Cell Wall Proteome of Candida albicans Reveals Proteins Associated with Tolerance to Antibiofilm Activity of a Lippia graveolens Kunth Stem Extract
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethical Aspects
2.2. Collection of Plant Material
2.3. Extract Preparation
2.4. Strains and Culture Conditions
2.5. Biofilm Formation Capacity of Clinical Isolates of C. albicans
2.6. Evaluation of the Effect of L. graveolens Kunth Stem Extract on the Planktonic Growth of C. albicans
2.7. Evaluation of the Effect of L. graveolens Kunth Stem Extract on Biofilm Formation in C. albicans Isolates
2.8. Evaluation of the Effect of L. graveolens Kunth Stem Extract on Cell Wall Proteins of C. albicans 2400H Isolate
2.8.1. Cell Lysis and Protein Extraction
2.8.2. Sample Preparation and Liquid Chromatography Coupled to Mass Spectrometry (LC-MS) Analysis
2.8.3. Protein Database Search and Protein Quantification
2.9. Evaluation of the Effect of the Stem Extract on the Expression Profile of the FOB64_005423, ALS3, TSA1, and RBT1 Genes
2.10. Statistical Analysis
3. Results
3.1. Biofilm Formation Capacity in C. albicans Isolates
3.2. Effect of L. graveolens Kunth Stem Extract on the Planktonic Growth of Isolates of C. albicans
3.3. Effect of L. graveolens Kunth Stem Extract on Biofilm Formation in C. albicans 2400H Isolate
3.4. Effect of L. graveolens Kunth Stem Extract in Cell Wall Proteins of C. albicans 2400H Isolate
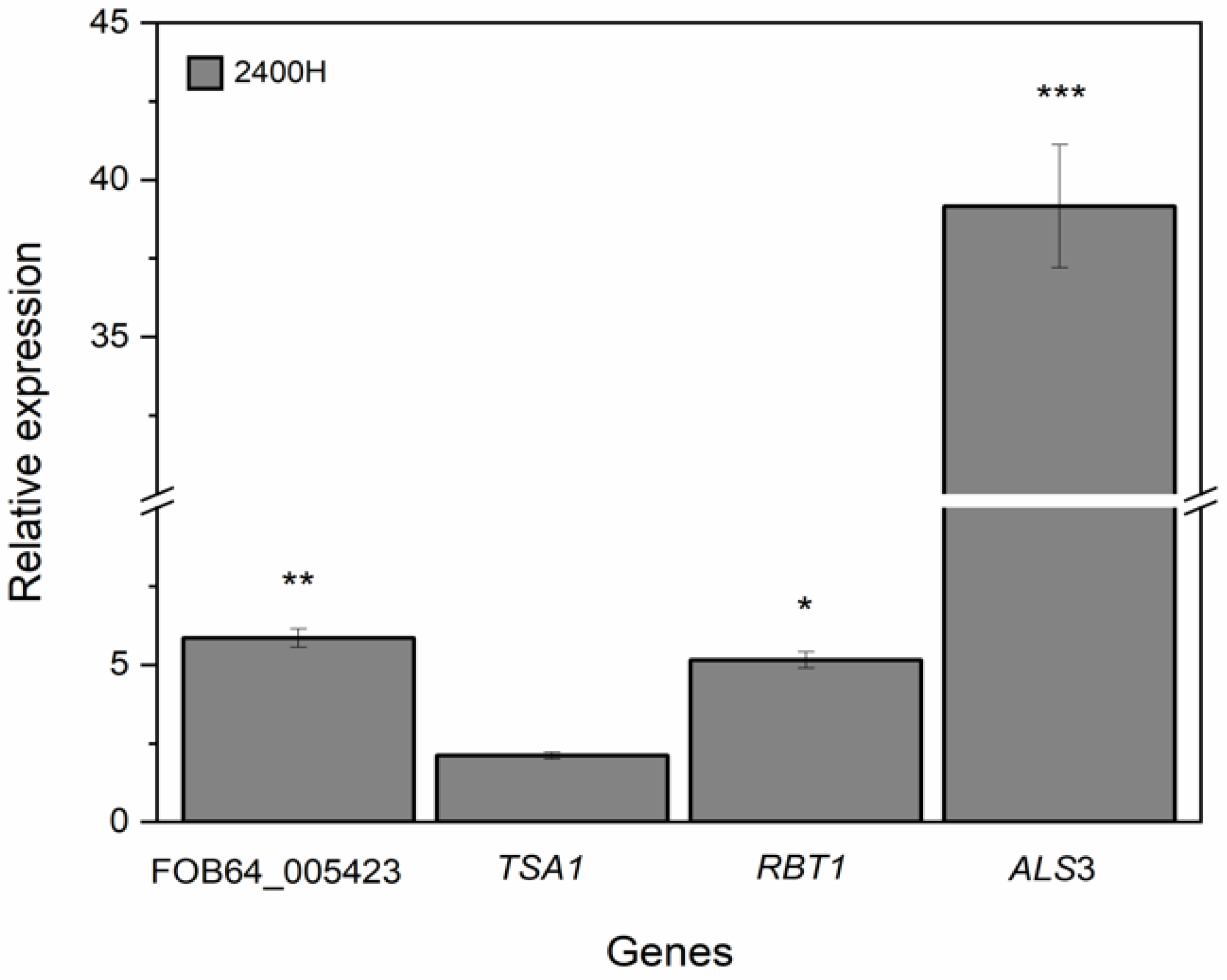
3.5. Expression Levels of FOB64_005423, RBT1, ALS3, and TSA1 Genes of C. albicans 2400H Isolate Exposed to L. graveolens Kunth Stem Extract
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pu, S.; Niu, S.; Zhang, C.; Xu, X.; Qin, M.; Huang, S.; Zhang, L. Epidemiology, antifungal susceptibilities, and risk factors for invasive candidiasis from 2011 to 2013 in a teaching hospital in southwest China. J. Microbiol. Immunol. Infect. 2017, 50, 97–103. [Google Scholar] [CrossRef]
- Cavalheiro, M.; Teixeira, M.C. Candida biofilms: Threats, challenges, and promising strategies. Front. Med. 2018, 5, 28. [Google Scholar] [CrossRef]
- Finkel, J.S.; Mitchell, A.P. Genetic control of Candida albicans biofilm development. Nat. Rev. Microbiol. 2011, 9, 109–118. [Google Scholar] [CrossRef]
- Pereira, R.; dos Santos-Fontenelle, R.O.; de Brito, E.H.S.; de Morais, S.M. Biofilm of Candida albicans: Formation, regulation and resistance. J. Appl. Microbiol. 2021, 131, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Nobile, C.J.; Johnson, A.D. Candida albicans biofilms and human disease. Annu. Rev. Microbiol. 2015, 69, 71–92. [Google Scholar] [CrossRef]
