A Compact Closed Genome of Orientia tsutsugamushi from Hainan Island, China Provides a TA763_A Reference and Reveals Repeat-Driven Remodeling
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Case Identification and Laboratory Confirmation
2.3. Isolation and Culture of O. tsutsugamushi from Blood Clot
2.4. Immuno-Electron Microscopy
2.5. Growth Kinetics by qPCR
2.6. Genomic DNA Preparation and Whole-Genome Sequencing
2.7. Genome Assembly, Annotation and Comparative Analysis
2.8. Data Availability
3. Results
3.1. Clinical Presentation and Laboratory Findings
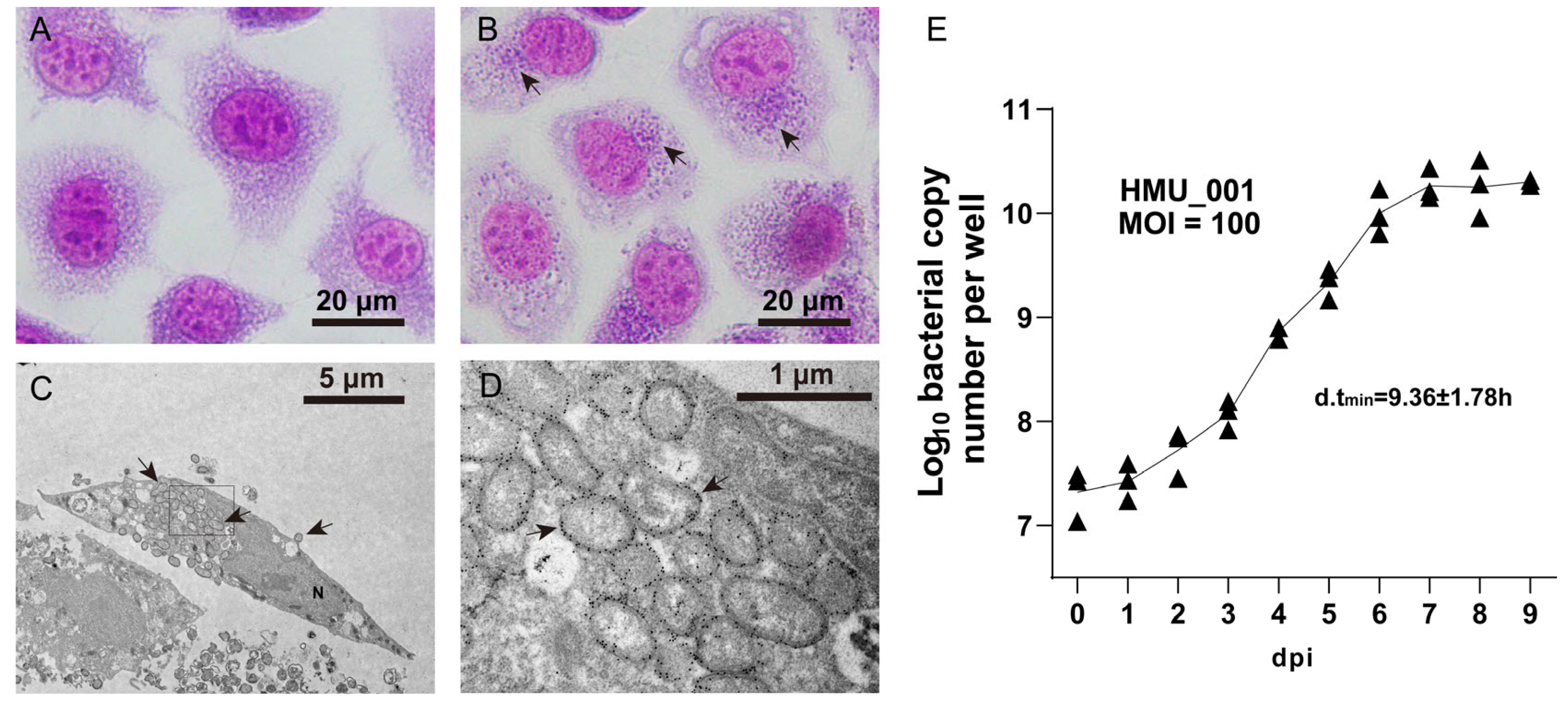
3.2. Bacterial Morphology and Growth Kinetics
3.3. Genome Composition and Functional Overview of O. tsutsugamushi HMU_001
3.4. Repeated Sequence Analysis
3.5. Orthologous Groups, and Phylogenetic Analysis and MLST Classification
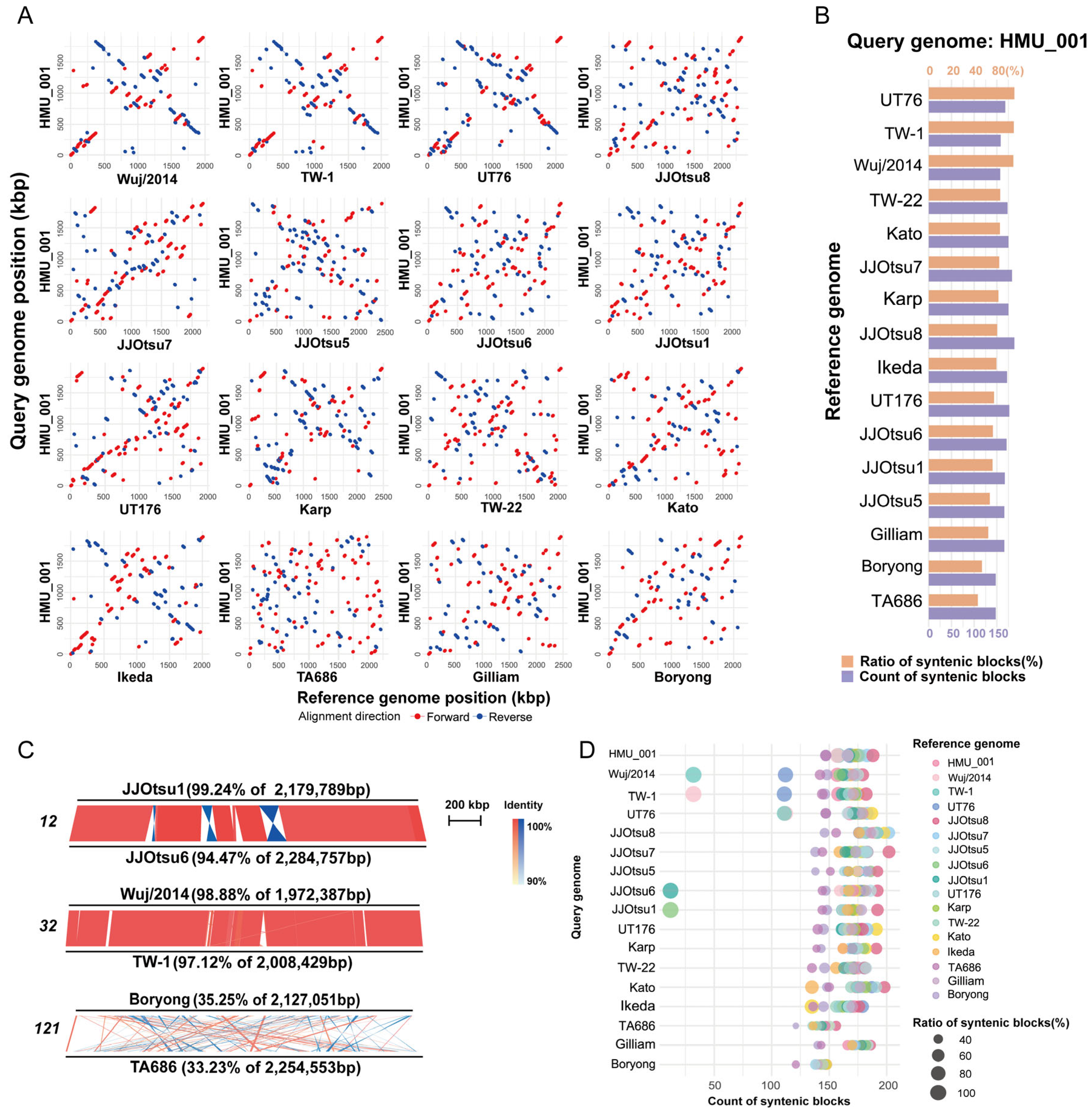
3.6. Genome Synteny Analysis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bhandari, M.; Singh, R.K.; Laishevtcev, A.; Mohapatra, T.M.; Nigam, M.; Mori, E.; Vasconcelos de Lacerda, B.C.G.; Coutinho, H.D.M.; Mishra, A.P. Revisiting scrub typhus: A neglected tropical disease. Comp. Immunol. Microbiol. Infect. Dis. 2022, 90–91, 101888. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Walker, D.H.; Jupiter, D.; Melby, P.C.; Arcari, C.M. A review of the global epidemiology of scrub typhus. PLoS Negl. Trop. Dis. 2017, 11, e0006062. [Google Scholar] [CrossRef]
- Walker, D.H. Scrub Typhus-Scientific Neglect, Ever-Widening Impact. N. Engl. J. Med. 2016, 375, 913–915. [Google Scholar] [CrossRef]
- Jiang, J.; Richards, A.L. Scrub Typhus: No Longer Restricted to the Tsutsugamushi Triangle. Trop. Med. Infect. Dis. 2018, 3, 11. [Google Scholar] [CrossRef]
- Alkathiry, H.A.; Alghamdi, S.Q.; Morgan, H.E.J.; Noll, M.E.; Khoo, J.J.; Alagaili, A.N.; Makepeace, B.L. Molecular Detection of Candidatus Orientia chuto in Wildlife, Saudi Arabia. Emerg. Infect. Dis. 2023, 29, 402–406. [Google Scholar] [CrossRef] [PubMed]
- Abarca, K.; Martínez-Valdebenito, C.; Angulo, J.; Jiang, J.; Farris, C.M.; Richards, A.L.; Acosta-Jamett, G.; Weitzel, T. Molecular Description of a Novel Orientia Species Causing Scrub Typhus in Chile. Emerg. Infect. Dis. 2020, 26, 2148–2156. [Google Scholar] [CrossRef]
- Dasgupta, S.; Asish, P.R.; Rachel, G.; Bagepally, B.S.; Chethrapilly Purushothaman, G.K. Global seroprevalence of scrub typhus: A systematic review and meta-analysis. Sci. Rep. 2024, 14, 10895. [Google Scholar] [CrossRef]
