Scanning Electron Microscopy Reveals New Ultrastructural Features in Metacercariae of Clinostomum cutaneum (Digenea: Clinostomidae) Infecting Oreochromis niloticus (Actinopterygii: Cichlidae) in Kenya
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area and Sample Collection
2.2. Parasitological Examination
2.3. Morphological Identification
2.4. Scanning Electron Microscopy
2.5. DNA Extraction, Amplification and Sequencing
2.6. Sequence Analysis
3. Results
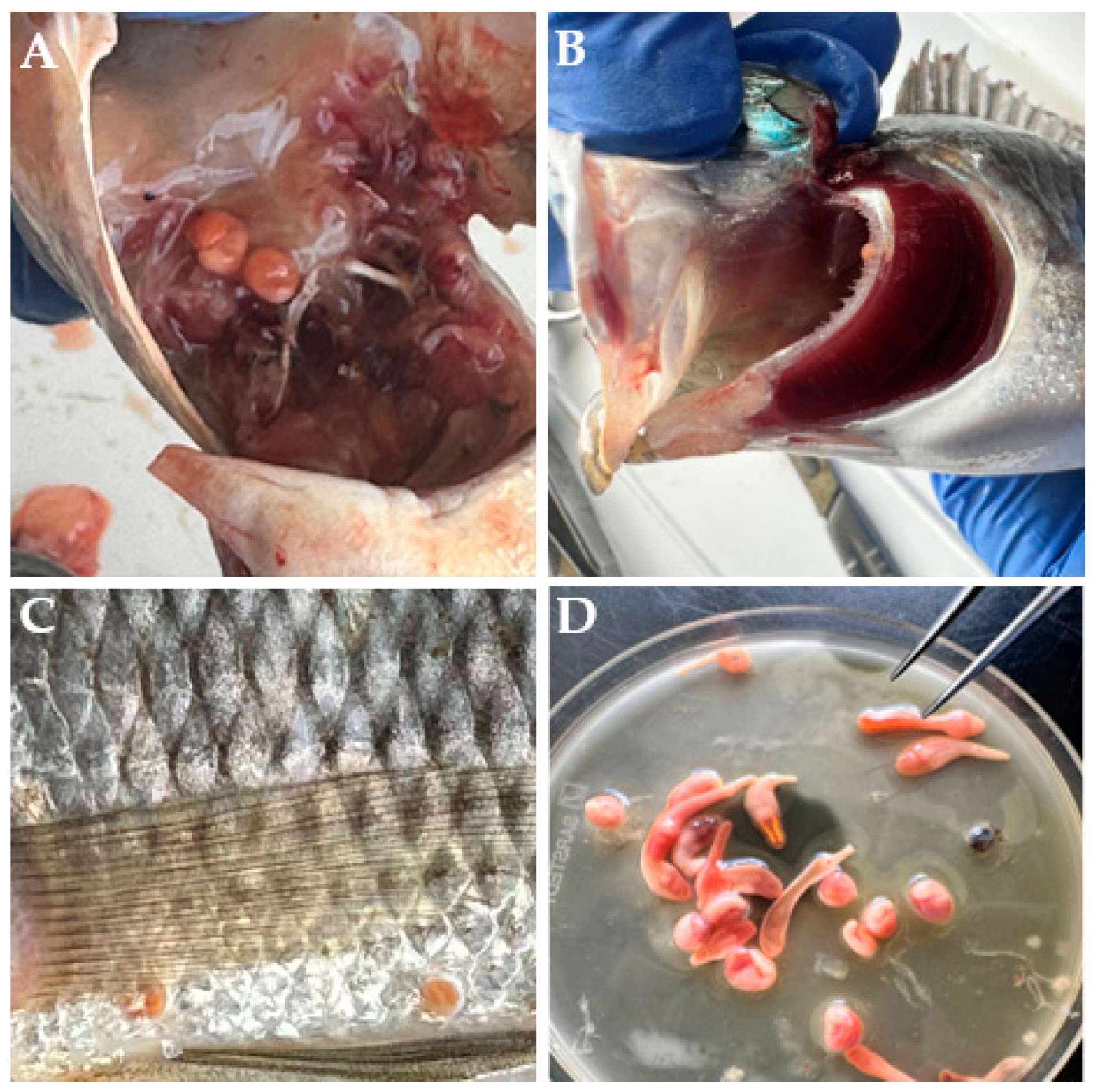
3.1. Infection Parameters and Isolated Metacercariae
3.2. Morphological Observations in C. cutaneum
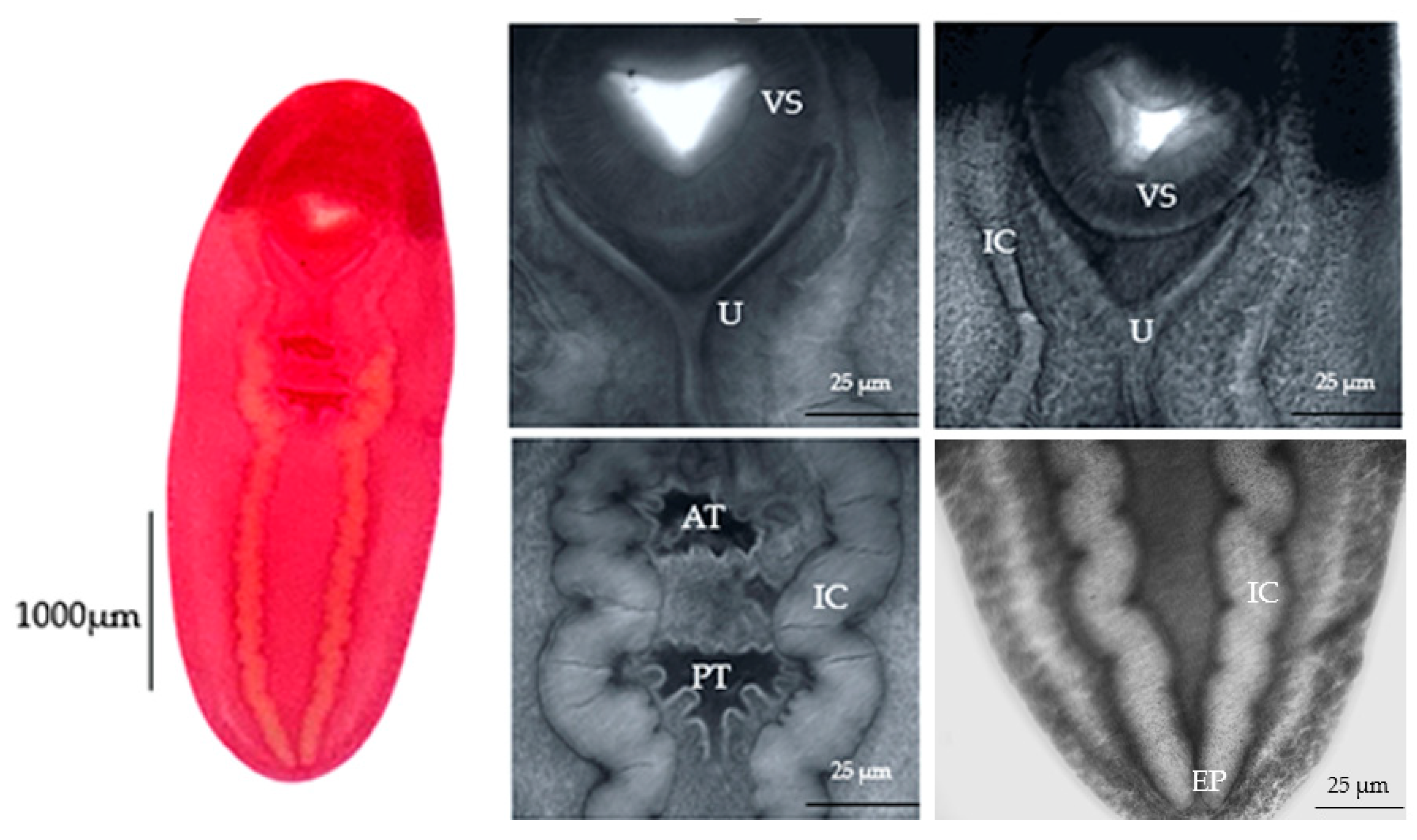
3.3. Scanning Electron Microscopy Results
3.4. Molecular Analyses
4. Discussion
5. Conclusion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Béné, C.; Barange, M.; Subasinghe, R.; Pinstrup-Andersen, P.; Merino, G.; Hemre, G.I.; Williams, M. Feeding 9 Billion by 2050—Putting Fish Back on the Menu. Food Secur. 2015, 7, 261–274. [Google Scholar] [CrossRef]
- Béné, C.; Arthur, R.; Norbury, H.; Allison, E.H.; Beveridge, M.; Bush, S.; Campling, L.; Leschen, W.; Little, D.; Squires, D.; et al. Contribution of Fisheries and Aquaculture to Food Security and Poverty Reduction: Assessing the Current Evidence. World Dev. 2016, 79, 177–196. [Google Scholar] [CrossRef]
- Munguti, J.; Obiero, K.; Orina, P.; Mirera, D.; Kyule, D.; Mwaluma, J.; Opiyo, M.; Musa, S.; Ochiewo, J.; Njiru, J.; et al. State of Aquaculture Report 2021: Towards Nutrition Sensitive Fish Food Production Systems; Techplus Media House: Nairobi, Kenya, 2021. [Google Scholar]
- KMFRI. Kenya’s Aquaculture Brief 2017: Status, Trends, Challenges and Future Outlook; Kenya Marine and Fisheries Research Institute: Mombasa, Kenya, 2017. [Google Scholar]