- Ji, H.F.; Li, X.J.; Zhang, H.Y. Natural products and drug discovery. EMBO Rep. 2009, 10, 194–200. [Google Scholar] [CrossRef]
- Abdul-Wahab, S.M.; Jantan, I.; Haque, M.A.; Arshad, L. Exploring the leaves of Annona muricata L. as a source of potential anti-inflammatory and anticancer agents. Front. Pharmacol. 2018, 9, 661. [Google Scholar] [CrossRef]
- Hadisaputri, Y.E.; Habibah, U.; Abdullah, F.F.; Halimah, E.; Mutakin, M.; Megantara, S.; Diantini, A. Antiproliferation activity and apoptotic mechanism of soursop (Annona muricata L.) leaves extract and fractions on MCF7 breast cancer cells. BCTT 2021, 13, 447–457. [Google Scholar] [CrossRef] [PubMed]
- Cordoba, S.; Vivot, W.; Szusz, W.; Albo, G. Antifungal activity of essential oils against Candida species isolated from clinical samples. Mycopathologia 2019, 184, 615–623. [Google Scholar] [CrossRef]
- Ruiz-Duran, J.; Torres, R.; Stashenko, E.E.; Ortiz, C. Antifungal and antibiofilm activity of Colombian essential oils against different Candida strains. Antibiotics 2023, 12, 668. [Google Scholar] [CrossRef] [PubMed]
- Funari, C.S.; Gullo, F.P.; Napolitano, A.; Carneiro, R.L.; Mendes-Giannini, M.J.S.; Fusco-Almeida, A.M.; Piacente, S.; Pizza, C.; Silva, D.H.S. Chemical and antifungal investigations of six lippia species (Verbenaceae) from Brazil. Food Chem. 2012, 135, 2086–2094. [Google Scholar] [CrossRef] [PubMed]
- Prado, J.C.S.; de Aguiar, F.L.L.; Prado, G.M.; do Nascimento, J.F.; de Souza, N.V.; Barbosa, F.C.B.; Lima, D.M.; Rodrigues, T.H.S.; Bessa, N.U.C.; Abreu, F.O.M.S.; et al. Development and characterization of nanoemulsions containing Lippia origanoides Kunth essential oil and their antifungal potential against Candida albicans. J. Appl. Microbiol. 2024, 135, Ixae271. [Google Scholar] [CrossRef]
- Ghasempour, M.; Omran, S.M.; Moghadamnia, A.A.; Shafiee, F. Effect of aqueous and ethanolic extracts of Lippia citriodora on Candida albicans. Electron. Physician 2008, 8, 2752. [Google Scholar] [CrossRef] [PubMed]
- Pozatti, P.; Scheid, L.A.; Spader, T.B.; Atayde, M.L.; Santurio, J.M.; Alves, S.H. In vitro activity of essential oils extracted from plants used as spices against fluconazole-resistant and fluconazole-susceptible Candida spp. Can. J. Microbiol. 2008, 54, 950–956. [Google Scholar] [CrossRef]
- Freire, I.A.; Bueno-Silva, B.; Galvao, L.C.D.C.; Duarte, M.C.T.; Sartoratto, A.; Figueira, G.M.; de Alencar, S.M.; Rosalen, P.L. The effect of essential oils and bioactive fractions on Streptococcus mutans and Candida albicans biofilms; a confocal analysis. Evid. Based. Complement. Alternat. Med. 2015, 1, 871316. [Google Scholar]
