Identification of Neopestalotiopsis spp. from Strawberry Leaf, Fruit, and Crown Tissues in North Carolina
Abstract
1. Introduction
2. Materials and Methods
2.1. Pathogen Isolation and Morphological Characterization
2.2. Genomic DNA Isolation and Amplification
2.3. Phylogenetic Analysis
2.4. Restriction Enzyme Digestion Assay
2.5. High-Resolution Melting (HRM) Analysis
2.6. Whole-Plant Pathogenicity Studies in the Greenhouse
2.7. Detached-Leaf Pathogenicity Tests
3. Results
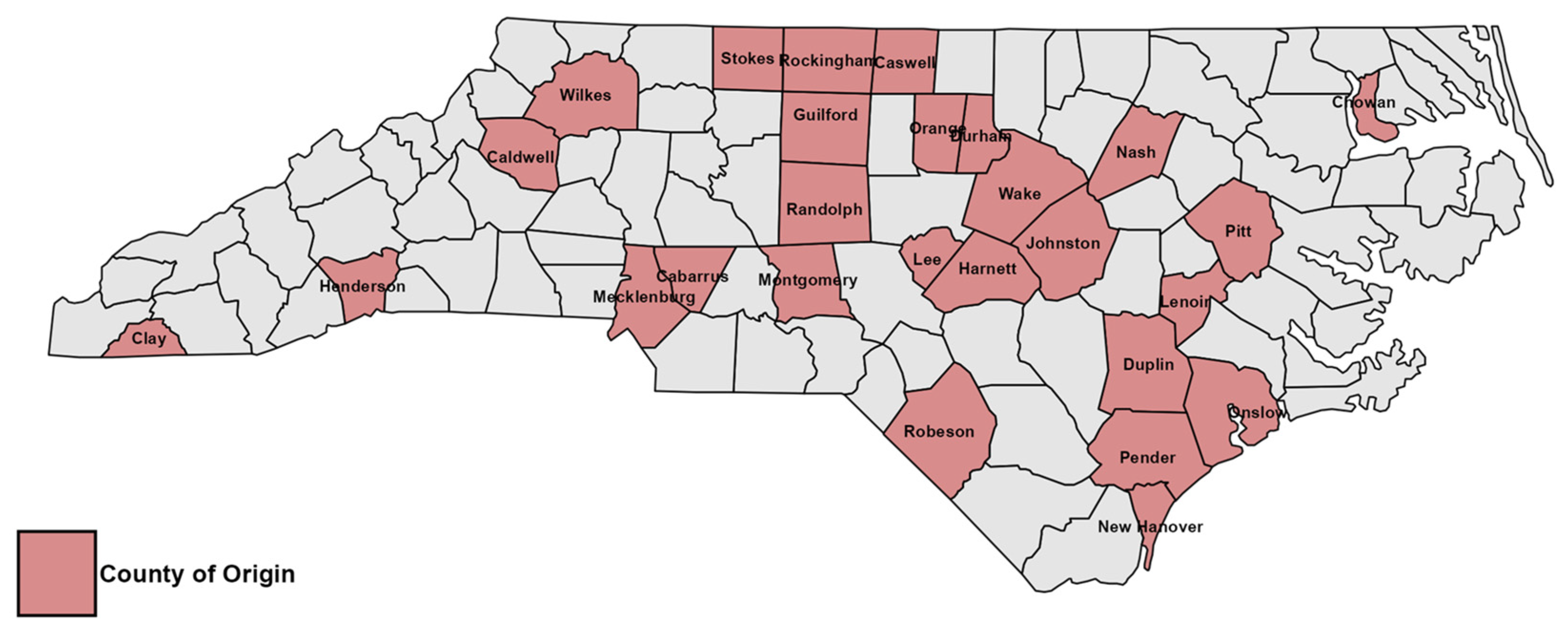
3.1. Prevalence of Neopestalotiopsis Species and Morphological Characteristics
3.2. Restriction Fragment Length Polymorphism
3.3. High-Resolution Melting Analysis
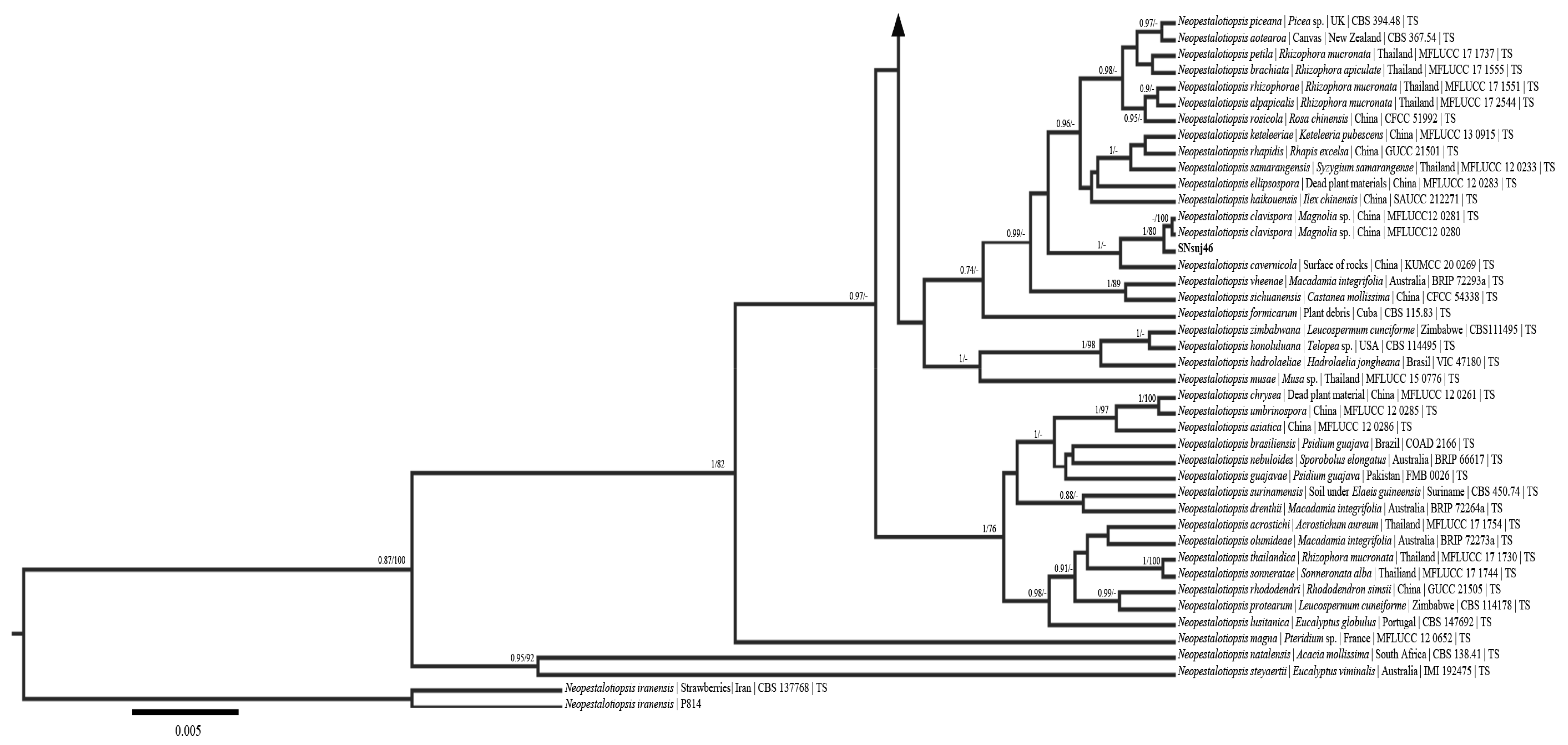
3.4. Phylogenetic Analysis
3.5. Pathogenicity Tests
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rusková, M.; Opálková Šišková, A.; Mosnáčková, K.; Gago, C.; Guerreiro, A.; Bučková, M.; Puškárová, A.; Pangallo, D.; Antunes, M.D. Biodegradable active packaging enriched with essential oils for enhancing the shelf life of strawberries. Antioxidants 2023, 12, 755. [Google Scholar] [CrossRef] [PubMed]
- Milosavljević, D.M.; Maksimović, V.M.; Milivojević, J.M.; Nakarada, Đ.J.; Mojović, M.D.; Dragišić Maksimović, J.J. Rich in Phenolics—Strong Antioxidant Fruit? Comparative Study of 25 Strawberry Cultivars. Plants 2022, 11, 3566. [Google Scholar] [CrossRef]
- Hodgdon, E.A.; Conner, D.S.; McDermott, L.G.; Pritts, M.P.; Handley, D.T.; Orde, K.M.; Sideman, R.G. A current view on strawberry production practices and trends in the Northeastern United States and Canada. HortTechnology 2024, 34, 574–584. [Google Scholar] [CrossRef]
- Trejo-Téllez, L.I.; Gómez-Merino, F.C. Nutrient management in strawberry: Effects on yield, quality and plant health. In Strawberries: Cultivation, Antioxidant Properties and Health Benefits; Nova Science Publishers: Hauppauge, NY, USA, 2014; pp. 239–267. [Google Scholar]
- Priyadarshi, R.; Jayakumar, A.; de Souza, C.K.; Rhim, J.W.; Kim, J.T. Advances in strawberry postharvest preservation and packaging: A comprehensive review. Compr. Rev. Food Sci. Food Saf. 2024, 23, e13417. [Google Scholar] [CrossRef]
- Şener, S. Abiotic stress factors and strawberry cultivation. In Impact of Climate Change on Agriculture; Holistence Publications: Çanakkale, Turkey, 2021; pp. 111–125. [Google Scholar]
- Staudt, G. Taxonomic studies in the genus Fragaria: Typification of Fragaria species known at the time of Linnaeus. Can. J. Bot. 1962, 40, 869–886. [Google Scholar] [CrossRef]
