4. Discussion
Our study represents a pioneering effort as the first study in India to utilize shotgun metagenomics to comprehensively map the bacterial microbiota and associated resistome in hospitalized sick cirrhosis patients. By integrating this high-resolution metagenomic data with detailed immunophenotyping (host response) and clinical outcomes, we delineated the specific microbial and genetic drivers of inflammation, immune exhaustion, and mortality in this high-risk population. In cirrhosis progression, the gut microbiome undergoes a pathological shift termed “dysbiosis.” This state is characterized by a reduction in beneficial autochthonous taxa, particularly from the families
Lachnospiraceae and
Ruminococcaceae (Class Clostridia), which are crucial for producing short-chain fatty acids like butyrate that maintain barrier integrity and immune homeostasis. In their place, there is a massive expansion of potentially pathogenic “pathobionts,” predominantly from the families
Enterobacteriaceae and
Enterococcaceae. This phenomenon, often described as the “Gram-negative bloom,” is clinically catastrophic [
9].
A pivotal finding in our cohort is the predominance of MASLD (55.1%) as the leading etiology of hospitalized decompensated cirrhosis, surpassing ALD. This mirrors the rapid epidemiological transition reported in South Asia, where the adoption of Westernized diets and sedentary lifestyles has accelerated the prevalence of metabolic liver disease. The distinct microbiome signatures we observed in MASLD patients [enrichment of tet(D) and catA1] versus ALD patients (enrichment of QnrB4) suggest that etiology drives distinct resistome acquisition pathways, likely influencing response to prophylactic antibiotics.
The divergent resistome profiles between MASLD and ALD patients may reflect fundamentally different pathways of resistance gene acquisition. In ALD, the predominance of QnrB4 (a plasmid-mediated quinolone resistance gene typically associated with Klebsiella and other Enterobacteriaceae) may be driven by the synergistic effects of alcohol on the gut barrier: ethanol directly increases intestinal permeability, promotes small intestinal bacterial overgrowth, and creates a bile acid milieu that selects for Gram-negative pathobionts intrinsically carrying quinolone resistance plasmids. Furthermore, ALD patients are frequently hospitalized with recurrent episodes requiring repeated antibiotic courses, creating sustained selective pressure favouring quinolone-resistant clones. In contrast, the enrichment of tet(D) and catA1 (tetracycline and chloramphenicol resistance, respectively) in MASLD may reflect a different ecological niche: the metabolic environment of insulin resistance, hyperglycaemia, and altered bile acid composition characteristic of MASLD selects for a distinct microbial community. Tetracycline resistance genes are commonly found in agricultural and dietary microbiota, and the dietary patterns associated with metabolic syndrome (high-fat, processed foods) may introduce or select for organisms carrying these environmental resistance determinants. These etiology-specific resistome profiles have direct implications for prophylactic antibiotic selection, suggesting that norfloxacin prophylaxis may be particularly ineffective in ALD patients harboring QnrB4-enriched flora.
In our cohort of 78 hospitalized Indian patients with cirrhosis characterized by high disease severity and long-term mortality, the results revealed a profound gut microbiome dysbiosis marked by a significant expansion of Bacilli and Gammaproteobacteria, specifically Enterobacteriaceae and Enterococcaceae, in subjects with advanced hepatic dysfunction and complications. This pathological shift, driven by the enrichment of pathobionts such as Klebsiella pneumoniae, Escherichia coli, and Enterococcus faecium, mirrored profiles of systemic inflammation (elevated IL-6, TNF-alpha) and immune paralysis (low HLA-DR). Resistome profiling demonstrated that while a background of tetracycline and macrolide resistance was pervasive, critical illness and poor outcomes (including mortality, infection, and ACLF presentations) were distinctly characterized by the enrichment of the quinolone resistance gene QnrB4 and multidrug efflux pumps like efmA. Notably, QnrB4 expression plausibly correlated with a pro-inflammatory cytokine storm and immune paresis, whereas tet(D) displayed a complex duality, associated with both hepatic decompensation markers and patient survival.