- Gutiérrez-Grijalva, E.P.; Antunes-Ricardo, M.; Acosta-Estrada, B.A.; Gutiérrez-Uribe, J.A.; Heredia, J.B. Cellular antioxidant activity and in vitro inhibition of α-glucosidase, α-amylase and pancreatic lipase of oregano polyphenols under simulated gastrointestinal digestion. Food Res. Int. 2019, 116, 676–686. [Google Scholar]
- Frías-Zepeda, M.E.; Rosales-Castro, M. Effect of extraction conditions on the concentration of phenolic compounds in Mexican oregano (Lippia graveolens Kunth) residues. Rev. Chapingo Ser. Cienc. For. Ambiente 2021, 27, 368–381. [Google Scholar]
- Morshedloo, M.R.; Alireza, S.S.; Nazeri, V.; Maggi, F.; Craker, L. Essential oil profile of oregano (Origanum vulgare L.) populations grown under similar soil and climate conditions. Ind. Crop. Prod. 2018, 119, 183–190. [Google Scholar] [CrossRef]
- Loaiza-Oliva, M.; Morales-Uchima, S.M.; Puerta-Suárez, J.; Mesa-Arango, A.C.; Martínez Pabón, M.C. Lippia origanoides derivatives in vitro evaluation on polymicrobial biofilms: Streptococcus mutans, Lactobacillus rhamnosus and Candida albicans. Arch. Oral Biol. 2023, 148, 105656. [Google Scholar] [CrossRef]
- Cabral-Miramontes, J.P.; Martínez-Rocha, A.L.; Rosales-Castro, M.; López-Rodríguez, A.; Meneses-Morales, I.; Del Campo-Quinteros, E.; Herrera-Ocelotl, K.K.; Gándara-Moreno, G.; Velázquez-Huizar, S.J.; Ibarra-Sánchez, L.; et al. Antifungal Activity of Mexican Oregano (Lippia graveolens Kunth) Extracts from Industrial Waste Residues on Fusarium spp. in Bean Seeds (Phaseolus vulgaris L.). Agriculture 2024, 14, 1975. [Google Scholar] [CrossRef]
- Frías-Zepeda, M.E.; Rosales-Castro, M.; Escalona-Cardoso, G.N.; Paniagua-Castro, N. Ethanolic extract of Lippia graveolens stem reduce biochemical markers in a murine model with metabolic syndrome. Saudi J. Biol. Sci. 2022, 29, 103422. [Google Scholar] [CrossRef]
- Frías-Zepeda, M.E.; Ibarra-Berumen, J.; Ordaz-Pichardo, C.; Rosales-Castro, M. Cytotoxic activity of ethanolic extracts of Lippia graveolens HBK leaves and stem against lung cancer cell line SK-LU-1. Blacpma 2022, 21, 646–653. [Google Scholar] [CrossRef]
- Secretaría de Salud. Reglamento de la ley General de Salud en Materia de Investigación Para la Salud. Diario Oficial de la Federación, México 2014. Available online: https://www.dof.gob.mx/ (accessed on 12 February 2023).
- White, T.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. Acad. Pre. Inc. 1990, 38, 315–322. [Google Scholar]
- Nikawa, H.; Jin, C.; Makihira, S.; Egusa, H.; Hamada, T.; Kumagai, H. Biofilm formation of Candida albicans on the surfaces of deteriorated soft denture lining materials caused by denture cleansers in vitro. J. Oral Rehabil. 2003, 30, 243–250. [Google Scholar] [CrossRef]
