AI-Enhanced FT-IR Spectroscopy: Evaluation of a Novel Tool for High-Throughput Serovar Typing of Salmonella enterica subsp. enterica in Croatia
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Collection
2.2. Strain Preparation
2.3. Spectra Acquisition and Analysis
3. Results
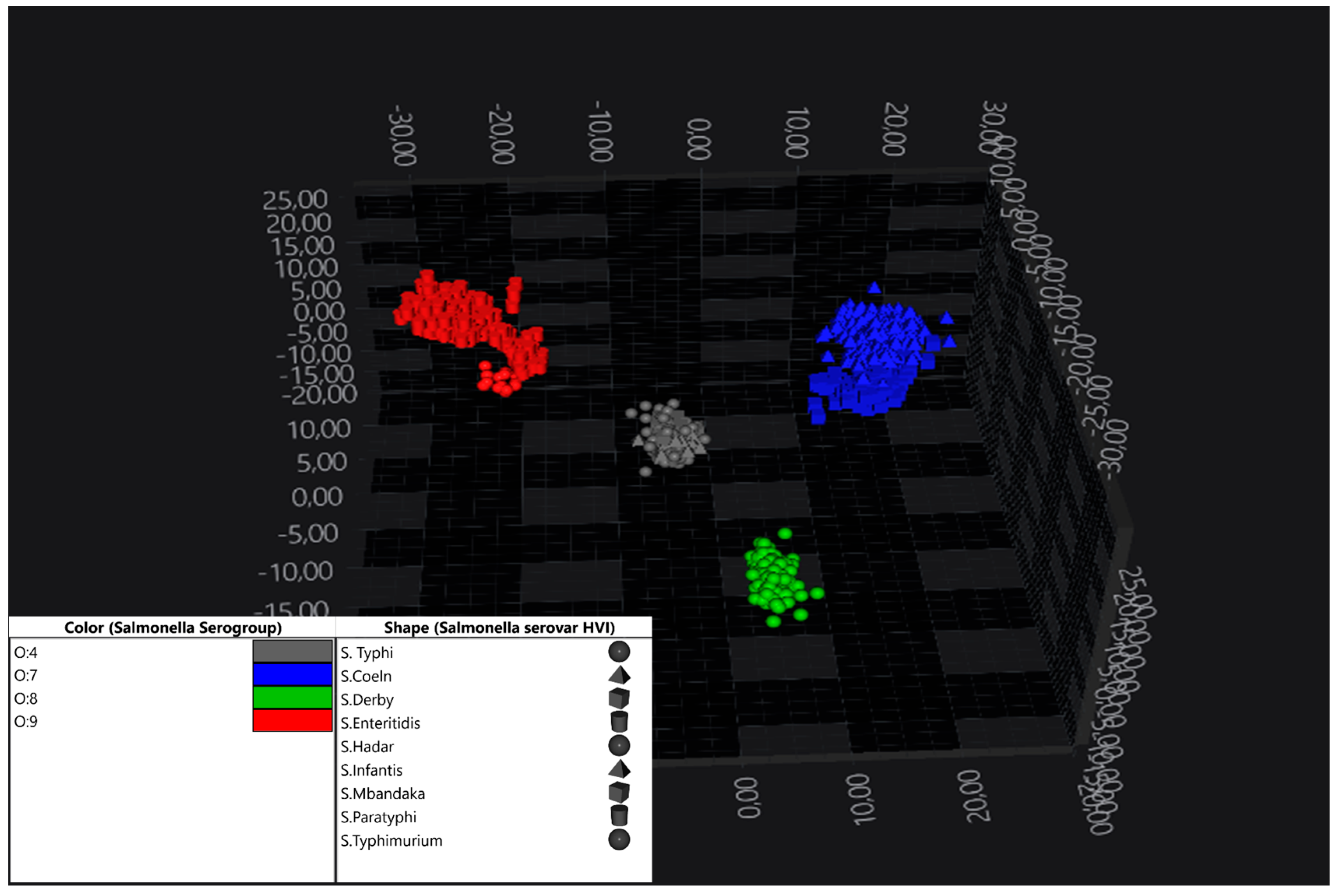
3.1. Discrimination Between Serogroups of S. enterica subsp. enterica
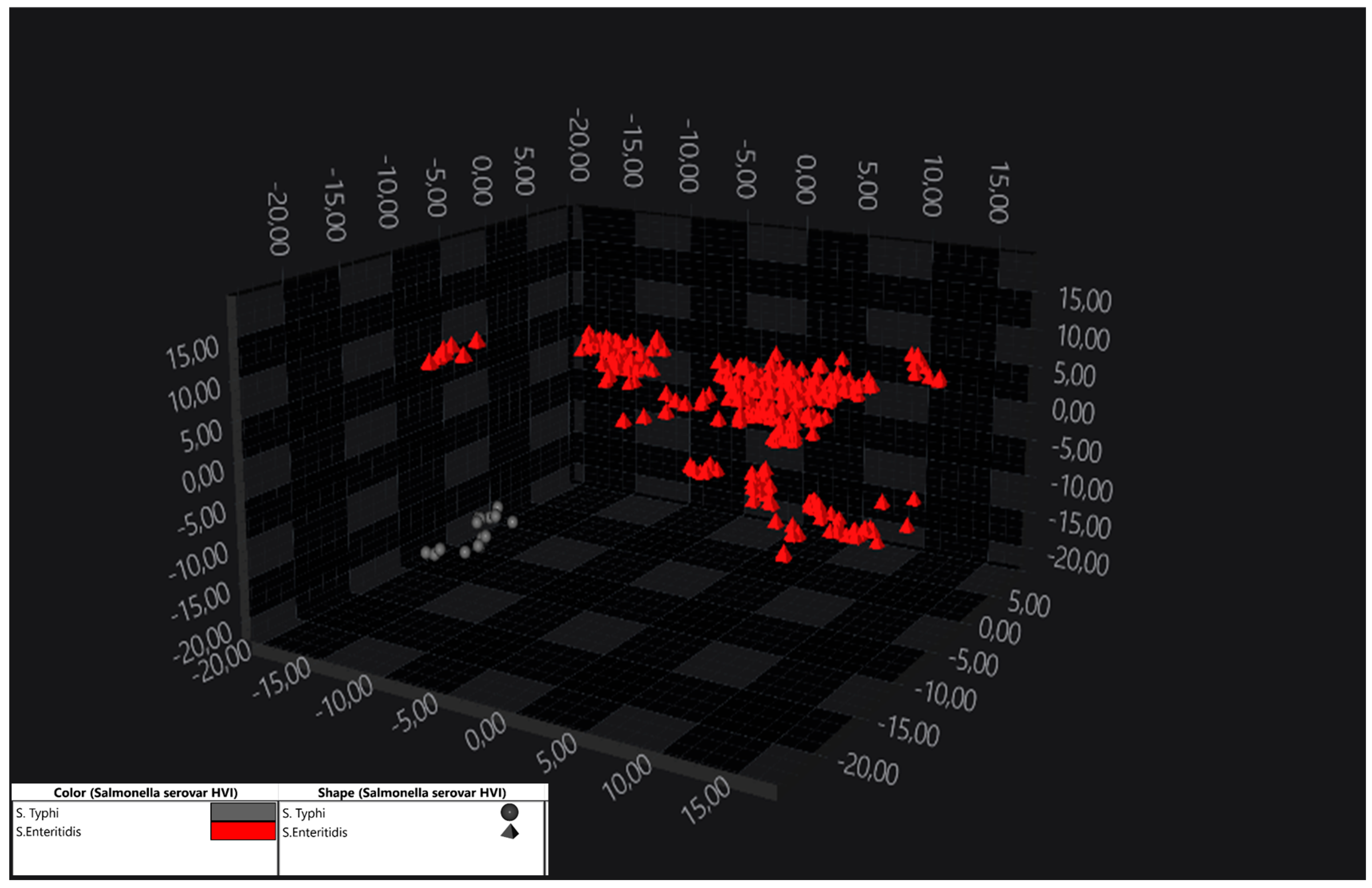
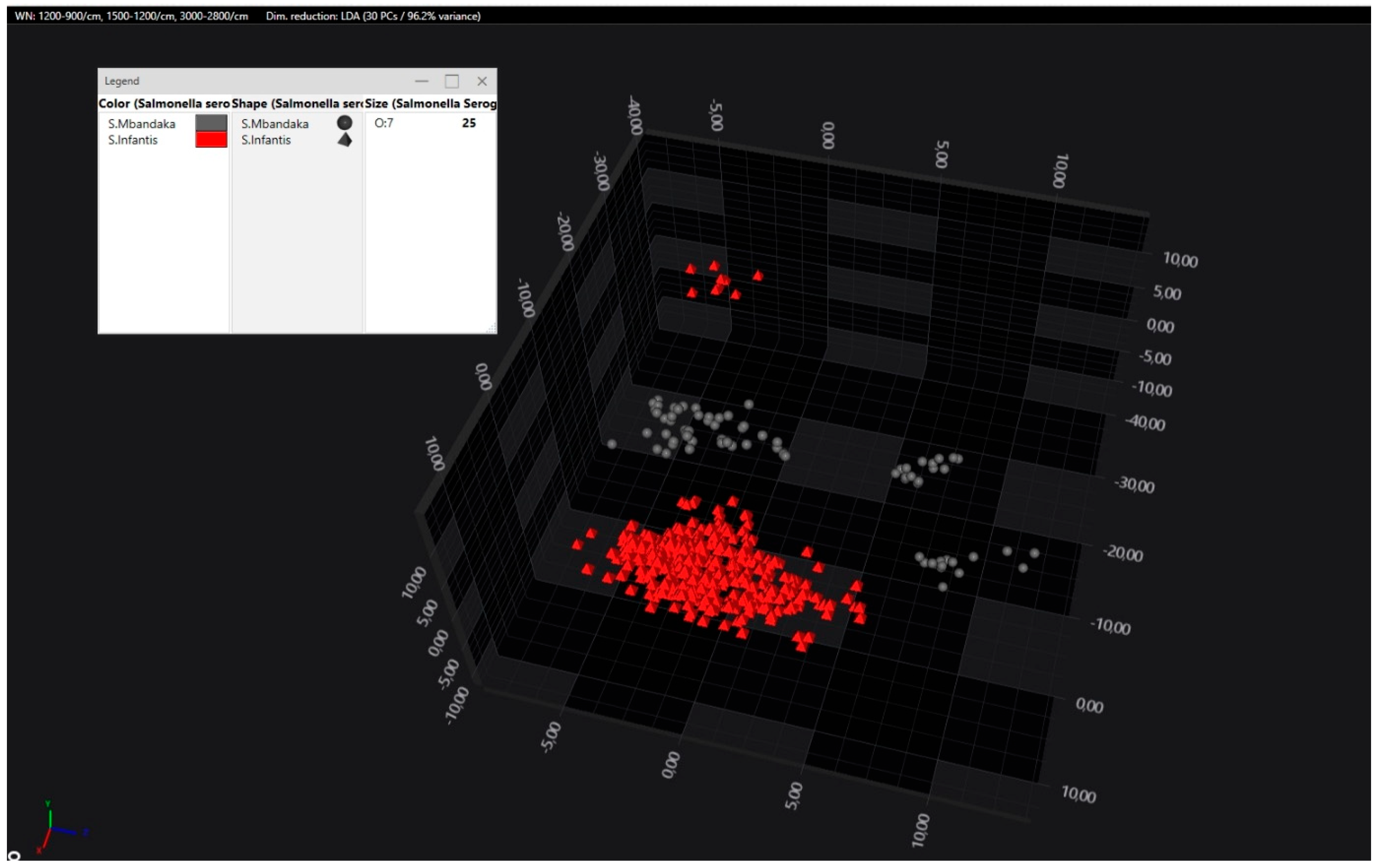
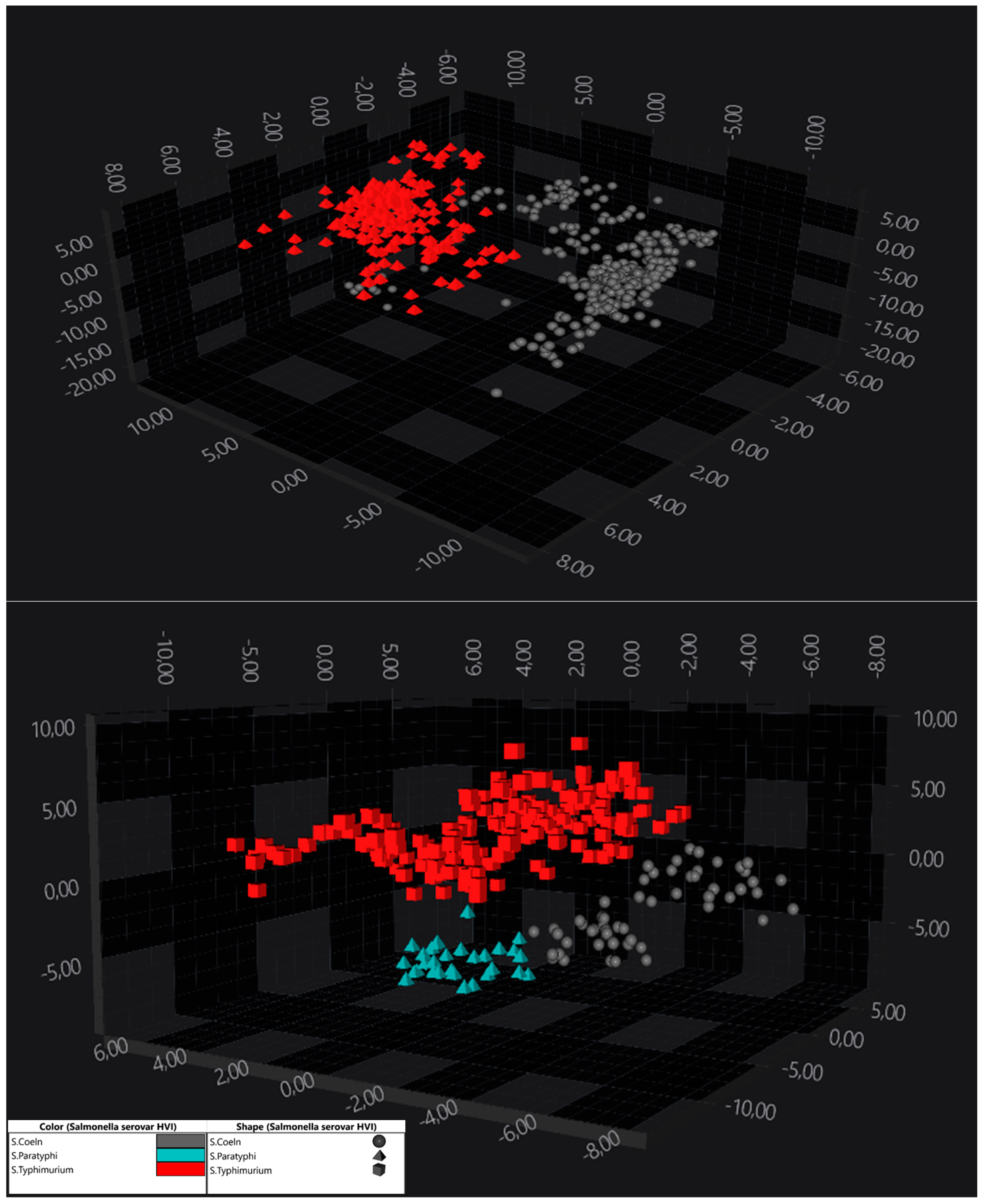
3.2. Discrimination Within S. enterica subsp. enterica Serogroups (Serovar Identification)
4. Discussion
5. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Alvarez-Ordóñez, A.; Halisch, J.; Prieto, M. Changes in Fourier transform infrared spectra of Salmonella enterica serovars Typhimurium and Enteritidis after adaptation to stressful growth conditions. Int. J. Food Microbiol. 2010, 142, 97–105. [Google Scholar] [CrossRef]
- Preisner, O.E.; Menezes, J.C.; Guiomar, R.; Machado, J.; Lopes, J.A. Discrimination of Salmonella enterica serotypes by Fourier transform infrared spectroscopy. Food Res. Int. 2012, 45, 1058–1064. [Google Scholar] [CrossRef]
- Cordovana, M.; Mauder, N.; Join-Lambert, O.; Gravey, F.; LeHello, S.; Auzou, M.; Pitti, M.; Zoppi, S.; Buhl, M.; Steinmann, J.; et al. Machine learning-based typing of Salmonella enterica O-serogroups by the Fourier-Transform Infrared (FTIR) Spectroscopy-based IR Biotyper system. J. Microbiol. Methods 2022, 201, 106564. [Google Scholar] [CrossRef]
- Chattaway, M.A.; Langridge, G.C.; Wain, J. Salmonella nomenclature in the genomic era: A time for change. Sci. Rep. 2021, 11, 7494. [Google Scholar] [CrossRef]