The microbial landscape in Indian hospitals differs significantly from the West. While Western ICUs often struggle with Vancomycin-Resistant
Enterococci (VRE) and
Clostridioides difficile, the Indian landscape is dominated by Gram-negative bacteria, particularly
Klebsiella pneumoniae,
Escherichia coli, and
Acinetobacter baumannii, which exhibit alarmingly high rates of resistance to third-generation cephalosporins (up to 84%) and carbapenems (over 47%). These organisms frequently carry potent resistance determinants such as
blaNDM-1 (New Delhi Metallo-beta-lactamase) and various
blaOXA carbapenemase. Consequently, when bacterial translocation occurs in Indian patients with cirrhosis, it often involves organisms that are resistant to standard empiric antibiotics (e.g., ceftriaxone or piperacillin-tazobactam), leading to rapid treatment failure, clinical deterioration, and high mortality [
10]. The 12–24-month mortality rate in our study was substantially high at 43.6%, with 38.5% of patients requiring direct ICU admission. This high mortality rate underscores the fact that the physiological reserve was depleted, and the window for therapeutic intervention is narrow. The distribution of complications, particularly the high rates of ascites and infection, points toward our cohort as one with severe portal hypertension and significant immune paresis.
Notably, elevated neutrophil CD64 expression (a validated marker of neutrophil activation in sepsis) was observed in some patients who did not have clinically or microbiologically confirmed infections (i.e., culture-negative). This apparent paradox likely reflects a state of ‘sterile’ systemic inflammation, wherein subclinical bacterial translocation from the gut (passage of viable bacteria or bacterial products such as LPS and bacterial DNA across the compromised intestinal barrier) triggers innate immune activation without progressing to overt bacteremia detectable by standard blood cultures. In cirrhosis, this phenomenon is well-described: portal hypertensive enteropathy and increased intestinal permeability allow continuous low-grade translocation of gut-derived PAMPs that activate neutrophils and monocytes via toll-like receptors, resulting in elevated nCD64, raised inflammatory cytokines, and clinical features mimicking sepsis in the absence of a positive culture. This ‘culture-negative inflammatory state’ is clinically significant because it may progress to overt sepsis and contributes to the hemodynamic instability characteristic of decompensated cirrhosis.
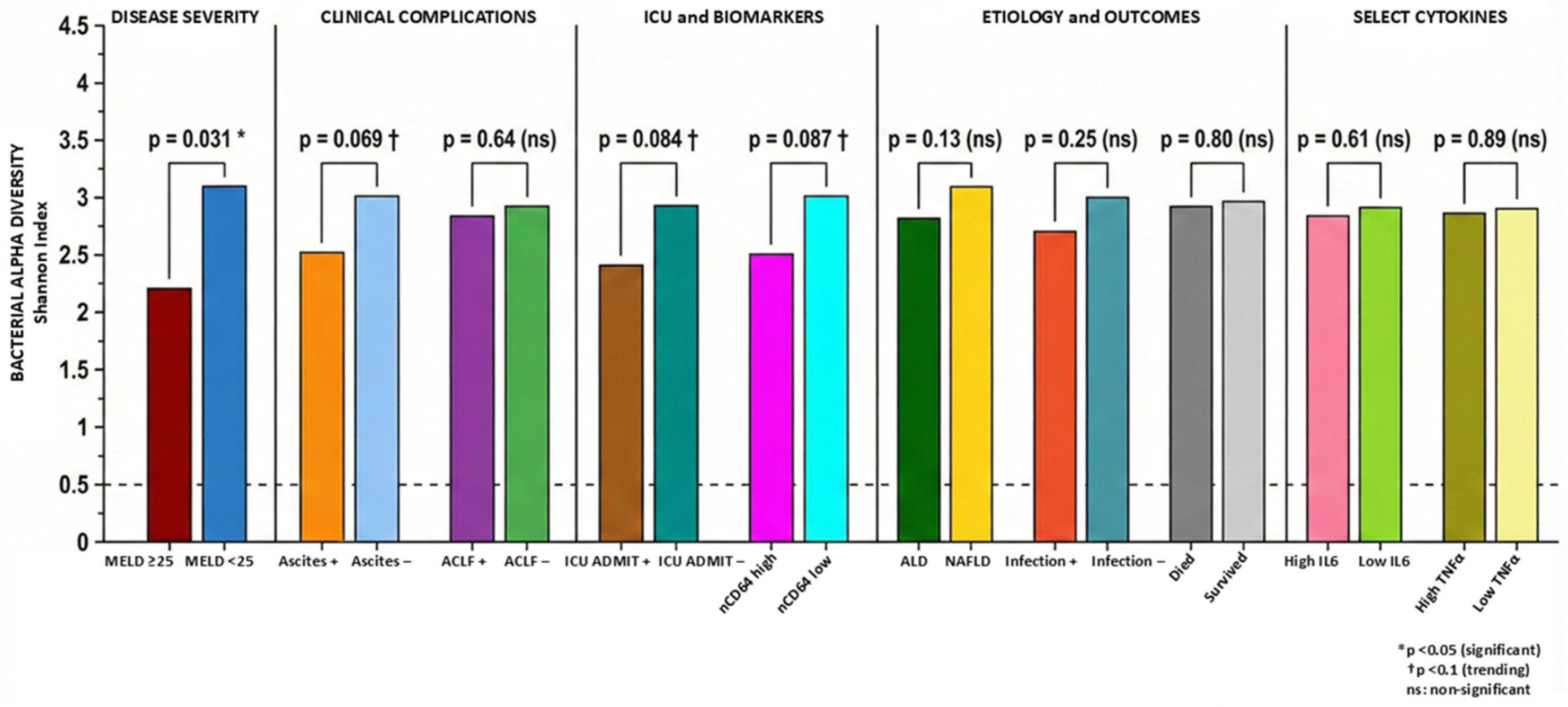
While high MELD scores (≥25) were associated with significantly reduced Shannon (alpha) diversity, several clinically important subgroups including ACLF showed no significant difference. These finding warrants explanation. First, dysbiosis in cirrhosis may manifest primarily as compositional shifts (changes in which taxa dominate) rather than overall diversity collapse. The Shannon Index captures both richness and evenness but may miss clinically relevant compositional changes when pathobiont expansion occurs alongside retained (but suppressed) commensals. Our data suggest that even in ACLF, patients retain detectable commensal populations while experiencing pathobiont expansion—a pattern of ‘partial dysbiosis’ rather than complete community collapse. Second, with only 15 ACLF patients (19.2% of cohort), statistical power to detect modest diversity differences was limited in our study. Third, ACLF encompasses heterogeneous precipitating events (infection, GI bleeding, hepatotoxins) that may have variable effects on microbiome diversity, potentially attenuating overall group differences. Thus, beta-diversity metrics examining community composition differences, rather than alpha-diversity, may be more informative for differentiating clinical phenotypes in this population.
Similarly, ICU admission, ascites, and elevated nCD64 showed only trending associations with reduced Shannon diversity that did not reach statistical significance. ICU admission is a heterogeneous endpoint reflecting various acute indications (GI bleeding, encephalopathy, respiratory failure) with potentially divergent microbiome effects. The near-significant trend for ascites supports a relationship with reduced diversity, though our binary classification (present/absent) rather than severity grading may have attenuated the signal. For nCD64, dichotomization of a continuous marker at the median may have reduced discriminatory power. Importantly, the absence of statistically significant alpha-diversity differences does not exclude important compositional differences. Indeed, differential abundance analyses revealed significant taxonomic and resistome differences in these groups, suggesting that community structure changes meaningfully even when overall diversity metrics are preserved.
The expansion of
Enterobacteriaceae in our cohort with severe disease is consistent with the established literature documenting this phenomenon in cirrhosis. While the expansion of
Enterobacteriaceae is a known hallmark of cirrhosis, our use of shotgun metagenomics allowed for the identification of species-level associations that provide new mechanistic insights. We identified
Sutterella sp. AM11-39 as a possible marker for ascites and higher levels of inflammation. Sutterella species are generally considered commensals with mild pro-inflammatory potential. However, our findings align with animal models of cirrhosis where
Sutterella translocation to ascitic fluid was linked to the onset of decompensation. This suggests
Sutterella may possess specific bile-resistance traits allowing it to survive the altered bile acid milieu of cirrhotic patients, acting as a driver of peritoneal inflammation [
11].
The association of
Clostridium sp. C5-48 with severe decompensation and jaundice is a critical finding. This organism has been recently reclassified as
Enterocloster alcoholdehydrogenati, a bacterium capable of producing high levels of carcinogenic acetaldehyde from ethanol. Its enrichment in our severe cohort could suggest a mechanism for “abstinential progression,” where the gut microbiota continues to generate hepatotoxic metabolites endogenously, perpetuating liver injury even in the absence of alcohol consumption [
12].
Our study is among the first in India to map the gut resistome in decompensated cirrhosis, revealing a high burden of
QnrB4,
OXA-833, and
efmA. The possible correlation between
QnrB4 (plasmid-mediated quinolone resistance) and mortality, ACLF, and immune paralysis (low HLA-DR) was striking.