- Shrief, R.; Zaki, M.E.S.; El-Sehsah, E.M.; Ghaleb, S.; Mofreh, M. Study of antifungal susceptibility, virulence genes and biofilm formation in Candida albicans. Open Microbiol. J. 2019, 13, 241–248. [Google Scholar] [CrossRef]
- Ruiz-Baca, E.; Leyva-Sánchez, H.; Calderón-Barraza, B.; Esquivel-Naranjo, U.; López-Romero, E.; López-Rodríguez, A.; Cuéllar-Cruz, M. Identification of proteins in Sporothrix schenckii sensu stricto in response to oxidative stress induced by hydrogen peroxide. Rev. Iberoam. Micol. 2019, 36, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Félix-Contreras, C.; Alba-Fierro, C.A.; Ríos-Castro, E.; Luna-Martínez, F.; Cuéllar-Cruz, M.; Ruiz-Baca, E. Proteomic analysis of Sporothrix schenckii cell wall reveals proteins involved in oxidative stress response induced by menadione. Microb. Pathog. 2020, 141, 103987. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Wisniewski, J.R.; Zougman, A.; Nagaraj, N.; Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 2009, 6, 359–362. [Google Scholar] [CrossRef] [PubMed]
- Erde, J.; Loo, R.R.O.; Loo, J.A. Enhanced FASP (eFASP) to increase proteome coverage and sample recovery for quantitative proteomic experiments. J. Proteome Res. 2014, 13, 1885–1895. [Google Scholar] [CrossRef] [PubMed]
- Ríos-Castro, E.; Souza, G.H.M.F.; Delgadillo-Álvarez, D.M.; Ramírez-Reyes, L.; Torres-Huerta, A.L.; Velasco-Suárez, A.; Tapia-Ramírez, J. Quantitative proteomic analysis of MARC-145 cells infected with a Mexican porcine reproductive and respiratory syndrome virus strain using a label-free based DIA approach. J. Am. Soc. Mass Spectrom. 2020, 31, 1302–1312. [Google Scholar] [CrossRef]
- Li, H.; Rose, M.J.; Tran, L.; Zhang, J.; Miranda, L.P.; James, C.A.; Sasu, B.J. Development of a method for the sensitive and quantitative determination of hepcidin in human serum using LC-MS/MS. J. Pharmacol. Toxicol. Methods 2009, 59, 171–180. [Google Scholar] [CrossRef]
- Silva, J.C.; Gorenstein, M.V.; Li, G.Z.; Vissers, J.P.; Geromanos, S.J. Absolute quantification of proteins by LCMSE: A virtue of parallel MS acquisition* S. Mol. Cell. Proteom. 2006, 5, 144–156. [Google Scholar] [CrossRef]
- Arnaud-Franco, G.; Ríos-Castro, E.; Velasco-Suárez, A.; García-de León, F.J.; Beltrán, L.F.; Carbajal-Saucedo, A. Venom comparisons of endemic and micro-endemic speckled rattlesnakes Crotalus mitchellii, C. polisi and C. thalassoporus from Baja California Peninsula. Toxicon 2023, 224, 107030. [Google Scholar] [CrossRef]
- Li, Q.Q.; Skinner, J.; Bennett, J.E. Evaluation of reference genes for real-time quantitative PCR studies in Candida glabrata following azole treatment. BMC Mol. Biol. 2012, 13, 22. [Google Scholar] [CrossRef]
