-like protein | 24.68 | Meiotic recombination protein SPO11, putative | −21.61 |
| Pyrroline-5-carboxylate dehydrogenase | −148.22 | Protein painting of fourth | 12.88 |
| Vitellogenin | −40.34 | Elongation factor 2 | −23.81 |
| Reverse transcriptase-like protein | −83.13 | Phosphate carrier protein, mitochondrial | 17.1 |
| Putative small heat shock protein | −23.9 | Retrovirus-related Pol polyprotein from transposon opus | 11.87 |
| SARTTc1 ORF2 protein | −29.89 | Probable RNA-directed DNA polymerase from transposon BS | 10.07 |
| Tubulin alpha chain | −121.15 | Cuticular protein 97Ea | 9.96 |
| Meiotic recombination protein SPO11, putative | −42.53 | Enolase | −17.11 |
| ATP synthase subunit alpha, mitochondrial | −25.5 | F-element protein | −19.39 |
| 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase epsilon-1 | 11.15 | Putative small heat shock protein | −15.97 |
| Peritrophin-1, putative | −39.63 | ATP-dependent RNA helicase p62 | −14.72 |
| Insecticide resistance protein CYP4G70 | −96.66 | Cathepsin B preproprotein-like protein | −11.99 |
2.2.2. Immune System-Related Transcripts
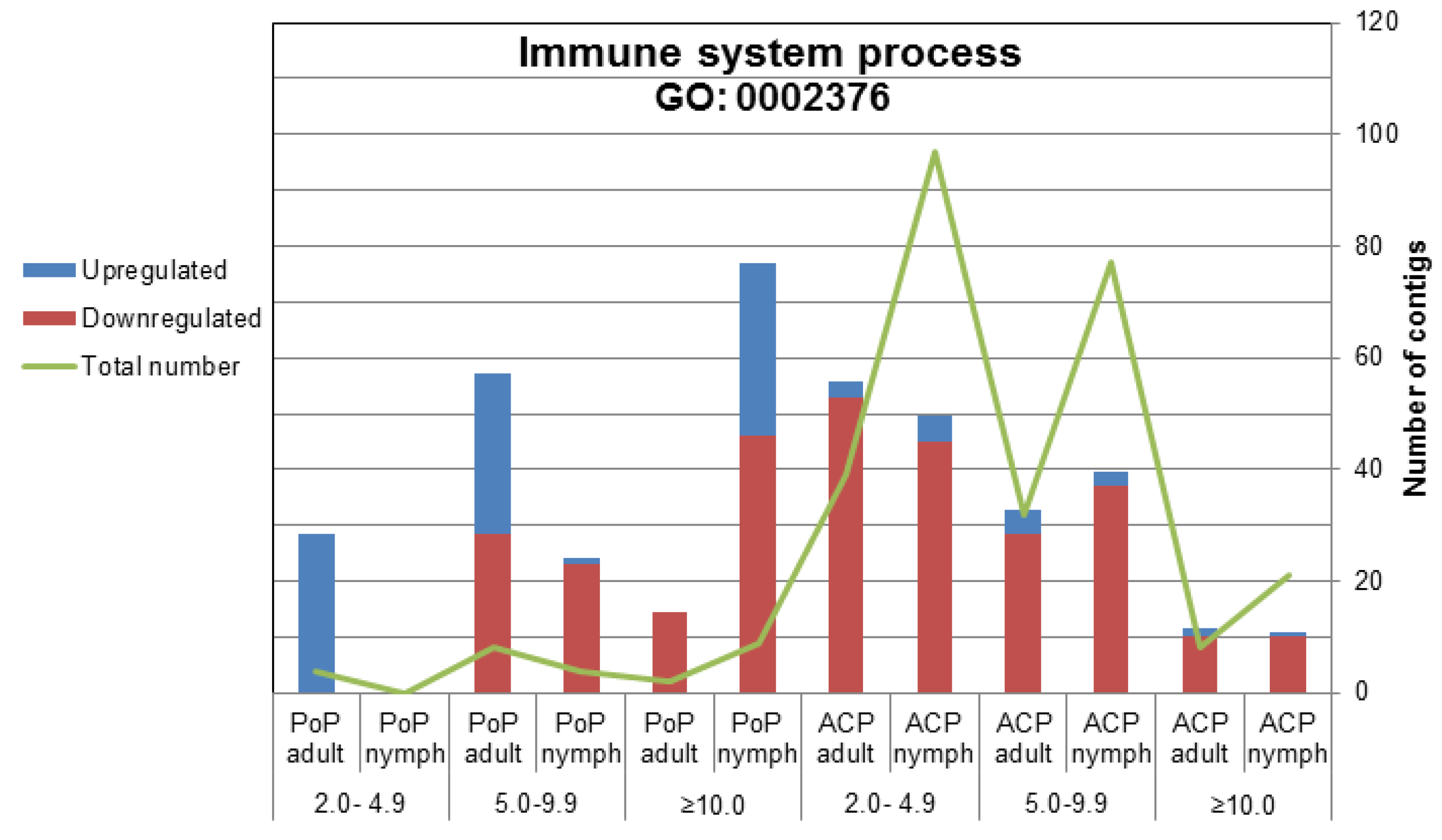
2.2.2.1. IMD Pathway
2.2.2.2. Toll Pathway
2.2.2.3. Phagocytosis
2.2.3. Metabolic-Related Transcripts
Iron Metabolism
2.3. Orthologs in the Potato and Asian Citrus Psyllid Transcriptomes
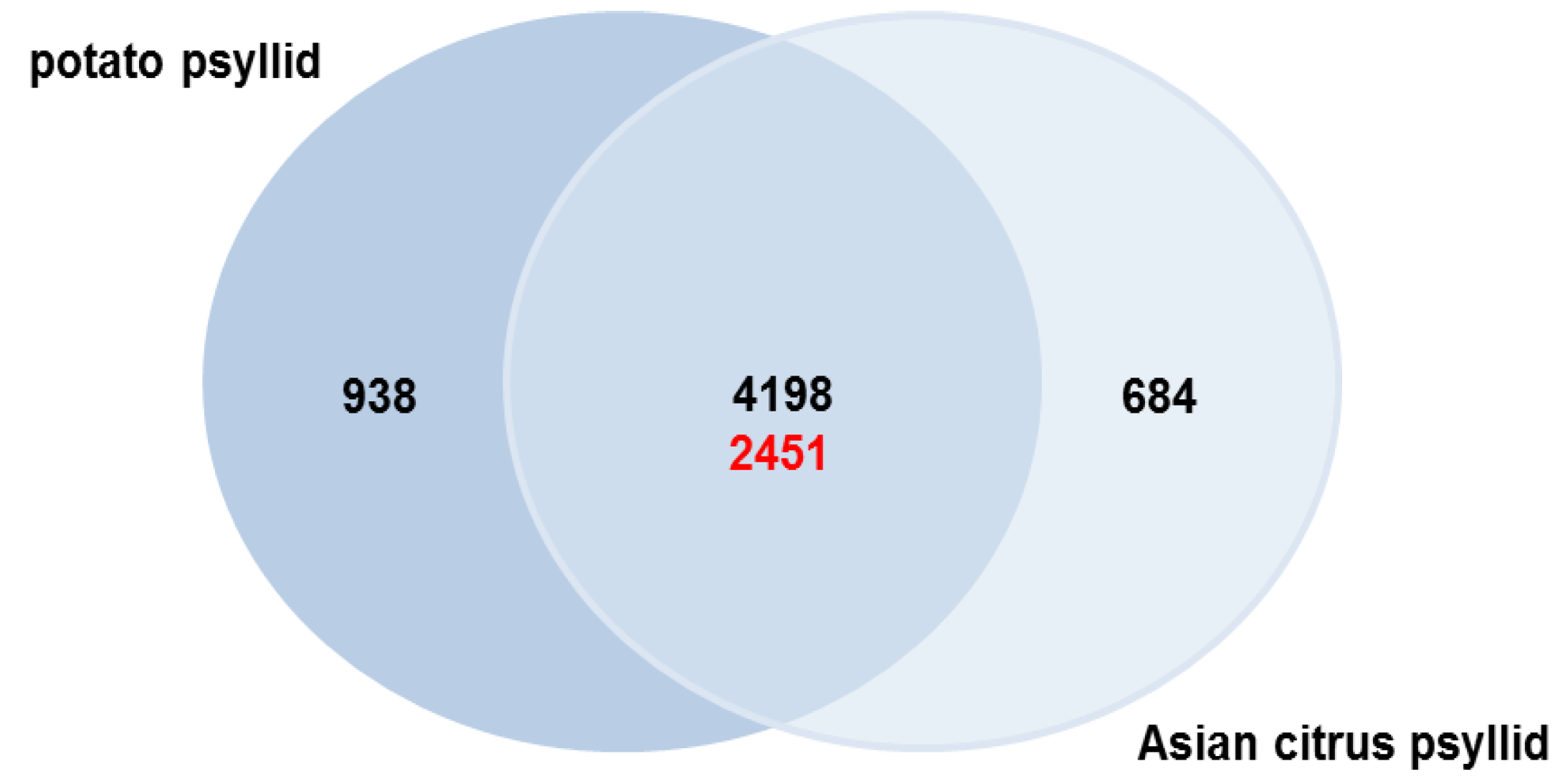
2.3.1. Potato Psyllid Paralogous Gene Families
2.3.2. Asian Citrus Psyllid Paralogous Gene Families
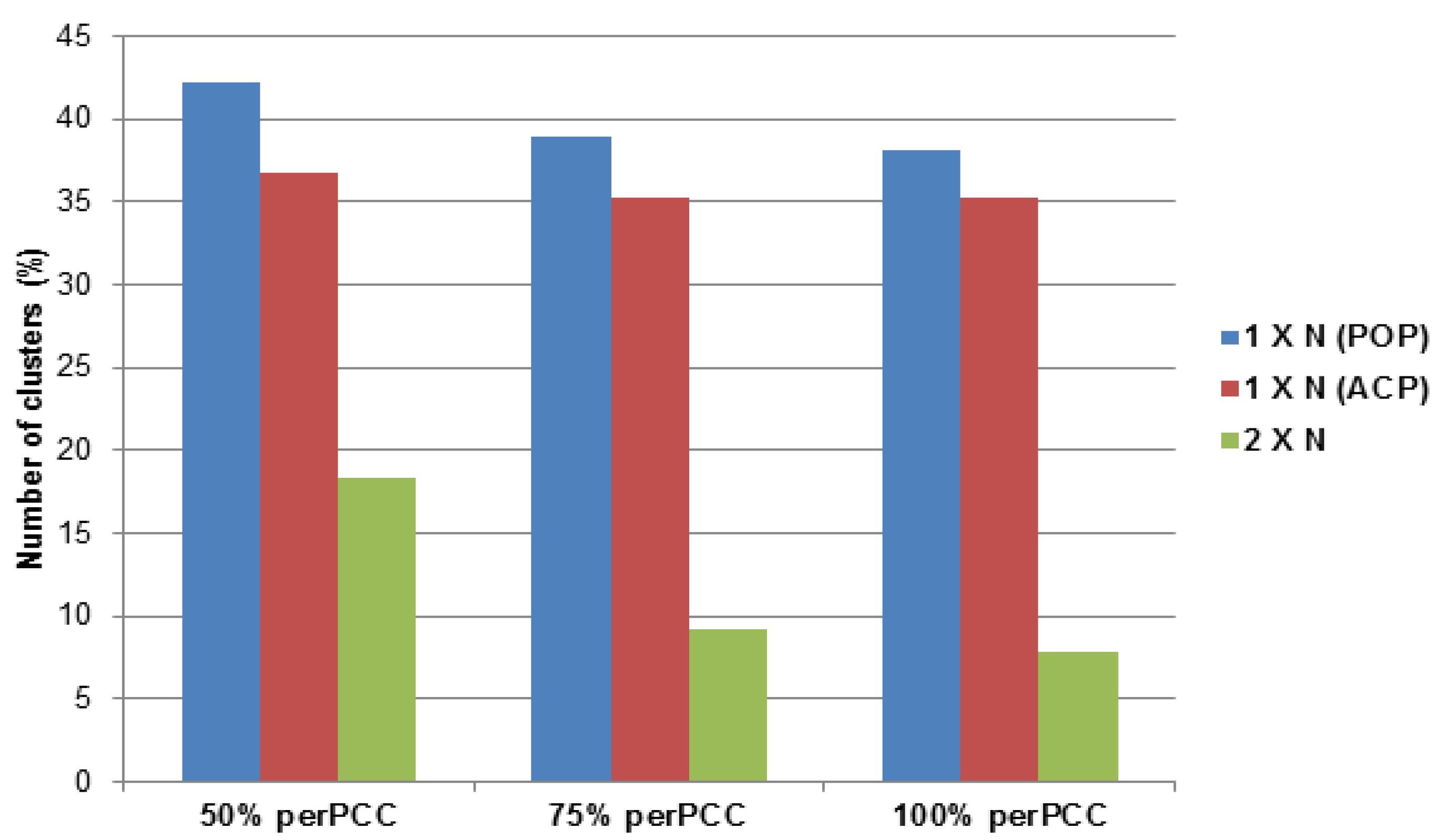
2.3.3. Asian Citrus and Potato Psyllid Orthogolous Gene Families
| Cluster ID | Size | Description | perPCC |
|---|---|---|---|
| TR_000786 | 3 | 3-hydroxy-3-methylglutaryl-coenzyme A reductase | 100 |
| TR_003785 | 3 | Abc transporter | 100 |
| TR_000835 | 3 | Atlastin | 100 |
| TR_001975 | 3 | Calmodulin | 100 |
| TR_001355 | 3 | cAMP-dependent protein kinase catalytic subunit, putative | 100 |
| TR_000708 | 4 | Chitin synthase 1 variant b | 100 |
| TR_000290 | 3 | Cleavage and polyadenylation specificity factor subunit 1 | 100 |
| TR_001473 | 3 | Cytochrome P450, putative | 100 |
| TR_000343 | 4 | DNA replication licensing factor MCM4 | 100 |
| TR_000489 | 3 | DNA-binding protein Ewg, putative | 100 |
| TR_001764 | 3 | Dual oxidase maturation factor 1 | 100 |
| TR_000565 | 3 | Dynamin-like 120 kDa protein, mitochondrial | 100 |
| TR_000908 | 3 | E78 nuclear receptor | 100 |
| TR_000324 | 3 | EH domain-binding protein 1 | 100 |
| TR_000905 | 3 | F-box only protein, putative | 100 |
| TR_000639 | 3 | Forkhead box protein K1 | 100 |
| TR_000486 | 3 | Hedgehog protein | 100 |
| TR_000141 | 3 | Inactive ubiquitin carboxyl-terminal hydrolase 54 | 100 |
| TR_000345 | 3 | Kinesin-like protein KIF21B | 100 |
| TR_000433 | 3 | Merlin | 100 |
| TR_001523 | 3 | Mitogen-activated protein kinase ERK-A, putative | 100 |