- Han, J.; Ljahdali, N.; Zhao, S.; Tang, H.; Harbottle, H.; Hoffmann, M.; Frye, J.G.; Foley, S.L. Infection biology of Salmonella enterica. EcoSal Plus 2024, 12, eesp-0001-2023. [Google Scholar] [CrossRef]
- Lamichhane, B.; Mawad, A.M.M.; Saleh, M.; Kelley, W.G.; Harrington, P.J.; Lovestad, C.W.; Amezcua, J.; Sarhan, M.M.; El Zowalaty, M.E.; Ramadan, H.; et al. Salmonellosis: An Overview of Epidemiology, Pathogenesis, and Innovative Approaches to Mitigate the Antimicrobial Resistant Infections. Antibiotics 2024, 13, 76. [Google Scholar] [CrossRef]
- Antunes, P.; Mourão, J.; Campos, J.; Peixe, L. Salmonellosis: The role of poultry meat. Clin. Microbiol. Infect. 2016, 22, 110–121. [Google Scholar] [CrossRef]
- Ferrari, R.G.; Rosario, D.K.A.; Cunha-Neto, A.; Mano, S.B.; Figueiredo, E.E.S.; Conte-Junior, C.A. Worldwide Epidemiology of Salmonella Serovars in Animal-Based Foods: A Meta-analysis. Appl. Environ. Microbiol. 2019, 85, e00591-19. [Google Scholar] [CrossRef]
- Abuhasna, S.; Al Jundi, A.; Rahman, M.U.; Said, W. Non-typhoidal Salmonella group D bacteremia and urosepsis in a patient diagnosed with HIV Infection. J. Glob. Infect. Dis. 2012, 4, 218–219. [Google Scholar] [CrossRef]
- Andino, A.; Hanning, I. Salmonella enterica: Survival, Colonization, and Virulence Differences among Serovars. Sci. World J. 2015, 1, 520179. [Google Scholar] [CrossRef]
- Kaavya, R.; Pandiselvam, R.; Abdullah, S.; Sruthi, N.U.; Jayanath, Y.; Ashokkumar, C.; Chandra Khanashyam, A.; Kothakota, A.; Ramesh, S.V. Emerging non-thermal technologies for decontamination of Salmonella in food. Trends Food Sci. Technol. 2021, 112, 400–418. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA); European Centre for Disease Prevention and Control (ECDC). The European Union One Health 2022 Zoonoses Report. EFSA J. 2023, 21, e8442. [Google Scholar] [CrossRef]