- Liu, H. Transcriptional control of dimorphism in Candida albicans. Curr. Opin. Microbiol. 2001, 4, 728–735. [Google Scholar] [CrossRef]
- Buakaew, W. Proteomic analysis reveals proteins involved in the mode of action of β-citronellol identified from citrus hystrix DC. leaf against Candida albicans. Front. Microbiol. 2022, 13, 894637. [Google Scholar] [CrossRef]
- Kim, Y.G.; Lee, J.H.; Park, S.; Khadke, S.K.; Shim, J.J.; Lee, J. Hydroquinones including tetrachlorohydroquinone inhibit Candida albicans biofilm formation by repressing hyphae-related genes. Microbiol. Spectr. 2022, 10, e0253622. [Google Scholar] [CrossRef]
- Tan, J.; Zhang, Z.; Zheng, D.; Mu, Y.; Cao, B.; Yang, J.; Han, L.; Huang, X. Structure-activity relationship and biofilm formation-related gene targets of oleanolic acid-type saponins from Pulsatilla chinensis against Candida albicans. Bioorg. Chem. 2024, 146, 107311. [Google Scholar] [CrossRef]
- Li, L.; Wei, M.P.; Yu, H.; Xie, Y.F.; Guo, Y.H.; Cheng, Y.L. Antifungal activity of Sapindus saponins against Candida albicans: Interruption of biofilm formation. J. Herb. Med. 2023, 42, 100776. [Google Scholar] [CrossRef]
- Norouzi, N.; Alizadeh, F.; Khodavani, A.; Jahangiri, M. Antifungal activity of menthol alone and in combination on growth inhibition and biofilm formation of Candida albicans. J. Herb. Med. 2021, 29, 100495. [Google Scholar] [CrossRef]
- Willaert, R.G. Adhesins of yeasts: Protein structure and interactions. J. Fungi 2018, 4, 119. [Google Scholar] [CrossRef]
- Willaert, R.G.; Kayacan, Y.; Devreese, B. The Flo adhesin family. Pathogens 2021, 10, 1397. [Google Scholar] [CrossRef]
- Gutiérrez-Escobedo, G.; Hernández-Carreón, O.; Morales-Rojano, B.; Revuelta-Rodríguez, B.; Vázquez-Franco, N.; Castaño, I.; De Las Peñas, A. Candida glabrata peroxiredoxins, Tsa1 and Tsa2, and sulfiredoxin, Srx1, protect against oxidative damage and are necessary for virulence. Fungal Genet. Biol. 2020, 135, 103287. [Google Scholar] [CrossRef]
- Ramírez-Quijas, M.D.; López-Romero, E.; Cuéllar-Cruz, M. Proteomic analysis of cell wall in four pathogenic species of Candida exposed to oxidative stress. Microb. Pathog. 2015, 87, 1–12. [Google Scholar] [CrossRef]
- Ksouri, S.; Djebir, S.; Bentorki, A.A.; Gouri, A.; Hadef, Y.; Benakhla, A. Antifungal activity of essential oils extract from Origanum floribundum Munby, Rosmarinus officinalis L. and Thymus ciliatus Desf. against Candida albicans isolated from bovine clinical mastitis. J. Mycol. Med. 2017, 27, 245–249. [Google Scholar]