| TR_000402 | 3 | Phosphatidylserine synthase 1 | 100 |
| TR_002280 | 4 | Probable aconitate hydratase, mitochondrial | 100 |
| TR_000043 | 3 | Probable phosphorylase b kinase regulatory subunit alpha | 100 |
| TR_000068 | 4 | Probable pre-mRNA-splicing factor ATP-dependent RNA helicase mog-5 | 100 |
| TR_000493 | 3 | Protein turtle | 100 |
| TR_000691 | 3 | Protein Wnt | 100 |
| TR_000597 | 3 | Proton-coupled amino acid transporter 4 | 100 |
| TR_001033 | 3 | Putative AMP dependent CoA ligase | 100 |
| TR_000510 | 3 | Pyruvate dehydrogenase E1 component subunit beta, mitochondrial | 100 |
| TR_000027 | 3 | SH3 and multiple ankyrin repeat domains protein 3 | 100 |
| TR_001667 | 3 | Shc transforming protein, putative | 100 |
| TR_001186 | 3 | Techylectin-5B | 100 |
| TR_000532 | 3 | Transient receptor potential cation channel trpm | 100 |
| TR_000454 | 3 | Triacylglycerol lipase, pancreatic, putative | 100 |
| TR_000971 | 4 | Actin-binding protein IPP | 83.33 |
| TR_001171 | 4 | Putative GPCR-type octopamine beta receptor (Fragment) | 83.33 |
| TR_000754 | 4 | Sodium/hydrogen exchanger | 83.33 |
| TR_000042 | 4 | Tyrosine-protein kinase transforming protein FPS, putative | 83.33 |
| TR_000375 | 5 | Ephrin receptor | 80 |
3. Methods
3.1. Psyllids
3.2. RNA Isolation and Quality Control
3.3. Library Construction and Illumina Paired-End Sequencing
3.4. Assembly, Annotation and Comparison of Illumina Sequences
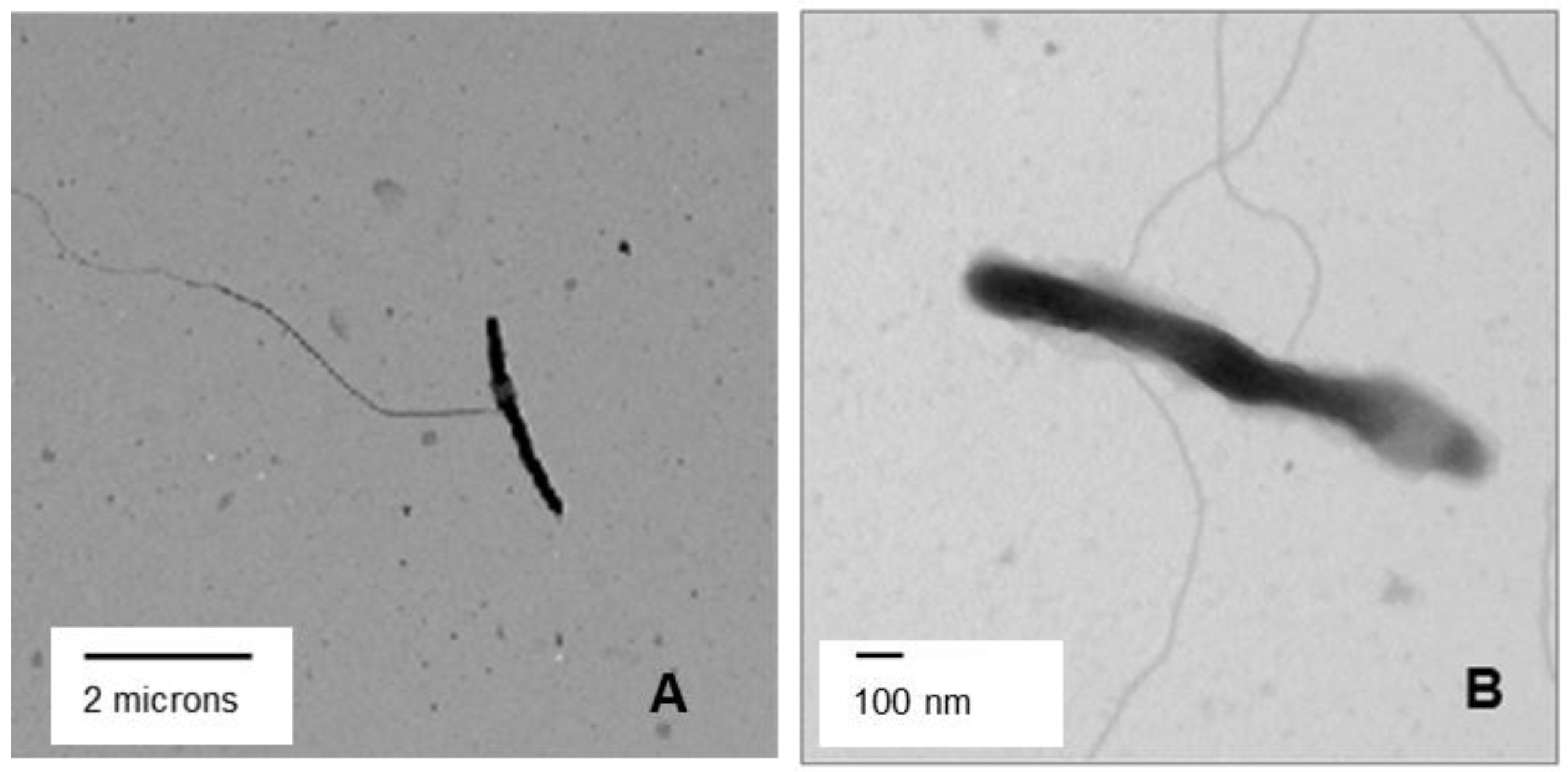
3.5. Transmission Electron Microscopy
4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Abbreviations
| ACP | Asian citrus psyllid |
| CLas | Candidatus Liberibacter asiaticus |
| CLso | Candidatus Liberibacter solanacearum |
| GO | Gene Ontology |
| HLB | Huanglongbing |
| PCC | Pearson Correlation Coefficient |
| perPCC | average PCC per cluster |
| PoP | potato psyllid |
| RPKM | Reads per KB per million reads |
| ZC | Zebra chip disease |
Conflicts of Interests
References and Notes
- Munyaneza, J.E. Psyllids as vectors of emerging bacterial diseases of annual crops. Southwest. Entomol. 2010, 35, 471–477. [Google Scholar] [CrossRef]
- Brown, J.K.; Rehman, M.; Rogan, D.; Martin, R.R.; Idris, A.M. First Report of “Candidatus Liberibacter psyllaurous” (synonym “Ca. L. solanacearum”) Associated with “Tomato Vein-Greening” and “Tomato Psyllid Yellows” Diseases in Commercial Greenhouses in Arizona. Plant Dis. 2010, 94, 376–376. [Google Scholar]
- Liefting, L.W.; Sutherland, P.W.; Ward, L.I.; Paice, K.L.; Weir, B.S.; Clover, G.R. G. A New “Candidatus Liberibacter” Species Associated with Diseases of Solanaceous Crops. Plant Dis. 2009, 93, 208–214. [Google Scholar] [CrossRef]
- Hansen, A.K.; Trumble, J.T.; Stouthamer, R.; Paine, T.D. A new Huanglongbing Species, “Candidatus Liberibacter psyllaurous”, found to infect tomato and potato, is vectored by the psyllid Bactericera cockerelli (Sulc). Appl. Environ. Microbiol. 2008, 74, 5862–5865. [Google Scholar] [CrossRef] [PubMed]