- Liston, A.; Cronn, R.; Ashman, T.L. Fragaria: A genus with deep historical roots and ripe for evolutionary and ecological insights. Am. J. Bot. 2014, 101, 1686–1699. [Google Scholar] [CrossRef] [PubMed]
- Simpson, D. The economic importance of strawberry crops. In The Genomes of Rosaceous Berries and Their Wild Relatives; Springer International Publishing: Cham, Switzerland, 2018; pp. 1–7. [Google Scholar] [CrossRef]
- FAOSTAT. Crops and livestock products. In FAOSTAT Statistical Database; Food and Agriculture Organization of the United Nations (FAO): Rome, Italy, 2023; Available online: https://www.fao.org/faostat/en/#data (accessed on 7 July 2025).
- Holmes, G.J. The California strawberry industry: Current trends and future prospects. Int. J. Fruit Sci. 2024, 24, 115–129. [Google Scholar] [CrossRef]
- Bandara, S.J.H. Understanding the economic sustainability of strawberry farming in North Carolina. Int. J. Food Agric. Econ. (IJFAEC) 2021, 9, 191–202. [Google Scholar] [CrossRef]
- Johnson, J. Evaluation of Yield, Fruit Chemistry, and Firmness of Seven Strawberry (Fragaria × ananassa) Cultivars and Four NCSU Advanced Strawberry Selections in an Eastern North Carolina Greenhouse; North Carolina State University: Raleigh, NC, USA, 2024. [Google Scholar] [CrossRef]
- Adhikari, T.B.; Muzhinji, N.; Philbrick, A.N.; Louws, F.J. Genomic insights and comparative analysis of Colletotrichum species associated with anthracnose fruit rot and crown rot of strawberry in North Carolina. Front. Microbiol. 2025, 16, 1515894. [Google Scholar] [CrossRef]
- Adhikari, T.B.; Muzhinji, N.; Philbrick, A.N.; Schnabel, G.; Louws, F.J. Genome Sequence Resource of Four Botrytis cinerea Isolates from North Carolina. PhytoFrontiers™ 2025, 5, PHYTOFR-05. [Google Scholar] [CrossRef]
- Marin, M.V.; Taylor, J.B.; Rebello, C.S.; Peres, N.A. Targeted Chemical Strategies in Planting Holes: A Proactive Approach to Managing Phytophthora Crown Rot of Strawberry. Plant Health Prog. 2024, 25, 454–459. [Google Scholar] [CrossRef]
- Baggio, J.S.; Cordova, L.G.; Seijo, T.E.; Noling, J.W.; Whitaker, V.M.; Peres, N.A. Cultivar selection is an effective and economic strategy for managing charcoal rot of strawberry in Florida. Plant Dis. 2021, 105, 2071–2077. [Google Scholar] [CrossRef] [PubMed]
- Burkhardt, A.; Henry, P.M.; Koike, S.T.; Gordon, T.R.; Martin, F. Detection of Fusarium oxysporum f. sp. fragariae from infected strawberry plants. Plant Dis. 2019, 103, 1006–1013. [Google Scholar] [CrossRef]
- Gordon, T.R.; Kirkpatrick, S.C.; Hansen, J.; Shaw, D.V. Response of strawberry genotypes to inoculation with isolates of Verticillium dahliae differing in host origin. Plant Pathol. 2006, 55, 766–769. [Google Scholar] [CrossRef]