- Gaba, S.; Gupta, M.; Singla, N.; Singh, R. Clinical outcome and predictors of severity in scrub typhus patients at a tertiary care hospital in Chandigarh, India. J. Vector Borne Dis. 2019, 56, 367–372. [Google Scholar] [CrossRef]
- Kumar, V.; Kumar, V.; Yadav, A.K.; Iyengar, S.; Bhalla, A.; Sharma, N.; Aggarwal, R.; Jain, S.; Jha, V. Scrub typhus is an under-recognized cause of acute febrile illness with acute kidney injury in India. PLoS Negl. Trop. Dis. 2014, 8, e2605. [Google Scholar] [CrossRef] [PubMed]
- Saraswati, K.; Maguire, B.J.; McLean, A.R.D.; Singh-Phulgenda, S.; Ngu, R.C.; Newton, P.N.; Day, N.P.J.; Guérin, P.J. Systematic review of the scrub typhus treatment landscape: Assessing the feasibility of an individual participant-level data (IPD) platform. PLoS Negl. Trop. Dis. 2021, 15, e0009858. [Google Scholar] [CrossRef]
- Blacksell, S.D.; Bryant, N.J.; Paris, D.H.; Doust, J.A.; Sakoda, Y.; Day, N.P. Scrub typhus serologic testing with the indirect immunofluorescence method as a diagnostic gold standard: A lack of consensus leads to a lot of confusion. Clin. Infect. Dis. 2007, 44, 391–401. [Google Scholar] [CrossRef]
- Kannan, K.; John, R.; Kundu, D.; Dayanand, D.; Abhilash, K.P.P.; Mathuram, A.J.; Zachariah, A.; Sathyendra, S.; Hansdak, S.G.; Abraham, O.C.; et al. Performance of molecular and serologic tests for the diagnosis of scrub typhus. PLoS Negl. Trop. Dis. 2020, 14, e0008747. [Google Scholar] [CrossRef] [PubMed]
- Taylor, A.J.; Paris, D.H.; Newton, P.N. A Systematic Review of Mortality from Untreated Scrub Typhus (Orientia tsutsugamushi). PLoS Negl. Trop. Dis. 2015, 9, e0003971. [Google Scholar] [CrossRef]
- Wang, Q.; Ma, T.; Ding, F.; Lim, A.; Takaya, S.; Saraswati, K.; Sartorius, B.; Day, N.P.J.; Maude, R.J. Global and regional seroprevalence, incidence, mortality of, and risk factors for scrub typhus: A systematic review and meta-analysis. Int. J. Infect. Dis. 2024, 146, 107151. [Google Scholar] [CrossRef]
- Kelly, D.J.; Fuerst, P.A.; Richards, A.L. Origins, Importance and Genetic Stability of the Prototype Strains Gilliam, Karp and Kato of Orientia tsutsugamushi. Trop. Med. Infect. Dis. 2019, 4, 75. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.S.; Chang, W.H. Antigenic relationship among the eight prototype and new serotype strains of Orientia tsutsugamushi revealed by monoclonal antibodies. Microbiol. Immunol. 1999, 43, 229–234. [Google Scholar] [CrossRef]
- Furuya, Y.; Yoshida, Y.; Katayama, T.; Yamamoto, S.; Kawamura, A., Jr. Serotype-specific amplification of Rickettsia tsutsugamushi DNA by nested polymerase chain reaction. J. Clin. Microbiol. 1993, 31, 1637–1640. [Google Scholar] [CrossRef]