- Crosslin, J.M.; Munyaneza, J.E.; Brown, J.K.; Liefting, L.W. A history in the making: Potato zebra chip disease associated with a new psyllid-borne bacterium—A tale of striped potatoes. APSnet Feature 2010. [Google Scholar] [CrossRef]
- Sengoda, V.G.; Munyaneza, J.E.; Crosslin, J.M.; Buchman, J.L.; Pappu, H.R. Phenotypic and etiological differences between psyllid yellows and zebra chip diseases of potato. Am. J. Potato Res. 2010, 87, 41–49. [Google Scholar] [CrossRef]
- Gottwald, T.R. Current Epidemiological Understanding of Citrus Huanglongbing. Annu. Rev. Phytopathol. 2010, 48, 119–139. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Zhou, L.; Hall, D.G.; Li, W.; Doddapaneni, H.; Lin, H.; Liu, L.; Vahling, C.M.; Gabriel, D.W.; Williams, K.P.; Dickerman, A.; Sun, Y.; Gottwald, T. Complete genome sequence of citrus huanglongbing bacterium, “Candidatus Liberibacter asiaticus” obtained through metagenomics. Mol. Plant Microbe Interact. 2009, 22, 1011–1020. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.; Gudmestad, N.C. Aspects of Pathogen Genomics, Diversity, Epidemiology, Vector Dynamics, and Disease Management for a Newly Emerged Disease of Potato: Zebra Chip. Phytopathology 2013, 103, 524–537. [Google Scholar] [CrossRef] [PubMed]
- Jagoueix, S.; Bove, J.; Garnier, M. The phloem-limited bacterium of greening disease of citrus is a member of the α subdivision of the Proteobacteria. Int. J. Syst. Bacteriol. 1994, 44, 379–386. [Google Scholar] [CrossRef] [PubMed]
- Inoue, H.; Ohnishi, J.; Ito, T.; Tomimura, K.; Miyata, S.; Iwanami, T.; Ashihara, W. Enhanced proliferation and efficient transmission of Candidatus Liberibacter asiaticus by adult Diaphorina citri after acquisition feeding in the nymphal stage. Ann. Appl. Biol. 2009, 155, 29–36. [Google Scholar] [CrossRef]
- Ammar, E.; Shatters, R.G., Jr.; Hall, D.G. Localization of Candidatus Liberibacter asiaticus, Associated with Citrus Huanglongbing Disease, in its Psyllid Vector using Fluorescence in situ Hybridization. J. Phytopathol. 2011, 159, 726–734. [Google Scholar] [CrossRef]
- Ammar, E.; Shatters, R.G., Jr.; Lynch, C.; Hall, D.G. Detection and Relative Titer of Candidatus Liberibacter asiaticus in the Salivary Glands and Alimentary Canal of Diaphorina citri (Hemiptera: Psyllidae) Vector of Citrus Huanglongbing Disease. Ann. Entomol. Soc. Am. 2011, 104, 526–533. [Google Scholar] [CrossRef]
- Sengoda, V.G.; Buchman, J.L.; Henne, D.C.; Pappu, H.R.; Munyaneza, J.E. “Candidatus Liberibacter solanacearum” Titer Over Time in Bactericera cockerelli (Hemiptera: Triozidae) after Acquisition from Infected Potato and Tomato Plants. J. Econ. Entomol. 2013, 106, 1964–1972. [Google Scholar] [CrossRef] [PubMed]