- Kaur, H.; Wesche, J.; Gelain, J.; Minzheng, C.; Luo, C.X.; Schnabel, G. Characterization and fungicide sensitivity of Gnomoniopsis fructicola causing Gnomonia Leaf Blotch of strawberry in the Carolinas. Plant Dis. 2025, 109, 107–114. [Google Scholar] [CrossRef]
- Koike, S.T.; Gordon, T.R.; Daugovish, O.; Ajwa, H.; Bolda, M.; Subbarao, K. Recent developments on strawberry plant collapse problems in California caused by Fusarium and Macrophomina. Int. J. Fruit Sci. 2013, 13, 76–83. [Google Scholar] [CrossRef]
- Husaini, A.M.; Xu, Y.W. Challenges of climate change to strawberry cultivation: Uncertainty and beyond. In Strawberry: Growth, Development and Diseases; CABI: Wallingford, UK, 2016; pp. 262–287. [Google Scholar]
- Fang, X.; Phillips, D.; Verheyen, G.; Li, H.; Sivasithamparam, K.; Barbetti, M.J. Yields and resistance of strawberry cultivars to crown and root diseases in the field, and cultivar responses to pathogens under controlled environment conditions. Phytopathol. Mediterr. 2012, 51, 69–84. [Google Scholar]
- Baggio, J.S.; Forcelini, B.B.; Wang, N.Y.; Ruschel, R.G.; Mertely, J.C.; Peres, N.A. Outbreak of leaf spot and fruit rot in Florida strawberry caused by Neopestalotiopsis spp. Plant Dis. 2021, 105, 305–315. [Google Scholar] [CrossRef]
- Cline, W.; Moparthi, S.; Hoffmann, M. Neopestalotiopsis Leaf, Fruit, and Crown Rot of Strawberry. North Carolina State University Extension. Available online: https://content.ces.ncsu.edu/neopestalotiopsis-leaf-fruit-and-crown-rot-of-strawberry (accessed on 5 December 2025).
- Shi, J.; Li, B.; Wang, S.; Zhang, W.; Shang, M.; Wang, Y.; Liu, B. Occurrence of Neopestalotiopsis clavispora causing apple leaf spot in China. Agronomy 2024, 14, 1658. [Google Scholar] [CrossRef]
- Gerardo-Lugo, S.S.; Tovar-Pedraza, J.M.; Maharachchikumbura, S.S.N.; Apodaca-Sánchez, M.A.; Correia, K.C.; Sauceda-Acosta, C.P.; Camacho-Tapia, M.; Hyde, K.D.; Marraiki, N.; Elgorban, A.M.; et al. Characterization of Neopestalotiopsis species associated with mango grey leaf spot disease in Sinaloa, Mexico. Pathogens 2020, 9, 788. [Google Scholar] [CrossRef] [PubMed]
- Beg, M.A.; Oliver, J.E. Georgia blueberry Neopestalotiopsis isolates, which are phylogenetically indistinguishable from the emerging novel strawberry Neopestalotiopsis sp., are pathogenic to both blueberry and strawberry. Plant Dis. 2025. [Google Scholar] [CrossRef]