- Novak, A.; Dzelalija, M.; Goic-Barisic, I.; Kovacic, A.; Pirija, M.; Maravic, A.; Radic, M.; Marinovic, J.; Rubic, Z.; Carev, M.; et al. Phenotypic and Molecular Characterization of a Hospital Outbreak Clonal Lineage of Salmonella enterica Subspecies enterica serovar Mikawasima Containing blaTEM-1B and blaSHV-2 That Emerged on a Neonatal Ward, During the COVID-19 Pandemic. Microb. Drug Resist. 2024, 30, 118–126. [Google Scholar] [CrossRef]
- Hugas, M.; Beloeil, P.A. Controlling Salmonella along the food chain in the European Union—Progress over the last ten years. Eurosurveillance 2014, 19, 20804. [Google Scholar] [CrossRef]
- Tang, S.; Orsi, R.H.; Luo, H.; Ge, C.; Zhang, G.; Baker, R.C.; Stevenson, A.; Wiedmann, M. Assessment and Comparison of Molecular Subtyping and Characterization Methods for Salmonella. Front. Microbiol. 2019, 10, 1591. [Google Scholar] [CrossRef]
- Gómez-Montaño, F.J.; Orduña-Díaz, A.; Avelino-Flores, M.C.G.; Avelino-Flores, F.; Ramos-Collazo, F.; Reyes-Betanzo, C.; López-Gayou, V.; Instituto Politécnico Nacional. Detection of Salmonella enterica on silicon substrates biofunctionalized with anti-Salmonella IgG, analyzed by FTIR spectroscopy. Rev. Mex. De Ing. Química 2020, 19, 1175–1185. [Google Scholar] [CrossRef]
- Hu, L.; Cao, G.; Brown, E.W.; Allard, M.W.; Ma, L.M.; Zhang, G. Whole genome sequencing and protein structure analyses of target genes for the detection of Salmonella. Sci. Rep. 2021, 11, 20887. [Google Scholar] [CrossRef]