- Cooper, W.; Sengoda, V.; Munyaneza, J. Localization of “Candidatus Liberibacter solanacearum” in Bactericera cockerelli (Hemiptera: Triozidae). Ann. Entomol. Soc. Am. 2014, 107, 204–210. [Google Scholar] [CrossRef]
- Pelz-Stelinski, K.S.; Brlansky, R.H.; Ebert, T.A.; Rogers, M.E. Transmission parameters for Candidatus liberibacter asiaticus by Asian citrus psyllid (Hemiptera: Psyllidae). J. Econ. Entomol. 2010, 103, 1531–1541. [Google Scholar] [CrossRef] [PubMed]
- Mann, R.S.; Pelz-Stelinski, K.; Hermann, S.L.; Tiwari, S.; Stelinski, L.L. Sexual transmission of a plant pathogenic bacterium, Candidatus Liberibacter asiaticus, between conspecific insect vectors during mating. PLoS One 2011, 6, e29197. [Google Scholar] [CrossRef]
- Cicero, J.M.; Fisher, T.W.; Stansly, P.A.; Brown, J.K. are preparing the manuscript with title “Anatomical evidence for circulation and propagation of Candidatus Liberibacter solanacearum in the potato psyllid vector, Bactericera cockerelli (Sulc) (Hemiptera: Triozidae)”. 2014. [Google Scholar]
- Nachappa, P.; Levy, J.; Pierson, E.; Tamborindeguy, C. Correlation between “Candidatus Liberibacter solanacearum” infection levels and fecundity in its psyllid vector. J. Invertebr. Pathol. 2013, 115, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Nachappa, P.; Shapiro, A.A.; Tamborindeguy, C. Effect of “Candidatus Liberibacter solanacearum” on Fitness of Its Insect Vector, Bactericera cockerelli (Hemiptera: Triozidae), on Tomato. Phytopathology 2012, 102, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Thao, M.L.; Clark, M.A.; Burckhardt, D.H.; Moran, N.A.; Baumann, P. Phylogenetic analysis of vertically transmitted psyllid endosymbionts (Candidatus Carsonella ruddii) based on atpAGD and rpoC: Comparisons with 16S–23S rDNA-derived phylogeny. Curr. Microbiol. 2001, 42, 419–421. [Google Scholar] [CrossRef] [PubMed]
- Nachappa, P.; Levy, J.; Pierson, E.; Tamborindeguy, C. Diversity of endosymbionts in the potato psyllid, Bactericera cockerelli (Triozidae), vector of zebra chip disease of potato. Curr. Microbiol. 2011, 62, 1510–1520. [Google Scholar] [CrossRef] [PubMed]
- Kageyama, D.; Narita, S.; Watanabe, M. Insect sex determination manipulated by their endosymbionts: Incidences, mechanisms and implications. Insects 2012, 3, 161–199. [Google Scholar] [CrossRef]
- Subandiyah, S.; Nikoh, N.; Tsuyumu, S.; Somowiyarjo, S.; Fukatsu, T. Complex endosymbiotic microbiota of the citrus psyllid Diaphorina citri (Homoptera: Psylloidea). Zool. Sci. 2000, 17, 983–989. [Google Scholar] [CrossRef]
- Lemaitre, B.; Hoffmann, J. The host defense of Drosophila melanogaster. Annu. Rev. Immunol. 2007, 25, 697–743. [Google Scholar] [CrossRef] [PubMed]
- International Aphid Genomics Consortium Genome sequence of the pea aphid Acyrthosiphon pisum. PLoS Biol. 2010, 8, e1000313.