- Opiyo, M.A.; Marijani, E.; Muendo, P.; Odede, R.; Leschen, W.; Charo-Karisa, H. A Review of Aquaculture Production and Health Management Practices of Farmed Fish in Kenya. Int. J. Vet. Sci. Med. 2018, 6, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Pillay, T.V.R.; Kutty, M.N. Aquaculture Principles and Practices, 2nd ed.; Blackwell Publishing: Oxford, UK, 2005; ISBN 9781780648972. [Google Scholar]
- Kibet, C.J.; Donde, O.O.; Okwiri, B.; Otachi, E.O. Taxonomic Status of Fish Parasites in Kenyan Inland Water Systems and Their Significance on the Freshwater Fisheries and Aquaculture Productivity within the Region. Lakes Reserv. Res. Manag. 2019, 24, 402–410. [Google Scholar] [CrossRef]
- Shamsi, S.; Halajian, A.; Tavakol, S.; Mortazavi, P.; Boulton, J. Pathogenicity of Clinostomum complanatum (Digenea: Clinostomidae) in Piscivorous Birds. Res. Vet. Sci. 2013, 95, 537–539. [Google Scholar] [CrossRef]
- Gustinelli, A.; Caffara, M.; Florio, D.; Otachi, E.O.; Wathuta, E.M.; Fioravanti, M.L. First Description of the Adult Stage of Clinostomum cutaneum Paperna, 1964 (Digenea: Clinostomidae) from Grey Herons Ardea cinerea L. and a Redescription of the Metacercaria from the Nile Tilapia Oreochromis niloticus niloticus (L.) in Kenya. Syst. Parasitol. 2010, 76, 39–51. [Google Scholar] [CrossRef]
- Wang, M.L.; Chen, H.Y.; Shih, H.H. Occurrence and Distribution of Yellow Grub Trematodes (Clinostomum complanatum) Infection in Taiwan. Parasitol. Res. 2017, 116, 1761–1771. [Google Scholar] [CrossRef]
- Park, C.W.; Kim, J.S.; Joo, H.S.; Kim, J. A Human Case of Clinostomum complanatum Infection in Korea. Korean J. Parasitol. 2009, 47, 401–404. [Google Scholar] [CrossRef]
- Ermakova, L.; Kozlov, S.; Nagorny, S.; Golovchenko, N.; Telicheva, V.; Kiosova, J.; Zotova, M.; Pshenichnaya, N. The First Case of Human Invasion by Clinostomum complanatum in the European Part of Russia. IJID Reg. 2024, 11, 100346. [Google Scholar] [CrossRef]