- Sihag, K.K.; Arif, W.; Srirama, S.; Chandrasekaran, A.K.; Raveendran, V.; Chandrakumar, A.B.; Kasirajan, A.; Thavaraj, S.A.P.; Srinivasan, L.; Choolayil, A.C.; et al. A longitudinal molecular surveillance of genetic heterogeneity of Orientia tsutsugamushi in humans, reservoir animals, and vectors in Puducherry, India. Front. Microbiol. 2025, 16, 1634394. [Google Scholar] [CrossRef]
- Wang, G.; Fu, R.; Zhang, L.; Xue, L.; Al-Mahdi, A.Y.; Xie, X.; Qin, A.; Tang, C.; Du, J.; Huang, Y.; et al. Genomic bacterial load associated with bacterial genotypes and clinical characteristics in patients with scrub typhus in Hainan Island, Southern China. PLoS Negl. Trop. Dis. 2023, 17, e0011243. [Google Scholar] [CrossRef]
- Tang, C.; Huang, Y.; Wang, G.; Xue, L.; Hu, X.; Peng, R.; Du, J.; Yang, J.; Niu, Y.; Deng, W.; et al. Patient-centric analysis of Orientia tsutsugamushi spatial diversity patterns across Hainan Island, China. PLoS Negl. Trop. Dis. 2025, 19, e0012909. [Google Scholar] [CrossRef]
- Chaichana, P.; Satapoomin, N.; Kullapanich, C.; Chuenklin, S.; Mohammad, A.; Inthawong, M.; Ball, E.E.; Burke, T.P.; Sunyakumthorn, P.; Salje, J. Comparative virulence analysis of seven diverse strains of Orientia tsutsugamushi reveals a multifaceted and complex interplay of virulence factors responsible for disease. PLoS Pathog. 2025, 21, e1012833. [Google Scholar] [CrossRef] [PubMed]
- Batty, E.M.; Chaemchuen, S.; Blacksell, S.; Richards, A.L.; Paris, D.; Bowden, R.; Chan, C.; Lachumanan, R.; Day, N.; Donnelly, P.; et al. Long-read whole genome sequencing and comparative analysis of six strains of the human pathogen Orientia tsutsugamushi. PLoS Negl. Trop. Dis. 2018, 12, e0006566. [Google Scholar] [CrossRef]
- Sonthayanon, P.; Peacock, S.J.; Chierakul, W.; Wuthiekanun, V.; Blacksell, S.D.; Holden, M.T.; Bentley, S.D.; Feil, E.J.; Day, N.P. High rates of homologous recombination in the mite endosymbiont and opportunistic human pathogen Orientia tsutsugamushi. PLoS Negl. Trop. Dis. 2010, 4, e752. [Google Scholar] [CrossRef]
- Phetsouvanh, R.; Sonthayanon, P.; Pukrittayakamee, S.; Paris, D.H.; Newton, P.N.; Feil, E.J.; Day, N.P. The Diversity and Geographical Structure of Orientia tsutsugamushi Strains from Scrub Typhus Patients in Laos. PLoS Negl. Trop. Dis. 2015, 9, e0004024. [Google Scholar] [CrossRef]