- Baumann, P.; Moran, N.A.; Baumann, L.; Dworkin, M. Bacteriocyte-associated endosymbionts of insects. Prokaryotes 2006, 1, 403–438. [Google Scholar]
- Buchner, P. Endosymbiosis of Animals with Plant Microorganisms; Interscience Publishers: New York, NY, USA, 1965. [Google Scholar]
- Haine, E.R. Symbiont-mediated protection. Proc. R. Soc. B Biol. Sci. 2008, 275, 353–361. [Google Scholar] [CrossRef]
- Engelstädter, J.; Hurst, G.D. The ecology and evolution of microbes that manipulate host reproduction. Annu. Rev. Ecol. Evol. Syst. 2009, 40, 127–149. [Google Scholar] [CrossRef]
- Buchman, J.L.; Sengoda, V.G.; Munyaneza, J.E. Vector Transmission Efficiency of Liberibacter by Bactericera cockerelli (Hemiptera: Triozidae) in Zebra Chip Potato Disease: Effects of Psyllid Life Stage and Inoculation Access Period. J. Econ. Entomol. 2011, 104, 1486–1495. [Google Scholar] [CrossRef] [PubMed]
- Wallis, C.M.; Rashed, A.; Wallingford, A.K.; Rush, C.M. Relationship of potato biochemical responses to “Candidatus Liberibacter solanacearum”, causal agent of zebra chip, to disease progression. Phytopathology 2013, 103, 154–154. [Google Scholar]
- Casteel, C.L.; Hansen, A.K.; Walling, L.L.; Paine, T.D. Manipulation of plant defense responses by the Tomato Psyllid (Bactericerca cockerelli) and its associated endosymbiont Candidatus Liberibacter Psyllaurous. PLoS One 2012, 7, e35191. [Google Scholar] [CrossRef]
- Grafton-Cardwell, E.E.; Stelinski, L.L.; Stansly, P.A. Biology and management of Asian citrus psyllid, vector of the Huanglongbing Pathogens. Annu. Rev. Entomol. 2013, 58, 413–432. [Google Scholar] [CrossRef] [PubMed]
- Munyaneza, J.E. Zebra chip disease of potato: Biology, epidemiology, and management. Am. J. Potato Res. 2012, 89, 329–350. [Google Scholar] [CrossRef]
- Wuriyanghan, H.; Rosa, C.; Falk, B.W. Oral delivery of double-stranded RNAs and siRNAs induces RNAi effects in the potato/tomato psyllid, Bactericerca cockerelli. PLoS One 2011, 6, e27736. [Google Scholar] [CrossRef]
- Wuriyanghan, H.; Falk, B.W. RNA Interference towards the Potato Psyllid, Bactericera cockerelli, is induced in plants infected with recombinant Tobacco mosaic virus (TMV). PLoS One 2013, 8, e66050. [Google Scholar] [CrossRef]
- Lu, H.; Zhang, C.; Albrecht, U.; Shimizu, R.; Wang, G.; Bowman, K.D. Overexpression of a citrus NDR1 ortholog increases disease resistance in Arabidopsis. Front. Plant Sci. 2013, 4. [Google Scholar] [CrossRef]
- Soderlund, C.; Nelson, W.; Willer, M.; Gang, D.R. TCW: Transcriptome computational workbench. PLoS One 2013, 8, e69401. [Google Scholar] [CrossRef]
- Vyas, M.; Fisher, T.W.; Nelson, W.; Yin, G.; Cicero, J.M.; Willer, M.; Kim, R.; Kramer, R.; May, G.A.; Crow, J.A.; Soderlund, C.A.; Gang, D.R.; Brown, J.K. Stage-specific Ca. Liberibacter-infected Asian citrus psyllid expression profiles reveal exploitation nutrition and energy metabolism in adults and perturbed innate immunity and development in nymphs. PLoS ONE 2014. submitted. [Google Scholar]
- Cicero, J.M.; Fisher, T.W.; Brown, J.K. are preparing the manuscript with title “Validation of the Candidatus Liberibacter solanacearum-morphotype by colloidal gold in situ hybridization and direct evidence of a flagellated form”. 2014. [Google Scholar]
- Hung, T.H.; Hung, S.C.; Chen, C.N.; Hsu, M.H.; Su, H.J. Detection by PCR of Candidatus Liberibacter asiaticus, the bacterium causing citrus huanglongbing in vector psyllids: Application to the study of vector-pathogen relationships. Plant Pathol. 2004, 53, 96–102. [Google Scholar] [CrossRef]
- Cicero, J.M.; Brown, J.K.; Roberts, P.D.; Stansly, P.A. The Digestive System of Diaphorina citri and Bactericera cockerelli (Hemiptera: Psyllidae). Ann. Entomol. Soc. Am. 2009, 102, 650–665. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.R.; Koren, S.; Sutton, G. Assembly algorithms for next-generation sequencing data. Genomics 2010, 95, 315–327. [Google Scholar] [CrossRef] [PubMed]
- Apweiler, R.; Martin, M.J.; O’Donovan, C.; Magrane, M.; Alam-Faruque, Y.; Antunes, R.; Barrell, D.; Bely, B.; Bingley, M.; Binns, D.; et al. UniProt Consortium The Universal Protein Resource (UniProt) in 2010. Nucleic Acids Res. 2010, 38, D142–D148. [Google Scholar]
- Apweiler, R.; Martin, M.J.; O’Donovan, C.; Magrane, M.; Alam-Faruque, Y.; Antunes, R.; Barrell, D.; Bely, B.; Bingley, M.; Binns, D.; et al. UniProt Consortium Ongoing and future developments at the Universal Protein Resource. Nucleic Acids Res. 2011, 39, D214–D219. [Google Scholar]
- Milius, S. Malaria mosquito dosed with disease-fighting bacteria, after thousands of tries lab gets parasite-carrying insect to catch Wolbachia. Available online: https://www.sciencenews.org/article/malaria-mosquito-dosed-disease-fighting-bacteria (accessed on 23 October 2014).
- Thao, M.L.; Moran, N.A.; Abbot, P.; Brennan, E.B.; Burckhardt, D.H.; Baumann, P. Cospeciation of psyllids and their primary prokaryotic endosymbionts. Appl. Environ. Microbiol. 2000, 66, 2898–2905. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.; Doddapaneni, H.; Chen, C.; Duan, Y.; Zhou, L.; Stenger, D.C.; Civerolo, E.L. Draft genome sequence of potato “Zebra Chip” associated bacterium “Candidatus Liberibacter solanacearum”. Phytopathology 2009, 99, S73–S73. [Google Scholar]
- Zou, H.; Gowda, S.; Zhou, L.; Hajeri, S.; Chen, G.; Duan, Y. The destructive citrus pathogen, “Candidatus Liberibacter asiaticus” encodes a functional flagellin characteristic of a pathogen-associated molecular pattern. PLoS One 2012, e46447. [Google Scholar] [CrossRef]
- Wairuri, C.K.; Van der Waals, Jacquie E; Van Schalkwyk, A.; Theron, J. Ralstonia solanacearum needs Flp pili for virulence on potato. Mol. Plant-Microbe Interact. 2012, 25, 546–556. [Google Scholar] [CrossRef]
- Chiancone, E.; Ceci, P.; Ilari, A.; Ribacchi, F.; Stefanini, S. Iron and proteins for iron storage and detoxification. BioMetals 2004, 17, 197–202. [Google Scholar] [CrossRef] [PubMed]
- Flipsen, J.; Mans, R.; Kleefsman, A.; Knebel-Mörsdorf, D.; Vlak, J.M. Deletion of the baculovirus ecdysteroid UDP-glucosyltransferase gene induces early degeneration of Malpighian tubules in infected insects. J. Virol. 1995, 69, 4529–4532. [Google Scholar] [PubMed]
