Detection of HIV-1 Resistance Mutations to Antiretroviral Therapy and Cell Tropism in Russian Patients Using Next-Generation Sequencing
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinical Samples
2.2. Extraction and Viral Load Quantification
2.3. Obtaining Consensus Sequences of the HIV-1 pol and env Genes
2.4. HIV-1 Subtyping Using pol Gene Sequences
2.5. Analysis of Drug Resistance Mutations in HIV-1 pol Gene
2.6. Analysis of Consensus Sequences of the HIV-1 env Gene and CXCR4 Cell Tropism Prediction
2.7. Statistical Analysis
3. Results
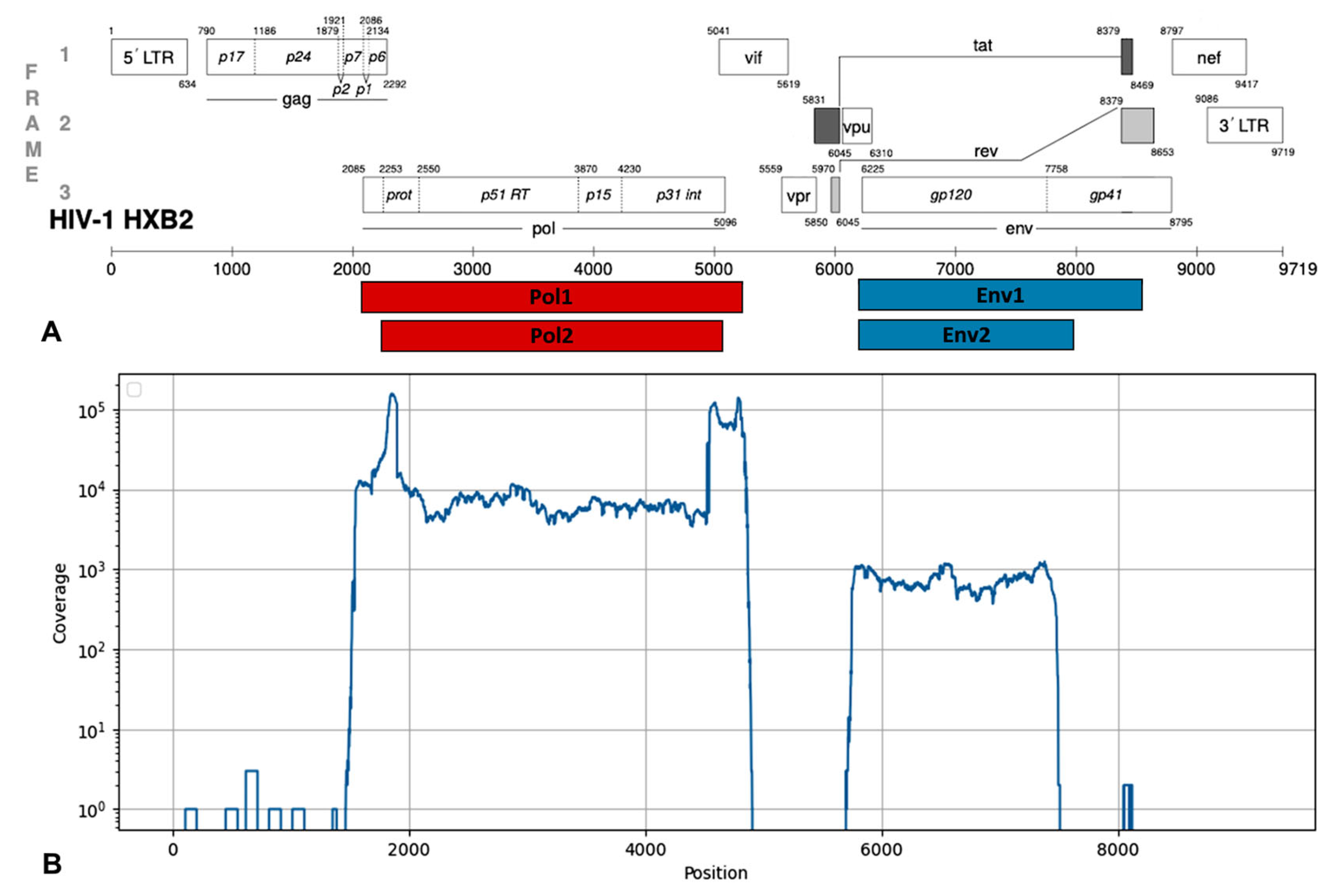
3.1. Testing of the NGS Protocol for HIV-1 pol and env Genes
3.2. Subtyping of HIV-1 Viruses
3.3. Analysis of Drug Resistance Mutations
3.4. Cell Co-Receptor Tropism Prediction
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Keller, S.C.; Yehia, B.R.; Eberhart, M.G.; Brady, K.A. Accuracy of definitions for linkage to care in persons living with HIV. J. Acquir. Immune Defic. Syndr. 2013, 63, 622–630. [Google Scholar] [CrossRef][Green Version]
- Samji, H.; Cescon, A.; Hogg, R.S.; Modur, S.P.; Althoff, K.N.; Buchacz, K.; Burchell, A.N.; Cohen, M.; Gebo, K.A.; Gill, M.J.; et al. Closing the gap: Increases in life expectancy among treated HIV-positive individuals in the United States and Canada. PLoS ONE 2013, 8, e81355. [Google Scholar] [CrossRef] [PubMed]
- Ogarkova, D.; Antonova, A.; Kuznetsova, A.; Adgamov, R.; Pochtovyi, A.; Kleimenov, D.; Tsyganova, E.; Gushchin, V.; Gintsburg, A.; Mazus, A. Current Trends of HIV Infection in the Russian Federation. Viruses 2023, 15, 2156. [Google Scholar] [CrossRef] [PubMed]
- National Virological Association, Moscow Oncology Society. Clinical guidelines. HIV Infection in Adults. Rubricator of Clinical Guidelines. 2024. Available online: https://cr.minzdrav.gov.ru/clin-rec/79_2 (accessed on 20 October 2025).
- Ahmed, I.; Tefera, F.; Bekele, A.; Ayalew, J.; Tessema, F.; Abera, G.; Ahmed, J.; Mekonnen, A.; Haile, A.; Yohannes, F.; et al. Suboptimal adherence to antiretroviral treatment and its predictors among people living with HIV in the era of test and treat. Sci. Rep. 2025, 15, 12666. [Google Scholar] [CrossRef] [PubMed]
- de Los Rios, P.; Okoli, C.; Punekar, Y.; Allan, B.; Muchenje, M.; Castellanos, E.; Richman, B.; Corbelli, G.M.; Hardy, W.D.; Young, B.; et al. Prevalence, determinants, and impact of suboptimal adherence to HIV medication in 25 countries. Prev. Med. 2020, 139, 106182. [Google Scholar] [CrossRef]
- Kirichenko, A.; Kireev, D.; Lapovok, I.; Shlykova, A.; Lopatukhin, A.; Pokrovskaya, A.; Bobkova, M.; Antonova, A.; Kuznetsova, A.; Ozhmegova, E.; et al. HIV-1 Drug Resistance among Treatment-Naïve Patients in Russia: Analysis of the National Database, 2006–2022. Viruses 2023, 15, 991. [Google Scholar] [CrossRef]