- Minahan, N.T.; Yen, T.Y.; Guo, Y.L.; Shu, P.Y.; Tsai, K.H. Concatenated ScaA and TSA56 Surface Antigen Sequences Reflect Genome-Scale Phylogeny of Orientia tsutsugamushi: An Analysis Including Two Genomes from Taiwan. Pathogens 2024, 13, 299. [Google Scholar] [CrossRef]
- Cho, N.H.; Kim, H.R.; Lee, J.H.; Kim, S.Y.; Kim, J.; Cha, S.; Kim, S.Y.; Darby, A.C.; Fuxelius, H.H.; Yin, J.; et al. The Orientia tsutsugamushi genome reveals massive proliferation of conjugative type IV secretion system and host-cell interaction genes. Proc. Natl. Acad. Sci. USA 2007, 104, 7981–7986. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, K.; Yamashita, A.; Kurokawa, K.; Morimoto, T.; Ogawa, M.; Fukuhara, M.; Urakami, H.; Ohnishi, M.; Uchiyama, I.; Ogura, Y.; et al. The Whole-genome sequencing of the obligate intracellular bacterium Orientia tsutsugamushi revealed massive gene amplification during reductive genome evolution. DNA Res. 2008, 15, 185–199. [Google Scholar] [CrossRef] [PubMed]
- Giengkam, S.; Kullapanich, C.; Wongsantichon, J.; Adcox, H.E.; Gillespie, J.J.; Salje, J. Orientia tsutsugamushi: Comprehensive analysis of the mobilome of a highly fragmented and repetitive genome reveals the capacity for ongoing lateral gene transfer in an obligate intracellular bacterium. mSphere 2023, 8, e0026823. [Google Scholar] [CrossRef]
- Nakano, K.; Shiroma, A.; Shimoji, M.; Tamotsu, H.; Ashimine, N.; Ohki, S.; Shinzato, M.; Minami, M.; Nakanishi, T.; Teruya, K.; et al. Advantages of genome sequencing by long-read sequencer using SMRT technology in medical area. Hum. Cell 2017, 30, 149–161. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.Y.; Jia, P.B.; Chen, L.; Jin, Y.M.; Feng, F.L.; Pan, B.Y.; Shen, Y.M.; Lin, X.Z.; He, Y.N. Epidemiological characteristics and trend prediction of scrub typhus in Hainan Province from 2008 to 2023. China Trop. Med. 2024, 24, 1042–1046. [Google Scholar] [CrossRef]
- Xie, X.; Zhang, Y.; Teng, Z.; Duan, B.; Hai, Y.; Wang, M.; Shao, Z.; Liang, W.; Kan, B.; Yin, F.; et al. The Prevalence of Rickettsial and Rickettsial-Like Diseases in Patients with Undifferentiated Febrile Illness-Hainan Province, China, 2018–2021. China CDC Wkly. 2024, 6, 734–739. [Google Scholar] [CrossRef]
- Kim, D.M.; Park, G.; Kim, H.S.; Lee, J.Y.; Neupane, G.P.; Graves, S.; Stenos, J. Comparison of conventional, nested, and real-time quantitative PCR for diagnosis of scrub typhus. J. Clin. Microbiol. 2011, 49, 607–612. [Google Scholar] [CrossRef] [PubMed]