- Slack, J.M.; Kuzio, J.; Faulkner, P. Characterization of v-cath, a cathepsin L-like proteinase expressed by the baculovirus Autographa californica multiple nuclear polyhedrosis virus. J. Gen. Virol. 1995, 76, 1091–1098. [Google Scholar] [CrossRef] [PubMed]
- Cook, P.; McMeniman, C.; O Neill, S. Modifying Insect Population Age Structure to Control Vector-Borne Disease; Aksoy, S., Ed.; Springer: New York, NY, USA, 2008; Volume 627; pp. 126–140. [Google Scholar]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene Ontology Consortium Gene Ontology: Tool for the unification of biology. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Shen, Z.; Jacobs-Lorena, M. A Type I Peritrophic matrix protein from the Malaria vector Anopheles gambiae binds to chitin: Cloning, Expression, and Characterization. J. Biol. Chem. 1998, 273, 17665–17670. [Google Scholar] [CrossRef] [PubMed]
- Hegedus, D.; Erlandson, M.; Gillott, C.; Toprak, U. New insights into peritrophic matrix synthesis, architecture, and function. Annu. Rev. Entomol. 2009, 54, 285–302. [Google Scholar] [CrossRef] [PubMed]
- Schulte, R.D.; Makus, C.; Hasert, B.; Michiels, N.K.; Schulenburg, H. Multiple reciprocal adaptations and rapid genetic change upon experimental coevolution of an animal host and its microbial parasite. Proc. Natl. Acad. Sci. USA 2010, 107, 7359–7364. [Google Scholar] [CrossRef] [PubMed]
- Raddadi, N.; Gonella, E.; Camerota, C.; Pizzinat, A.; Tedeschi, R.; Crotti, E.; Mandrioli, M.; Bianco, P.A.; Daffonchio, D.; Alma, A. “Candidatus Liberibacter europaeus” sp. nov. that is associated with and transmitted by the psyllid Cacopsylla pyri apparently behaves as an endophyte rather than a pathogen. Environ. Microbiol. 2011, 13, 414–426. [Google Scholar] [CrossRef] [PubMed]
- Doucet, D.; Retnakaran, A. Insect Chitin: Metabolism, Genomics and Pest Management. Insect Growth Disruptors 2012, 43, 437–511. [Google Scholar]
- Killiny, N.; Prado, S.S.; Almeida, R.P. Chitin utilization by the insect-transmitted bacterium Xylella fastidiosa. Appl. Environ. Microbiol. 2010, 76, 6134–6140. [Google Scholar] [CrossRef] [PubMed]
- Ortiz-Rodriguez, T.; de la Fuente-Salcido, N.; Bideshi, D.K.; Salcedo-Hernandez, R.; Barboza-Corona, J.E. Generation of chitin-derived oligosaccharides toxic to pathogenic bacteria using ChiA74, an endochitinase native to Bacillus thuringiensis. Lett. Appl. Microbiol. 2010, 51, 184–190. [Google Scholar] [PubMed]
- Gerke, V.; Moss, S.E. Annexins and membrane dynamics. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 1997, 1357, 129–154. [Google Scholar] [CrossRef]
- Tjota, M.; Lee, S.K.; Wu, J.; Williams, J.A.; Khanna, M.R.; Thomas, G.H. Annexin B9 binds to beta(H)-spectrin and is required for multivesicular body function in Drosophila. J. Cell. Sci. 2011, 124, 2914–2926. [Google Scholar] [CrossRef] [PubMed]
- Kotsyfakis, M.; Ehret‐Sabatier, L.; Siden‐Kiamos, I.; Mendoza, J.; Sinden, R.E.; Louis, C. Plasmodium berghei ookinetes bind to Anopheles gambiae and Drosophila melanogaster annexins. Mol. Microbiol. 2005, 57, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Lavine, M.; Strand, M. Insect hemocytes and their role in immunity. Insect Biochem. Mol. Biol. 2002, 32, 1295–1309. [Google Scholar] [CrossRef] [PubMed]
- Ratzka, C.; Gross, R.; Feldhaar, H. Endosymbiont Tolerance and Control within Insect Hosts. Insects 2012, 3, 553–572. [Google Scholar] [CrossRef]
- Wang, N.; Trivedi, P. Citrus huanglongbing: A newly relevant disease presents unprecedented challenges. Phytopathology 2013, 103, 652–665. [Google Scholar] [CrossRef] [PubMed]
- Kovács, Á.T.; van Gestel, J.; Kuipers, O.P. The protective layer of biofilm: A repellent function for a new class of amphiphilic proteins. Mol. Microbiol. 2012, 85, 8–11. [Google Scholar] [CrossRef] [PubMed]
- Bastian, F.O.; Elzer, P.H.; Wu, X. Spiroplasma spp. biofilm formation is instrumental for their role in the pathogenesis of plant, insect and animal diseases. Exp. Mol. Pathol. 2012, 93, 116–128. [Google Scholar] [CrossRef] [PubMed]
- Nachappa, P.; Levy, J.; Tamborindeguy, C. Transcriptome analyses of Bactericera cockerelli adults in response to “Candidatus Liberibacter solanacearum” infection. Mol. Genet. Genomics 2012, 287, 803–817. [Google Scholar] [CrossRef] [PubMed]
- Leulier, F.; Rodriguez, A.; Khush, R.S.; Abrams, J.M.; Lemaitre, B. The Drosophila caspase Dredd is required to resist gram-negative bacterial infection. EMBO Rep. 2000, 1, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Doyle, S.E.; O’Connell, R.M.; Miranda, G.A.; Vaidya, S.A.; Chow, E.K.; Liu, P.T.; Suzuki, S.; Suzuki, N.; Modlin, R.L.; Yeh, W. Toll-like receptors induce a phagocytic gene program through p38. J. Exp. Med. 2004, 199, 81–90. [Google Scholar] [PubMed]
- Valanne, S.; Wang, J.; Rämet, M. The Drosophila Toll Signaling Pathway. J. Immunol. 2011, 186, 649–656. [Google Scholar] [CrossRef] [PubMed]
- Flannagan, R.S.; Jaumouillé, V.; Grinstein, S. The cell biology of phagocytosis. Annu. Rev. Pathol. Mech. Dis. 2012, 7, 61–98. [Google Scholar] [CrossRef]
- Mansueto, P.; Vitale, G.; Cascio, A.; Seidita, A.; Pepe, I.; Carroccio, A.; di Rosa, S.; Rini, G.B.; Cillari, E.; Walker, D.H. New insight into immunity and immunopathology of Rickettsial diseases. Clin. Dev. Immunol. 2012, 2–26. [Google Scholar]
- Cossart, P.; Veiga, E. Non-classical use of clathrin during bacterial infections. J. Microsc. 2008, 231, 524–528. [Google Scholar] [CrossRef] [PubMed]
- Sarantis, H.; Grinstein, S. Subversion of phagocytosis for pathogen survival. Cell Host Microbe 2012, 12, 419–431. [Google Scholar] [CrossRef] [PubMed]
- Hudson, B.I.; Kalea, A.Z.; del Mar Arriero, M.; Harja, E.; Boulanger, E.; D’Agati, V.; Schmidt, A.M. Interaction of the RAGE cytoplasmic domain with diaphanous-1 is required for ligand-stimulated cellular migration through activation of Rac1 and Cdc42. J. Biol. Chem. 2008, 283, 34457–34468. [Google Scholar] [CrossRef] [PubMed]
- Castellano, F.; Montcourrier, P.; Chavrier, P. Membrane recruitment of Rac1 triggers phagocytosis. J. Cell. Sci. 2000, 113, 2955–2961. [Google Scholar] [PubMed]
- Fu, Y.; Galán, J.E. A Salmonella protein antagonizes Rac-1 and Cdc42 to mediate host-cell recovery after bacterial invasion. Nature 1999, 401, 293–297. [Google Scholar] [CrossRef] [PubMed]