- Monakhov, N.E.; Sharipova, E.A.; Lapunova, S.V.; Vinogradova, T.N.; Lioznov, D.A. Acquired drug resistance of HIV in patients with virological failure of first-line art. HIV Infect. Immunosuppr. Disord. 2025, 17, 42–53. (In Russian) [Google Scholar] [CrossRef]
- Kapustin, D.V.; Nalimova, T.M.; Ekushov, V.E.; Kriklivaya, N.P.; Halikov, M.R.; Krasnova, E.I.; Khokhlova, N.I.; Demchenko, S.V.; Pozdnaykova, L.L.; Sivay, M.V.; et al. Patterns of HIV-1 drug resistance among HIV-infected patients receiving first-line antiretroviral therapy in Novosibirsk Region, Russia. J. Glob. Antimicrob. Resist. 2023, 35, 1–5. [Google Scholar] [CrossRef]
- Kirichenko, A.A.; Kireev, D.E.; Shlykova, A.V.; Lopatukhin, A.E.; Lapovok, I.A.; Saleeva, D.V.; Kravchenko, A.V.; Pokrovsky, V.V. HIV-1 drug resistance in patients with virological inefficiency on ART in Russia in 2013–2021. Epidemiol. Infect. Dis. 2021, 11, 53–62. (In Russian) [Google Scholar] [CrossRef]
- Wensing, A.M.; Calvez, V.; Ceccherini-Silberstein, F.; Charpentier, C.; Günthard, H.F.; Paredes, R.; Shafer, R.W.; Richman, D.D. 2022 update of the drug resistance mutations in HIV-1. Top. Antivir. Med. 2022, 30, 559–574. [Google Scholar]
- Puertas, M.C.; Ploumidis, G.; Ploumidis, M.; Fumero, E.; Clotet, B.; Walworth, C.M.; Petropoulos, C.J.; Martinez-Picado, J. Pan-resistant HIV-1 emergence in the era of integrase strand-transfer inhibitors: A case report. Lancet Microbe 2020, 1, e130–e135. [Google Scholar] [CrossRef]
- Mazzuti, L.; Melengu, T.; Falasca, F.; Calabretto, M.; Cella, E.; Ciccozzi, M.; Mezzaroma, I.; Iaiani, G.; Spaziante, M.; d’Ettorre, G.; et al. Transmitted drug resistance mutations and trends of HIV-1 subtypes in treatment-naïve patients: A single-centre experience. J. Glob. Antimicrob. Resist. 2020, 20, 298–303. [Google Scholar] [CrossRef] [PubMed]
- Larder, B. Mechanisms of HIV-1 drug resistance. AIDS 2001, 15, S27–S34. [Google Scholar] [CrossRef] [PubMed]
- Carr, A.; Mackie, N.E.; Paredes, R.; Ruxrungtham, K. HIV drug resistance in the era of contemporary antiretroviral therapy: A clinical perspective. Antivir. Ther. 2023, 28, 13596535231201162. [Google Scholar] [CrossRef] [PubMed]
- Finzi, D.; Hermankova, M.; Pierson, T.; Carruth, L.M.; Buck, C.; Chaisson, R.E.; Quinn, T.C.; Chadwick, K.; Margolick, J.; Brookmeyer, R.; et al. Identification of a reservoir for HIV-1 in patients on highly active antiretroviral therapy. Science 1997, 278, 1295–1300. [Google Scholar] [CrossRef]
- Bruner, K.M.; Wang, Z.; Simonetti, F.R.; Bender, A.M.; Kwon, K.J.; Sengupta, S.; Fray, E.J.; Beg, S.A.; Antar, A.A.R.; Jenike, K.M.; et al. A quantitative approach for measuring the reservoir of latent HIV-1 proviruses. Nature 2019, 566, 120–125. [Google Scholar] [CrossRef]
- Kulpa, D.A.; Paiardini, M.; Silvestri, G. Immune-mediated strategies to solving the HIV reservoir problem. Nat. Rev. Immunol. 2025, 25, 542–553. [Google Scholar] [CrossRef]