- Giengkam, S.; Blakes, A.; Utsahajit, P.; Chaemchuen, S.; Atwal, S.; Blacksell, S.D.; Paris, D.H.; Day, N.P.; Salje, J. Improved Quantification, Propagation, Purification and Storage of the Obligate Intracellular Human Pathogen Orientia tsutsugamushi. PLoS Negl. Trop. Dis. 2015, 9, e0004009. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Concepcion, G.T.; Feng, X.; Zhang, H.; Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 2021, 18, 170–175. [Google Scholar] [CrossRef]
- Hunt, M.; Silva, N.D.; Otto, T.D.; Parkhill, J.; Keane, J.A.; Harris, S.R. Circlator: Automated circularization of genome assemblies using long sequencing reads. Genome Biol. 2015, 16, 294. [Google Scholar] [CrossRef] [PubMed]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Nawrocki, E.P.; Eddy, S.R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 2013, 29, 2933–2935. [Google Scholar] [CrossRef]
- Chan, P.P.; Lowe, T.M. tRNAscan-SE: Searching for tRNA Genes in Genomic Sequences. In Gene Prediction; Methods in Molecular Biology; Humana Press: New York, NY, USA, 2019; Volume 1962, pp. 1–14. [Google Scholar] [CrossRef]
- Syberg-Olsen, M.J.; Garber, A.I.; Keeling, P.J.; McCutcheon, J.P.; Husnik, F. Pseudofinder: Detection of Pseudogenes in Prokaryotic Genomes. Mol. Biol. Evol. 2022, 39, msac153. [Google Scholar] [CrossRef] [PubMed]
- Cantalapiedra, C.P.; Hernández-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. eggNOG-mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Mol. Biol. Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef]
- Emms, D.M.; Kelly, S. OrthoFinder: Phylogenetic orthology inference for comparative genomics. Genome Biol. 2019, 20, 238. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Jolley, K.A.; Bray, J.E.; Maiden, M.C.J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018, 3, 124. [Google Scholar] [CrossRef]
- Marçais, G.; Delcher, A.L.; Phillippy, A.M.; Coston, R.; Salzberg, S.L.; Zimin, A. MUMmer4: A fast and versatile genome alignment system. PLoS Comput. Biol. 2018, 14, e1005944. [Google Scholar] [CrossRef]
- Phuklia, W.; Panyanivong, P.; Sengdetka, D.; Sonthayanon, P.; Newton, P.N.; Paris, D.H.; Day, N.P.J.; Dittrich, S. Novel high-throughput screening method using quantitative PCR to determine the antimicrobial susceptibility of Orientia tsutsugamushi clinical isolates. J. Antimicrob. Chemother. 2019, 74, 74–81. [Google Scholar] [CrossRef] [PubMed]
- Inthawong, M.; Sunyakumthorn, P.; Wongwairot, S.; Anantatat, T.; Dunachie, S.J.; Im-Erbsin, R.; Jones, J.W.; Mason, C.J.; Lugo, L.A.; Blacksell, S.D.; et al. A time-course comparative clinical and immune response evaluation study between the human pathogenic Orientia tsutsugamushi strains: Karp and Gilliam in a rhesus macaque (Macaca mulatta) model. PLoS Negl. Trop. Dis. 2022, 16, e0010611. [Google Scholar] [CrossRef]