- Caffara, M.; Locke, S.A.; Gustinelli, A.; Marcogliese, D.J.; Fioravanti, M.L. Morphological and Molecular Differentiation of Clinostomum complanatum and Clinostomum marginatum (Digenea: Clinostomidae) Metacercariae and Adults. J. Parasitol. 2011, 97, 884–891. [Google Scholar] [CrossRef]
- Locke, S.A.; Caffara, M.; Marcogliese, D.J.; Fioravanti, M.L. A Large-Scale Molecular Survey of Clinostomum (Digenea, Clinostomidae). Zool. Scr. 2015, 44, 203–217. [Google Scholar] [CrossRef]
- Caffara, M.; Locke, S.A.; Echi, P.C.; Halajian, A.; Benini, D.; Luus-Powell, W.J.; Tavakol, S.; Fioravanti, M.L. A Morphological and Molecular Study of Clinostomid Metacercariae from African Fish with a Redescription of Clinostomum tilapiae. Parasitology 2017, 144, 1519–1529. [Google Scholar] [CrossRef] [PubMed]
- Li, B.F.; Liu, X.H.; Ge, H.L.; Xie, C.Y.; Cai, R.Y.; Hu, Z.C.; Zhang, Y.G.; Wang, Z.J. The Discovery of Clinostomum complanatum Metacercariae in Farmed Chinese Sucker, Myxocyprinus asiaticus. Aquaculture 2018, 495, 273–280. [Google Scholar] [CrossRef]
- Caffara, M.; Locke, S.A.; Echi, P.C.; Halajian, A.; Luus-Powell, W.J.; Benini, D.; Tedesco, P.; Fioravanti, M.L. A New Species of Clinostomum Leidy, 1856 Based on Molecular and Morphological Analysis of Metacercariae from African Siluriform Fishes. Parasitol. Res. 2020, 119, 885–892. [Google Scholar] [CrossRef]
- Sereno-Uribe, A.L.; García-Varela, M.; Pinacho-Pinacho, C.D.; Pérez-Ponce de León, G. Three New Species of Clinostomum Leidy, 1856 (Trematoda) from Middle American Fish-Eating Birds. Parasitol. Res. 2018, 117, 2171–2185. [Google Scholar] [CrossRef]
- Sereno-Uribe, A.L.; Pinacho-Pinacho, C.D.; García-Varela, M.; De León, G.P.P. Using Mitochondrial and Ribosomal DNA Sequences to Test the Taxonomic Validity of Clinostomum complanatum Rudolphi, 1814 in Fish-Eating Birds and Freshwater Fishes in Mexico, with the Description of a New Species. Parasitol. Res. 2013, 112, 2855–2870. [Google Scholar] [CrossRef]