- McGalliard, R.; Muhamadali, H.; AlMasoud, N.; Haldenby, S.; Romero-Soriano, V.; Allman, E.; Xu, Y.; Roberts, A.P.; Paterson, S.; Carrol, E.D.; et al. Bacterial discrimination by Fourier transform infrared spectroscopy, MALDI-mass spectrometry and whole-genome sequencing. Future Microbiol. 2024, 19, 795–810. [Google Scholar] [CrossRef]
- Wang, Y.; Zhou, Q.; Li, B.; Liu, B.; Wu, G.; Ibrahim, M.; Xie, G.; Li, H.; Sun, G. Differentiation in MALDI-TOF MS and FTIR spectra between two closely related species Acidovorax oryzae and Acidovorax citrulli. BMC Microbiol. 2012, 12, 182. [Google Scholar] [CrossRef]
- Cordovana, M.; Mauder, N.; Kostrzewa, M.; Wille, A.; Rojak, S.; Hagen, R.M.; Ambretti, S.; Pongolini, S.; Soliani, L.; Justesen, U.S.; et al. Classification of Salmonella enterica of the (Para-)Typhoid Fever Group by Fourier-Transform Infrared (FTIR) Spectroscopy. Microorganisms 2021, 9, 853. [Google Scholar] [CrossRef]
- Campos, J.; Sousa, C.; Mourão, J.; Lopes, J.; Antunes, P.; Peixe, L. Discrimination of non-typhoid Salmonella serogroups and serotypes by Fourier Transform Infrared Spectroscopy: A comprehensive analysis. Int. J. Food Microbiol. 2018, 285, 34–41. [Google Scholar] [CrossRef]
- Curtoni, A.; Pastrone, L.; Cordovana, M.; Bondi, A.; Piccinini, G.; Genco, M.; Bottino, P.; Polizzi, C.; Cavallo, L.; Mandras, N.; et al. Fourier Transform Infrared Spectroscopy Application for Candida auris Outbreak Typing in a Referral Intensive Care Unit: Phylogenetic Analysis and Clustering Cut-Off Definition. Microorganisms 2024, 12, 1312. [Google Scholar] [CrossRef]
- Passaris, I.; Mauder, N.; Kostrzewa, M.; Burckhardt, I.; Zimmermann, S.; van Sorge, N.M.; Slotved, H.C.; Desmet, S.; Ceyssens, P.J. Validation of Fourier Transform Infrared Spectroscopy for Serotyping of Streptococcus pneumoniae. J. Clin. Microbiol. 2022, 60, e00325-22. [Google Scholar] [CrossRef]