- Turriziani, O.; Andreoni, M.; Antonelli, G. Resistant viral variants in cellular reservoirs of human immunodeficiency virus infection. Clin. Microbiol. Infect. 2010, 16, 1518–1524. [Google Scholar] [CrossRef] [PubMed]
- Manyana, S.; Gounder, L.; Pillay, M.; Manasa, J.; Naidoo, K.; Chimukangara, B. HIV-1 Drug Resistance Genotyping in Resource Limited Settings: Current and Future Perspectives in Sequencing Technologies. Viruses 2021, 13, 1125. [Google Scholar] [CrossRef]
- Nannyonjo, M.; Omooja, J.; Bugembe, D.L.; Bbosa, N.; Lunkuse, S.; Nabirye, S.E.; Nassolo, F.; Namagembe, H.; Abaasa, A.; Kazibwe, A.; et al. Next-Generation Sequencing Reveals a High Frequency of HIV-1 Minority Variants and an Expanded Drug Resistance Profile Among Individuals on First-Line ART. Viruses 2024, 16, 1454. [Google Scholar] [CrossRef]
- Gaitan, N.C.; D’Antoni, M.L.; Acosta, R.K.; Gianella, S.; Little, S.J.; Chaillon, A. Brief Report: Comparative Analysis of Pre-existing HIV Drug Resistance Mutations in Proviral DNA Using Next-Generation Sequencing and Routine HIV RNA Genotyping. J. Acquir. Immune Defic. Syndr. 2023, 93, 213–218. [Google Scholar] [CrossRef]
- Zhang, C.; Li, H.; Poluektova, L.Y.; Gendelman, H.E.; Dash, P.K. Unique molecular signatures in rebound viruses from antiretroviral drug and CRISPR-treated HIV-1-infected humanized mice. Commun. Biol. 2025, 8, 1077. [Google Scholar] [CrossRef] [PubMed]
- Ávila-Ríos, S.; Parkin, N.; Swanstrom, R.; Paredes, R.; Shafer, R.; Ji, H.; Kantor, R. Next-Generation Sequencing for HIV Drug Resistance Testing: Laboratory, Clinical, and Implementation Considerations. Viruses 2020, 12, 617. [Google Scholar] [CrossRef] [PubMed]
- Weber, J.; Volkova, I.; Sahoo, M.K.; Tzou, P.L.; Shafer, R.W.; Pinsky, B.A. Prospective Evaluation of the Vela Diagnostics Next-Generation Sequencing Platform for HIV-1 Genotypic Resistance Testing. J. Mol. Diagn. 2019, 21, 961–970. [Google Scholar] [CrossRef] [PubMed]
- Raymond, S.; Nicot, F.; Abravanel, F.; Minier, L.; Carcenac, R.; Lefebvre, C.; Harter, A.; Martin-Blondel, G.; Delobel, P.; Izopet, J. Performance evaluation of the Vela Dx Sentosa next-generation sequencing system for HIV-1 DNA genotypic resistance. J. Clin. Virol. 2020, 122, 104229. [Google Scholar] [CrossRef]
- Wymant, C.; Blanquart, F.; Golubchik, T.; Gall, A.; Bakker, M.; Bezemer, D.; Croucher, N.J.; Hall, M.; Hillebregt, M.; Ong, S.H.; et al. Easy and accurate reconstruction of whole HIV genomes from short-read sequence data with shiver. Virus Evol. 2018, 4, vey007. [Google Scholar] [CrossRef]
- Chen, S. fastp 1.0: An ultra-fast all-round tool for FASTQ data quality control and preprocessing. iMeta 2025, 4, e70078. [Google Scholar] [CrossRef]