- Cortés-Chitala, M.D.C.; Flores-Martínez, H.; Orozco-Ávila, I.; León-Campos, C.; Suárez-Jacobo, Á.; Estarrón-Espinosa, M.; López-Muraira, I. Identification and quantification of phenolic compounds from Mexican oregano (Lippia graveolens HBK) hydroethanolic extracts and evaluation of its antioxidant capacity. Molecules 2021, 26, 702. [Google Scholar] [CrossRef]
- Herrera-Rodríguez, S.E.; López-Rivera, R.J.; García-Márquez, E.; Estarrón-Espinoza, M.; Espinoza-Andrews, H. Mexican Oregano (Lippia graveolens) essential oil-in-water emulsions: Impact of emulsifier type on the antifungal activity of Candida albicans. Food Sci. Biotechnol. 2018, 28, 441–448. [Google Scholar]
- Gil, G.A.; Kakuda, L.; Tonani, L.; von Zeska Kress, M.R.; Oliveira, W.P. Surfactant-driven effects on the antifungal activity of Lippia origanoides Kunth essential oil encapsulated in lipid-based nanosystems. ACS Omega 2025, 10, 7876–7887. [Google Scholar]
- Singh, R.; Kumari, A.; Kaur, K.; Kaur, R. Relevance of antifungal penetration in biofilm-associated resistance of Candida albicans and non-albicans Candida species. J. Med. Microbiol. 2018, 67, 922–926. [Google Scholar] [CrossRef]
- Muthamil, S.; Prasath, K.G.; Priya, A.; Precille, P.; Pandian, S.K. Global proteomic analysis deciphers the mechanism of action of plant derived oleic acid against Candida albicans virulence and biofilm formation. Sci. Rep. 2022, 10, 5113. [Google Scholar] [CrossRef]
- Ortiz-Ramírez, J.A.; Cuéllar-Cruz, M.; López-Romero, E. Cell compensatory responses of fungi to damage of the cell wall induced by calcofluor white and congo red with emphasis on Sporothrix schenckii and Sporothrix globosa. A review. Front. Cell. Infect. Microbiol. 2022, 12, 976924. [Google Scholar] [CrossRef]
- Iyer, K.R.; Robbins, N.; Cowen, L.E. The role of Candida albicans stress response pathways in antifungal tolerance and resistance. iScience 2022, 25, 103953. [Google Scholar] [CrossRef]




| Biofilm Forming | Isolate | Number of Isolates | Percentage (%) |
|---|---|---|---|
| High | 22400H, 1887H, 2127H, and 332H | 4 | 22.2 |
| Moderate | 1573H, 920H, 3468H, 2420H, 2517H, and 1670H | 6 | 33.3 |
| Low | ATCC 10231, 2948H, 2422H, and 727H | 3 | 16.6 |
| Non-forming | 316H, 389H, 701H, 1620H, and 2749H | 5 | 27.9 |
| Biofilm Forming | Isolates | Extract-MIC90 (mg/mL) | Fluconazole-MIC90 (µg/mL) |
|---|---|---|---|
| High | 2400H | 16 | <0.125 |
| High | 1887H | 16 | 0.125 |
| High | 2127H | 16 | <0.125 |
| High | 332H | 16 | 0.125 |
| Moderate | 1573H | 16 | 0.25 |
| Moderate | 920H | 16 | 0.25 |
| Moderate | 3468H | 16 | <0.125 |
| Moderate | 2420H | 16 | 0.125 |
| Moderate | 2517H | 16 | <0.125 |
| Moderate | 1670H | 16 | 0.5 |
| Low | ATCC 10231 | 16 | 0.25 |
| Uniprot No. Access | ENA Gene ID | Protein Putative Function | Log2 Fold Change | p |