- Wang-Wang, J.H.; Bordoy, A.E.; Martró, E.; Quesada, M.D.; Pérez-Vázquez, M.; Guerrero-Murillo, M.; Tiburcio, A.; Navarro, M.; Castellà, L.; Sopena, N.; et al. Evaluation of Fourier Transform Infrared Spectroscopy as a First-Line Typing Tool for the Identification of Extended-Spectrum β-Lactamase-Producing Klebsiella pneumoniae Outbreaks in the Hospital Setting. Front. Microbiol. 2022, 13, 897161. [Google Scholar] [CrossRef]
- Martak, D.; Valot, B.; Sauget, M.; Cholley, P.; Thouverez, M.; Bertrand, X.; Hocquet, D. Fourier-Transform InfraRed Spectroscopy Can Quickly Type Gram-Negative Bacilli Responsible for Hospital Outbreaks. Front. Microbiol. 2019, 10, 1440. [Google Scholar] [CrossRef]
- Vogt, S.; Löffler, K.; Dinkelacker, A.G.; Bader, B.; Autenrieth, I.B.; Peter, S.; Liese, J. Fourier-Transform Infrared (FTIR) Spectroscopy for Typing of Clinical Enterobacter cloacae Complex Isolates. Front. Microbiol. 2019, 10, 2582. [Google Scholar] [CrossRef]
- Muchaamba, F.; Stephan, R.A. Comprehensive Methodology for Microbial Strain Typing Using Fourier-Transform Infrared Spectroscopy. Methods Protoc. 2024, 7, 48. [Google Scholar] [CrossRef]
- Dourou, D.; Grounta, A.; Argyri, A.A.; Froutis, G.; Tsakanikas, P.; Nychas, G.J.E.; Doulgeraki, A.I.; Chorianopoulos, N.G.; Tassou, C.C. Rapid Microbial Quality Assessment of Chicken Liver Inoculated or Not With Salmonella Using FTIR Spectroscopy and Machine Learning. Front. Microbiol. 2021, 11, 623788. [Google Scholar] [CrossRef]
- Yu, C.; Irudayaraj, J. Identification of Pathogenic Bacteria in Mixed Cultures by FTIR Spectroscopy. Trans. ASABE 2006, 49, 1623–1632. [Google Scholar] [CrossRef]