- Li, D.; Liu, C.M.; Luo, R.; Sadakane, K.; Lam, T.W. MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 2015, 31, 1674–1676. [Google Scholar] [CrossRef]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar] [CrossRef]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve years of SAMtools and BCFtools. GigaScience 2021, 10, giab008. [Google Scholar] [CrossRef]
- Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef] [PubMed]
- Grubaugh, N.D.; Gangavarapu, K.; Quick, J.; Matteson, N.L.; De Jesus, J.G.; Main, B.J.; Tan, A.L.; Paul, L.M.; Brackney, D.E.; Grewal, S.; et al. An amplicon-based sequencing framework for accurately measuring intrahost virus diversity using PrimalSeq and iVar. Genome Biol. 2019, 20, 8. [Google Scholar] [CrossRef] [PubMed]
- Struck, D.; Lawyer, G.; Ternes, A.M.; Schmit, J.C.; Bercoff, D.P. COMET: Adaptive context-based modeling for ultrafast HIV-1 subtype identification. Nucleic Acids Res. 2014, 42, e144. [Google Scholar] [CrossRef] [PubMed]
- Shafer, R.W. Rationale and uses of a public HIV drug-resistance database. J. Infect. Dis. 2006, 194, S51–S58. [Google Scholar] [CrossRef]
- Aksamentov, I.; Roemer, C.; Hodcroft, E.B.; Neher, R.A. Nextclade: Clade assignment, mutation calling and quality control for viral genomes. J. Open Source Softw. 2021, 6, 3773. [Google Scholar] [CrossRef]
- HIVDB Genotypic Resistance Interpretation Program. Available online: https://github.com/hivdb/sierra (accessed on 23 December 2025).
- Bennett, D.E.; Camacho, R.J.; Otelea, D.; Kuritzkes, D.R.; Fleury, H.; Kiuchi, M.; Heneine, W.; Kantor, R.; Jordan, M.R.; Schapiro, J.M.; et al. Drug resistance mutations for surveillance of transmitted HIV-1 drug-resistance: 2009 update. PLoS ONE 2009, 4, e4724. [Google Scholar] [CrossRef]
- Bailey, A.J.; Rhee, S.Y.; Shafer, R.W. Integrase Strand Transfer Inhibitor Resistance in Integrase Strand Transfer Inhibitor-Naive Persons. AIDS Res. Hum. Retrovir. 2021, 37, 736–743. [Google Scholar] [CrossRef]
- Riemenschneider, M.; Cashin, K.Y.; Budeus, B.; Sierra, S.; Shirvani-Dastgerdi, E.; Bayanolhagh, S.; Kaiser, R.; Gorry, P.R.; Heider, D. Genotypic Prediction of Co-receptor Tropism of HIV-1 Subtypes A and C. Sci. Rep. 2016, 6, 24883. [Google Scholar] [CrossRef]
- Esbjörnsson, J.; Månsson, F.; Martínez-Arias, W.; Vincic, E.; Biague, A.J.; da Silva, Z.J.; Fenyö, E.M.; Norrgren, H.; Medstrand, P. Frequent CXCR4 tropism of HIV-1 subtype A and CRF02_AG during late-stage disease—Indication of an evolving epidemic in West Africa. Retrovirology 2010, 7, 23. [Google Scholar] [CrossRef]
- Raymond, S.; Delobel, P.; Mavigner, M.; Cazabat, M.; Souyris, C.; Sandres-Sauné, K.; Cuzin, L.; Marchou, B.; Massip, P.; Izopet, J. Correlation between genotypic predictions based on V3 sequences and phenotypic determination of HIV-1 tropism. AIDS 2008, 22, F11–F16. [Google Scholar] [CrossRef]
- HIV French Resistance. Available online: https://hivfrenchresistance.org/hiv-french-resistance-hiv-tropism/ (accessed on 29 December 2025).