- Thiriot, J.D.; Liang, Y.; Gonzales, C.; Sun, J.; Yu, X.; Soong, L. Differential cellular immune responses against Orientia tsutsugamushi Karp and Gilliam strains following acute infection in mice. PLoS Negl. Trop. Dis. 2023, 17, e0011445. [Google Scholar] [CrossRef]
- Hagen, R.; Verhoeve, V.I.; Gillespie, J.J.; Driscoll, T.P. Conjugative Transposons and Their Cargo Genes Vary across Natural Populations of Rickettsia buchneri Infecting the Tick Ixodes scapularis. Genome Biol. Evol. 2018, 10, 3218–3229. [Google Scholar] [CrossRef]
- Wu, M.; Sun, L.V.; Vamathevan, J.; Riegler, M.; Deboy, R.; Brownlie, J.C.; McGraw, E.A.; Martin, W.; Esser, C.; Ahmadinejad, N.; et al. Phylogenomics of the reproductive parasite Wolbachia pipientis wMel: A streamlined genome overrun by mobile genetic elements. PLoS Biol. 2004, 2, E69. [Google Scholar] [CrossRef]
- Fleshman, A.; Mullins, K.; Sahl, J.; Hepp, C.; Nieto, N.; Wiggins, K.; Hornstra, H.; Kelly, D.; Chan, T.C.; Phetsouvanh, R.; et al. Comparative pan-genomic analyses of Orientia tsutsugamushi reveal an exceptional model of bacterial evolution driving genomic diversity. Microb. Genom. 2018, 4, e000199. [Google Scholar] [CrossRef]
- Gillespie, J.J.; Joardar, V.; Williams, K.P.; Driscoll, T.; Hostetler, J.B.; Nordberg, E.; Shukla, M.; Walenz, B.; Hill, C.A.; Nene, V.M.; et al. A Rickettsia genome overrun by mobile genetic elements provides insight into the acquisition of genes characteristic of an obligate intracellular lifestyle. J. Bacteriol. 2012, 194, 376–394. [Google Scholar] [CrossRef]
- Adcox, H.E.; Hunt, J.R.; Allen, P.E.; Siff, T.E.; Rodino, K.G.; Ottens, A.K.; Carlyon, J.A. Orientia tsutsugamushi Ank5 promotes NLRC5 cytoplasmic retention and degradation to inhibit MHC class I expression. Nat. Commun. 2024, 15, 8069. [Google Scholar] [CrossRef]
- Evans, S.M.; Rodino, K.G.; Adcox, H.E.; Carlyon, J.A. Orientia tsutsugamushi uses two Ank effectors to modulate NF-κB p65 nuclear transport and inhibit NF-κB transcriptional activation. PLoS Pathog. 2018, 14, e1007023. [Google Scholar] [CrossRef]
- Bang, S.; Min, C.K.; Ha, N.Y.; Choi, M.S.; Kim, I.S.; Kim, Y.S.; Cho, N.H. Inhibition of eukaryotic translation by tetratricopeptide-repeat proteins of Orientia tsutsugamushi. J. Microbiol. 2016, 54, 136–144. [Google Scholar] [CrossRef]