- Daltonics, Bruker GmbH & Co. KG (Bremen, Germany). Instructions for Use—IR Biotyper Kit; REF 1851760, (Doc. no. 5023357 Revision B); Daltonics, Bruker GmbH & Co. KG: Bremen, Germany, 2021. [Google Scholar]
- Daltonics, Bruker GmbH & Co. KG (Bremen, Germany). IR Biotyper Software User Manual; REF 1845471, (Doc. no. 5025119 Revision F); Daltonics, Bruker GmbH & Co. KG: Bremen, Germany, 2022. [Google Scholar]
- Oberreuter, H.; Cordovana, M.; Dyk, M.; Rau, J. Establishment and Thorough External Validation of a FTIR Spectroscopy Classifier for Salmonella Serogroup Differentiation. bioRxiv 2025, 643663. [Google Scholar] [CrossRef]
- Fredes-García, D.; Jiménez-Rodríguez, J.; Piña-Iturbe, A.; Caballero-Díaz, P.; González-Villarroel, T.; Dueñas, F.; Wozniak, A.; Adell, A.D.; Moreno-Switt, A.I.; García, P. Development of a robust FT-IR typing system for Salmonella enterica, enhancing performance through hierarchical classification. Microbiol. Spectr. 2025, 13, e00159-25. [Google Scholar] [CrossRef]
- Oberreuter, H.; Rau, J. Artificial neural network-assisted Fourier transform infrared spectroscopy for differentiation of Salmonella serogroups and its application on epidemiological tracing of Salmonella Bovismorbificans outbreak isolates from fresh sprouts. FEMS Microbiol. Lett. 2019, 366, fnz193. [Google Scholar] [CrossRef]
- Oberreuter, H.; Dyk, M.; Rau, J. Validated differentiation of Listeria monocytogenes serogroups by FTIR spectroscopy using an Artificial Neural Network based classifier in an accredited official food control laboratory. Clin. Spectrosc. 2023, 5, 100030. [Google Scholar] [CrossRef]