- Fiorenza, A.; Gusella, G.; Aiello, D.; Polizzi, G.; Voglmayr, H. Neopestalotiopsis siciliana sp. nov. and N. rosae causing stem lesion and dieback on avocado plants in Italy. J. Fungi 2022, 8, 562. [Google Scholar] [CrossRef]
- Di Francesco, A.; Gallo, G.; Lanzarini, M.; Cignola, R. First report of Neopestalotiopsis clavispora, causal agent of grape trunk disease in grapevine in Northeastern Italy. Plant Dis. 2025, 109, PDIS-05. [Google Scholar] [CrossRef]
- O’Donnell, K.; Cigelnik, E.; Nirenberg, H.I. Molecular systematics and phylogeography of Gibberella fujikuroi species complex. Proc. Natl. Acad. Sci. USA 1998, 95, 2044–2049. [Google Scholar] [CrossRef] [PubMed]
- Glass, N.L.; Donaldson, G.C. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microbiol. 1995, 61, 1323–1330. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar] [CrossRef]
- Van der Vyver, L.S.; De Bruin, W.; Siyoum, N.; Nsibo, D.L.; Groenewald, J.Z.; Crous, P.W.; Korsten, L. Exploring Neopestalotiopsis diversity associated with blueberry leaf and twig blight in South African nurseries. Fungal Syst. Evol. 2025, 16, 41–53. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Abadi, S.; Azouri, D.; Mayrose, I.; Pupko, T. Model selection may not be a mandatory step for phylogeny reconstruction. Nat. Commun. 2019, 10, 934. [Google Scholar] [CrossRef] [PubMed]
- Gernhard, T. The conditioned reconstructed process. J. Theor. Biol. 2008, 253, 769–778. [Google Scholar] [CrossRef]
- Suchard, M.A.; Lemey, P.; Baele, G.; Ayres, D.L.; Drummond, A.J.; Rambaut, A. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol. 2018, 4, Vey016. [Google Scholar] [CrossRef] [PubMed]
- Rambaut, A. Fig Tree Ver. 1.3.1. Available online: http://tree.bio.ed.ac.uk/software/figtree (accessed on 2 November 2025).
- Silvestro, D.; Michalak, I. RaxmlGUI: A graphical front-end for RAxML. Org. Divers. Evol. 2012, 12, 335–337. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Kaur, H.; Gelain, J.; Marin, M.V.; Peres, N.A.; Schnabel, G. Development of a molecular tool for identification of a new Neopestalotiopsis sp. associated with disease outbreaks on strawberry. Plant Dis. 2023, 107, 1544–1549. [Google Scholar] [CrossRef]
- Rebello, C.S.; Wang, N.Y.; Marin, M.V.; Baggio, J.S.; Peres, N.A. Detection and species differentiation of Neopestalotiopsis spp. from strawberry (Fragaria × ananassa) in Florida using a high-resolution melting analysis. PhytoFrontiers™ 2023, 3, 156–163. [Google Scholar] [CrossRef]
- Madrid, A.J.; Munoz, G.; Collins, C.; Brannen, P. First report of the new Neopestalotiopsis species causing strawberry leaf spot and fruit rot in Georgia. Plant Dis. 2024, 108, 2574. [Google Scholar] [CrossRef]