- Lengauer, T.; Sander, O.; Sierra, S.; Thielen, A.; Kaiser, R. Bioinformatics prediction of HIV coreceptor usage. Nat. Biotechnol. 2007, 25, 1407–1410. [Google Scholar] [CrossRef] [PubMed]
- Landmarks of the HIV Genome. Available online: https://www.hiv.lanl.gov/content/sequence/HIV/MAP/landmark.html (accessed on 26 December 2025).
- Wensing, A.M.; Calvez, V.; Ceccherini-Silberstein, F.; Charpentier, C.; Günthard, H.F.; Jacobsen, D.M.; Paredes, R.; Shafer, R.W.; Richman, D.D. 2025 update of the drug resistance mutations in HIV-1. Top. Antivir. Med. 2025, 33, 457–473. [Google Scholar] [PubMed]
- Sede, M.M.; Moretti, F.A.; Laufer, N.L.; Jones, L.R.; Quarleri, J.F. HIV-1 Tropism Dynamics and Phylogenetic Analysis from Longitudinal Ultra-Deep Sequencing Data of CCR5- and CXCR4-Using Variants. PLoS ONE 2014, 9, e102857. [Google Scholar] [CrossRef] [PubMed]
- Connell, B.J.; Hermans, L.; Wensing, A.M.J.; Schellens, I.; Schipper, P.I.; van Ham, P.M.; de Jong, D.T.C.M.; Otto, S.; Mathe, T.; Moraba, R.; et al. Immune activation correlates with and predicts CXCR4 co-receptor tropism switch in HIV-1 infection. Sci. Rep. 2020, 10, 15866. [Google Scholar] [CrossRef]
- Mosier, D.E. How HIV changes its tropism: Evolution and adaptation? Curr. Opin. HIV AIDS 2009, 4, 125–130. [Google Scholar] [CrossRef]
- Kirichenko, A.A.; Kireev, D.E.; Lopatukhin, A.E.; Murzakova, A.V.; Lapovok, I.A.; Ladnaya, N.N.; Pokrovsky, V.V. Prevalence and Structure of Hiv-1 Drug Resistance Among Treatment Naïve Patients Since the Introduction of Antiretroviral Therapy In The Russian Federation. HIV Infect. Immunosuppr. Disord. 2019, 11, 75–83. (In Russian) [Google Scholar] [CrossRef]
- Pimentel, V.; Pingarilho, M.; Sebastião, C.S.; Miranda, M.; Gonçalves, F.; Cabanas, J.; Costa, I.; Diogo, I.; Fernandes, S.; Costa, O.; et al. Applying Next-Generation Sequencing to Track HIV-1 Drug Resistance Mutations Circulating in Portugal. Viruses 2024, 16, 622. [Google Scholar] [CrossRef]
- Pipitò, L.; Cannella, S.; Mascarella, C.; Graceffa, D.; Trizzino, M.; Iaria, C.; Colletti, P.; Mazzola, G.; Giammanco, G.M.; Cascio, A.; et al. Next-Generation Sequencing Analysis for HIV-1 Genotyping and Drug Resistance Mutations Mapping in Sicily, Italy. Viruses 2025, 17, 1129. [Google Scholar] [CrossRef]
- Lapovok, I.; Murzakova, A.; Lopatukhin, A.; Kireev, D.; Kazennova, E.; Bobkova, M. Prevalence of HIV-1 drug resistance mutations among ART-naïve patients in Russia from 2005 to 2015. In Proceedings of the 14th European Meeting on HIV & Hepatitis, Rome, Italy, 25–27 May 2016; pp. 25–27. [Google Scholar]
- Zhang, J.; Sun, B.; Sheng, Z.; Ding, X.; Fan, Q.; Huang, G.; Guo, Z.; Zhong, P.; Liao, L.; Xing, H.; et al. Full-Spectrum Surveillance of Pre-Treatment HIV Drug Resistance in Southeastern China. Pharmaceuticals 2024, 17, 900. [Google Scholar] [CrossRef]