- Napoleoni, M.; Ceschia, S.; Mitri, E.; Beneitez, E.E.; Silenzi, V.; Staffolani, M.; Rocchegiani, E.; Blasi, G.; Gurian, E. Identification of Salmonella Serogroups and Distinction Between Typhoidal and Non-Typhoidal Salmonella Based on ATR-FTIR Spectroscopy. Microorganisms 2024, 12, 2318. [Google Scholar] [CrossRef]
- Kassem, A.; Abbas, L.; Coutinho, O.; Opara, S.; Najaf, H.; Kasperek, D.; Pokhrel, K.; Li, X.; Tiquia-Arashiro, S. Applications of Fourier Transform-Infrared spectroscopy in microbial cell biology and environmental microbiology: Advances, challenges, and future perspectives. Front. Microbiol. 2023, 14, 1304081. [Google Scholar] [CrossRef]
- Al-Fraihat, E.; Barker, K.R.; Tadros, M. The IR Biotyper as a tool for typing organisms of significance for hospital epidemiology- A subject review. Diagn. Microbiol. Infect. Dis. 2025, 111, 116676. [Google Scholar] [CrossRef]
- Lurie-Weinberger, M.N.; Temkin, E.; Kastel, O.; Bechor, M.; Bychenko-Banyas, D.; Efrati-Epchtien, R.; Levi, G.D.; Rakovitsky, N.; Keren-Paz, A.; Carmeli, Y. Use of a national repository of Fourier-transform infrared spectroscopy spectra enables fast detection of silent outbreaks and prevention of spread of new antibiotic-resistant sequence types. Antimicrob. Resist. Infect. Control. 2025, 14, 34. [Google Scholar] [CrossRef]
- Kon, H.; Lurie-Weinberger, M.N.; Lugassy, C.; Chen, D.; Schechner, V.; Schwaber, M.J.; Hussein, K.; Alon, T.; Tarabeia, J.; Hamo, M.; et al. Use of Fourier-transform infrared spectroscopy for real-time outbreak investigation of OXA-48-producing Escherichia coli. J. Antimicrob. Chemother. 2024, 79, 349–353. [Google Scholar] [CrossRef]