- Gangwar, P.; Xu, X.; Pollok, J.; Sedghi, N.; Flanagan, R.D.; Zeng, Y. First report of Neopestalotiopsis vaccinii in Delaware and Neopestalotiopsis rosae in Virginia associated with similar disease symptoms in strawberry (Fragaria × ananassa). Plant Dis. 2025, 109. [Google Scholar] [CrossRef]
- Lawrence, D.P.; Brittain, G.D.; Aglave, B.; Sances, F.V. First Report of Neopestalotiopsis rosae causing crown and root rot of strawberry in California. Plant Dis. 2023, 107, 566. [Google Scholar] [CrossRef]
- Dardani, G.; Martino, I.; Aloi, F.; Carli, C.; Giordano, R.; Spadaro, D.; Guarnaccia, V. Characterization of Neopestalotiopsis species associated with strawberry crown rot in Italy. Agronomy 2025, 15, 422. [Google Scholar] [CrossRef]
- Chamorro, M.; Aguado, A.; De los Santos, B. First report of root and crown rot caused by Pestalotiopsis clavispora (Neopestalotiopsis clavispora) on strawberry in Spain. Plant Dis. 2016, 100, 1495. [Google Scholar] [CrossRef]
- Mahapatra, S.; Banerjee, J.; Kumar, K.; Pramanik, S.; Pramanik, K.; Islam, S.; Das, S. Leaf spot and fruit rot of strawberry caused by Neopestalotiopsis clavispora in Indo-Gangetic plains of India. Indian Phytopathol. 2018, 71, 279–283. [Google Scholar] [CrossRef]
- Gilardi, G.; Bergeretti, F.; Gullino, M.L.; Garibaldi, A. First report of Neopestalotiopsis clavispora causing root and crown rot on strawberry in Italy. Plant Dis. 2019, 103, 2959. [Google Scholar] [CrossRef]
- Shi, J.; Zhang, X.; Liu, Y.; Zhang, Z.; Wang, Z.; Xue, C.; Ma, Y.; Wang, F. First report of Neopestalotiopsis clavispora causing calyx and receptacle blight on strawberry in China. Plant Dis. 2022, 106, 1307. [Google Scholar] [CrossRef] [PubMed]
- Maharachchikumbura, S.S.N.; Hyde, K.D.; Groenewald, J.Z.; Xu, J.; Crous, P.W. Pestalotiopsis revisited. Stud. Mycol. 2014, 79, 121–186. [Google Scholar] [CrossRef]
- Yang, Q.; Zeng, X.Y.; Yuan, J.; Zhang, Q.; He, Y.K.; Wang, Y. Two new species of Neopestalotiopsis from southern China. Biodivers. Data J. 2021, 9, e70446. [Google Scholar] [CrossRef]
- Fernández-Ozuna, Y.A.; Álvarez, A.G.; Lopez-Nicora, H.D.; Alvarenga, A.A.; Colmán, A.A. First report of Neopestalotiopsis rosae causing leaf spot and crown rot on strawberry (Fragaria × ananassa) in Paraguay. New Dis. Rep. 2023, 48, 2. [Google Scholar] [CrossRef]




| GenBank Accession Number | ||||
|---|---|---|---|---|
| Neopestalotiopsis spp. | Isolate | ITS | TEF1-α | β-tubulin |
| N. rosae | SNcah118 | PX563605 | PX711530 | PX711490 |
| N. rosae | SNcc08 | PX563606 | PX711531 | PX711491 |