- Shigoley, M.I.; Antoine-Moussiaux, N.; Jauniaux, T.; Vanhove, M.P.M. Parasitology of One of the World’s Foremost Fisheries Target Species Lacks a One Health Approach. Hydrobiologia 2024, 1–16. [Google Scholar] [CrossRef]
- Ochieng, V.O.; Matolla, G.K.; Khyria, S.K. A Study of Clinostomum Affecting Oreochromis niloticus in Small Water Bodies in Eldoret-Kenya. Int. J. Sci. Eng. Res. 2012, 3, 1–6. [Google Scholar]
- Bello, M.A.; Ruiz-León, Y.; Sandoval-Sierra, J.V.; Rezinciuc, S.; Diéguez-Uribeondo, J. Scanning Electron Microscopy (SEM) Protocols for Problematic Plant, Oomycete, and Fungal Samples. J. Vis. Exp. 2017, 2017, 55031. [Google Scholar] [CrossRef]
- Lange, K.; Mogoi, S.; Van Weert, F. The Economics of Ecosystem Services of the Tana River Basin; Wetlands International Kenya: Nairobi, Kenya, 2015. [Google Scholar]
- Bush, A.O.; Lafferty, K.D.; Lotz, J.M.; Shostak, A.W. Parasitology Meets Ecology on Its Own Terms: Margolis et al. Revisited. J. Parasitol. 1997, 83, 575–583. [Google Scholar] [CrossRef]
- Hillis, D.M.; Dixon, M.T. Ribosomal DNA: Molecular Evolution and Phylogenetic Inference. Q. Rev. Biol. 1991, 66, 411–453. [Google Scholar] [CrossRef] [PubMed]
- Vanhove, M.P.M.; Tessens, B.; Schoelinck, C.; Jondelius, U.; Littlewood, D.T.J.; Artois, T.; Huyse, T. Problematic Barcoding in Flatworms: A Case-Study on Monogeneans and Rhabdocoels (Platyhelminthes). Zookeys 2013, 365, 355–379. [Google Scholar] [CrossRef] [PubMed]
- Kmentová, N.; Koblmüller, S.; Van Steenberge, M.; Raeymaekers, J.A.M.; Artois, T.; De Keyzer, E.L.R.; Milec, L.; Muterezi Bukinga, F.; Mulimbwa N’sibula, T.; Masilya Mulungula, P.; et al. Weak Population Structure and Recent Demographic Expansion of the Monogenean Parasite Kapentagyrus spp. Infecting Clupeid Fishes of Lake Tanganyika, East Africa. Int. J. Parasitol. 2020, 50, 471–486. [Google Scholar] [CrossRef]