- Tao, K.; Rhee, S.Y.; Chu, C.; Avalos, A.; Ahluwalia, A.K.; Gupta, R.K.; Jordan, M.R.; Shafer, R.W. Treatment Emergent Dolutegravir Resistance Mutations in Individuals Naïve to HIV-1 Integrase Inhibitors: A Rapid Scoping Review. Viruses 2023, 15, 1932. [Google Scholar] [CrossRef]
- Liu, J.; Li, C.; Sun, Y.; Fu, C.; Wei, S.; Zhang, X.; Ma, J.; Zhao, Q.; Huo, Y. Characteristics of drug resistance mutations in ART-experienced HIV-1 patients with low-level viremia in Zhengzhou City, China. Sci. Rep. 2024, 14, 10620. [Google Scholar] [CrossRef]
- Krishnan, S.; Lippincott, C.K.; Bjerrum, S.; Martinez Rivera, M.B.; Shah, M. A National HIV Provider Survey of Antiretroviral Therapy Preferences for Management of Treatment-Naive and Experienced Individuals with Drug Resistance. Open Forum Infect. Dis. 2023, 10, ofad541. [Google Scholar] [CrossRef]
- Diallo, K.; Götte, M.; Wainberg, M.A. Molecular impact of the M184V mutation in human immunodeficiency virus type 1 reverse transcriptase. Antimicrob. Agents Chemother. 2003, 47, 3377–3383. [Google Scholar] [CrossRef]
- Antonova, A.; Kazennova, E.; Lebedev, A.; Ozhmegova, E.; Kuznetsova, A.; Tumanov, A.; Bobkova, M. Recombinant Forms of HIV-1 in the Last Decade of the Epidemic in the Russian Federation. Viruses 2023, 15, 2312. [Google Scholar] [CrossRef]
- Singh, A.; Sunpath, H.; Green, T.N.; Padayachi, N.; Hiramen, K.; Lie, Y.; Anton, E.D.; Murphy, R.; Reeves, J.D.; Kuritzkes, D.R.; et al. Drug resistance and viral tropism in HIV-1 subtype C-infected patients in KwaZulu-Natal, South Africa: Implications for future treatment options. J. Acquir. Immune Defic. Syndr. 2011, 58, 233–240. [Google Scholar] [CrossRef]
- Williams, A.; Menon, S.; Crowe, M.; Agarwal, N.; Biccler, J.; Bbosa, N.; Ssemwanga, D.; Adungo, F.; Moecklinghoff, C.; Macartney, M.; et al. Geographic and Population Distributions of Human Immunodeficiency Virus (HIV)-1 and HIV-2 Circulating Subtypes: A Systematic Literature Review and Meta-analysis (2010–2021). J. Infect. Dis. 2023, 228, 1583–1591. [Google Scholar] [CrossRef]



| Federal District | Total Samples (n = 1888) Abs./% |
|---|---|
| Siberian | 732/38.77 |
| Southern | 551/29.18 |
| Volga region | 258/13.67 |
| Northwestern | 175/9.27 |
| Far Eastern | 155/8.21 |
| Central | 17/0.90 |
| Primer Name | Primer Sequence |
|---|---|
| Pol-F-1 | GGGCCCCTAGGAAAAAGGG |
| Pol-R-1 | CCTGTATGCAGACCCCAATATGTT |
| Pol-F-2 | CCCTCARATCACTCTTTGGCA |
| Pol-R-2 | TGCCACACAATCATCACCTG |
| Env-F-1 | GAGCAGAAGAYAGTGGMAATGA |
| Env-R-1 | GMKGAARAGGCACAGGYTCC |
| Env-F-2 | GAGCAGAAGAYAGTGGMAATGA |
| Env-R-2 | GAGCTGYTTRATGCCCCAGAC |
| Mutation | Total (n = 1888) Abs./% | ART Experience (n = 1466) | ||
|---|---|---|---|---|
| No (n = 411) Abs./% | Yes (n = 1055) Abs./% | p | ||
| M46I 1 | 15/0.79 | 4/0.97 | 10/0.95 | 1.000 |
| K43T | 10/0.53 | 2/0.49 | 5/0.47 | 1.000 |
| L33F | 10/0.53 | 1/0.24 | 4/0.38 | 1.000 |
| Q58E | 6/0.32 | 0/0.00 | 5/0.47 | 1.000 |
| V11I | 4/0.21 | 0/0.00 | 2/0.19 | 1.000 |
| I54S 1 | 4/0.21 | 1/0.24 | 2/0.19 | 1.000 |