- Kontani, K.; Tada, M.; Ogawa, T.; Okai, T.; Saito, K.; Araki, Y.; Katada, T. Di-Ras, a distinct subgroup of ras family GTPases with unique biochemical properties. J. Biol. Chem. 2002, 277, 41070–41078. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Moon, M.; Kim, H.; Li, Y.; Song, D.; Kim, J.; Lee, J.; Kim, J.; Kim, S.; Park, J. Ras-related GTPases Rap1 and RhoA collectively induce the phagocytosis of serum-opsonized zymosan particles in macrophages. J. Biol. Chem. 2012, 287, 5145–5155. [Google Scholar] [CrossRef] [PubMed]
- Akbar, M.A.; Tracy, C.; Kahr, W.H.; Krämer, H. The full-of-bacteria gene is required for phagosome maturation during immune defense in Drosophila. J. Cell Biol. 2011, 192, 383–390. [Google Scholar] [CrossRef]
- Parker, H.; Chitcholtan, K.; Hampton, M.B.; Keenan, J.I. Uptake of Helicobacter pylori outer membrane vesicles by gastric epithelial cells. Infect. Immun. 2010, 78, 5054–5061. [Google Scholar] [CrossRef] [PubMed]
- Jin, T.; Satoh, T.; Liao, Y.; Song, C.; Gao, X.; Kariya, K.; Hu, C.; Kataoka, T. Role of the CDC25 homology domain of phospholipase Cε in amplification of Rap1-dependent signaling. J. Biol. Chem. 2001, 276, 30301–30307. [Google Scholar] [CrossRef] [PubMed]
- Lopez, I.; Mak, E.C.; Ding, J.; Hamm, H.E.; Lomasney, J.W. A novel bifunctional phospholipase C that is regulated by Gα12 and stimulates the Ras/mitogen-activated protein kinase pathway. J. Biol. Chem. 2001, 276, 2758–2765. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.; Lou, B.; Glynn, J.M.; Doddapaneni, H.; Civerolo, E.L.; Chen, C.; Duan, Y.; Zhou, L.; Vahling, C.M. The complete genome sequence of “Candidatus Liberibacter solanacearum”, the bacterium associated with potato zebra chip disease. PLoS One 2011, 6, e19135. [Google Scholar] [CrossRef]
- Battaile, K.P.; McBurney, M.; Van Veldhoven, P.P.; Vockley, J. Human long chain, very long chain and medium chain acyl-CoA dehydrogenases are specific for the S-enantiomer of 2-methylpentadecanoyl-CoA. Biochim. Biophys. Acta-Lipids Lipid Metab. 1998, 1390, 333–338. [Google Scholar] [CrossRef]
- Kazakov, A.E.; Rodionov, D.A.; Alm, E.; Arkin, A.P.; Dubchak, I.; Gelfand, M.S. Comparative Genomics of Regulation of Fatty Acid and Branched-Chain Amino Acid Utilization in Proteobacteria. J. Bacteriol. 2009, 191, 52–64. [Google Scholar] [CrossRef] [PubMed]
- Falabella, P.; Riviello, L.; Stradis, M.D.; Stigliano, C.; Varricchio, P.; Grimaldi, A.; Eguileor, M.D.; Graziani, F.; Gigliotti, S.; Pennacchio, F. Aphidius ervi teratocytes release an extracellular enolase. Insect Biochem. Mol. Biol. 2009, 39, 801–813. [Google Scholar] [CrossRef] [PubMed]
- Mitra, S.; Cui, J.; Robbins, P.W.; Samuelson, J. A deeply divergent phosphoglucomutase (PGM) of Giardia lamblia has both PGM and phosphomannomutase activities. Glycobiology 2010, 20, 1233–1240. [Google Scholar] [CrossRef] [PubMed]
- Lazarevic, V.; Soldo, B.; Médico, N.; Pooley, H.; Bron, S.; Karamata, D. Bacillus subtilis α-Phosphoglucomutase Is Required for Normal Cell Morphology and Biofilm Formation. Appl. Environ. Microbiol. 2005, 71, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Runyen-Janecky, L.J.; Brown, A.N.; Ott, B.; Tujuba, H.G.; Rio, R.V. Regulation of high-affinity iron acquisition homologues in the tsetse fly symbiont Sodalis glossinidius. J. Bacteriol. 2010, 192, 3780–3787. [Google Scholar] [CrossRef] [PubMed]
- Meyron-Holtz, E.G.; Moshe-Belizowski, S.; Cohen, L.A. A possible role for secreted ferritin in tissue iron distribution. J. Neural Transm. 2011, 118, 337–347. [Google Scholar] [CrossRef] [PubMed]
- Tran, S.L.; Guillemet, E.; Lereclus, D.; Ramarao, N. Iron regulates Bacillus thuringiensis haemolysin hlyII gene expression during insect infection. J. Invertebr. Pathol. 2013, 113, 205–208. [Google Scholar] [CrossRef] [PubMed]
- Arosio, P.; Ingrassia, R.; Cavadini, P. Ferritins: A family of molecules for iron storage, antioxidation and more. Biochim. Biophys. Acta (BBA)-Gen. Subj. 2009, 1790, 589–599. [Google Scholar] [CrossRef]
- Kim, B.Y.; Lee, K.S.; Choo, Y.M.; Kim, I.; Je, Y.H.; Woo, S.D.; Lee, S.M.; Park, H.C.; Sohn, H.D.; Jin, B.R. Insect transferrin functions as an antioxidant protein in a beetle larva. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2008, 150, 161–169. [Google Scholar] [CrossRef]
- Gabaldon, T.; Koonin, E.V. Functional and evolutionary implications of gene orthology. Nat. Rev. Genet. 2013, 14, 360–366. [Google Scholar] [CrossRef] [PubMed]
- Iseli, C.; Jongeneel, C.V.; Bucher, P. ESTScan: A program for detecting, evaluating, and reconstructing potential coding regions in EST sequences. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1999, 138–148. [Google Scholar]
- Li, L.; Stoeckert, C.J., Jr.; Roos, D.S. OrthoMCL: Identification of ortholog groups for eukaryotic genomes. Genome Res. 2003, 13, 2178–2189. [Google Scholar] [CrossRef] [PubMed]
- Lee Rodgers, J.; Nicewander, W.A. Thirteen ways to look at the correlation coefficient. Am. Stat. 1988, 42, 59–66. [Google Scholar] [CrossRef]
- Goh, K.; Cusick, M.E.; Valle, D.; Childs, B.; Vidal, M.; Barabasi, A. The human disease network. Proc. Natl. Acad. Sci. USA 2007, 104, 8685–8690. [Google Scholar] [CrossRef] [PubMed]
- Aruni, W.; Vanterpool, E.; Osbourne, D.; Roy, F.; Muthiah, A.; Dou, Y.; Fletcher, H.M. Sialidase and Sialoglycoproteases Can Modulate Virulence in Porphyromonas gingivalis. Infect. Immun. 2011, 79, 2779–2791. [Google Scholar] [CrossRef] [PubMed]
- Severi, E.; Hood, D.W.; Thomas, G.H. Sialic acid utilization by bacterial pathogens. Microbiology 2007, 153, 2817–2822. [Google Scholar] [CrossRef] [PubMed]
- Wallace, A.J.; Stillman, T.J.; Atkins, A.; Jamieson, S.J.; Bullough, P.A.; Green, J.; Artymiuk, P.J. E. coli hemolysin E (HlyE, ClyA, SheA): X-ray crystal structure of the toxin and observation of membrane pores by electron microscopy. Cell 2000, 100, 265–276. [Google Scholar] [CrossRef] [PubMed]
- Zitzer, A.; Zitzer, O.; Bhakdi, S.; Palmer, M. Oligomerization of Vibrio cholerae cytolysin yields a pentameric pore and has a dual specificity for cholesterol and sphingolipids in the target membrane. J. Biol. Chem. 1999, 274, 1375–1380. [Google Scholar] [CrossRef] [PubMed]