- Kmentová, N.; Hahn, C.; Koblmüller, S.; Zimmermann, H.; Vorel, J.; Artois, T.; Gelnar, M.; Vanhove, M.P.M. Contrasting Host-Parasite Population Structure: Morphology and Mitogenomics of a Parasitic Flatworm on Pelagic Deepwater Cichlid Fishes from Lake Tanganyika. Biology 2021, 10, 797. [Google Scholar] [CrossRef]
- Newton, L.A.; Chilton, N.B.; Beveridge, I.; Hoste, H.; Nansen, P.; Gasser, R.B. Genetic Markers for Strongylid Nematodes of Livestock Defined by PCR-Based Restriction Analysis of Spacer RDNA. Acta Trop. 1998, 69, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Littlewood, D.T.J.; Rohde, K.; Clough, K.A. Parasite Speciation within or between Host Species?—Phylogenetic Evidence from Site-Specific Polystome Monogeneans. Int. J. Parasitol. 1997, 27, 1289–1297. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic Local Alignment Search Tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Kimura, M. A Simple Method for Estimating Evolutionary Rates of Base Substitutions through Comparative Studies of Nucleotide Sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Tamura, K.; Nei, M. Estimation of the Number of Nucleotide Substitutions in the Control Region of Mitochondrial DNA in Humans and Chimpanzees. Mol. Biol. Evol. 1993, 10, 512–526. [Google Scholar] [CrossRef] [PubMed]
- Matschiner, M. Fitchi: Haplotype Genealogy Graphs Based on the Fitch Algorithm. Bioinformatics 2016, 32, 1250–1252. [Google Scholar] [CrossRef]
- Caffara, M.; Davidovich, N.; Falk, R.; Smirnov, M.; Ofek, T.; Cummings, D.; Gustinelli, A.; Fioravanti, M.L. Redescription of Clinostomum phalacrocoracis Metacercariae (Digenea: Clinostomidae) in Cichlids from Lake Kinneret, Israel. Parasite 2014, 21, 32. [Google Scholar] [CrossRef] [PubMed]
- Caffara, M.; Locke, S.A.; Cristanini, C.; Davidovich, N.; Markovich, M.P.; Fioravanti, M.L. A Combined Morphometric and Molecular Approach to Identifying Metacercariae of Euclinostomum heterostomum (Digenea: Clinostomidae). J. Parasitol. 2016, 102, 239–248. [Google Scholar] [CrossRef] [PubMed]