QnrB4 is often co-located on multidrug-resistance plasmids with other virulence factors. Its presence likely signifies colonization by high-risk MDR clones (e.g.,
Klebsiella ST37 or ST11) that are endemic in India. The correlation with immune paralysis supports the hypothesis that chronic endotoxin exposure from these specific MDR pathobionts drives monocytes into a state of exhaustion (endotoxin tolerance), rendering the host defenseless against secondary infections [
13,
14].
The specific elevation of the carbapenemase gene
OXA-
833 in patients with GI bleeding is a novel observation.
OXA-
833 is an emerging resistance gene in South Asia, often found in
Proteus and
Klebsiella species [
15]. This probably suggests that the nutrient-rich environment created by intraluminal blood (iron and protein) may exert a selective pressure favouring specific proteolytic, carbapenem-resistant organisms, complicating prophylactic antibiotic strategies in bleeders.
The multidrug efflux pump gene
efmA was linked to Enterococcus faecium abundance. Interestingly, it was enriched in milder disease states (Child B) compared to genes like
ErmX but was universally upregulated in non-survivors. The
efmA is an intrinsic major facilitator superfamily (MFS) efflux pump in
E. faecium that confers low-level resistance to fluoroquinolones and macrolides. The omnipresence of
efmA in
E. faecium strains confirms that the expanding
Enterococcus populations in these patients are intrinsically equipped to survive antibiotic pressure [
16]. The shift from
efmA dominance to other genes like
ErmX or
tet(O) in specific complications (like HE) suggests a dynamic restructuring of the resistome under the selective pressure of different treatment regimens (e.g., rifaximin, which targets RNA polymerase but may select for other resistance mechanisms via plasmid co-selection).
The identification of Enterococcus faecium as a key pathobiont in our cohort warrants specific discussion, given that certain strains of this species are widely used as probiotics in commercially available formulations. It is essential to recognize that the species E. faecium encompasses genetically distinct lineages with fundamentally different clinical implications. Probiotic E. faecium strains typically belong to commensal-associated Clade B (e.g., ST296, ST67), which lack the virulence and resistance gene repertoire of hospital-adapted strains. In contrast, the E. faecium populations expanding in our decompensated cirrhosis patients were characterized by the enrichment of the efmA multidrug efflux pump, a hallmark of the hospital-adapted Clade A1 lineage (e.g., ST17, ST18, ST78) that carries intrinsic resistance to fluoroquinolones and macrolides. The co-existence of efmA enrichment with elevated TNF-alpha and markers of immune paralysis in our cohort suggests colonization by these pathogenic lineages rather than probiotic-type commensals. This distinction has critical clinical implications: the empirical use of E. faecium-containing probiotics in patients with advanced cirrhosis may potentially and inadvertently introduce organisms capable of acquiring vancomycin resistance (vanA/vanB) via horizontal gene transfer from co-resident pathobionts, potentially worsening the resistome burden. We therefore opine that clinicians exercise caution regarding probiotic formulations containing E. faecium in decompensated cirrhosis, pending species- and strain-level safety evaluation in this population.
Hepatic encephalopathy was associated with the highest intensity of AMR gene expression, specifically the lincosamide (e.g., clindamycin) resistance gene
lnuB.
lnuB encodes a lincosamide nucleotidyltransferase and is often found on mobile genetic elements (transposons and plasmids) in
Streptococcus and
Enterococcus species [
17]. Overt HE treatment relies heavily on rifaximin (a rifamycin) and lactulose. While rifaximin resistance is a concern, the expansion of
lnuB (which confers resistance to clindamycin/lincomycin) might be a collateral marker of the specific dysbiosis associated with HE (e.g.,
Streptococcus salivarius or
Enterococcus overgrowth). It indicates that the gut flora in HE is not just ammoniagenic but also a dense reservoir of mobile resistance elements that could be transferred to pathogens. Additionally, HE linked to the highest AMR gene expression may also be due to several other factors: severe portal hypertension and portosystemic shunting in HE lead to reduced hepatic clearance of gut bacteria, while a distinct dysbiosis with urease-producing, ammonia-generating organisms could be common. Hepatic encephalopathy patients also have prolonged antibiotic exposure within hospitalizations, fostering resistant strains and gene transfer. Additionally, slower gut motility allows more time for plasmid and resistance gene exchange. Together, these factors possibly create a “resistance amplification niche,” accounting for the high AMR gene burden seen in HE.