| N. rosae | SNcc09 | PX563607 | PX711532 | PX711492 |
| N. rosae | SNcc10 | PX563608 | PX711533 | PX711493 |
| N. rosae | SNcc11 | PX563609 | PX711534 | PX711494 |
| N. rosae | SNcc117 | PX563610 | PX711535 | PX711495 |
| N. rosae | SNccd02 | PX563611 | PX711536 | PX711496 |
| N. rosae | SNccd16 | PX563612 | PX711537 | PX711497 |
| N. rosae | SNccg04 | PX563613 | PX711538 | PX711498 |
| N. rosae | SNccm94 | PX563614 | PX711539 | PX711499 |
| N. rosae | SNccm95 | PX563615 | PX711540 | PX711500 |
| N. rosae | SNcco03 | PX563616 | PX711541 | PX711501 |
| N. rosae | SNcco72 | PX563617 | PX711542 | PX711502 |
| N. rosae | SNccr112 | PX563618 | PX711543 | PX711503 |
| N. rosae | SNccrr57 | PX563619 | PX711544 | PX711504 |
| N. rosae | SNclh124 | PX563620 | PX711545 | NA 1 |
| N. rosae | SNcm93 | PX563621 | PX711546 | PX711505 |
| N. rosae | SNcnc100 | PX563622 | PX711547 | PX711506 |
| N. rosae | SNcnc99 | PX563623 | PX711548 | PX711507 |
| N. rosae | SNcnd17 | PX563624 | PX711549 | PX711508 |
| N. rosae | SNcnd53 | PX563625 | PX711550 | PX711509 |
| N. rosae | SNcnd54 | PX563626 | PX711551 | PX711510 |
| N. rosae | SNcnd55 | PX563627 | PX711552 | PX711511 |
| N. rosae | SNcnd56 | PX563628 | PX711553 | PX711512 |
| N. rosae | SNcnh86 | PX563629 | PX711554 | PX711513 |
| N. rosae | SNcnn81 | PX563630 | PX711555 | PX711514 |
| N. rosae | SNcnrjl52 | PX563631 | PX711556 | PX711515 |
| N. rosae | SNco73 | PX563632 | PX711557 | PX711516 |
| N. rosae | SNcph125 | PX563633 | PX711558 | NA 1 |
| N. rosae | SNcrj108 | PX563634 | PX711559 | PX711517 |
| N. rosae | SNcrj109 | PX563635 | PX711560 | PX711518 |
| N. rosae | SNcrjp63 | PX563636 | PX711561 | PX711519 |
| N. rosae | SNcrjp64 | PX563637 | PX711562 | PX711520 |
| N. rosae | SNcrjr114 | PX563638 | PX711563 | PX711521 |
| N. rosae | SNcscr58 | PX563639 | PX711564 | PX711522 |
| N. rosae | SNcscr59 | PX563640 | PX711565 | PX711523 |
| N. rosae | SNcw12 | PX563641 | PX711566 | PX711524 |
| N. rosae | SNcw13 | PX563642 | PX711567 | PX711525 |
| N. rosae | SNar79 | PX563643 | PX711568 | PX711526 |
| N. rosae | SNsnn60 | PX563644 | PX711569 | PX711527 |
| N. rosae | SNsrjp65 | PX563645 | PX711570 | PX711528 |
| N. rosae | SNrjl01 | PX563646 | PX711571 | PX711529 |
| N. clavispora | SNsuj46 | PX586188 | PX699155 | PX699157 |
| N. hispanica | SNah119 | PX619695 | PX699089 | PX711486 |
| N. hispanica | SNah120 | PX619696 | PX699090 | PX711485 |
| N. hispanica | SNas104 | PX619697 | PX699091 | PX711484 |
| N. hispanica | SNcno51 | PX619698 | PX699092 | PX711487 |
| N. hispanica | SNcrh69 | PX619699 | PX699093 | PX711489 |