- Fuxelius, H.H.; Darby, A.C.; Cho, N.H.; Andersson, S.G. Visualization of pseudogenes in intracellular bacteria reveals the different tracks to gene destruction. Genome Biol. 2008, 9, R42. [Google Scholar] [CrossRef] [PubMed]
- Blanc, G.; Ogata, H.; Robert, C.; Audic, S.; Suhre, K.; Vestris, G.; Claverie, J.M.; Raoult, D. Reductive genome evolution from the mother of Rickettsia. PLoS Genet. 2007, 3, e14. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Zhang, L.; Hu, H.; Liu, X.; Zhang, C.; Xin, Y.; Liu, B.; Chen, Z.; Xu, K.; Liu, Y. Complete genome sequencing and comparative genomic analyses of a new spotted-fever Rickettsia heilongjiangensis strain B8. Emerg. Microbes Infect. 2023, 12, 2153085. [Google Scholar] [CrossRef]
- Elliott, I.; Thangnimitchok, N.; de Cesare, M.; Linsuwanon, P.; Paris, D.H.; Day, N.P.J.; Newton, P.N.; Bowden, R.; Batty, E.M. Targeted capture and sequencing of Orientia tsutsugamushi genomes from chiggers and humans. Infect. Genet. Evol. 2021, 91, 104818. [Google Scholar] [CrossRef] [PubMed]



| No. | Strain Name | Geographic Location | Host Source | Collection Date | Release Date | Size (bp) | Gene * | CDS * | tRNA * | rRNA * | Pseudogene * | tsa56 Genotype # | Sequence Type (ST) | Reference |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HMU_001 | Hainan | Homo sapiens | 2023 | / | 1,895,724 | 2075 | 2038 | 34 | 3 | 765 | TA763_A | NA † | this study |
| 2 | JJOtsu7 | Vellore | Homo sapiens | 2024 | 22 August 2024 | 2,183,885 | 2462 | 2425 | 34 | 3 | 987 | Karp_A | 133 | unpublished |
| 3 | JJOtsu8 | Vellore | Homo sapiens | 2023 | 22 August 2024 | 2,344,138 | 2592 | 2554 | 35 | 3 | 914 | TA763_B | NA † | unpublished |
| 4 | JJOtsu5 | Vellore | Homo sapiens | 2023 | 22 August 2024 | 2,446,845 | 2746 | 2709 | 34 | 3 | 1019 | Karp_A | 135 | unpublished |
| 5 | JJOtsu6 | Vellore | Homo sapiens | 2023 | 22 August 2024 | 2,284,757 | 2523 | 2487 | 33 | 3 | 955 | Kato_A | 132 | unpublished |
| 6 | JJOtsu1 | Vellore | Homo sapiens | 2022 | 22 August 2024 | 2,179,789 | 2447 | 2410 | 34 | 3 | 963 | Kato_A | 132 | unpublished |
| 7 | Wuj/2014 | Zhejiang | Homo sapiens | 2014 | 25 September 2019 | 1,972,387 | 2148 | 2111 | 34 | 3 | 831 | Karp_A | NA † | unpublished |
| 8 | TW-1 | Taiwan | Homo sapiens | 2007 | 1 May 2024 | 2,008,429 | 2150 | 2113 | 34 | 3 | 806 | Karp_A | NA † | [25] Minahan NT et al. 2024. |
| 9 | TW-22 | Taiwan | Homo sapiens | 2007 | 1 May 2024 | 2,044,475 | 2315 | 2278 | 34 | 3 | 869 | Kato_A | NA † | [25] Minahan NT et al. 2024. |