|---|---|---|---|---|
| A0A8H6F377 | FOB64_004736 | Lysophospholipase | 2.50 | 0.0000 |
| A0A8H6F4R1 | RBT1 | Cell wall protein RBT1 | 2.32 | 0.0006 |
| A0A8H6BX16 | CHT3 | Chitinase 3 | 1.41 | 0.0003 |
| A0A8H6BUX2 | GAM1 | Glucoamylase 1 | 1.40 | 0.0000 |
| A0A8H6BYS4 | ALS3 | Agglutinin-like protein 3 | 1.39 | 0.0000 |
| A0A8H6BUN5 | NCP1 | NADPH-cytochrome P450 reductase | 1.17 | 0.0007 |
| A0A8H6BT95 | FOB64_005423 | Flocculin type 3 repeat family protein | 1.14 | 0.0062 |
| A0A8H6BQY9 | FOB64_006659 | Thioredoxin reductase | 1.02 | 0.0000 |
| A0A8H6BYJ9 | FOB64_004041 | Ribosomal protein S10 | 0.97 | 0.0073 |
| A0A8H6BXS5 | FOB64_002993 | Nascent polypeptide-associated complex subunit beta | 0.93 | 0.0004 |
| A0A8H6BQW6 | CHT2 | Chitinase 2 | 0.902 | 0.0399 |
| A0A8H6F6X4 | FOB64_000863 | Translationally controlled tumor protein homolog | 0.867 | 0.0030 |
| A0A8H6F124 | MP65 | Cell surface mannoprotein MP65 domain protein | 0.83 | 0.0057 |
| A0A8H6F3U2 | FOB64_002340 | Arginase | 0.82 | 0.0057 |
| A0A8H6BWF6 | FOB64_004094 | Phosphotransferase | 0.81 | 0.0301 |
| A0A8H6F501 | ADO1 | Adenosine kinase | 0.80 | 0.0004 |
| A0A8H6F4L7 | TSA1 | Peroxiredoxin TSA1-A | 0.77 | 0.0220 |
| A0A8H6F3P9 | VPS21 | Vacuolar protein sorting-associated protein 21 | 0.75 | 0.0029 |
| A0A8H6BXS5 | FOB64_002993 | Nascent polypeptide-associated complex subunit alpha | 0.72 | 0.0073 |
| A0A8H6BZX4 | FOB64_001976 | Glycerol-3-phosphate dehydrogenase [NAD(+)] | 0.72 | 0.0108 |
| A0A8H6F2S3 | HSP78 | Heat shock protein 78_mitocondrial | 0.70 | 0.0074 |
| A0A8H6BU09 | FOB64_004742 | Adenine phosphoribosyltransferase | 0.68 | 0.0064 |
| A0A8H6BQY7 | swoH | Nucleoside diphosphate kinase | 0.60 | 0.0257 |
| A0A8H6C397 | FOB64_002507 | 14-3-3 domain-containing protein | 0.60 | 0.0029 |
| A0A8H6C3Z6 | RPL19B | Ribosomal protein L19 | −0.59 | 0.0098 |
| A0A8H6F362 | FOB64_003210 | 6-phosphogluconate dehydrogenase | −0.59 | 0.0082 |
| A0A8H6BUX0 | IPP1 | Inorganic diphosphatase | −0.64 | 0.0073 |
| A0A8H6F4H4 | RPL4B | 60S ribosomal protein L4-B | −0.66 | 0.0027 |
| A0A8H6BZ37 | RPS6A | 40S ribosomal protein S6 | −0.68 | 0.0008 |
| A0A8H6BYQ8 | PGI1 | Glucose-6-phosphate isomerase | −0.75 | 0.0004 |
| A0A8H6C486 | MET6 | 5-methyltetrahydropteroyltriglutamate--homocysteine S-methyltransferase | −0.77 | 0.0000 |
| A0A8H6F4K9 | FOB64_002438 | Ribosomal protein L10 | −0.97 | 0.0015 |
| A0A8H6BRV2 | FOB64_006313 | Phosphoglycerate mutase | −1.05 | 0.0019 |
| A0A8H6BU78 | FOB64_006449 | Cytochrome P450 | −1.11 | 0.0003 |
| A0A8H6F012 | FOB64_005613 | Methionine adenosyltransferase | −1.16 | 0.0141 |