- Hu, Y.; Zhou, H.; Lu, J.; Sun, Q.; Liu, C.; Zeng, Y.; Zhang, R. Evaluation of the IR Biotyper for Klebsiella pneumoniae typing and its potentials in hospital hygiene management. Microb. Biotechnol. 2021, 14, 1343–1352. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dopuđ, M.; Šuto, S.; Tomašković, D.; Hlebić, L.; Peinović, L.; Špičić, S.; Zdelar Tuk, M.; Reil, I.; Humski, A.; Kompes, G.; et al. AI-Enhanced FT-IR Spectroscopy: Evaluation of a Novel Tool for High-Throughput Serovar Typing of Salmonella enterica subsp. enterica in Croatia. Pathogens 2025, 14, 856. https://doi.org/10.3390/pathogens14090856
Dopuđ M, Šuto S, Tomašković D, Hlebić L, Peinović L, Špičić S, Zdelar Tuk M, Reil I, Humski A, Kompes G, et al. AI-Enhanced FT-IR Spectroscopy: Evaluation of a Novel Tool for High-Throughput Serovar Typing of Salmonella enterica subsp. enterica in Croatia. Pathogens. 2025; 14(9):856. https://doi.org/10.3390/pathogens14090856
Chicago/Turabian StyleDopuđ, Maja, Sandra Šuto, Dora Tomašković, Lucija Hlebić, Lovran Peinović, Silvio Špičić, Maja Zdelar Tuk, Irena Reil, Andrea Humski, Gordan Kompes, and et al. 2025. "AI-Enhanced FT-IR Spectroscopy: Evaluation of a Novel Tool for High-Throughput Serovar Typing of Salmonella enterica subsp. enterica in Croatia" Pathogens 14, no. 9: 856. https://doi.org/10.3390/pathogens14090856
APA StyleDopuđ, M., Šuto, S., Tomašković, D., Hlebić, L., Peinović, L., Špičić, S., Zdelar Tuk, M., Reil, I., Humski, A., Kompes, G., Strugar, S. Š., Papić, B., Avberšek, J., Mikolajczak, A., & Duvnjak, S. (2025). AI-Enhanced FT-IR Spectroscopy: Evaluation of a Novel Tool for High-Throughput Serovar Typing of Salmonella enterica subsp. enterica in Croatia. Pathogens, 14(9), 856. https://doi.org/10.3390/pathogens14090856