| F53L 1 | 3/0.16 | 0/0.00 | 2/0.19 | 1.000 |
| I47V 1 | 2/0.11 | 0/0.00 | 2/0.19 | 1.000 |
| I54V 1 | 2/0.11 | 0/0.00 | 1/0.09 | 1.000 |
| M46L 1 | 2/0.11 | 1/0.24 | 1/0.09 | 1.000 |
| Mutation | Total (n = 1888) Abs./% | ART Experience (n = 1466) | ||
|---|---|---|---|---|
| No (n = 411) Abs./% | Yes (n = 1055) Abs./% | p | ||
| A62V | 506/26.80 | 64/15.57 | 334/31.66 | <0.001 |
| M184V 1 | 234/12.39 | 6/1.46 | 210/19.91 | <0.001 |
| K103N 1 | 206/10.91 | 28/6.81 | 149/14.12 | 0.013 |
| S68G | 176/9.32 | 32/7.79 | 114/10.81 | 1.000 |
| V90I | 162/8.58 | 20/4.87 | 115/10.90 | 0.037 |
| E138A | 158/8.37 | 31/7.54 | 91/8.63 | 1.000 |
| G190S 1 | 135/7.15 | 7/1.70 | 116/11.00 | <0.001 |
| K65R 1 | 114/6.04 | 0/0.00 | 105/9.95 | <0.001 |
| V106I | 113/5.99 | 9/2.19 | 93/8.82 | 0.001 |
| K101E 1 | 108/5.72 | 5/1.22 | 95/9.00 | <0.001 |
| Y181C 1 | 87/4.61 | 2/0.49 | 76/7.20 | <0.001 |
| M184I 1 | 58/3.07 | 1/0.24 | 54/5.12 | <0.001 |
| H221Y | 46/2.44 | 1/0.24 | 41/3.89 | 0.002 |
| V179E | 42/2.22 | 4/0.97 | 29/2.75 | 1.000 |
| E138K | 36/1.91 | 0/0.00 | 31/2.94 | 0.004 |
| P225H 1 | 34/1.80 | 1/0.24 | 31/2.94 | 0.037 |
| Y115F 1 | 31/1.64 | 0/0.00 | 28/2.65 | 0.012 |
| D67N 1 | 29/1.54 | 2/0.49 | 26/2.46 | 0.675 |
| Y318F | 28/1.48 | 0/0.00 | 26/2.46 | 0.019 |
| Mutation | Total (n = 1888) Abs./% | ART Experience (n = 1466) | ||
|---|---|---|---|---|
| No (n = 411) Abs./% | Yes (n = 1055) Abs./% | p | ||
| L74I | 1422/75.32 | 275/66.91 | 802/76.02 | 0.013 |
| E157Q | 30/1.59 | 3/0.73 | 21/1.99 | 1.000 |
| T97A | 12/0.64 | 0/0.00 | 9/0.85 | 1.000 |
| Y143R 1 | 7/0.37 | 0/0.00 | 7/0.66 | 1.000 |
| G163R | 4/0.21 | 1/0.24 | 3/0.28 | 1.000 |
| E138K 1 | 4/0.21 | 0/0.00 | 4/0.38 | 1.000 |
| Q148R 1 | 4/0.21 | 0/0.00 | 3/0.28 | 1.000 |
| D232N | 4/0.21 | 0/0.00 | 3/0.28 | 1.000 |
| L74M | 4/0.21 | 0/0.00 | 2/0.19 | 1.000 |
| N155H 1 | 3/0.16 | 0/0.00 | 3/0.28 | 1.000 |
| R263K 1 | 3/0.16 | 0/0.00 | 2/0.19 | 1.000 |
| E92G 1 | 3/0.16 | 2/0.49 | 1/0.09 | 1.000 |
| G140A 1 | 2/0.11 | 0/0.00 | 1/0.09 | 1.000 |
| Y143H 1 | 2/0.11 | 0/0.00 | 2/0.19 | 1.000 |
| G118R 1 | 2/0.11 | 0/0.00 | 2/0.19 | 1.000 |
| T66I 1 | 2/0.11 | 1/0.24 | 1/0.09 | 1.000 |
| E92Q 1 | 2/0.11 | 0/0.00 | 2/0.19 | 1.000 |
| S147G 1 | 2/0.11 | 0/0.00 | 2/0.19 | 1.000 |
| Protein | Drug Class | Drug Name | Resistance Mutations |
|---|---|---|---|
| Protease | PIs | Atazanavir | L33F, M46I |
| Tipranavir | L33F, K43T, M46I | ||
| Reverse transcriptase | NRTIs | Abacavir | K65R, M184V |
| Emtricitabine/Lamivudine | K65R, M184V | ||
| Tenofovir | K65R | ||
| NNRTIs | Doravirine | V106I, G190S | |
| Efavirenz | K101E, K103N, V106I, G190S | ||
| Etravirine | V90I, K101E, E138A, G190S | ||
| Nevirapine | K101E, K103N, V106I, G190S | ||
| Rilpivirine | K101E, E138A | ||
| Integrase | INSTIs | Cabotegravir | L74I, T97A |
| Rule | Number of Positive Samples (Proportion from the Total Number of Samples) | Disease Stages 1 of Studied Patients | Used Antiretroviral Therapy |