| A0A8H6F3P4 | FOB64_004035 | Aconitate hydratase_mitocondrial | −1.31 | 0.0042 |
| A0A8H6BVM4 | FOB64_006072 | Catalase family protein | −1.34 | 0.0001 |
| A0A8H6BWI4 | FOB64_005268 | Flavodoxin-like domain-containing protein | −1.69 | 0.0085 |
| A0A8H6C4N6 | FDH1 | Formate dehydrogenase | −1.91 | 0.0003 |
| A0A8H6BZT5 | SOD5 | Cell surface Cu-only superoxide dismutase 5 | −2.40 | 0.0002 |
| Name | Function | Log2 Fold Change | References |
|---|---|---|---|
| Cell wall protein, Rbt1 | GPI-anchored cell wall protein required for virulence, mating efficiency, biofilm formation, and dimorphism. | 2.32 | [37,38,39] |
| Agglutinin-like protein 3, Als3 | Crucial role in the adhesion stage, biofilm formation, and dimorphism. | 1.39 | [37,40,41,42,43,44] |
| Flocculin type 3 repeat family protein, FOB64_005423 | Adhesion of the pathogen to host tissue cells or abiotic surfaces such as catheters and implants. | 1.14 | [43,44] |
| Peroxiredoxin Tsa1 | Ability to protect biomolecules from oxidative damage. | 0.77 | [45,46] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
García-Núñez, A.; Martínez-Rocha, A.L.; Alba-Fierro, C.A.; Ríos-Fránquez, F.J.; Cabral-Miramontes, J.P.; Frías-Zepeda, M.E.; Escobedo-Bretado, M.Á.; Ruiz-Baca, E. Cell Wall Proteome of Candida albicans Reveals Proteins Associated with Tolerance to Antibiofilm Activity of a Lippia graveolens Kunth Stem Extract. Pathogens 2026, 15, 216. https://doi.org/10.3390/pathogens15020216
García-Núñez A, Martínez-Rocha AL, Alba-Fierro CA, Ríos-Fránquez FJ, Cabral-Miramontes JP, Frías-Zepeda ME, Escobedo-Bretado MÁ, Ruiz-Baca E. Cell Wall Proteome of Candida albicans Reveals Proteins Associated with Tolerance to Antibiofilm Activity of a Lippia graveolens Kunth Stem Extract. Pathogens. 2026; 15(2):216. https://doi.org/10.3390/pathogens15020216
Chicago/Turabian StyleGarcía-Núñez, Alejandra, Ana Lilia Martínez-Rocha, Carlos Antonio Alba-Fierro, Francisco Javier Ríos-Fránquez, Juan Pablo Cabral-Miramontes, María Estela Frías-Zepeda, Miguel Ángel Escobedo-Bretado, and Estela Ruiz-Baca. 2026. "Cell Wall Proteome of Candida albicans Reveals Proteins Associated with Tolerance to Antibiofilm Activity of a Lippia graveolens Kunth Stem Extract" Pathogens 15, no. 2: 216. https://doi.org/10.3390/pathogens15020216
APA StyleGarcía-Núñez, A., Martínez-Rocha, A. L., Alba-Fierro, C. A., Ríos-Fránquez, F. J., Cabral-Miramontes, J. P., Frías-Zepeda, M. E., Escobedo-Bretado, M. Á., & Ruiz-Baca, E. (2026). Cell Wall Proteome of Candida albicans Reveals Proteins Associated with Tolerance to Antibiofilm Activity of a Lippia graveolens Kunth Stem Extract. Pathogens, 15(2), 216. https://doi.org/10.3390/pathogens15020216