- Amino, R.; Martins, R.M.; Procopio, J.; Hirata, I.Y.; Juliano, M.A.; Schenkman, S. Trialysin, a novel pore-forming protein from saliva of hematophagous insects activated by limited proteolysis. J. Biol. Chem. 2002, 277, 6207–6213. [Google Scholar] [CrossRef] [PubMed]
- Petersen, U.; Kadalayil, L.; Rehorn, K.; Hoshizaki, D.K.; Reuter, R.; Engström, Y. Serpent regulates Drosophila immunity genes in the larval fat body through an essential GATA motif. EMBO J. 1999, 18, 4013–4022. [Google Scholar] [CrossRef] [PubMed]
- Ennes-Vidal, V.; Menna-Barreto, R.F. S.; Santos, A.L. S.; Branquinha, M.H.; d’Avila-Levy, C.M. MDL28170, a Calpain Inhibitor, Affects Trypanosoma cruzi metacyclogenesis, ultrastructure and attachment to Rhodnius prolixus midgut. PLoS One 2011, 6, e18371. [Google Scholar] [CrossRef] [PubMed]
- Patt, J.; Sétamou, M. Responses of the Asian citrus psyllid to volatiles emitted by the flushing shoots of its rutaceous host plants. Environ. Entomol. 2010, 39, 618–624. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Broder, C.C.; Nikolov, D.B. Ephrin-B2 and Ephrin-B3 as Functional Henipavirus Receptors. Semin. Cell Dev. Biol. 2012, 23, 116–123. [Google Scholar] [CrossRef] [PubMed]
- Nilius, B.; Owsianik, G. The transient receptor potential family of ion channels. Genome Biol. 2011. [Google Scholar] [CrossRef]
- Schroder, H.C.; Ushijima, H.; Krasko, A.; Gamulin, V.; Thakur, N.L.; Diehl-Seifert, B.; Muller, I.M.; Muller, W.E. Emergence and disappearance of an immune molecule, an antimicrobial lectin, in basal metazoa. A tachylectin-related protein in the sponge Suberites domuncula. J. Biol. Chem. 2003, 278, 32810–32817. [Google Scholar] [CrossRef] [PubMed]
- Reiko, F.; Shusaku, I.; Yoshimitsu, K.; Toshitsugu, U.; Yoshiko, M. Characterization of tectonins I and II from Physarum polycephalum. Adv. Biol. Chem. 2012. [Google Scholar] [CrossRef]
- Heung, L.J.; Luberto, C.; del Poeta, M. Role of sphingolipids in microbial pathogenesis. Infect. Immun. 2006, 74, 28–39. [Google Scholar] [CrossRef] [PubMed]
- Rajakumari, S.; Rajasekharan, R.; Daum, G. Triacylglycerol lipolysis is linked to sphingolipid and phospholipid metabolism of the yeast Saccharomyces cerevisiae. Biochim. Biophys. Acta (BBA)-Mol. Cell Biol. Lipids 2010, 1801, 1314–1322. [Google Scholar] [CrossRef]
- Lattif, A.A.; Mukherjee, P.K.; Chandra, J.; Roth, M.R.; Welti, R.; Rouabhia, M.; Ghannoum, M.A. Lipidomics of Candida albicans biofilms reveals phase-dependent production of phospholipid molecular classes and role for lipid rafts in biofilm formation. Microbiology 2011, 157, 3232–3242. [Google Scholar] [CrossRef] [PubMed]
- Swisher, K.D.; Munyaneza, J.E.; Crosslin, J.M. High resolution melting analysis of the cytochrome oxidase I gene identifies three haplotypes of the potato psyllid in the United States. Environ. Entomol. 2012, 41, 1019–1028. [Google Scholar] [CrossRef]
- Jagoueix, S.; Bové, J.M.; Garnier, M. PCR detection of the two Candidatus liberobacter species associated with greening disease of citrus. Mol. Cell. Probes 1996, 10, 43–50. [Google Scholar] [CrossRef] [PubMed]
- He, R.; Kim, M.J.; Nelson, W.; Balbuena, T.S.; Kim, R.; Kramer, R.; Crow, J.A.; May, G.D.; Thelen, J.J.; Soderlund, C.A.; et al. Next-generation sequencing-based transcriptomic and proteomic analysis of the common reed, Phragmites australis (Poaceae), reveals genes involved in invasiveness and rhizome specificity. Am. J. Bot. 2012, 99, 232–247. [Google Scholar] [CrossRef] [PubMed]
- Birol, I.; Jackman, S.D.; Nielsen, C.B.; Qian, J.Q.; Varhol, R.; Stazyk, G.; Morin, R.D.; Zhao, Y.; Hirst, M.; Schein, J.E.; et al. De novo transcriptome assembly with ABySS. Bioinformatics 2009, 25, 2872–2877. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Yu, C.; Li, Y.; Lam, T.; Yiu, S.; Kristiansen, K.; Wang, J. SOAP2: An improved ultrafast tool for short read alignment. Bioinformatics 2009, 25, 1966–1967. [Google Scholar] [CrossRef] [PubMed]
- Chevreux, B.; Pfisterer, T.; Drescher, B.; Driesel, A.J.; Muller, W.E. G.; Wetter, T.; Suhai, S. Using the miraEST assembler for reliable and automated mRNA transcript assembly and SNP detection in sequenced ESTs. Genome Res. 2004, 14, 1147–1159. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 2006, 22, 1658–1659. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Young, M.; Wakefield, M.; Smyth, G.; Oshlack, A. Gene ontology analysis for RNA-seq: Accounting for selection bias. Genome Biol. 2010. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fisher, T.W.; Vyas, M.; He, R.; Nelson, W.; Cicero, J.M.; Willer, M.; Kim, R.; Kramer, R.; May, G.A.; Crow, J.A.; et al. Comparison of Potato and Asian Citrus Psyllid Adult and Nymph Transcriptomes Identified Vector Transcripts with Potential Involvement in Circulative, Propagative Liberibacter Transmission. Pathogens 2014, 3, 875-907. https://doi.org/10.3390/pathogens3040875
Fisher TW, Vyas M, He R, Nelson W, Cicero JM, Willer M, Kim R, Kramer R, May GA, Crow JA, et al. Comparison of Potato and Asian Citrus Psyllid Adult and Nymph Transcriptomes Identified Vector Transcripts with Potential Involvement in Circulative, Propagative Liberibacter Transmission. Pathogens. 2014; 3(4):875-907. https://doi.org/10.3390/pathogens3040875
Chicago/Turabian StyleFisher, Tonja W., Meenal Vyas, Ruifeng He, William Nelson, Joseph M. Cicero, Mark Willer, Ryan Kim, Robin Kramer, Greg A. May, John A. Crow, and et al. 2014. "Comparison of Potato and Asian Citrus Psyllid Adult and Nymph Transcriptomes Identified Vector Transcripts with Potential Involvement in Circulative, Propagative Liberibacter Transmission" Pathogens 3, no. 4: 875-907. https://doi.org/10.3390/pathogens3040875
APA StyleFisher, T. W., Vyas, M., He, R., Nelson, W., Cicero, J. M., Willer, M., Kim, R., Kramer, R., May, G. A., Crow, J. A., Soderlund, C. A., Gang, D. R., & Brown, J. K. (2014). Comparison of Potato and Asian Citrus Psyllid Adult and Nymph Transcriptomes Identified Vector Transcripts with Potential Involvement in Circulative, Propagative Liberibacter Transmission. Pathogens, 3(4), 875-907. https://doi.org/10.3390/pathogens3040875