|---|---|---|---|
| A net charge rule of ≥+5 and total number of charged amino acids in the V3-loop of ≥8 | 23 (2.5%) | 3, 4A, 4B, 4C | 3TC:ABC:LPV:RTV; 3TC:AZT:ATV:RTV; 3TC:LPV:RTV:TDF; 3TC:DTG:TDF; 3TC:EFV:TDF; FTC:RPV:TDF |
| Loss of the N-linked glycosylation site in the V3-loop and a net charge of ≥+4 | 11 (1.2%) | 3, 4A | 3TC:ATV:RTV:TDF; 3TC:AZT:LPV:RTV; 3TC:EFV:TDF |
| R or K at position 11 and/or K at position 25 of the V3-loop (11/25 rule) | 10 (1.1%) | 3, 4A, 4C | 3TC:EFV:TDF 3TC:LPV:RTV:TDF |
| R at position 25 of the V3-loop and a net charge of ≥+5 | 5 (0.5%) | 4A, 4B | 3TC:ABC:DTG; 3TC:ABC:ESV; 3TC:ABC:LPV:RTV; 3TC:DTG:TDF |
| Net charge ≥+6 | 1 (0.1%) | 2B | no data |
| Rule | Number of Samples in Group | Number of Geno2Pheno Results Matching Empirical Predictions | Percentage of Concordance Between Geno2Pheno and Empirical Predictions |
|---|---|---|---|
| No CXCR4 co-receptor tropism predicted | 866 | 728 | 84.0% |
| A net charge rule of ≥+5 and total number of charged amino acids in the V3-loop of ≥8 | 23 | 21 | 91.3% |
| Loss of the N-linked glycosylation site in the V3-loop and a net charge of ≥+4 | 11 | 9 | 81.8% |
| R or K at position 11 and/or K at position 25 of the V3-loop (11/25 rule) | 10 | 4 | 40% |
| R at position 25 of the V3-loop and a net charge of ≥+5 | 5 | 3 | 60% |
| Net charge ≥+6 | 1 | 0 | 0% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Fadeev, A.; Eder, V.; Pisareva, M.; Tsvetkov, V.; Masharskiy, A.; Komissarova, K.; Ivanova, A.; Yolshin, N.; Komissarov, A.; Mazus, A.; et al. Detection of HIV-1 Resistance Mutations to Antiretroviral Therapy and Cell Tropism in Russian Patients Using Next-Generation Sequencing. Pathogens 2026, 15, 144. https://doi.org/10.3390/pathogens15020144
Fadeev A, Eder V, Pisareva M, Tsvetkov V, Masharskiy A, Komissarova K, Ivanova A, Yolshin N, Komissarov A, Mazus A, et al. Detection of HIV-1 Resistance Mutations to Antiretroviral Therapy and Cell Tropism in Russian Patients Using Next-Generation Sequencing. Pathogens. 2026; 15(2):144. https://doi.org/10.3390/pathogens15020144
Chicago/Turabian StyleFadeev, Artem, Veronika Eder, Maria Pisareva, Valery Tsvetkov, Alexey Masharskiy, Kseniya Komissarova, Anna Ivanova, Nikita Yolshin, Andrey Komissarov, Alexey Mazus, and et al. 2026. "Detection of HIV-1 Resistance Mutations to Antiretroviral Therapy and Cell Tropism in Russian Patients Using Next-Generation Sequencing" Pathogens 15, no. 2: 144. https://doi.org/10.3390/pathogens15020144
APA StyleFadeev, A., Eder, V., Pisareva, M., Tsvetkov, V., Masharskiy, A., Komissarova, K., Ivanova, A., Yolshin, N., Komissarov, A., Mazus, A., & Lioznov, D. (2026). Detection of HIV-1 Resistance Mutations to Antiretroviral Therapy and Cell Tropism in Russian Patients Using Next-Generation Sequencing. Pathogens, 15(2), 144. https://doi.org/10.3390/pathogens15020144