| N. hispanica | SNcrp06 | PX619700 | PX699094 | PX711483 |
| N. hispanica | SNcrp07 | PX619701 | PX699095 | PX711482 |
| N. hispanica | SNht14 | PX619702 | PX699154 | PX711481 |
| N. hispanica | SNmh122 | PX619703 | PX699096 | PX711480 |
| N. hispanica | SNmh123 | PX619704 | PX699097 | PX711479 |
| N. hispanica | SNnac19 | PX619705 | PX699098 | PX711478 |
| N. hispanica | SNnc18 | PX619706 | PX699099 | PX711477 |
| N. hispanica | SNnh15 | PX619707 | PX699100 | PX711476 |
| N. hispanica | SNnh85 | PX619708 | PX699101 | PX711475 |
| N. hispanica | SNnh88 | PX619709 | PX699102 | PX711474 |
| N. hispanica | SNnl98 | PX619710 | PX699103 | PX711473 |
| N. hispanica | SNnn78 | PX619711 | PX699104 | PX711472 |
| N. hispanica | SNnn90 | PX619712 | PX699105 | PX711471 |
| N. hispanica | SNnn91 | PX619713 | PX699106 | PX711470 |
| N. hispanica | SNnn92 | PX619714 | PX699107 | PX711469 |
| N. hispanica | SNnn101 | PX619715 | PX699108 | PX711468 |
| N. hispanica | SNnnh83 | PX619716 | PX699109 | PX711467 |
| N. hispanica | SNnnh84 | PX619717 | PX699110 | PX711466 |
| N. hispanica | SNp132 | PX619718 | PX699111 | PX711465 |
| N. hispanica | SNrjh121 | PX619719 | PX699112 | PX711464 |
| N. hispanica | SNrn134 | PX619720 | PX699113 | PX711463 |
| N. hispanica | SNsajj36 | PX619721 | PX699114 | PX711462 |
| N. hispanica | SNsc82 | PX619722 | PX699115 | PX711461 |
| N. hispanica | SNscc20 | PX619723 | PX699116 | PX711460 |
| N. hispanica | SNscc21 | PX619724 | PX699117 | PX711459 |
| N. hispanica | SNscm67 | PX619725 | PX699118 | PX711458 |
| N. hispanica | SNscm68 | PX619726 | PX699119 | PX711457 |
| N. hispanica | SNscr61 | PX619727 | PX699120 | PX711456 |
| N. hispanica | SNscrh70 | PX619728 | PX699121 | PX711455 |
| N. hispanica | SNscrh71 | PX619729 | PX699122 | PX711454 |
| N. hispanica | SNscrj28 | PX619730 | PX699123 | PX711453 |
| N. hispanica | SNscrj34 | PX619731 | PX699124 | PX711452 |
| N. hispanica | SNscrj45 | PX619732 | PX699125 | PX711451 |
| N. hispanica | SNsfj24 | PX619733 | PX699126 | PX711450 |
| N. hispanica | SNsfj35 | PX619734 | PX699127 | PX711449 |
| N. hispanica | SNsmjj37 | PX619735 | PX699128 | PX711448 |
| N. hispanica | SNsn135 | PX619736 | PX699129 | PX711447 |
| N. hispanica | SNsnh87 | PX619737 | PX699130 | PX711446 |
| N. hispanica | SNsnj22 | PX619738 | PX699131 | PX711445 |
| N. hispanica | SNsnj23 | PX619739 | PX699132 | PX711444 |
| N. hispanica | SNso50 | PX619740 | PX699133 | PX711443 |
| N. hispanica | SNsp129 | PX619741 | PX699134 | PX711442 |
| N. hispanica | SNssj25 | PX619742 | PX699135 | PX711441 |