- Monnens, M.; Halajian, A.; Littlewood, D.T.J.; Briscoe, A.G.; Artois, T.; Vanhove, M.P.M. Can Avian Flyways Reflect Dispersal Barriers of Clinostomid Parasites? First Evidence from the Mitogenome of Clinostomum complanatum. Gene 2023, 851, 146952. [Google Scholar] [CrossRef]
- Paradis, E. Analysis of Haplotype Networks: The Randomized Minimum Spanning Tree Method. Methods Ecol. Evol. 2018, 9, 1308–1317. [Google Scholar] [CrossRef]
- Nguyen, J.A.; Woodyard, E.T.; McAllister, C.T.; Marcquenski, S.V.; Rose, D.; Slifka, C.M.; Robison, L.R.S.; Griffin, M.J.; Rosser, T.G. Morphological and Molecular Data Establishes Clinostomum dolichorchum n. sp. (Digenea: Clinostomidae) in the Great Blue Heron Ardea herodias L. and American Bullfrog Rana catesbeiana Shaw. Syst. Parasitol. 2024, 101, 1–26. [Google Scholar] [CrossRef]
- Hamouda, A.H.; Younis, A.E. Characterization of Clinostomum cutaneum and Clinostomum phalacrocoracis in Tilapia Species of Aswan Governorate, Egypt: A Morphological, Molecular and Histopathological Study. Aquac. Res. 2021, 52, 6726–6740. [Google Scholar] [CrossRef]
- Mahdy, O.A.; Attia, M.M.; Shaheed, I.B.; Abdelsalam, M.; Elgendy, M.Y.; Salem, M.A. Evaluation of Praziquantel Effectiveness in Treating Nile Tilapia Clinostomid Infections and Its Relationships to Fish Health and Water Quality. BMC Vet. Res. 2024, 20, 449. [Google Scholar] [CrossRef]
- Echi, P.C.; Eyo, J.E.; Okafor, F.C.; Onyishi, G.C.; Ivoke, N. First Record of Co-Infection of Three Clinostomatid Parasites in Cichlids (Osteichthyes: Cichlidae) in a Tropical Freshwater Lake. Iran. J. Public Health 2012, 41, 86–90. [Google Scholar]
- Lo, C.F.; Chen, S.C.; Wang, C.H. The Study of Clinostomum complanatum (RUD., 1814) V. The Influences of Metacercaria of Clinostomum complanatum on Fish. Fish Pathol. 1985, 20, 305–312. [Google Scholar] [CrossRef]