Our study differentiated between the tetracycline resistance genes,
tet(M) and
tet(O), revealing distinct clinical associations. The
tet(M) gene expression abundance was associated with milder disease (Child B), survival, and higher regenerative markers (EGF/VEGF). The
tet(M) is a ribosomal protection protein often found in commensals like Firmicutes. Its presence in healthier patients likely reflects a preserved commensal flora (which naturally carries some resistance) rather than a pathogen bloom. Conversely,
tet(O) was associated with severe disease (Child C, ACLF), ALD, and mortality. The
tet(O) gene is frequently associated with pathogens like
Campylobacter and
Streptococcus [
10]. The switch from
tet(M) to
tet(O) may represent a “resistome shift” from commensal-driven resistance to pathogen-driven resistance as cirrhosis progresses.
Our study provided compelling evidence for a direct mechanistic link between the gut resistome and host immune function. High AMR genes burden was not a silent passenger but correlated satisfactorily with systemic inflammation (high IL-6, TNF-alpha) and immune paralysis (low HLA-DR, high Sepsis Index). Our work also highlighted that not all dysbiosis was created equal and that different pathobionts drive different states within the systemic inflammation. For example, we observed that
Ligilactobacillus salivarius abundance was significantly linked to IFN-gamma and IL-6, suggesting a Th1/Th17-type inflammatory response. This aligns with the potential for
L. salivarius to act as an opportunistic pathogen driving cellular immunity. Similarly,
Enterococcus faecium was ubiquitous in high TNF-alpha states.
E. faecium lipoteichoic acid is a potent inducer of TNF-alpha via TLR2, contributing to the hemodynamic instability (vasodilation) seen in decompensated patients [
18,
19].
While this exploratory study requires prospective validation, several findings have potential clinical implications. First, the high burden of quinolone resistance genes (QnrB4) documented in our cohort suggests that norfloxacin, the current standard for spontaneous bacterial peritonitis prophylaxis, may be deleterious in patients with advanced decompensation. Second, the association of QnrB4 with mortality and immune paralysis suggests potential utility as a prognostic biomarker; pending validation, stool-based resistome screening could identify patients requiring early ICU triage. Third, the association of Ligilactobacillus salivarius with pro-inflammatory cytokines (IL-6, IFN-gamma) challenges empirical use of generic Lactobacillus-containing probiotics in decompensated cirrhosis, warranting species-specific safety evaluation. Fourth, the enrichment of OXA-833 (carbapenemase) in patients with gastrointestinal bleeding suggests that standard ceftriaxone prophylaxis may be insufficient in this subgroup. Finally, the identification of specific pathobionts (Clostridium sp. C5-48, Enterococcus faecium) provides targets for precision microbiome interventions including bacteriophage therapy or targeted faecal microbiota transplantation.
From a translational perspective, the feasibility and cost of resistome-based screening merit discussion. Current shotgun metagenomic sequencing, as employed in our study, costs approximately USD 150–300 per sample (including DNA extraction, library preparation, and sequencing on Illumina platforms), with turnaround times of 5–7 days, which limits its utility for acute clinical decision-making. However, rapid technological advances are narrowing this gap. Targeted amplicon-based panels for specific resistance genes (such as QnrB4, blaOXA, and blaNDM) using real-time PCR can provide results within 4–6 h at a fraction of the cost (USD 30–50 per panel). In resource-limited settings like India, a tiered approach may be most practical: targeted PCR panels for high-risk resistance genes (QnrB4, OXA-833) as a rapid bedside screen, with comprehensive shotgun metagenomics reserved for research cohorts or patients with refractory infections. The cost-effectiveness of such screening relative to the economic burden of treating MDR infections in ICU settings (estimated at USD 5000–15,000 per episode in Indian tertiary centers) warrants formal health economic evaluation.
This study’s primary strength lies in its integration of shotgun metagenomics with deep immunophenotyping in an Indian cohort, allowing us to link specific resistome elements like QnrB4 to immune states such as HLA-DR expression and revealing region-specific patterns distinct from Western cohorts. The detailed clinical stratification further enabled the identification of niche-specific associations, such as the unique resistome signature of GI bleeding versus hepatic encephalopathy. However, limitations include a modest sample size (n = 78) that restricts multivariate adjustments, a cross-sectional design that precludes causal inference regarding the resistome role in decompensation, and the analysis of luminal rather than mucosal microbiota. Dietary factors are important modulators of gut microbiome composition. While all patients were on oral/enteral nutrition at sample collection, detailed nutritional assessment (caloric intake, macronutrient distribution, fiber content) was not performed. Future studies should incorporate dietary questionnaires or food frequency data to assess diet-microbiome interactions in this population. Additionally, while current antibiotic use was recorded, the retrospective quantification of lifetime antibiotic exposure remains a challenge. It is imperative to note that the etiological distribution in our cohort (MASLD 55.1%, ALD 38.5%, with only 3.8% viral hepatitis) reflects the contemporary epidemiological landscape of cirrhosis at tertiary care centers in South India, where successful antiviral treatment programs have reduced viral hepatitis as a cause of decompensated cirrhosis. However, this limits generalizability to populations where viral hepatitis predominates. Some findings, such as Enterobacteriaceae expansion and resistome burden, likely relate to portal hypertension and gut barrier dysfunction common across etiologies, while others (e.g., Clostridium sp. C5-48 associations with ALD) may be etiology-specific. Validation in viral hepatitis cohorts is warranted before broader generalization.