| 10 | UT176 | Udon Thani | Homo sapiens | 2004 | 12 May 2018 | 1,932,116 | 2124 | 2086 | 35 | 3 | 838 | Karp_B | 10 | [22] Batty EM et al. 2018. |
| 11 | UT76 | Udon Thani | Homo sapiens | 2003 | 12 May 2018 | 2,078,193 | 2284 | 2247 | 34 | 3 | 863 | Karp_A | 1 | [22] Batty EM et al. 2018. |
| 12 | Boryong | South Korea | Homo sapiens | 1980s | 15 May 2007 | 2,127,051 | 2480 | 2443 | 34 | 3 | 1088 | Boryong/Karp | 48 | [26] Cho NH et al. 2007. |
| 13 | Ikeda | Japan | Homo sapiens | 1979 | 30 May 2008 | 2,008,987 | 2222 | 2185 | 34 | 3 | 882 | JG_A/Gilliam | 49 | [27] Nakayama K et al. 2008. |
| 14 | TA686 | Thailand | Tupaia glis | 1963 | 12 May 2018 | 2,254,553 | 2583 | 2546 | 34 | 3 | 1122 | Shimokoshi | NA † | [22] Batty EM et al. 2018. |
| 15 | Kato | Niigata | Homo sapiens | 1955 | 12 May 2018 | 2,319,449 | 2444 | 2406 | 35 | 3 | 863 | Kato_B | 20 | [22] Batty EM et al. 2018. |
| 16 | Karp | New Guinea | Homo sapiens | 1943 | 12 May 2018 | 2,469,803 | 2615 | 2578 | 34 | 3 | 943 | Karp_C | 45 | [22] Batty EM et al. 2018. |
| 17 | Gilliam | Indian-Burmese border | Homo sapiens | 1943 | 12 May 2018 | 2,465,012 | 2746 | 2709 | 34 | 3 | 1030 | Gilliam | 46 | [22] Batty EM et al. 2018. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Niu, Y.; Guo, Y.; Xu, Z.; Chen, S.; Zhang, L.; Cui, X.; Lin, D.; Yuen, K.-Y.; Chan, J.F.-W.; Tang, C.; et al. A Compact Closed Genome of Orientia tsutsugamushi from Hainan Island, China Provides a TA763_A Reference and Reveals Repeat-Driven Remodeling. Pathogens 2026, 15, 318. https://doi.org/10.3390/pathogens15030318
Niu Y, Guo Y, Xu Z, Chen S, Zhang L, Cui X, Lin D, Yuen K-Y, Chan JF-W, Tang C, et al. A Compact Closed Genome of Orientia tsutsugamushi from Hainan Island, China Provides a TA763_A Reference and Reveals Repeat-Driven Remodeling. Pathogens. 2026; 15(3):318. https://doi.org/10.3390/pathogens15030318
Chicago/Turabian StyleNiu, Yi, Yijia Guo, Zhao Xu, Siqi Chen, Liyuan Zhang, Xiuji Cui, Dachuan Lin, Kwok-Yung Yuen, Jasper Fuk-Woo Chan, Chuanning Tang, and et al. 2026. "A Compact Closed Genome of Orientia tsutsugamushi from Hainan Island, China Provides a TA763_A Reference and Reveals Repeat-Driven Remodeling" Pathogens 15, no. 3: 318. https://doi.org/10.3390/pathogens15030318
APA StyleNiu, Y., Guo, Y., Xu, Z., Chen, S., Zhang, L., Cui, X., Lin, D., Yuen, K.-Y., Chan, J. F.-W., Tang, C., & Yin, F. (2026). A Compact Closed Genome of Orientia tsutsugamushi from Hainan Island, China Provides a TA763_A Reference and Reveals Repeat-Driven Remodeling. Pathogens, 15(3), 318. https://doi.org/10.3390/pathogens15030318