- Eiras, C.; Luiza, M.; Goulart, G.; Pavanelli, G.C.; Machado, M.H. Histological Studies on the Effects of Clinostomum marginatum (Digenea, Clinostomidae) in Its Second Intermediate Host Loricariichthys platymetopon (Osteichthyes, Loricariidae) of the Upper Paraná River, Brazil. Acta Sci. Biol. Sci. 1999, 21, 237–241. [Google Scholar]


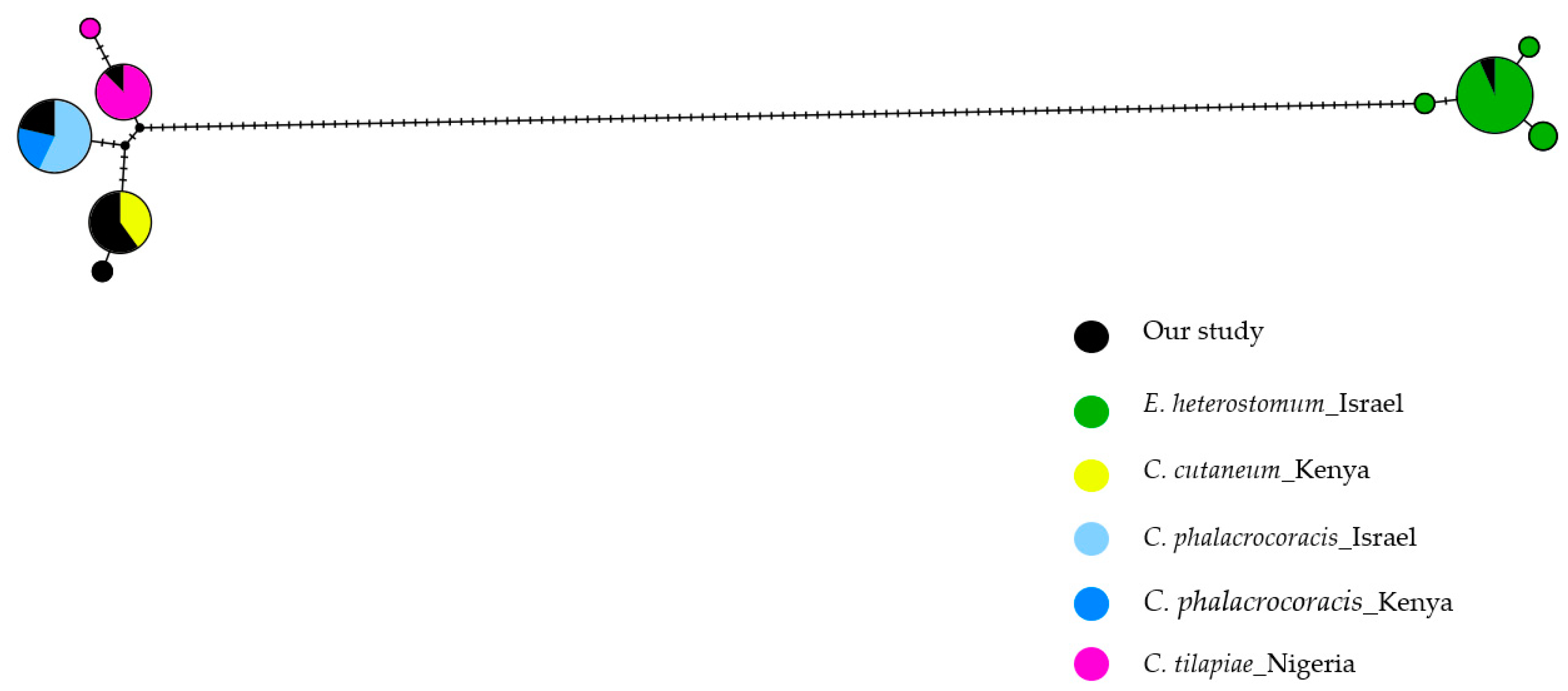
| Parasite Species | Host Species | Locality | Accession Number | Reference |
|---|---|---|---|---|
| C. cutaneum | Oreochromis niloticus | Sagana, Kenya | PV123689-91, PV123693-95, PV123697, PV123699 | Present study |
| C. phalacrocoracis | Oreochromis niloticus | Sagana, Kenya | PV123696, PV123698, PV123700 | Present study |
| C. tilapiae | Oreochromis niloticus | Sagana, Kenya | PV123701 | Present study |
| E. heterostomum | Oreochromis niloticus | Sagana. Kenya | PV123692 | Present study |
| C. cutaneum | Oreochromis niloticus | Sagana, Kenya | KP110564-65 | [14] |
| C. cutaneum | Oreochromis niloticus | Sagana, Kenya | FJ609421 | [9] |
| C. cutaneum | Ardea cinerea | Sagana, Kenya | GQ339114 | [9] |
| C. phalacrocoracis | Ardea cinerea | Sagana, Kenya | FJ609423 | [9] |
| C. phalacrocoracis | Oreochromis niloticus | Sagana, Kenya | FJ609422 | [9] |
| C. phalacrocoracis | Oreochromis niloticus | Sagana, Kenya | KP110567-69 | [14] |
| C. phalacrocoracis | Cichlids | Lake Kinneret, Israel | KJ786975-982 | [37] |
| C. tilapiae | Synodontis batensoda | Anambra basin, Nigeria | KY649349-356 | [15] |
| E. heterostomum | Cichlids | Lake Kinneret, Israel | KP721422-439 | [38] |
| Interspecific | Intraspecific | |||||
|---|---|---|---|---|---|---|
| C. tilapiae | C. cutaneum | C. phalacrocoracis | E. heterostomum | # Differences | Distance | |
| C. tilapiae | 7–11 | 6–10 | 120–126 | 0–3 | 0.000–0.006 | |
| C. cutaneum | 0.008–0.015 | 7–9 | 121–125 | 0–1 | 0.000–0.002 | |
| C. phalacrocoracis | 0.006–0.012 | 0.004–0.014 | 122–126 | 0–1 * | 0.000 | |