Additionally, several potential confounders of the resistome were incompletely captured. While current antibiotic use (within 7 days of admission) and history of prior SBP were documented, we lack systematic data on: hospitalizations within the preceding 90 days, prior drug-resistant colonization/infection status, cumulative duration of antibiotic prophylaxis, and socioeconomic status. The omission of these factors is a limitation, as they may act as hidden drivers of the resistome. For instance, patients with frequent prior hospitalizations (unmeasured) may have a higher baseline load of multidrug-resistant organisms due to nosocomial exposure, potentially inflating the association between resistance genes and disease severity. Similarly, socioeconomic factors could influence environmental exposure to resistance determinants. Furthermore, due to the modest sample size relative to the high dimensionality of metagenomic data, we did not perform multivariate regression adjusting for age or these unmeasured confounders to avoid overfitting. Consequently, the associations reported herein should be interpreted as descriptive of the clinical phenotype rather than causally isolated.
An important consideration in interpreting our resistome findings is that the presence of an antimicrobial resistance gene in metagenomic data does not automatically confer phenotypic resistance. Our study detected DNA (gene presence) rather than RNA (gene expression), and several factors determine whether detected genes result in functional resistance: (1) Gene expression—many ARGs may be present but transcriptionally silent or expressed below clinically relevant levels; (2) Genetic context—ARGs require appropriate promoters, regulatory elements, and in some cases accessory genes for functional expression; truncated genes or those lacking proper regulatory contexts may not confer resistance; (3) Host organism—an ARG in a non-pathogenic commensal may have different clinical implications than the same gene in a pathogen, though horizontal gene transfer potential remains a concern; (4) Gene variants—some variants of resistance genes may have reduced or absent functional activity. However, the genes we identified (QnrB4, OXA-833, efmA, tetracycline resistance genes) are well-characterized resistance determinants with established functional roles documented in CARD and peer-reviewed literature. Furthermore, the clinical correlations observed—QnrB4 with mortality in patients likely receiving prophylactic quinolones, OXA-833 with GI bleeding where cephalosporin prophylaxis is standard—provide circumstantial support for functional relevance. Nevertheless, phenotypic susceptibility testing of cultured clinical isolates would provide definitive confirmation, representing an inherent limitation of purely metagenomic approaches. Therefore, the resistance genes identified in this study (QnrB4, OXA-833, efmA) should be interpreted primarily as markers of colonization by resistant flora rather than as direct evidence of phenotypic resistance in bloodstream infections. Their clinical correlations, while biologically plausible, require prospective validation through paired metagenomic and phenotypic susceptibility testing of cultured isolates.
Furthermore, given that differential abundance testing was performed across 30 clinical comparisons without formal multiple-comparison correction, the risk of false-positive associations is non-trivial. Although a dual threshold of nominal p < 0.05 combined with log2 fold change ≥ 1 was employed to mitigate this risk, some of the reported gene-level associations may represent chance findings. Accordingly, readers should interpret individual associations—particularly those involving genes with borderline significance or smaller effect sizes—with appropriate caution. Prospective validation studies with pre-specified hypotheses, adequate sample sizes, and formal FDR correction are warranted before any of these associations are considered for clinical application.
These findings pave the way for resistome-based risk stratification, where rapid screening for genes like QnrB4 could guide early ICU transfer or antibiotic escalation. The identification of specific pathobionts such as Clostridium sp. C5-48 and Enterococcus faecium highlights the potential for precision microbiome editing via bacteriophage therapy to reduce toxin and AMR burdens. Moreover, the high burden of quinolone resistance calls for revised prophylaxis guidelines, potentially replacing norfloxacin with non-antibiotic alternatives like Lachnospiraceae-enriched Faecal Microbiota Transplantation (FMT) or dietary interventions targeting metabolic substrates to restore colonization resistance.