| N. hispanica | SNssn133 | PX619743 | PX699136 | PX711440 |
| N. hispanica | SNssuj38 | PX619744 | PX699137 | PX711439 |
| N. hispanica | SNssuj39 | PX619745 | PX699138 | PX711438 |
| N. hispanica | SNsuj26 | PX619746 | PX699139 | PX711437 |
| N. hispanica | SNsuj27 | PX619747 | PX699140 | PX711436 |
| N. hispanica | SNsuj29 | PX619748 | PX699141 | PX711435 |
| N. hispanica | SNsuj30 | PX619749 | PX699142 | PX711434 |
| N. hispanica | SNsuj31 | PX619750 | PX699143 | PX711433 |
| N. hispanica | SNsuj32 | PX619751 | PX699144 | PX711432 |
| N. hispanica | SNsuj33 | PX619752 | PX699145 | PX711431 |
| N. hispanica | SNsuj40 | PX619753 | PX699146 | PX711430 |
| N. hispanica | SNsuj41 | PX619754 | PX699147 | PX711429 |
| N. hispanica | SNsuj42 | PX619755 | PX699148 | PX711428 |
| N. hispanica | SNsuj43 | PX619756 | PX699149 | PX711427 |
| N. hispanica | SNsuj44 | PX619757 | PX699150 | PX711426 |
| N. hispanica | SNsuj47 | PX619758 | PX699151 | PX711425 |
| N. hispanica | SNsuj48 | PX619759 | PX699152 | PX711424 |
| N. hispanica | SNucnh05 | PX619760 | PX699153 | PX711486 |
| N. longiappendiculata | SNccrl62 | PX643237 | PX699156 | PX699159 |
| N. scalabiensis | SNYh127 | PX643238 | NA 1 | PX699158 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Moparthi, S.; Bradshaw, M.J.; Cline, W.; Munster, M.J.; Hoffmann, M.; Segovia, D.R.; Crouch, U.; Li, C.; Almeyda, C.; Salazar, J.; et al. Identification of Neopestalotiopsis spp. from Strawberry Leaf, Fruit, and Crown Tissues in North Carolina. Pathogens 2026, 15, 10. https://doi.org/10.3390/pathogens15010010
Moparthi S, Bradshaw MJ, Cline W, Munster MJ, Hoffmann M, Segovia DR, Crouch U, Li C, Almeyda C, Salazar J, et al. Identification of Neopestalotiopsis spp. from Strawberry Leaf, Fruit, and Crown Tissues in North Carolina. Pathogens. 2026; 15(1):10. https://doi.org/10.3390/pathogens15010010
Chicago/Turabian StyleMoparthi, Swarnalatha, Michael J. Bradshaw, William Cline, Michael J. Munster, Mark Hoffmann, Diana Ramirez Segovia, Uma Crouch, Chunying Li, Christie Almeyda, Jhoselin Salazar, and et al. 2026. "Identification of Neopestalotiopsis spp. from Strawberry Leaf, Fruit, and Crown Tissues in North Carolina" Pathogens 15, no. 1: 10. https://doi.org/10.3390/pathogens15010010
APA StyleMoparthi, S., Bradshaw, M. J., Cline, W., Munster, M. J., Hoffmann, M., Segovia, D. R., Crouch, U., Li, C., Almeyda, C., Salazar, J., & Bertone, M. A. (2026). Identification of Neopestalotiopsis spp. from Strawberry Leaf, Fruit, and Crown Tissues in North Carolina. Pathogens, 15(1), 10. https://doi.org/10.3390/pathogens15010010