| E. heterostomum | 0.169–0.215 | 0.203–0.220 | 0.199–0.220 | 0–3 | 0.000–0.014 |
| Interspecific | Intraspecific | |||
|---|---|---|---|---|
| C. cutaneum | C. tilapiae | # Differences | Distance | |
| C. cutaneum | 0–1 | 0.000–0.009 | ||
| C. tilapiae | 0.114–0.131 | - | - | |
| C. phalacrocoracis | 0.124–0.143 | 0.081–0.127 | 0 | 0.000 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shigoley, M.I.; Kmentová, N.; Ndegwa, D.M.; Topić, M.; Thys, K.J.M.; Vanhove, M.P.M. Scanning Electron Microscopy Reveals New Ultrastructural Features in Metacercariae of Clinostomum cutaneum (Digenea: Clinostomidae) Infecting Oreochromis niloticus (Actinopterygii: Cichlidae) in Kenya. Pathogens 2025, 14, 249. https://doi.org/10.3390/pathogens14030249
Shigoley MI, Kmentová N, Ndegwa DM, Topić M, Thys KJM, Vanhove MPM. Scanning Electron Microscopy Reveals New Ultrastructural Features in Metacercariae of Clinostomum cutaneum (Digenea: Clinostomidae) Infecting Oreochromis niloticus (Actinopterygii: Cichlidae) in Kenya. Pathogens. 2025; 14(3):249. https://doi.org/10.3390/pathogens14030249
Chicago/Turabian StyleShigoley, Miriam Isoyi, Nikol Kmentová, Daniel Mungai Ndegwa, Martina Topić, Kelly J. M. Thys, and Maarten P. M. Vanhove. 2025. "Scanning Electron Microscopy Reveals New Ultrastructural Features in Metacercariae of Clinostomum cutaneum (Digenea: Clinostomidae) Infecting Oreochromis niloticus (Actinopterygii: Cichlidae) in Kenya" Pathogens 14, no. 3: 249. https://doi.org/10.3390/pathogens14030249
APA StyleShigoley, M. I., Kmentová, N., Ndegwa, D. M., Topić, M., Thys, K. J. M., & Vanhove, M. P. M. (2025). Scanning Electron Microscopy Reveals New Ultrastructural Features in Metacercariae of Clinostomum cutaneum (Digenea: Clinostomidae) Infecting Oreochromis niloticus (Actinopterygii: Cichlidae) in Kenya. Pathogens, 14(3), 249. https://doi.org/10.3390/pathogens14030249






